Abstract
Lathyrus sativus is considered as an important economical source of high-quality proteins and adaptability to global climate change. To detect accessions useful for future breeding strategies, the genetic variability among and within seven Lathyrus sativus accessions was assessed by morphological characters and the in vitro digestibility study. An important genetic diversity was found using various morphological traits. The significant heterogeneity revealed by variance analysis (ANOVA) and correlation test could be elucidated by the different geographical origins of the studied accessions and the natural selection of grass pea. The Principal Component Analysis (PCA) and Hierarchical Cluster Analysis (HCA) exhibited the accumulation of several genotypes in the same cluster which confirm that these genotypes are not dispersed according to their origin. Whereas, the individuals of grass pea from Ethiopia were collected in distinct group indicating that these samples are close each other’s and are different from the other accessions. In the same aim, the in vitro digestion model showed that most of the proteins were digested after 60-min incubation except the β-lathyrin protein. The different digestion kinetics found could be used as a method to distinguish different crop accessions. In fact, the densitometry analysis of β-lathyrin protein band showed that grass pea from Ethiopia, Germany, Bangladesh, and Morocco were more digested than grass pea from Canada, Afghanistan, and Greece in gastric phase, and Grass pea from Afghanistan and Germany were more digested than the rest of the samples in duodenal phase.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The cultivated Lathyrus species are considered as orphan crops. Widely cultivated in poor countries, they constitute a reliable resource in marginal low-input agricultural ecosystems [1]. Grass pea (Lathyrus sativus L.) is one of the most cultivated species in the genus Lathyrus because of their economical source of high-quality proteins and adaptability to global climate change [2]. It is considered as pasture crop, green fodder, animal feed, and human food [3]. Grass pea has a special advantage over other species: its hardy and penetrating root system allows cultivation in a wide spectrum of soil types, including very poor soils. Besides, grass pea roots fix their nitrogen in the soil to provide their own nitrogen needs which affects positively the soil nitrogen balance [4, 5]. These characteristics defined this crop as the most potential genetic resources for feeding future breeding programs to face the ongoing major world challenges and climate changes [6].
The origin of L. sativus is unknown; however, their supposed origin center is Southwest and Central Asia [7].
Different ways have been used to assess genetic diversity like pedigree analysis, morphological traits, or using molecular markers [8]. Many studies based on morphological characters, cytology, and enzyme electrophoresis have been used to study the diversity of grass pea [9, 10]. In this scenario, the assessment of digestibility and physiological effects of proteins is becoming progressively pertinent as novel dietary protein sources to being explored, driven by food technological advances, changing consumer awareness, and factors related to cost and sustainability [11]. In vitro digestibility is a rapid method, less labor intensive, and having significantly less bioethical restrictions.
This study aimed to assess the genetic diversity among and within seven populations of Lathyrus sativus using morphological traits and to study the in vitro digestibility of the studied grass pea flour for planning future breeding programs.
Materials and methods
Morphological study
The experimental material comprised seven accessions of Lathyrus sativus obtained from the International Center for Agricultural Research in the Dry Areas (ICARDA) Syria and Lebanon, and originating from seven countries: Canada, Greece, Germany, Morocco, Ethiopia, Afghanistan, and Bangladesh (Fig. 1); the seeds were kept in carton bags with 100 seeds/bag. In this study, the accessions were chosen according to the variability of the origins of each accession itself. From a database of seed accessions of Lathyrus sativus present in our laboratory, we selected all the accessions having a different geographical origin. The seeds were sown in Tunisia under a greenhouse in 25 cm diameter pots by randomized block designed by two replicates of 10 plants each, utilizing as substrate peat (Pindstrup plus LV10336, GHS, Tunisia) and sand (1:1 in volume). The germination period was from 16 November 2017 to 30 May 2018. No mineral fertilization was applied during the growth and development of plants. The plants were watered regularly with 200 mL of tap water every two days. Ten plants from each population were tagged randomly and evaluated for days to first flowering (d), days to 50% flowering (d), maturation period (d), days to first mature pod (d), number of pods/plant, plant height (cm), leaflet length, length and width pod, length of primary branch (cm), nodes number to the first bearing pod, pod curvature, pod shape, flower color, pod curvature, seed shape, seed coat color, and plant growth habit. All the morphological traits mentioned above were scored according to the suggestions given in the ‘‘Descriptors for Lathyrus ssp.’’ by IPGRI [12]. Additionally, the leaf area (mm2) trait was calculated using the Mesurim software.
Biochemical study
Flour preparation
To prepare the flour from the seven Grass pea populations, a step of grounding seeds for 5 min was performed using a variable speed laboratory blender (LB20ES, Waring Commercial, Torrington, Connecticut, USA) to let the flour pass through a 425-μm stainless steel sieve (Octagon Digital Endecotts Limited, Lombard Road, London, UK). Then, the flour samples were collected and stored in polyethylene bags at 4 °C until used for analysis.
Protein content
The Kjeldahl method was used to extract and determine the protein content in Grass pea flour. The amount of proteins was determined by measuring the nitrogen content of the material and multiplying that value by the factor of 6.25 [13]. To this aim, the samples were digested in concentrated sulfuric acid, and then the acid was neutralized with concentrated sodium hydroxide, followed by distillation of the ammonia (derived from the nitrogen of proteins) with HCl 1 N. Briefly, 0.25 g of grass pea flour were let to pass through a sieve (cut-off 2 mm) and dried at 105 °C until the constant weight was reached. When dried, the flour was poured into the tubes with the following digestive reagent: 7 g of potassium sulfate, 5 mg of selenium powder, 7 mL of concentrated sulfuric acid, 5 mL of hydrogen peroxide (35%), and 2–3 glass beads. The solution was heated at 420 °C for 20 min and then was cooled to 50 °C, adding at the end 50 mL of distilled water. A mattress with 25 mL of boric acid (5%) was assembled on the distillation unit and then 50 mL of sodium hydroxide (35%) were poured into the tubes. The distillation continued until 100 mL of solutions were collected.
The concentration of organic nitrogen was obtained by pH titration of solution with HCl 0.1 N until the pH reached the value of 4.7. The amount of HCl (1 mL of HCl 1N is equal to 1.4 mg of N-NH3) used was directly proportional to nitrogen content by multiplying with the factor value of 6.25 [13].
In vitro protein digestion
Grass pea flour was subjected to a three-stage in vitro digestion by using an adult model [14,15,16], under simulated oral, gastric and duodenal physiological conditions. To mimic the oral phase digestion, 4 mL of grass pea flour (25 mg mL−1) boiled at 100 °C for 15 min to simulate the cooking process. Then, it was incubated for 2 min at 170 rpm and 37 °C with amylase (150 U/mL) dissolved in 4 mL of simulated salivary fluid (SSF, 0.15 M of NaCl, 3 mM of urea, pH 6.9). Afterward, the samples were subjected to gastric and duodenal digestion as described by Giosafatto et al. [14]. Briefly, aliquots (100 µL) of simulated gastric fluid (SGF, 0.15 M of NaCl, pH 2.5) were placed in 1.5-mL microcentrifuge tubes and incubated at 37 °C. 100 µL of oral samples, the pH of which was adjusted to 2.5 with HCl 6 M, was added together with pepsin (1:20 w/w respect to grass pea protein content) to each of the SGF vials to start the digestion reaction. The ratio of pepsin to test proteins was 20:1 (w/w). At intervals of 1, 2, 5, 10, 20, 40, 60 min, 40 µL of 0.5 M of ammonium bicarbonate (NH4HCO3) were added to each vial to stop the pepsin reaction. The control was set up by incubating the sample for 60 min without the protease. Duodenal digestions were performed using, as the starting material, the gastric digests after the 60 min, adjusted to pH 6.5 with 0.5 M Bis–Tris HCl pH 6.5. Bile salts (sodium taurocholate and sodium glycodeoxycholate) dissolved in simulated duodenal fluid (SDF, 0.15 M of NaCl at pH 6.5) were added to a final concentration of 4 mM. After equilibrating at 37 °C for 10 min, trypsin and chymotrypsin (the ratio of trypsin and chymotrypsin with test proteins was 1:400 (w/w) and 1:100 (w/w), respectively) were added to the duodenal mix. Aliquots were removed over the 60-min digestion time course and proteolysis was stopped by the addition of a twofold excess of soybean Bowman–Birk trypsin–chymotrypsin inhibitor above that calculated to inhibit trypsin and chymotrypsin of the digestion mix. The control was carried out by incubating the sample without the proteases for 60 min. The samples were then analyzed using the SDS-PAGE (12%) procedure described below.
Sodium dodecylsulfate polyacrylamide gel electrophoresis (SDS-PAGE)
2.5 µL of sample buffer (15 mM of Tris–HCl, pH6.8, containing 0.5% (w/v) of SDS, 2.5% (v/v) of glycerol, 200 mM of β-mercaptoethanol, and 0.003% (w/v) of bromophenol blue) was added to aliquots of 10 µL of each protolyzed sample and analyzed by 12% SDS-PAGE, as described by Laemmli [17]. Electrophoresis was performed at constant voltage (80 V for 2–3 h), and the proteins were stained with Coomassie Brilliant Blue R250. Bio-Rad Precision Protein Standards were used as molecular weight markers.
Statistical analyses
Morphological analysis
Several statistical procedures were estimated to study the morphological diversity in seven Lathyrus sativus accessions and to elucidate relationships between them. For the morphological characterization by regions, the mean, maximum and minimum values of different quantitative traits were calculated. The qualitative morphological traits of grass pea accessions were estimated. Analysis of variance (ANOVA) was treated by XLSTAT software version 2021.01.24 (Addinsoft, Paris, France), at a significant level of p < 0.01 and 95% of confidence limits. The differences between the means values were compared using the Tukey test (Table 3 in letters). The descriptive traits and the Pearson correlation analysis between all quantitative traits were performed using the IBM SPSS Statistics software version 20 [18]. A Hierarchical Cluster Analysis (HCA) using Euclidean distance of Bray and Curtis following Ward’s method and Principal Component Analysis (PCA) were performed using the XLSTAT Software version 2021.01.24.
Densitometry analysis of protein digestion
Densitometry analysis was carried out using Image Lab software (version 5.2.1) from Bio-Rad Laboratories. Each SDS-PAGE image was analyzed detecting all the lanes and protein bands. The protein band having a Mr equal to ~ 45 kDa was used to determine band intensity of gastric and duodenal protein digestion of Grass pea flour with respect to the control that was not subjected to any treatment with both gastric and duodenal proteases.
Results
Variation of morphological traits
The research results obtained from analyses and observation of seventy grass pea samples indicated wide and significant variations among and within themselves. The range, mean, and coefficient variation percent (CV %) showed the existence of considerable variability among genotypes (Table 1). The coefficient of variation ranged from 6.17% to 82.53% which indicated a large variation in the studied characters. Among the 12 traits, plant height, maturation period, leaf area, and length of primary branch were found to have a relatively higher coefficient of variation (37.44%, 61.38%, 64.45%, and 82.53%, respectively) than the other traits.
The differences between maximum and minimum values of the studied characters by region also exposed the existence of diversity among samples (Table 2). Genotype originating from Canada showed the tallest plant (183 cm), height primary branch (109), maximum nodes per plant (33), and maximum leaflet length and leaf area (3.8 cm, 80 mm2, respectively), while genotype from Afghanistan showed the least plant height and leaflet length (31 and 1.3 cm, respectively). The least primary branch is obtained from Canada accession (3 cm). Days to first flowering ranged from 99 to 153 days. Maximum days was observed in Canada accession and minimum in Morocco accession. Similarly, days to first fruiting varied from 118 to 158 days with maximum days obtained from the Afghanistan population and minimum from Ethiopia accession. Minimum and maximum days for 50% flowering were recorded at 125 and 152 days, respectively. Maturation period ranged from 4 to 46 days, maximum days detected in Morocco accession and minimum in Afghanistan accession. The highest pod length and broadest pod (3 and 2 cm, respectively) were found in Morocco accession (Table 2). Comparing the mean values of the studied traits, IFLA1720 and IFLA 274 accessions are presented by the highest adult plant with higher nodes number and leaf size, and later maturation pods. IFLA 242 is characterized by the least high plant but highest branch length, few nods, small leaves, and later flowering. IFLA 2636 and IFLA 134 are categorized by medium height plant and branch, some nodes, small leaves and pods, and few days to mature pods. IFLA 160 is presented by least height plant and branch, few nodes, small leaves and pods and few days to first flowering. IFLA 479 is described by highest plant and medium branch, more nodes, big leaves, and early flowering (Table 3). Those characteristics showed the high diversity among and within accessions.
Analysis of variance (ANOVA) test was used to determine the effect of geographic origin, pod shape, flower color, and pod curvature on all the quantitative morphological characters. This ANOVA test (Tukey Contrasts) confirmed the obtained results and revealed statistically significant differences between genotypes for all morphological traits based on the effect of geographic origin at p < 0.01 (Table 3). The effect of pod curvature and flower color were significantly different for PH, LPB, NFP, LL, PN, and PW. Significant effect of pod shape was observed on all quantitative traits except on LPB and PN (Table 3). These results revealed an important inter-population variability for the different studied quantitative traits.
Based on geographic origin, the Tukey test subdivided our populations into two groups for the pods number, the length of primary branch, the leaf length, and pod width. It subdivided them into three groups for the leaf area, the plant height, pod length, days to first mature pod, days to first flowering and maturation period, and into four groups for the nodes number (Table 3).
Regarding qualitative traits, Grass pea Morocco, Canada, and Ethiopia are characterized by curved pod, broad elliptical pod shape, and violet-blue flowers. Grass pea Afghanistan is distinguished by pink flowers, broad linear pod shape, and slightly curved pod. Grass pea Bangladesh and Germany have broad linear pod shape, curved pod, and violet-blue flowers. Grass pea Greece is presented by beaded pod shape, curved shape, and violet–blue flowers. All accessions are characterized by rhomboid seeds, red–purple seed coat, and semi-erect growth habit (Table 4).
Additionally, the Pearson correlation results had confirmed this diversity. The plant height was highly correlated with primary branch length (−0.610), nodes number (0.706), leaflet length (0.299), leaf area (0.246), pod width (0.262), days to first mature pod (−0.176), and days to 50% flowering (−0.316). Length of primary branch was positively associated with maturation period and negatively associated with nods number and pod width. A significant correlation of nods number with leaflet length, leaf area, pod length and width, days to first mature pod, and days to 50% flowering was also recorded. Leaf length was negatively correlated with days to first flowering, maturation period and days to first and 50% flowering, and positively correlated with leaf area and pod width. On the other hand, a significant correlation was observed between leaf area, pod length, and maturation period, between pods number, pod length, and days to first flowering. A negative correlation was also found among pod length, days to first flowering, and days to the first mature pod (Table 5). Also high correlation was observed among days to first flowering, first mature pod, 50% flowering and maturation period.
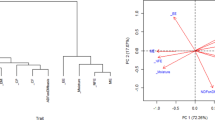
To study the genetic relationships between Lathyrus sativus accessions based on 12 quantitative morphological parameters, PCA and HCA were performed. The Principal Component Analysis (PCA) (Fig. 2) allowed the original variables to be condensed into a lower number of independent variables. All quantitative characteristics were selected to display the discriminative analysis of the PCA as presented in the factors map (Fig. 2b). The percentage of variability revealed that the first two PCs accounted for 48.21% of the whole variation observed in the dataset. The first axis explained 29.68% of the variability. The second axis explained 18.53% of the variability (Fig. 2a). The projection of genotypes in the PCA map revealed the presence of three major groups. The group I gathers all Ethiopian samples. Group II consists of the samples of grass pea originating from Canada and Morocco. The third group regrouped the rest of the genotypes originating from Afghanistan, Germany, Greece, and Bangladesh with one samples from Canada. In this study, all samples appeared distributed in the PCA plot without clear segregations.
However, the results obtained from the constructed dendrogram via the Hierarchical Ascending Classification identified three major groups (Fig. 3). The first group englobes the majority of grass pea Afghanistan with two samples from each Canada, Bangladesh, and Greece. The second group includes the majority of grass pea Ethiopia regrouped in one sub-clade, five samples of grass pea Morocco, five samples from Canada, two from Bangladesh, one from Greece, and one from Germany regrouped in another sub-clade. The third group contains a large number of samples belonging to six countries. It was subdivided into two clads, one clade regrouped the majority of grass pea Bangladesh, and Greece with some individuals from Morocco, Afghanistan and Canada. The second clade was formed by all samples of grass pea Germany with two samples from Ethiopia and one from Afghanistan.
Protein analysis: content and in vitro digestion
Using the Kjeldahl method, the protein content of the seven grass pea accessions ranged between 23% and 25% with standard deviation equal to1.050 (Table 6). Grass pea Germany was 23.04% and grass pea Morocco was 25.5%.
After checking that the flour has a protein tenor, the concentration of seven grass pea proteins was visualized by SDS-PAGE. Grass pea Canada and Afghanistan (25% protein) seem to be more condensed than the rest of accessions (Fig. 4).
To study the digestibility by the human gut of different accessions of grass pea, oral, gastric, and duodenal digestion experiments were performed under physiological conditions [14, 19].
The SDS-PAGE profiles of digestibility of all grass pea accessions showed that the most of proteins are digested after 60-min incubation at the gastric phase, except the ~ 45-kDa protein which is even in the duodenal phase and was not completely digested. For example, the SDS-PAGE profile of digestibility of grass pea Afghanistan (Fig. 5) exhibited clear digestion of all proteins for both phases except the ~ 45-kDa protein. The SDS-PAGE profile shown in Fig. 5 evidences that the 45-kDa protein band (as indicated by the arrow) resulted in rapid breakdown and proteolytic product appeared within 1-min incubation with pepsin. However after 1-min digestion with pepsin, the protein seems stable to be hydrolyzed by both gastric and duodenal proteases. The density of digestibility of this protein was different from one variety to another. In fact, grass pea Ethiopia, Germany, Bangladesh, and Morocco were more digested than grass pea, Canada, Afghanistan, and Greece (Fig. 6) in the gastric phase. On the other hand, grass pea Afghanistan and Germany were more digested than the rest of the samples in the duodenal phase (Fig. 7). The ~ 45-kDa protein is probably the β-Lathyrin (7S globulin) made up of several polypeptides of different molecular weights (14–66 kDa). The double band observed is due to the fact that at least two of the subunits of β-lathyrin are glycosylated [20].
Discussion
In the context of selection and conservation of the most promising accessions, the estimation of morphological and biochemical variability of seven grass pea accessions was analyzed. An important intra- and inter-population diversities were found in the studied L. sativus accessions using various quantitative and qualitative morphological analyses. Mean, range, and coefficient of variation in quantitative traits are widely used to determine variations within and between populations [21, 22]. In our study, the observed variability was confirmed by the minimum and maximum values, mean, and especially by the coefficient of variation. The greatest variability as expressed by the coefficient of variation for all the studied populations was founded in plant height, maturation period, leaf area, length of primary branch. The least variability was that relating to the dates of blooming and maturing first pod (Table 1). Similar results were obtained by Rybiñski et al. [23] using different morphological traits of grass pea accessions originating from European countries. However, according to De la Rosa and Martín [24], the lesser variation founded on grass pea Spain was due to the restriction of the geographic origin of the used collection. In fact, the high variability in plant, pod, and seed traits found by Jackson and Yunus [25] supports our results and assumes that the great variation obtained by the studied L. sativus accessions is due to their various geographical repartitions as well as selection by man.
According to qualitative traits, the pod shape character subdivided our populations into three groups: grass pea Morocco, Canada, and Ethiopia which were regrouped into a group with a broad elliptical pod. Afghanistan, Bangladesh, and Germany populations were collected in a group that has broad linear pod, and grass pea Greece in a distinct group has beaded pod. While, flower color and pod curvature divided our accessions into two groups: one has a pink flower and slightly curved, and one has violet-blue flower and curved pod (Table 4). In fact, the array of variation in the studied L. sativus based on qualitative traits was pod shape, flower color, and pod curvature. However, the pattern variation described by Jackson and Yunus [25] and Vavilov [26] was flower color and seed coat color.
Moreover, the obtained results of analysis of variance (ANOVA) based on the effect of geographic origin, pod shape and curvature, and flower color showed significant variability among and within accessions. This heterogeneity could be explained by the large distribution area of our plants and their adaptability to abiotic and biotic stress [27]. Similarly, the presence of a strong significant correlation among the studied traits could be due to natural selection between samples within a population [21].
To study the phylogenetic relationships between the studied grass pea, the clustering pattern by Principal Component Analysis (PCA) (Fig. 2) was in accordance with the grouping pattern by Hierarchical Cluster Analysis (HCA) (Fig. 3). These two models showed the accumulation of a large number of genotypes from different geographic origins in the same cluster which indicate that studied genotypes are not distributed according to their origin as reported in a previous study where 107 grass pea genotypes from different geographic origin were analyzed and the founded results confirmed that the natural distribution was completely disconnected from their cultivation [28]. Whereas, in PCA the individuals of grass pea Ethiopia were collected in distinct group (group I), which indicate that these samples were very close between themselves and diversified regarding the other accessions. The same results were obtained by Tsegaye [29] in which the samples of grass pea Ethiopia were regrouped into an individual group.
With the aim of selection and conservation of the most promising accessions, the in vitro digestion models were established to serve as research tools to characterize and clarify the structural and biochemical changes of food components under physiological conditions, caused by alimentary enzymes [30].
As a first step for biochemical characterization of our plants, the determination and visualization of different protein content in the studied Lathyrus sativus accessions were made. All samples possess a similar protein content with no significant differences and the average of protein ranged between 23% and 25%. Then the oral, gastric, and duodenal in vitro digestion protein was studied for seven accessions of grass pea. Most of the proteins were digested in the gastric environment after 60 min incubation with pepsin, except of the protein band having a Mr ~ 45 kDa likely corresponding to lathyrin protein [31]. In fact, the densitometry analysis of the 45-kDa protein band showed that grass pea Ethiopia, Germany, Bangladesh, and Morocco were more digested than grass pea Canada, Afghanistan, and Greece (Fig. 6). Another study focused on grass pea Italy found that this variety was completely digested during the gastric phase for both adult and elderly models [32]. Hence, the proteins treated for 60 min with the pepsin was subjected to duodenal digestion. The densitometry analysis of the seven grass pea accessions showed that the 45-kDa protein, the β-lathyrin, was slowly digested and even after 60-min duodenal digestion is still present. However, this protein from grass pea Afghanistan and Germany accessions seems to be more digested than the rest of samples. It is worthy to mention that β-lathyrin contains IgE-binding epitopes and hence is considered as an allergenic protein, and probably this is in line with its resistances to protease hydrolysis. β-lathyrin was found to be in common with other legume allergens and conservation of IgE-binding epitopes among the different legume allergens may be responsible for the observed phenomenon of cross-reactivity [31].
Conclusion
Morphological traits are affected by environmental factors and the stage of development of the plant, and, as a consequence, they do not entirely explain the genetic diversity among samples or accessions. However, it is important to use morphological data to evaluate qualitative and quantitative characters in plant breeding [33]. Due to the various geographic distributions of the studied L. sativus accessions and their selection by man, an important intra- and inter-population diversities were found using quantitative and qualitative morphological features. The phylogenetic relationships obtained among the seven accessions of grass pea were completely disconnected from their cultivation origin. The in vitro digestibility was similar showed among all the investigated samples even though different digestion kinetics were found, indicating that this kind of investigation could be used as a method to distinguish different crop accessions.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article.
References
Vaz Patto MC, Fernandez-Aparicio M, Moral A, Rubiales D (2006) Characterization of resistance to powdery mildew (Erysiphe pisi) in a germplasm collection of Lathyrus sativus. Plant Breed 125:308–310
Campbell CG, Mehra RB, Agrawal SK, Chen YZ, Abd El-Moneim AM, Khawaja HIT, Yadov CR, Tay JU, Araya WA (1993) Current status and future strategy in breeding grass pea (Lathyrus sativus). Euphytica 73:167–175
Kumar S (1997) Utilization of Lathyrus. Lathyrus Genetic Resources Network, New Delhi, India, pp 57–59
Ahlawat IPS, Singh A, Saraf CS (1981) Effects of winter legumes on the nitrogen economy and productivity of succeeding cereals. Exp Agric 17:57–62
Campbell CG (1997) Grass pea (Lathyrus sativus L.) promoting the conservation and use of underutilized and neglected crops, vol 18. International Plant Genetic Resources Institute, Italy
Aci MM, Lupini A, Badagliacca G, Mauceri A, Presti EL, Preiti G (2020) Genetic diversity among Lathyrus ssp. based on agronomic traits and molecular markers. Agronomy 10:1182
Smartt J (1990) Grain legumes: Evolution and genetic resources. Cambridge University Press, Cambridge, UK, p 200
Pejic I, Ajmone-Marsan P, Morgante M, Kozumplick V, Castiglioni P, Taramino G, Motto M (1998) Comparative analysis of genetic similarity among maize inbred lines detected by RFLPs, RAPDs, SSRs, and AFLPs. Theor Appl Genet 97:1248–1255
Hossaert M, Valero M (1986) Vegetative propagation and sexual reproduction in two perennial Lathyrus Species. In: Kaul AK, Combes D (eds) Lathyrus and Lathyrism. Third World Medical Research Foundation, New York, pp 175–185
Yunus AG, Jackson MT (1991) The gene pools of the grass pea (Lathyrus sativus L.). Plant Breed 106:319–328
Egbert WR, Payne CT (2009) Plant proteins. In: Tarte R (ed) Ingredients in meat products: properties, functionality and applications. Springer Science, New York, p 11129
IPGRI (2000) Descriptors for Lathyrus spp. International Plant Genetic Resource Institute, Rome, Italy
Kjeldahl J (1883) Neue Methode zur Bestimmung des Stickstoffs in organischen Körpern. Fresenius’ J Anal Chem 22:66–382
Giosafatto CVL, Rigby NM, Wellner N, Ridout M, Husband F, Mackie AR (2012) Microbial transglutaminase-mediated modification of ovalbumin. Food Hydrocoll 26:261–267
Minekus M, Alminger M, Alvito P, Balance S, Bohn T, Bourlieu C, Carrière F, Boutrou R, Corredig M, Dupont D, Dufour C, Egger L, Golding M, Karakaya S, Kirkhus B, Le Feunteun S, Lesmes U, Macierzanka A, Mackie A, Marze S, McClements DJ, Ménard O, Recio I, Santos CN, Singh RP, Vegarud GE, Wickham MSJ, Weitschies W, Brodkorb AA (2014) Standardised static in vitro digestion method suitable for food—an international consensus. Food Funct 5:1113–1124
Bourlieu C, Ménard O, Bouzerzour K, Mandalari G, Macierzanka A, Mackie AR (2014) Dupont, D. Specificity of infant digestive conditions: some clues for developing relevant in vitro models. Crit Rev Food Sci Nutr 54:1427–1457
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Nie N, Hull C, Bent D (2011) IBM statistical package for the social sciences (SPSS Version 20). Computer Software. SPSS, Chicago, IL
Giosafatto CVL, Di Pierro P, Gunning P, Mackie A, Porta R, Mariniello L (2014) Characterization of Citrus pectin edible films containing transglutaminase-modified phaseolin. Carbohydr Polym 106:200–208
Rosa MJ, Ferreira RB, Teixeira AR (2000) Storage proteins from Lathyrus sativus seeds. Agric Food Chem 48:5432–5439
Endashaw B (1996) Morphological analysis of Eragrotis teff: detection for regional patterns of variation. SINET Ethiopian J Sci 19:117–140
Sharma S, Chahota K, Lal C (1995) Genetic diversity and agronomic evaluation of microsperma and macrosperma lentils. Genet Resour Crop Evol 42:217–222
Rybiñski W, Szot B, Rusinek R (2008) Estimation of morphological traits and mechanical properties of grass pea seeds (Lathyrus sativus L.) originating from EU countries. Int Agrophys 22:261–275
De la Rosa LMI (2001) Morphological characterization of Spanish genetic resources of Lathyrus sativus L. Lathyrus Lathyrism Newslett 2:31–34
Jackson MT, Yunus AG (1984) Variation in the grass pea (Lathyrus sativus L.) and wild species. Euphytica 33:549–559
Vavilov NI (1951) Phytogeographic basis of plant breeding. In: Verdoorn F (ed) The origin, variation, immunity, and breeding of cultivated plants. Chronica Bot, vol 13. The Ronald Press Company, New York, pp 13–54
Sdouga D, Branca F, Kabtni S, Di Bella MC, Trifi-Farah N, Marghali S (2020) Morphological traits and phenolic compounds in Tunisian wild populations and cultivated accessions of Portulaca oleracea L. Agronomy 10:948
Ahamed KU, Akhter B, Islam SMAS, Moniruzzaman M, Alam MA (2012) Genetic variability of some morphological traits in grass pea (Lathyrus sativus) germoplasm. Bull Inst Trop Agr Kyushu Univ 35:61–68
Tsegaye M (2007) Assessment of diversity, morphological variation and description of grasspea (Lathyrus sativus) and other related species. Addis Ababa Univ (Ethiopia). School of Graduate Studies [Corporate Author]
Romano A, Giosafatto CVL, Di Pierro P, Romano R, Masi P, Mariniello L (2016) Impact of transglutaminase on properties and in vitro digestibility of white bean (Phaseolus vulgaris L.) flour. Food Res Int 88:239–246
Xu Q, Song B, Liu F, Song Y, Chen P, Liu S, Krishnan HB (2018) Identification and characterization of β-lathyrin, an abundant glycoprotein of grass pea (Lathyrus sativus L.), as a potential allergen. J Agric Food Chem 66:8496–8503
Romano A, Giosafatto CVL, Al-Asmar A, Masi P, Aponte M, Mariniello L (2018) Grass pea (Lathyrus sativus) flour: microstructure, physico-chemical properties and in vitro digestion. Eur Food Res Technol 8:244
Dey SS, Singh AK, Chandel D, Behera TK (2006) Genetic diversity of bitter gourd (Momordica charantina L.) genotypes revealed by RAPD markers and agronomic traits. Sci Hortic 109:21–28
Acknowledgements
This research work was financially supported by the Tunisian Ministry of Higher Education and Scientific Research (LR99ES12) and was carried in the University of Naples Federico II, Naples, Italy.
Funding
Open access funding provided by Università degli Studi di Napoli Federico II within the CRUI-CARE Agreement. This work was supported by the Tunisian Ministry of Higher Education and Scientific Research under a grant (LR99ES12).
Author information
Authors and Affiliations
Contributions
SL designed the study, interpreted the data, and drafted the article. NTF, CVLG and LM revised the article.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that there are no conflicts of interest.
Compliance with ethics requirements
This study does not contain any Research involving Human Participants and/or Animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lassoued, S., Giosafatto, C.V.L., Mariniello, L. et al. Morphological characterization and in vitro digestibility of seven Lathyrus sativus (grass pea) accessions originating from Eurasia, Africa, and Canada. Eur Food Res Technol 249, 2419–2432 (2023). https://doi.org/10.1007/s00217-023-04266-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-023-04266-1