Abstract
Saffron, a spice obtained by drying the scars of saffron flowers (Crocus sativus L.), is by far the most valuable spice in the world, and, not surprisingly, it often becomes an object of adulteration. In this study, non-target screening of volatile compounds in 38 authentic saffron samples and 25 samples of plant materials, potential saffron adulterants (safflower, calendula, capsicum and turmeric) was performed using headspace solid-phase microextraction for sample extraction followed by gas chromatography coupled to high-resolution mass spectrometry (HS-SPME-GC-HRMS) for non-target volatiles screening. The chemometric analysis of generated data by principal component analysis (PCA) and partial least squares discriminative analysis (PLS-DA) showed good separation of authentic saffron from potential plant adulterants. Significant compounds (‘markers’) for each group of these plants, as well as for saffron were tentatively identified. The target screening of selected ‘markers’ in model admixtures enabled simple and reliable detection levels as low as 2% w/w of safflower, calendula, capsicum or turmeric present in saffron.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Saffron, the most valuable spice in the world, is obtained by drying the stigma of saffron flowers (Crocus sativus L.) [1]. The high price of saffron is due to the traditional collection method, where the stigma must be collected by hand, and the collection yield is minimal [2, 3].
Considering the high economic value and the limited sources of raw material, saffron is vulnerable to fraud. One of the most frequent fraudulent practices is dilution by low-priced plant materials such as safflower petals (Carthamus tinctorius), calendula petals (Calendula officinalis), turmeric powder (Curcuma longa), beet fibers (Beta vulgaris), pomegranate fibers (Punica granatum), arnica flowers (Arnica montana) and achiote seeds (Bixa orellana) [4,5,6] with similar color and morphology to saffron [3, 5].
The chemical composition of saffron, like that of other spices, is very complex, containing a high number of secondary metabolites responsible for characteristic bioactivities and overall quality. The major phytochemicals characteristic for saffron are crocetin glycosidic esters associated with a typical golden-yellow tinge of flavored food [1, 5], safranal, the key component of volatiles fraction, and picrocrocin, which is responsible for the characteristic bitter taste of this spice. Picrocrocin, theunique secondary metabolite, is one of the known molecular markers of saffron [1, 3]. Regarding saffron aroma, isophorone related compounds (structurally similar to safranal), C13-norisoprenoids derived from lipophilic carotenoids, and saturated hydrocarbons such as octadecane, docosane, tricosane, and pentacosane are typically represented [5, 7, 8].
Quality and classification of saffron is defined by ISO 3632–2 2010 (microscopic analysis for detection of extraneous material in saffron) and ISO 362–1 2011 (UV–Vis), which is based on measurement of the absorption maxima values of aqueous saffron extracts at 257 nm (detection of picrocrocin), 330 nm (detection of safranal) and 440 nm (detection of crocins)[1, 6]. However, the above-mentioned UV–Vis spectrophotometric method has been shown as insufficient when saffron is adulterated by plant material with a similar color and morphology. The study performed by Sabatino et. al [9]. demonstrated that the UV–Vis method did not enable detection of the addition of adulterants (safflower, calendula, and turmeric) at a level lower than 20% w/w. Besides the molecular biology-based approach employing polymerase chain reaction (PCR) followed by sequencing [10, 11], a number of instrumental strategies such as nuclear magnetic resonance (NMR) [2, 12, 13], liquid chromatography coupled with mass spectrometry (LC–MS) [3, 14, 15], infrared spectroscopy (IR) [16,17,18] were used for saffron authentication.
In this study, we aimed to investigate in depth the application potential of head-space sampling of saffron volatiles by solid-phase microextraction extraction followed by gas chromatography coupled to high-resolution mass spectrometry (HS-SPME-GC-HRMS) for authentication purpose. There was only one study employing SPME-GC-HRMS for the detection of plant-derived adulterant [6] and we were wondering whether our approach using chemometric data handling would be able to identify similar markers and which lowest concentration of plant adulterant (safflower, calendula, capsicum and turmeric) in the admixture could be recognized.
Materials and methods
Samples
In total, 38 authentic saffron (Crocus sativus L.) samples from different origins (Iran, Spain, Greece and Italy) were obtained for this study in collaboration with the Czech Agriculture and Food Inspection Authority. Also, 25 samples of spices and plant material, which could be used for an adulteration of saffron, were collected for this study. They were Turmeric (Curcuma longa L., n = 6), Calendula (Calendula officinalis L., n = 9), Capsicum (Capsicum annum L., n = 6) and Safflower (Carthamus tinctorius L., n = 4). Some of the samples were supplied as ground material (powder). In some samples (mostly saffron obtained as dried stigma), a fine homogenization was required. To avoid the loss of volatiles, the homogenization was manually performed in the ceramic grinding bowl. All samples were stored in gas-tight containers in the freezer at – 18 °C.
Chemicals and materials
SPME fibers coated with divinylbenzene/carboxen/polydimethylsiloxane (DVB/CAR/PDMS, 50/30 μm, 1 cm) were purchased from Supelco (Bellefonte, USA). Sodium chloride was purchased from Penta (Prague, Czech Republic). Water purified by a Milli-Q® Integral system supplied by Merck was used throughout the study. C7–C30 saturated alkanes mixture (certified reference material, 1000 μg/mL each component in hexane), used for unknown compounds retention index (RI) based identification, was obtained from Sigma-Aldrich (Steinheim, Germany).
Sample preparation
Non-target screening of volatile profiles of saffron and adulterants
Finely ground samples (50 mg) were placed in SPME crimp cap vials (10 mL) and 2 mL of a saturated solution of NaCl (26.4% w/w) was added. The vials were immediately crimped with a gas-tight SPME cap. Each sample was prepared in one repetition. In addition, samples for background measurement (blanks) were prepared (2 mL of a saturated solution of NaCl) as well as quality control samples (QC). For the QC sample, 50 mg of the pooled samples were used, and the QC sample was prepared in six repetitions. It should be noted that during the development of the SPME-GC-HRMS method, the different amount of the sample (25 mg, 50 mg and 100 mg) were considered, as well as the addition of a saturated solution of NaCl. Finally, the combination of 50 mg of the sample with the 2 mL of saturated solution of NaCl was chosen.
Artificially adulterated saffron samples
In this part of the experiment, saffron was artificially adulterated by mixing it with selected plant materials in different proportions (percentage by mass – % w/w). Specifically, for the mixtures of saffron with turmeric, paprika, safflower and calendula, the amounts of saffron/adulterant were as follows: 80/20% w/w, 90/10% w/w, 96/4% w/w and 98/2% w/w. The artificially adulterated samples as well as authentic saffron samples and pure adulterant samples were prepared as described in Sect. "Non-target screening of volatile profiles of saffron and adulterants”. in two repetitions.
SPME-GC-HRMS
For the extraction of head-spaced volatiles, the HS-SPME technique was used. The SPME parameters were first optimized to achieve the most efficient extraction of volatiles to the SPME fiber, at the same time it was crucial to test different desorption times to ensure the complete desorption of volatiles from the fiber to avoid cross contamination. For the temperature of extraction 40 °C, 45 °C and 50 °C were tested. Furthermore, the extraction time was tested at 10 min, 15 min and 20 min. As for desorption time, 1 min and 3 min were considered. The final parameters of the SPME method are described below. The temperature of incubation and extraction: 45 °C, sample incubation time – 20 min, extraction time – 10 min, sample desorption to the GC system – 3 min (desorption temperature – 240 °C).
The Agilent 7200B system consists of an Agilent 7890B gas chromatograph equipped with a multimode inlet, PAL RSI 85 for automated headspace–solid phase microextraction (HS–SPME) and direct injection, and quadrupole – time of flight mass spectrometer (Q-TOF) (Agilent Technologies, Palo Alto, California, USA) was employed. For instrument control and data acquisition, Mass-Hunter GC/MS Acquisition (Agilent Technologies, Palo Alto, California, USA) software (B.10.0.384.1) was used. Sample components were separated on a 30 m HP–5MS capillary column (0.25 mm id, film thickness: 0.25 µm; Agilent Technologies, Palo Alto, California, USA).
Samples were injected using split mode (5:1), injector temperature was 240 °C and the oven temperature program was as follows: 40 °C (2 min), 2 °C/min to 64.5 °C, 0.5 °C/min to 70 °C, 2 °C/min to 80 °C, 5 °C/min to 100 °C, 0.5 °C/min to 114 °C, 2 °C/min to 120 °C, 5 °C/min to 184 °C, 20 °C/min to 240 °C (5 min).
The mass spectrometric detector was operated in the electron ionization (EI) mode. The temperature of the transferline was 280 °C and the temperature of the ion source was 230 °C. The mass range setting was 40–550 m/z, the acquisition speed was 5 specter/s and the resolution of the mass analyzer was set > 12,500 (FWHM).
GC–MS data processing and chemometric analysis
First of all, data obtained by SPME-GC-HRMS was transferred to the SureMass format. Then, our SureMass data was subjected to peak detection, deconvolution, and library search (NIST17.L). The data was then converted to CEF files (.CEF). All these data processing steps were performed in MassHunter Unknowns Analysis (Version B.10.1). Based on the converted QC CEF file in MassHunter Quantitative Analysis (Version.B.10.1) was created a data processing method and performed a recursive analysis. All software were from Agilent Technologies, Palo Alto, California, USA. After the performance of a recursive analysis, data were exported to the MetaboAnalyst 5.0 software (https://www.metaboanalyst.ca). Employing this software, the following data treatment operations were performed: sum-normalization (relative intensities of each signal obtained by division of the sum of all signals), log transformation and Pareto scaling. These data processing operations ensure the normal distribution and higher significance of low abundant compounds. In addition, in MetaboAnalyst, principal component analysis (PCA) was performed to provide oversight of exported data. Analysis of Variance (ANOVA) or t-test (False Discovery Rate (FDR), p-value < 0.05) were employed to filter out the statistically insignificant compounds. The filtered data were normalized by sum in MS Excel and loaded to SIMCA (Sartorius, Göttingen, Germany), where unsupervised principal component analysis (PCA) and supervised partial least squares discriminant analysis (PLS-DA) was performed. R2 (cumulative) and Q2 (cumulative) parameters were obtained for each model by fourfold internal cross-validation and were used to determine the validity of the models. R2 (cum) shows the variation described by all components in the model and Q2 (cum) is a criterion of how accurately the model can predict class membership. The prediction and recognition abilities of the models were also obtained by fourfold internal cross-validation. The significant volatile compounds (‘markers’) were selected based on their Variable Importance in Projection (VIP) score in PLS-DA models. In this study, the most significant compounds with VIP score of more than 1.0 were considered. The Pattern Search function of MetaboAnalyst software which allows to specify and search for specific patterns of interest in the data was also used for detection of characteristic ‘markers’. This function uses a template matching method and the results are expressed as a ranked list of variables with the Spearman correlation coefficient and p-value. In this study the compounds with the Spearman correlation coefficient ≥ 0.80 were contemplated.
Identification of selected ‘markers’ was performed using mass spectra NIST 17 library, for tentative identification only compounds with match factor ≥ 750 and relevant Kovats retention indexes (RI) relative to n-alkanes (C7–C30) were considered. The compliance of exact mass of detected ions (mass error < 5 ppm) and isotopic pattern were used to confirm the identification.
Results and discussion
Optimization of SPME-GC-HRMS
For the purpose of this study, HS-SPME was the technique of choice. Sample preparation, extraction temperature together with desorption conditions in GC injector are described in detail in the Sects. “Chemicals and Materials “Sample Preparation" " SPME-GC-HRMS". With regards to different patterns of volatiles in the analyzed plant species, in addition to saffron also turmeric was used for method optimization. To control the repeatability of measurements, QC sample (described in the Sect. "Non-target screening of volatile profiles of saffron and adulterants") containing all the tested plants was analyzed throughout the study (the retention time tolerance was 0.2 min and the repeatability of the peaks areas 5%).
Non-target screening of saffron and plant adulterants
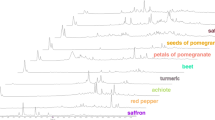
In the first phase of the experiments, 38 authentic saffron samples and 25 samples of different plant materials representing four plant species were analysed. The chromatograms shown in Fig. 1 demonstrate the difference in the main volatile profiles between the saffron samples and potential adulterants.
While volatiles with retention times not exceeding 45 min under used GC conditions on HP–5MS capillary column dominated in saffron (A), capsicum (D) and safflower (E), volatiles with relatively high intensities with retention times up to 60 min were present in turmeric (B) and calendula (C). When using the m/z value range 40–500 high number of compounds ranging in average from 150 (capsicum) to 550 (turmeric) was detected after application of deconvolution function.
Results of chemometric data evaluation for all samples
To investigate the differences and get an in-depth insight into the volatile profiles of saffron and its potential adulterants, chemometric analysis starting with PCA and followed by PLS-DA was performed. The score plot shown in Fig. 2 shows distinct separation of saffron samples from other plant matrices. In spite of the diverse geographic origin of measured saffron samples, their harvest time and types (both powders and stigmas were contained in the experimental set), the clustering was relatively tight. Regarding matrices of other botanical origin, samples of turmeric were well separated from all other plant adulterants which overlapped to some extent, no improvement was obtained by supervised PLS-DA analysis. The tight cluster of QC samples documents a good reproducibility of the measurements.
Chemometric analysis of saffron vs. single plant species (binary models)
Although PCA would allow identification of ‘pure’ authentic saffron, under real life conditions total replacement by other plant species is rather improbable, dilution is more common. On these terms, the knowledge of the adulterant’s specific ‘markers’ is important for fraud detection. To simplify their selection, binary PLS-DA classification models for saffron and potential adulterants were constructed (see Fig. 3) and then, internal fourfold cross-validation was performed.
Table 1. shows excellent (100%) prediction and recognition abilities of the models, as well as high values of Q2 and R2 parameters (the lowest but still very high was in case of the ‘Saffron and Safflower’ binary model).
Selection and identification of characteristic ‘markers’
In the next phase of experiments, deconvoluted peaks characterized by particular m/z value and retention time with VIP-scores higher than 1.0 were considered as candidate significant ‘markers’. The other (complementary) tool which was used for the potential ‘markers’ selection was Pattern search function of MetaboAnalyst software, in that case the criterion was the value of the Pearson correlation coefficient (≥ 0.8 required). The identification of compounds that met at least one of these selection criteria was based on the match of (i) measured spectra with the records on the NIST library, (ii) accurate mass of detected m/z (mass error < 5 ppm) corresponding to the expected elemental composition, and (iii) the compliance with Kovats indexes on the respective capillary column. It should be noted, that we identified number of unique volatile components [19,20,21] characteristic for saffron aroma (see Table 2), nevertheless, as the aim of this study was to find out ‘markers’ of adulterants, more emphasis is placed on the identification of volatiles occurring in potential plant adulterants. The results of their tentative identifications are summarized in Table 3. Some of them such as α-phellandrene, β-myrcene, α-terpinene, p-cymene, D-limonene and eucalyptol were reported to be contained also in saffron[19, 20], nevertheless, their concentration in samples available in this study did not exceed detection threshold.
Detection of plant adulterants in saffron samples
As mentioned in the Introduction, saffron is a high value commodity susceptible to fraud, typically by dilution by a cheaper plant material. To verify the function of ‘markers’ selected in the first phase of experiments for the detection of this type of fraud, several admixtures were prepared; the authentic saffron samples were artificially adulterated by turmeric, calendula, capsicum and safflower. The sample preparation procedure is described in Sect. "Artificially adulterated saffron samples". Using markers summarized in Tables 2 and 3 for PCA and PLS-DA models construction, additions of potential plant adulterants as low as 2% w/w could be reliably identified. The paragraphs below provide more detailed information.
The PLS-DA score plot (see Fig. 4A) obtained for saffron-turmeric admixtures shows a good separation of the sample groups. The classification model was built based on 27 volatile compounds (16 ‘markers’ of turmeric and 11 ‘markers’ of saffron). The specific ‘markers’ of turmeric were: α-tumerone, ar-turmerone, cis-γ-atlantone, β-turmerone, 6S,7R-bisabolone and trans-α-atlantone, an example of specificity of β-turmerone characteristic ‘marker’ ion m/z = 105.0695 ± 5 ppm is illustrated in Fig. 4B.
Another plant material used for artificial adulteration was calendula. The statistical evaluation was performed based on 19 volatiles (8 ‘markers’ of calendula and 11 ‘markers’ of saffron). In the case of this classification model the statistical difference was observed at the all % w/w levels of the addition of calendula, which could be observed from PLS-DA score plot (see Fig. 5A). The specific ‘markers’ of calendula were α-calacorene and thymoquinone. Interestingly, the presence of the latter compound has not been reported in Calendula officinalis yet. Figure 5B showing the extracted ion chromatograph (EIC) of thymoquinone characteristic ‘marker’ ion m/z = 164.0827 ± 5 ppm documents its specificity.
In Fig. 6A, PLS-DA score plot demonstrates the separation of the artificially adulterated saffron sample by capsicum. For the construction of classification model in this case were used 21 compounds (10 ‘markers’ of capsicum and 11 ‘markers’ of saffron). Among the capsicum specific ‘markers’ were 3-carene, D-carvone and 2,2,6-trimethylcyclohexanone. The EIC of specific ‘marker’ ion m/z = 108.0929 ± 5 ppm for D-carvone is shown in Fig. 6B.
The last plant material which were tested for artificial adulteration of saffron was safflower. From the PLS-DA score plot (see Fig. 7A) could be seen that all % w/w levels were separated. For the construction of this classification model, 20 volatiles (9 ‘safflower markers’ and 11 ‘markers’ of saffron) volatiles were used. Specific and characteristic ‘markers’ of safflower were as follows: 3-methyl butanal, 3-methyl-1-butanol and γ-terpinene. Figure 7B shows EIC of specific terpinenen ‘marker’ ion m/z = 136.1070 ± 5 ppm.
The critical comparison of the results with the studies previously performed
In the final phase of this study, we compared obtained results with similar studies concerned with the use of GC for saffron authentication. For instance, Morozzi et. al. [38] used flash HS-GC-FID for analysis of 28 saffron samples and admixtures with turmeric and calendula (20, 15, 10 and 5% w/w). Using the created chemometric models, it was possible to separate the mixed samples from the authentic ones in case of both adulterants even at the lowest level of contamination. However, with respect to the use of FID for volatile fingerprinting, no concrete ‘markers’ were identified. Di Donato et. al [6]. also studied the adulteration of authentic saffron by dilution with three plant materials (calendula, safflower, and turmeric). They used SPME–GC–MS (ion trap mass analyzer) for the screening of volatile substances. Alike in our study, the presence of adulterants at low levels, 2–3% w/w, could be detected, nevertheless, contrary to our results, only one (trans-anethole for safflower and ar-turmerone for turmeric), or in maximum two (γ-cadinene and δ-cadinene for calendula) ‘markers’ were identified for respective plants. These substances were also detected in our samples of adulterants, nevertheless, their VIP scores, with exception of ar-turmerone, were relatively low compared to those found in our study. The discrepancy in identified ‘markers’ might be due to fairly different chromatographic conditions, while starting temperature of GC separation in Di Donato et. al [6]. study was 120 °C, we started at 40 °C what enabled us detection and identification of more volatile ‘marker’ compounds with high VIP scores. It is worth mentioning that a complementary authentication study employing ultra-high performance reverse-phase liquid chromatography coupled to tandem high-resolution mass spectrometry (UHPLC-HRMS/MS) was performed in our lab on the same unique samples set by Ryparova Kvirencova et al.[39]. Comparable results in terms of Q2 and R2 parameters and the ability to detect the addition of adulterants at the level of 2% w/w were achieved.
In conclusion, implementation of SPME-GC-HRMS technique for detection of saffron dilution by the most common adulterants (turmeric, calendula, safflower and capsicum) enabled the distinguishing authentic saffron samples from all other potential plant adulterants, all constructed classification models showed 100% prediction and recognition abilities and excellent Q2 and R2 parameters (> 0.9). Another outcome of the study was identification of eleven unique ‘markers’ for saffron and twenty-seven volatile ‘markers’ for four potential plant adulterants This enables not only fraud detection but also recognition of the type of plant adulterant. When targeting identified ‘markers’ of plant adulterant their presence in admixtures with saffron even at the level of 2% w/w could be recognized. The knowledge of respective ‘markers’ allows the simplification of authentication procedures. For target analysis, common triple quadrupole mass analyzers can be used.
Data availability
Data available on request from the authors.
References
Moratalla-López N, Zalacain A, Bagur MJ, Salinas MR, Alonso GL (2018) Saffron. In: FoodIntegrity Handbook on Food Authenticity Issues and Related Analytical Techniques. [Internet]. Eurofins Analytics France; [193–204]
Dowlatabadi R, Farshidfar F, Zare Z, Pirali M, Rabiei M, Khoshayand M, Vogel H (2017) Detection of adulteration in Iranian saffron samples by 1H NMR spectroscopy and multivariate data analysis techniques. Metabolomics 13(2):19
Guijarro-Díez M, Nozal L, Marina ML, Crego AL (2015) Metabolomic fingerprinting of saffron by LC/MS: novel authenticity markers. Anal Bioanal Chem 407(23):7197–7213
Khilare V, Tiknaik A, Prakash B, Ughade B, Korhale G, Nalage D, Ahmed N, Khedkar C, Khedkar G (2019) Multiple tests on saffron find new adulterant materials and reveal that Ist grade saffron is rare in the market. Food Chem 272:635–642
Farag MA, Hegazi N, Dokhalahy E, Khattab AR (2020) Chemometrics based GC-MS aroma profiling for revealing freshness, origin and roasting indices in saffron spice and its adulteration. Food Chem 331:127358
Di Donato F, D’Archivio AA, Maggi MA, Rossi L (2021) Detection of plant-derived adulterants in saffron (Crocus sativus L.) by HS-SPME/GC-MS profiling of volatiles and chemometrics. Food Anal Methods 14(4):784–796
Jalali-Heravi M, Parastar H, Ebrahimi H (2009) Characterization of volatile components of Iranian saffron using factorial-based response surface modeling of ultrasonic extraction combined with gas chromatography-mass spectrometry analysis. J Chromatogr A 1216:6088–6097
Anastasaki E, Kanakis C, Pappas C, Maggi L, Campo C, Delgado M, Alonso G, Polissiou M (2009) Geographical differentiation of saffron by GC–MS/FID and chemometrics. Eur Food Res Technol 229:899–905
Sabatino L, Scordino M, Gargano M, Belligno A, Traulo P, Gagliano G (2011) HPLC/PDA/ESI-MS Evaluation of Saffron (Crocus sativus L.) Adulter Nat Prod Commun 6 (12):1934578X1100601220
Villa C, Costa J, Oliveira MB, Mafra I (2017) Novel quantitative real-time PCR approach to determine safflower (Carthamus tinctorius) adulteration in saffron (Crocus sativus). Food Chem 229:680–687
Soffritti G, Busconi M, Sánchez R, Thiercelin J, Polissiou M, Roldan-Medina M, Fernández J (2016) Genetic and epigenetic approaches for the possible detection of adulteration and auto-adulteration in saffron (Crocus sativus L.) spice. Molecules 21:343
Gunning Y, Davies KS, Kemsley EK (2023) Authentication of saffron using 60 MHz 1H NMR spectroscopy. Food Chem 404:134649
Musio B, Todisco S, Antonicelli M, Garino C, Arlorio M, Mastrorilli P, Latronico M, Gallo V (2022) Non-targeted NMR method to assess the authenticity of saffron and trace the agronomic practices applied for its production. Appl Sci 12(5):2583
Hegazi NM, Khattab AR, Frolov A, Wessjohann LA, Farag MA (2022) Authentication of saffron spice accessions from its common substitutes via a multiplex approach of UV/VIS fingerprints and UPLC/MS using molecular networking and chemometrics. Food Chem 367:130739
Liu JD, Chen N, Yang J, Yang B, Ouyang Z, Wu CX, Yuan Y, Wang WH, Chen M (2018) An integrated approach combining HPLC, GC/MS, NIRS, and chemometrics for the geographical discrimination and commercial categorization of saffron. Food Chem 253:284–292
Naim N, Ennahli N, Hanine H, Lahlali R, Tahiri A, Fauconnier M-L, Madani I, Ennahli S (2022) ATR-FTIR spectroscopy combined with DNA barcoding and GC-MS to assess the quality and purity of saffron (Crocus sativus L.). Vibr. Spectrosc. 123:103446
Amirvaresi A, Nikounezhad N, Amirahmadi M, Daraei B, Parastar H (2021) Comparison of near-infrared (NIR) and mid-infrared (MIR) spectroscopy based on chemometrics for saffron authentication and adulteration detection. Food Chem 344:128647
Foschi M, Tozzi L, Di Donato F, Biancolillo A, D’Archivio AA (2022) A Novel FTIR-based chemometric solution for the assessment of saffron adulteration with non-fresh stigmas. Molecules 28(1):33
Cozzolino R, Stocchero M, Perestrelo R, Camara JS (2022) Comprehensive evaluation of the volatomic fingerprint of saffron from campania towards its authenticity and quality. Foods 11(3):366
Zwane BN, Kamatou GP, Viljoen AM, Betti G, Schmidt M (2020) Variation in headspace volatiles of saffron determined by GCxGC-ToF-MS. Nat Prod Commun 15(11):1–9
Cid-Perez TS, Nevarez-Moorillon GV, Ochoa-Velasco CE, Navarro-Cruz AR, Hernandez-Carranza P, Avila-Sosa R (2021) The relation between drying conditions and the development of volatile compounds in saffron (Crocus sativus). Molecules 26(22):6954
Maggi L, Carmona M, Zalacain A, Kanakis CD, Anastasaki E, Tarantilis PA, Polissiou MG, Alonso GL (2010) Changes in saffron volatile profile according to its storage time. Food Res Int 43(5):1329–1334
Kabiri G, Hssaini L, Naim N, Houmanat K, Ennahli S, Fauconnier M-L, Hanine H (2023) Aromatic potential, quality and antioxidant activity of saffron grown in Morocco. Flavour Frag J 38(1):13–26
Wang L, Chen Z, Han B, Wu W, Zhao Q, Wei C, Liu W (2020) Comprehensive analysis of volatile compounds in cold-pressed safflower seed oil from Xinjiang. China Food Sci Nutr 8(2):903–914
Poudel DK, Ojha PK, Rokaya A, Satyal R, Satyal P, Setzer WN (2022) Analysis of volatile constituents in curcuma species, viz. C. aeruginosa, C. zedoaria, and C. longa, from Nepal. Plants (Basel, Switzerland) 11(15):1932
Chaaban A, Gomes EN, Richardi VS, Martins CEN, Brum JS, Navarro-Silva MA, Deschamps C, Molento MB (2019) Essential oil from Curcuma longa leaves: Can an overlooked by-product from turmeric industry be effective for myiasis control? Ind Crops Prod 132:352–364
Oyemitan I, Elusiyan C, Onifade A, Akanmu M, Oyedeji A, McDonald A (2017) Neuropharmacological profile and chemical analysis of fresh rhizome essential oil of Curcuma longa (turmeric) cultivated in Southwest Nigeria. Toxicol Rep 4:391–398
Sahingil D (2019) GC/MS-Olfactometric characterization of the volatile compounds, determination antimicrobial and antioxidant activity of essential oil from flowers of calendula (Calendula officinalis L.). J Essent Oil-Bear Plants 22(6):1571–1580
John R, Jan D (2017) Calendula officinalis-an important medicinal plant with potential biological properties. Proc Indian Natn Sci Acad 83(4):769–787
Olennikov DN, Kashchenko NI (2022) Marigold metabolites: diversity and separation methods of calendula genus phytochemicals from 1891 to 2022. Molecules 27(23):8626
Reale S, Biancolillo A, Gasparrini C, Di Martino L, Di Cecco V, Manzi A, Di Santo M, D’Archivio AA (2021) Geographical discrimination of bell pepper (Capsicum annuum) spices by (HS)-SPME/GC-MS aroma profiling and chemometrics. Molecules 26(20):6177
Kevrešan Ž, Mastilović J, Sinadinovic-Fiser S, Hrabovski N, Radusin T (2013) Spice paprika volatiles as affected by different postharvest ripening treatments of red pepper (Capsicum annuum L.) variety aleva NK. Acta Period Technol 44:75–86
Izcara S, Perestrelo R, Morante-Zarcero S, Sierra I, Câmara JS (2022) Spices volatilomic fingerprinting–a comprehensive approach to explore its authentication and bioactive properties. Molecules 27(19):6403
Attia H, Harrathi J, Alamer K, Alsalmi F, Magné C, Khalil M (2021) Effects of NaCl on antioxidant, antifungal, and antibacterial activities in safflower essential oils. Plants 10:2809
Laokuldilok N, Utama-ang N, Kopermsub P, Thakeow P (2017) Characterization of odor active compounds of fresh and dried turmeric by gas chromatography-mass spectrometry, gas chromatography olfactometry and sensory evaluation. Food Appl Biosci J 3(3):216–230
Ziarati P (2012) The essential oil composition of Carthamus tinctorius L. flowers growing in Iran. Afr J Biotechnol 11(65):12921–12924
Ak G, Zengin G, Ceylan R, Mahomoodally F, Jugreet S, Mollica A, Stefanucci A (2021) Chemical composition and biological activities of essential oils from Calendula officinalis L. flowers and leaves. Flavour Frag. J 36(3):554–563
Morozzi P, Zappi A, Gottardi F, Locatelli M, Melucci D (2019) A quick and efficient non-targeted screening test for saffron authentication: application of chemometrics to gas-chromatographic data. Molecules 24:2602
Ryparova Kvirencova J, Navratilova K, Hrbek V, Hajslova J (2023) Detection of botanical adulterantsin saffron powder. Anal Bioanal Chem. https://doi.org/10.1007/s00216-023-04853-x)
Acknowledgements
The authors thank the Department of Food Analysis and Nutrition of UCT Prague for the contribution for the research.
Funding
Open access publishing supported by the National Technical Library in Prague. This work was supported by METROFOOD-CZ research infrastructure project (MEYS Grant No: LM2023064) including access to its facilities. This work was financially supported from a specific university research A1_FPBT_2023_002.
Author information
Authors and Affiliations
Contributions
MF: Conceptualization, Methodology, Writing- Original draft preparation. MS: Conceptualization, Methodology, Writing—Review & Editing. JH: Supervision, Writing—Review & Editing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest, either real or potential, associated with this work.
Compliance with ethics requirements
This article does not contain any studies with human or animal subject.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Filatova, M., Hajslová, J. & Stupak, M. Detection of saffron adulteration by other plant species using SPME-GC-HRMS. Eur Food Res Technol 250, 911–922 (2024). https://doi.org/10.1007/s00217-023-04443-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-023-04443-2