Abstract
Amphipathic polymers known as “amphipols” provide a highly stabilizing environment for handling membrane proteins in aqueous solutions. A8-35, an amphipol with a polyacrylate backbone and hydrophobic grafts, has been extensively characterized and widely employed for structural and functional studies of membrane proteins using biochemical and biophysical approaches. Given the sensitivity of membrane proteins to their environment, it is important to examine what effects amphipols may have on the structure and dynamics of the proteins they complex. Here we present the first molecular dynamics study of an amphipol-stabilized membrane protein, using Escherichia coli OmpX as a model. We begin by describing the structure of the complexes formed by supplementing OmpX with increasing amounts of A8-35, in order to determine how the amphipol interacts with the transmembrane and extramembrane surfaces of the protein. We then compare the dynamics of the protein in either A8-35, a detergent, or a lipid bilayer. We find that protein dynamics on all accessible length scales is restrained by A8-35, which provides a basis to understanding some of the stabilizing and functional effects of amphipols that have been experimentally observed.









Similar content being viewed by others
Abbreviations
- 2D:
-
Two-dimensional
- A8-35:
-
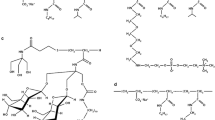
A poly(sodium acrylate)-based amphipol comprising ~35 % of free carboxylates, ~25 % of octyl chains, ~40 % of isopropyl groups
- APol:
-
Amphipol
- CG:
-
Coarse-grain
- DDM:
-
n-Dodecyl-β-d-maltopyranoside
- DHPC:
-
Dihexanoylphosphatidylcholine
- DOPC:
-
Dioleoylphosphatidylcholine
- EM:
-
Electron microscopy
- FWHM:
-
Full width at half-maximum
- MD:
-
Molecular dynamics
- MP:
-
Membrane protein
- OmpX:
-
Outer membrane protein X from Escherichia coli
- PCA:
-
Principal component analysis
- RCG:
-
Reverse coarse-grain
- RMSF:
-
Root mean squared fluctuations
- SERCA1a:
-
The fast twitch sarcoplasmic calcium pump
References
Althoff T, Mills DJ, Popot J-L, Kühlbrandt W (2011) Arrangement of electron transport chain components in bovine mitochondrial supercomplex I1III2IV1. EMBO J 30:4652–4664
Böckmann RA, Caflisch A (2005) Spontaneous formation of detergent micelles around the outer membrane protein OmpX. Biophys J 88:3191–3204
Catoire LJ, Zoonens M, van Heijenoort C et al (2009) Inter- and intramolecular contacts in a membrane protein/surfactant complex observed by heteronuclear dipole-to-dipole cross-relaxation. J Magn Reson 197:91–95
Catoire LJ, Zoonens M, van Heijenoort C et al (2010) Solution NMR mapping of water-accessible residues in the transmembrane beta-barrel of OmpX. Eur Biophys J 39:623–630
Champeil P, Menguy T, Tribet C et al (2000) Interaction of amphipols with sarcoplasmic reticulum Ca2+-ATPase. J Biol Chem 275:18623–18637
Charvolin D, Picard M, Huang L-S, et al. (2014) Solution behavior and crystallization of cytochrome bc1 in the presence of amphipols. J Membrane Biol. doi:10.1007/s00232-014-9694-4
Choutko A, Glättli A, Fernández C et al (2011) Membrane protein dynamics in different environments: simulation study of the outer membrane protein X in a lipid bilayer and in a micelle. Eur Biophys J 40:39–58
Dahmane T, Rappaport F, Popot J-L (2013) Amphipol-assisted folding of bacteriorhodopsin in the presence or absence of lipids: functional consequences. Eur Biophys J 42:85–101
Etzkorn M, Zoonens M, Catoire LJ et al (2014) How amphipols embed membrane proteins: global solvent accessibility and interaction with a flexible protein terminus. J Membr Biol. doi:10.1007/s00232-014-9657-9
Fernández C, Adeishvili K, Wüthrich K (2001) Transverse relaxation-optimized NMR spectroscopy with the outer membrane protein OmpX in dihexanoyl phosphatidylcholine micelles. Proc Natl Acad Sci USA 98:2358–2363
Giusti F, Popot J-L, Tribet C (2012) Well-defined critical association concentration and rapid adsorption at the air/water interface of a short amphiphilic polymer, amphipol A8-35: a study by Förster resonance energy transfer and dynamic surface tension measurements. Langmuir 28:10372–10380
Giusti F, Rieger J, Catoire LJ et al (2014) Synthesis, characterization and applications of a perdeuterated amphipol. J Membr Biol. doi: 10.1007/s00232-014-9656-x
Gohon Y, Pavlov G, Timmins P et al (2004) Partial specific volume and solvent interactions of amphipol A8-35. Anal Biochem 334:318–334
Gohon Y, Giusti F, Prata C et al (2006) Well-defined nanoparticles formed by hydrophobic assembly of a short and polydisperse random terpolymer, amphipol A8-35. Langmuir 22:1281–1290
Gohon Y, Dahmane T, Ruigrok RWH et al (2008) Bacteriorhodopsin/amphipol complexes: structural and functional properties. Biophys J 94:3523–3537
Hagn F, Etzkorn M, Raschle T, Wagner G (2013) Optimized phospholipid bilayer nanodiscs facilitate high-resolution structure determination of membrane proteins. J Am Chem Soc 135:1919–1925. doi:10.1021/ja310901f
Hess B, Kutzner C, van der Spoel D, Lindahl E (2008) GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J Chem Theory Comput 4:435–447
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38
Huynh KW, Cohen MR, Moiseenkova-Bell VY (2014) Application of amphipols for structure-functional analysis of TRP channels. J Membr Biol. doi:10.1007/s00232-014-9684-6
Kim PS, Baldwin RL (1982) Influence of charge on the rate of amide proton exchange. Biochemistry (Mosc) 21:1–5. doi:10.1021/bi00530a001
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132
Liao M, Cao E, Julius D, Cheng Y (2013) Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504:107–112
Liao M, Cao E, Julius D, Cheng Y (2014) Single particle electron cryo-microscopy of a mammalian ion channel. Curr Opin Struct Biol 27:1–7. doi:10.1016/j.sbi.2014.02.005
Marrink SJ, Risselada HJ, Yefimov S et al (2007) The MARTINI force field: coarse grained model for biomolecular simulations. J Phys Chem B 111:7812–7824
Martinez KL, Gohon Y, Corringer P-J et al (2002) Allosteric transitions of Torpedo acetylcholine receptor in lipids, detergent and amphipols: molecular interactions vs. physical constraints. FEBS Lett 528:251–256
Periole X, Cavalli M, Marrink S-J, Ceruso MA (2009) Combining an elastic network with a coarse-grained molecular force field: structure, dynamics, and intermolecular recognition. J Chem Theory Comput 5:2531–2543
Perlmutter JD, Drasler WJ II, Xie W et al (2011) All-atom and coarse-grained molecular dynamics simulations of a membrane protein stabilizing polymer. Langmuir 27:10523–10537
Picard M, Dahmane T, Garrigos M et al (2006) Protective and inhibitory effects of various types of amphipols on the Ca2+-ATPase from sarcoplasmic reticulum: a comparative study. Biochemistry 45:1861–1869
Planchard N, Point É, Dahmane T et al (2014) The use of amphipols for solution NMR studies of membrane proteins: advantages and constraints as compared to other solubilizing media. J Membr Biol. doi:10.1007/s00232-014-9654-z
Pocanschi CL, Popot J-L, Kleinschmidt JH (2013) Folding and stability of outer membrane protein A (OmpA) from Escherichia coli in an amphipathic polymer, amphipol A8-35. Eur Biophys J 42:103–118
Popot J-L (2010) Amphipols, nanodiscs, and fluorinated surfactants: three nonconventional approaches to studying membrane proteins in aqueous solutions. Annu Rev Biochem 79:737–775
Popot J-L, Berry EA, Charvolin D et al (2003) Amphipols: polymeric surfactants for membrane biology research. Cell Mol Life Sci 60:1559–1574
Popot J-L, Althoff T, Bagnard D et al (2011) Amphipols from A to Z. Annu Rev Biophys 40:379–408
Swift J (1726) Travels into several remote nations of the world. In four parts. By Lemuel Gulliver, first a surgeon, and then a captain of several ships. Benjamin Motte, London
Tehei M, Giusti F, Zaccai G, Popot J-L (2014) Thermal fluctuations in amphipol A8-35 particles measured by neutron scattering. J Membr Biol (under review)
Tribet C, Audebert R, Popot JL (1996) Amphipols: polymers that keep membrane proteins soluble in aqueous solutions. Proc Natl Acad Sci USA 93:15047–15050
Vogt J, Schulz GE (1999) The structure of the outer membrane protein OmpX from Escherichia coli reveals possible mechanisms of virulence. Structure 7:1301–1309
Zoonens M, Popot J-L (2014) Amphipols for each season. J Membr Biol. doi:10.1007/s00232-014-9666-8
Zoonens M, Catoire LJ, Giusti F, Popot J-L (2005) NMR study of a membrane protein in detergent-free aqueous solution. Proc Natl Acad Sci USA 102:8893–8898
Zoonens M, Giusti F, Zito F, Popot J-L (2007) Dynamics of membrane protein/amphipol association studied by Förster resonance energy transfer: implications for in vitro studies of amphipol-stabilized membrane proteins. Biochemistry 46:10392–10404
Acknowledgments
Particular thanks are due to L. J. Catoire, M. Tehei, and G. Zaccai for discussions about the manuscript, as well as to a referee for useful comments. Computational resources were provided by the Minnesota Supercomputing Institute (MSI). This work was also supported by the French Centre National de la Recherche Scientifique.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Perlmutter, J.D., Popot, JL. & Sachs, J.N. Molecular Dynamics Simulations of a Membrane Protein/Amphipol Complex. J Membrane Biol 247, 883–895 (2014). https://doi.org/10.1007/s00232-014-9690-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-014-9690-8