Abstract
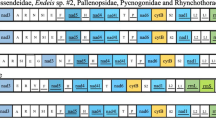
The phylogenetic relationships among all living families of sea spiders (Arthropoda: Pycnogonida) are investigated using nearly complete 18S rRNA sequences from 57 ingroup species and five chelicerates under the Bayesian and maximum likelihood methods. Monophyly of Colossendeidae, Pycnogonidae, Phoxichilidiidae, Endeidae and Pallenopsidae is consistently supported. However, the genera formerly classified in the family Ammotheidae are split up into two distantly related groups. The genera Ascorhynchus and Eurycyde (here recognized as Ascorhynchidae) are possibly an early offshoot of sea spiders, whereas other ammotheids constitute a robust terminal clade with Pallenopsidae, Phoxichilidiidae and Endeidae. This topology also opposes the prevalent assumption of successive losses and simplification of three kinds of cephalic appendages like in a previous cladistic analysis. At least three independent losses are suggested for palps by the inferred topology, and both chelifores and female ovigers may have been lost twice. Our knowledge of early ontogeny and internal anatomy is more congruent with the present 18S rRNA data. The families Callipallenidae and Nymphonidae with unique “attaching larvae” are grouped together in present molecular trees, suggesting that extended paternal care of offspring evolved only once in Pycnogonida. Confident clustering of Pycnogonidae and Rhynchothoracidae indicates that the number of female genital pores is an evolutionary conservative character.


Similar content being viewed by others
References
Arango CP (2002) Morphological phylogenetics of the sea spiders (Arthropoda: Pycnogonida). Org Divers Evol 2:107–125
Arango CP (2003) Molecular approach to the phylogenetics of sea spiders (Arthropoda: Pycnogonida) using partial sequences of nuclear ribosomal DNA. Mol Phylogenet Evol 28:588–600
Arango CP, Wheeler WC (2007) Phylogeny of the sea spiders (Arthropoda: Pycnogonida) based on direct optimization of six loci and morphology. Cladistics 23:255–293
Arnaud F, Bamber RN (1987) The biology of Pycnogonida. Adv Mar Biol 24:1–95
Brusca RC, Brusca GJ (2003) Invertebrates, 2nd edn. Sinauer Associates, Sunderland
Child CA (1979) Shallow-water Pycnogonida of the Isthmus of Panama and the coast of middle America. Smithson Contrib Zool 293:1–86
Child CA (1998) The marine fauna of New Zealand: Pycnogonida (Sea Spiders). NIWA Biodivers Mem 109:1–71
Child CA (2001) Resolution for a confusing problem in the classification of Pycnogonida by setting a new family Pallenopsidae. PhD Thesis, Hokkaido University
Dunlop JA, Arango CP (2004) Pycnogonid affinities: a review. J Zool Syst Evol Res 43:8–21
Fry WG (1978) A classification within the pycnogonids. Zool J Linn Soc 63:35–38
Gillespie JM, Bain BA (2006) Postembryonic development of Tanystylum bealensis (Pycnogonida, Ammotheidae) from Barkley Sound, British Columbia, Canada. J Morphol 267:308–317
Giribet G, Edgecombe GD, Wheeler WC (2001) Arthropod phylogeny based on eight molecular loci and morphology. Nature 413:157–161
Hassanin A (2006) Phylogeny of Arthropoda inferred from mitochondrial sequences: strategies for limiting the misleading effects of multiple changes in pattern and rates of substitution. Mol Phylogenet Evol 38:100–116
Hedgpeth JW (1947) On the evolutionary significance of the Pycnogonida. Smithson Misc Coll 106:1–53
Hedgpeth JW (1948) The Pycnogonida of the western North Atlantic and the Caribbean. Proc US Natl Mus 98:157–342
Hedgpeth JW (1982) Pycnogonida. In: Parker SP (ed) Synopsis and classification of living organisms, vol 2, pp 169–173
Hoek PPC (1881) Nouvelles études sur les Pycnogonides. Arch Zool Exp Gén 9:445–542
Jager M, Murienne J, Clabaut C, Deutsch J, Le Guyader H, Manuel M (2006) Homology of arthropod anterior appendages revealed by Hox gene expression in a sea spider. Nature 441:506–508
Löytynoja A, Milinkovitch MC (2003) A hidden Markov model for progressive multiple alignment. Bioinformatics 19:1505–1513
Maddison DR, Maddison WP (2005) MacClade 4: analysis of phylogeny and character evolution. Version 4.08. Sinauer Associates, Sunderland
Mallatt J, Giribet G (2006) Further use of nearly complete 28S and 18S rRNA genes to classify Ecdysozoa: 37 more arthropods and a kinorhynch. Mol Phylogenet Evol 40:772–794
Manuel M, Jager M, Murienne J, Clabaut C, Guyader HL (2006) Hox genes in sea spiders (Pycnogonida) and the homology of arthropod head segments. Dev Genes Evol 216:481–491
Maxmen A, Browne WE, Martindale MQ, Giribet G (2005) Neuroanatomy of sea spiders implies an appendicular origin of the protocerebral segment. Nature 437:1144–1148
Morgan TH (1891) A contribution to the embryology and phylogeny of the pycnogonids. Stud Biol Lab J Hopkins Univ 5:1–76
Munilla T (1999) Evolución y filogenia de los Picnogónidos. In: Melic A, de Haro JJ, Mendez M, Ribera I (eds) Evolución y filogenia de Arthropoda. Boletin de la Sociedad Entomológica Aragonesa 26:273–279
Nakamura K (1981) Post embryonic development of a pycnogonid, Propallene longiceps. J Nat Hist 15:49–62
Nakamura K, Child CA (1983) Shallow-water Pycnogonida from the Izu Peninsula, Japan. Smiths Contrib Zool 386:1–71
Nakamura K, Child CA (1991) Pycnogonida from waters adjacent to Japan. Smithson Contrib Zool 512:1–74
Okuda S (1940) Metamorphosis of a pycnogonid parasitic in a hydromedusa. J Fac Sci Hokkaido Univ 7:73–86
Podsiadlowski L, Braband A (2006) The complete mitochondrial genome of the sea spider Nymphon gracile (Arthropoda: Pycnogonida). BMC Genomics 7:284
Poschmann M, Dunlop JA (2006) A new sea spider (Arthropoda: Pycnogonida) with a flagelliform telson from Lower Devonian Hunsrück Slate, Germany. Palaeontology 49:983–989
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818
Regier JC, Shultz JW, Kambic RE (2005) Pancrustacean phylogeny: hexapods are terrestrial crustaceans and maxillopods are not monophyletic. Proc R Soc B 272:395–401
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Shimodaira H, Hasegawa M (1999) Multiple comparisons of log-likelihood with applications to phylogenetic inference. Mol Biol Evol 16:1114–1116
Siveter DJ, Sutton MD, Briggs DEG, Siveter DJ (2004) A Silurian sea spider. Nature 431:978–980
Stock JH (1955) Pycnogonida from the West Indies, Central America and the Pacific Coast of North America. Papers from Dr. Mortensen’s Pacific Expedition 1914–1916. Vidensk Medd fra Dansk Naturh Foren 117:209–266
Stock JH (1974) Pycnogonida from the continental shelf, slope, and deep sea of the tropical Atlantic and East Pacific. Bull Mar Sci 24:957–1092
Stock JH (1994) Indo-west pacific Pycnogonida collected by some major oceanographic expeditions. Beaufortia 44:17–77
Swofford DL (2002) PAUP*. Phylogenetic analysis using parsimony (*and other methods). Version 4. Sinauer Associates, Sunderland
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acid Res 24:4876–4882
Vilpoux K, Waloszek D (2003) Larval development and morphogenesis of the sea spider Pycnogonum litorale (Ström, 1762) and the tagmosis of the body of Pantopoda. Arthropod Struct Dev 32:349–383
Waloszek D, Dunlop JA (2002) A larval sea spider (Arthropoda: Pycnogonida) from the Upper Cambrian “Orsten” of Sweden, and the phylogenetic position of pycnogonids. Palaeontology 45:421–446
Wheeler WC (1996) Optimization alignment: the end of multiple sequence alignment in phylogenetics? Cladistics 12:1–9
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR Protocols: a guide to methods and applications. Academic, New York, pp 315–322
Zwickl DJ (2006) Genetic algorithm approaches for the phylogenetic analysis of large biological sequence datasets under the maximum likelihood criterion. PhD Thesis, University Texas, Austin
Acknowledgments
We are deeply indebted to the following colleagues for their help in obtaining pycnogonid material: Tadashi Akiyama (Okayama University), Kazunori Hasegawa, Katsumi Kubodera and Hiroshi Namikawa (National Science Museum, Tokyo), Isadora Kotuku (Aqua Scape Research Co.), Masaaki Ito (Tokyo), Dhugal Linsay and Hiroshi Miyake (JAMSTEC), Katumi Miyazaki (Kyoto University), Susumu Ohtsuka (Hiroshima University), Suguru Ohta and Takenori Sasaki (University of Tokyo), Hiroshi Oida (Chiba Prefectural Agriculture Research Center), Koji Tojo (Shinshu University), Hajime Ueda (Shizuoka), and Kensuke Yanagi (Natural History Museum and Institute, Chiba). We are also indebted to the crews of R.V. Tansei-Maru and Hakuho-Maru (University of Tokyo), R.V. Natsushima (JAMSTEC) and R.V. Toyoshio-Maru (Hiroshima University). Special thanks are given to Kazuki Kanakatsu and Kunikatsu Hamano (Tokyo University of Agriculture and Technology), Kazuhiro Nakaya (Hokkaido University), Shigeaki Kojima (University of Tokyo), Ryuichiro Machida and Seiji Tokumasu (Tsukuba University), Reiko Nakamura (Sequella Inc.), Hironobu Suzuki (Olympus Co.), and Yoshihisa Suyama (Tohoku University) for technical help and discussion. We also thank Claudia Arango (Queensland Museum), Amy Maxmen (Harvard University) and anonymous reviewers for their invaluable comments on earlier versions of this manuscript. Allan Child gave us suggestions in species identification. This study was partly supported by a research grant from Nikaido Gakuen, Tokyo.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Nishida.
Rights and permissions
About this article
Cite this article
Nakamura, K., Kano, Y., Suzuki, N. et al. 18S rRNA phylogeny of sea spiders with emphasis on the position of Rhynchothoracidae. Mar Biol 153, 213–223 (2007). https://doi.org/10.1007/s00227-007-0803-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00227-007-0803-0