Abstract
Some of health-protecting phytochemicals can be found in the sprouts in a much higher concentration than in the developed plants. However, the polyphenol composition of Lepidium sativum L. sprouts is still unresolved, especially with regard to flavonoids and phenolic acids. To develop a rapid and reliable UPLC-PDA-Q/TOF-MS method for the identification and quantification of main flavonoids and phenolic acids from L. sativum sprouts, the samples of L. sativum sprouts were grown with and without the exposition to light. Ten phenolic compounds were identified on the basis of their mass spectra in MS mode, MS/MS spectra, PDA spectrum, and retention times as compared with available reference substances. The major compounds were found to be sinapic acid derivatives. Acylated flavonoids in L. sativum sprouts were identified as kaempferol and quercetin with the same glycosidic substitution pattern as the major compounds, and varying degrees of acylation with sinapic and ferulic acids. Total content of phenolics in both samples was similar, but the content of flavonoids was higher and sinapine was lower in the sample of sprouts germinated in light. The determination and the evaluation of L. sativum extracts is very effective while using the UPLC-PDA-Q/TOF-MS. Even though the substances only occur in low concentrations, the combination of the retention time and the accurate molecular mass allows for their fast qualification.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lepidium sativum L. (Fam. Brassicaceae) is a small herb with 30–50 cm of height which bears laciniate pinnate entire leaves. The aqueous extracts of the plant have previously been screened for various biological activities: cardiotonic hypotensive, antimicrobial, bronchodilator, hypoglycemic, and allelopathic activity [1, 2]. The whole seed which is edible is known to have health-promoting properties. Hence, it was assumed that these seeds can be a functional food [3–8]. Although L. sativum is widely used as food and medicinal plant, only the composition of glucosinolate is known well, and it has been identified as a principle bioactive constituent of this plant [9]. L. sativum contains also phenolic compounds. These compounds exist widely in plants. They are plant secondary metabolites, and they play an important role as defense compounds, not only for plants but also their antioxidant activity is important for general human health. Sinapic acid and its derivatives, most notably sinapine, were found in L. sativum seeds [10]. Sinapic acid is widely distributed not only in L. sativum but also in other plants of Brassicaceae, such as B. juncea. A number of phenolic choline esters, known as sinapine, have also been reported [11]. Sinapine, choline ester of sinapic acid, being the most abundant was followed by sinapoyl glucose in canola and rapeseeds [12, 13]. Presence of sinapine and related phenolic compounds such as sinapic acid has been of concern to the oilseed breeders and processors as they impart bitter taste, astringency, and dark color to repressed-mustard products [14].
HPLC and HPTLC methods have been used for identification and quantitative determination of sinapic acid in L. sativum [15, 16]. Derivatives of flavonols, kaempferol and quercetin, are the main flavonoids reported in Brassica vegetables [16–19]. It is not known how flavonol composition and conditions of seed sprouting influence phenolic compounds in L. sativum. It was reported that the phenolic components in the sprouts vary according to growth conditions, and it was also established that the light can stimulate the production of phytochemicals including the higher isoflavone content in the soya sprouts [20]. Similar abiotic stress can be triggered also by the UV light, yet the concentration of the flavonoids (morin, myricetin, quercetin, and kaempferol) in case of the radish and alfalfa sprouts was higher.
Different conditions of seed sprouting have effect on the flavonol content. The highest myricetin, merin, quercetin, and kaempferol contents in the radish and lucerne sprouts were measured when the sprouting was done in darkness at 20 °C. Neither an increase of the germination temperature up to 30 °C nor a decrease down to 10 °C affected the efficiency of the flavonol synthesis. Similarly, neither a UV nor an IR radiation for between 20 min and 24 h increased significantly the flavonol content of the sprouts as compared to the seeds [21].
During germination, from the seed at rest, a new plant is developing provided that the moisture content is favorable, the temperature is appropriate, and oxygen is available for the respiration. In Japan, the sprouts are classified into different categories depending on whether they were grown in artificial or natural light or in darkness, out of which the sprouts produced in light are consumed raw, whereas those produced in darkness are consumed heat-treated. We therefore set an aim to determine the phenolic compounds in L. sativum sprouts during their germination in darkness and in light.
The purpose of this research was to identify phenolic compounds and quantify main flavonoids and phenolic acids that were found to be characteristic for L. sativum sprouts. A sensitive, accurate, and specific method of ultra performance liquid chromatography (UPLC) was applied for short run times combined with a photodiode array detector (PDA) and a quadrupole/time of flight-mass spectrometer (Q/TOF-MS) which offers high mass accuracy.
The aim of the study was the separation and identification of phenolic acids and flavonoid glycosides in the methanolic extract of L. sativum sprouts. Fragmentation behavior of flavonoid glycosides and phenolic acids was investigated using mass spectrometry in negative and positive mode. The MS, MS/MS, and UV data together with UPLC retention time (Rt) of phenolic acids and flavonoids allowed for structural characterization of these compounds. Furthermore, accurate mass measurement technique was successfully applied for the first time in L. sativum sprouts to elucidate the elemental composition of the polyphenols studied.
Experimental
Chemicals
Acetonitrile, formic acid, and methanol were purchased from Sigma-Aldrich (Steinheim, Germany). Quercetin, kaempferol, and ferulic and sinapic acid were purchased from Extrasynthese (Lyon, France).
Plant material
Lepidium sativum L. seeds were purchased from local market of Wroclaw city (Poland). Seeds (L. sativum) were germinated in Petri (r = 50 mm) dishes on two layers of filter paper with 10 mL water at 20 °C in day light and dark by three indecently grown.
After 8 day, each germination sample of sprouts was collected and frozen in liquid nitrogen and freeze-dried (24 h; Alpha 1–4 LSC; Christ, Osterode am Harz, Germany). The homogeneous powders were obtained by crushing the dried tissues using a closed laboratory mill to avoid hydration. Powders were kept in refrigerator (−80 °C) until extract preparation.
Extraction procedure
The sprouts powder (1 g) was extracted using 25 ml of 30 % methanol acidified with 1 % acetic acid and containing 1 % ascorbic acid as antioxidant. The extraction was performed by incubation for 20 min under sonication (Sonic 6D, Polsonic, Warsaw, Poland) and with occasional shaking. This method has proved to be adequate for complete extraction. Next, the slurry was centrifuged at 19,000g for 10 min and the supernatant was filtered through a 0.2 μm PTFE filter (Millex Samplicity Filter, Merck) and used for analysis. All extractions were carried out in triplicates.
Identification and quantification of polyphenols by the ultra performance liquid chromatography-photodiode detector mass spectrometry (UPLC-PDA-MS) method. Identification and quantification of polyphenol of sprout extracts was carried out with the use of an ACQUITY Ultra Performance LC system equipped with a photodiode array detector with a binary solvent manager (Waters Corporation, Milford, MA) series with a mass detector G2 Q/TOF Micro mass spectrometer (Waters, Manchester, U.K.) equipped with an electrospray ionization (ESI) source operating in negative and positive modes. Separations of individual polyphenols were carried out using a UPLC BEH C18 column (1.7 μm, 2.1 × 50 mm, Waters Corporation, Milford, MA) at 30 °C. The samples (10 μL) were injected, and the elution was completed in 15 min with a sequence of linear gradients and isocratic flow rates of 0.45 mL/min. The mobile phase consisted of solvent A (4.5 % formic acid, v/v) and solvent B (100 % of acetonitrile). The program began with isocratic elution with 99 % solvent A (0–1 min), and then a linear gradient was used until 12 min, lowering solvent A to 0 %; from 12.5 to 13.5 min, the gradient returned to the initial composition (99 % A), and then it was held constant to re-equilibrate the column. The analysis was carried out using full-scan, data-dependent MS scanning from m/z 100 to 1,500. Leucine enkephalin was used as the reference compound at a concentration of 500 pg/μL, at a flow rate of 2 μL/min, and the [M − H]− ion at 554.2615 Da was detected. The [M − H]− ion was detected during 15-min analysis performed within ESI–MS accurate mass experiments, which were permanently introduced via the LockSpray channel using a Hamilton pump. The lock mass correction was ±1.000 for the mass window. The mass spectrometer was operated in negative ion mode, set to the base peak intensity (BPI) chromatograms, and scaled to 12,400 counts per second (cps) (100 %). The optimized MS conditions were as follows: capillary voltage of 2,500 V, cone voltage of 30 V, source temperature of 100 °C, desolvation temperature of 300 °C, and desolvation gas (nitrogen) flow rate of 300 L/h. Collision-induced fragmentation experiments were performed using argon as collision gas, with voltage ramping cycles from 0.3 to 2 V. The characterization of the single components was carried out via the retention time and the accurate molecular masses. Each compound was optimized to its estimated molecular mass [M − H]− in the negative mode before and after fragmentation. The data obtained from UPLC–MS were subsequently entered into the MassLynx 4.0 ChromaLynx Application Manager software. On the basis of these data, the software is able to scan different samples for the characterized substances.
The runs were monitored at the following wavelength: flavonol glycosides at 360 nm and phenolic acids at 320 nm. The PDA spectra were measured over the wavelength range of 200–800 nm in steps of 2 nm. The retention times and spectra were compared to those of the pure standards. The calibration curves were run at 360 and 320 nm for the standard at concentrations ranging from 0.05 to 5 mg/mL (r 2 = 0.9998).
Results and discussion
The data of retention times (Rt), wavelengths of maximum absorbance (λmax) and protonated/deprotonated molecules ([M + H]+/[M − H]−), diagnostic fragments, and identification for each peak of the phenolic compounds detected in the L. sativum sprout extracts are listed in Table 1. The compounds were identified according to their retention times at 320 nm, mass spectra of LC–MS analyses. LC–MS chromatograms for selected data of phenolic compounds are shown in Figs. 1, 2, and 3, respectively.
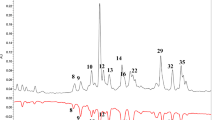
Chromatogram UPLC-PDA profile at 320 nm phenolic compounds of L. sativum L. sprouts. For peak labels, see Table 1
UPLC–MS chromatograms of phenolic compounds in L. sativum sprouts. For peak identification, see number in Table 1. The data presented are selected masses from the full-scan data of peak 1–10
Ten phenolic acid derivatives and flavonoids were detected in the L. sativum sprout extracts. Among them were six compounds of sinapic acid and their derivatives and four compounds of flavonol nonacylated and acylated glycosides. The sinapic acid was identified by UPLC-PDA-MS–MS method by comparison with standards. The molecular mass identified for sinapic acid by MS is the same as reported by Harbaum et al. [22]. This compound was previously reported in L. sativum by other researchers [16]. Sinapic acid is widely distributed in plants of Brassicaceae, such as B. juncea and L. sativum [11]. In selected mass (m/z = 223.0601), UPLC–MS chromatogram (Fig. 3) found two peaks corresponding to two isomers cis and trans. These compounds were identified only in L. sativum sprout germinated in the presence of light. Both cis and trans isomers of sinapic acid were found also by Thiyam et al. [23] in rapeseed isolate. 1-O-sinapoyl-β-glucose with a retention time of 5.0 min shows in the negative ion mode the signal of the deprotonated substance at m/z 385.1136 (Table 1). Fragments are at m/z 265.0716 (loss of a part of hexose ring) and at m/z 223.0598 (sinapic acid). The MS fragmentation observed for these compounds was similar to that described in a previous work by Ferreres et al. [24].
For the first time, two compounds, 9 and 10, with retention time of 10.88 and 12.60 min were identified as di-O-sinapoyl-hexose and its isomer with ion mass signal in negative ionization ([M − H]−) at m/z 591.1714. Fragments MS/MS were m/z 367.1029 (loss of a part of sinapic acid ring) and m/z 223.0603 (sinapic acid).
Sinapoyl malic acid isomers, probably cis and trans, that were found in L. sativum sprouts and their proposed structures (fragmentation pattern by MS/MS) and UV data are listed in Table 1. The mass [M − H]− at m/z 339.0719 and the resulting fragment mass MS/MS at m/z 223.0603 (sinapic acid) were detected and identified as an ester of malic acid with sinapic acid. Compounds 7 and 8 are coeluted with sinapic acid, but the use of UPLC–MS-Q/TOF allows for the analysis of these compounds according to m/z ions separately, as shown in Fig. 3.
The loss of 116 amu by MS/MS is characteristic for the malic acid derivatives, which is seen in Table 1. The fragmentation pattern corresponds with data in the literature [25].
To our knowledge, these sinapic acid derivatives of malic acid are described in L. sativum for the first time. Information about the presence of malic acid moieties of hydroxycinnamic acids is scarce in the literature. It has been reported that caffeoyl-malate occurs in lettuce as well as in herbs such as white horehound (Marrubium vulgare). In radishes, which also belong to the Brassicaceae family, different malic acid esters of caffeic acid, p-coumaric acid, sinapic acid, and ferulic acid have previously been found by Brandl et al. [26] and Nielsen et al. [27].
It is also known that sinapoyl malate occurs in Arabidopsis thaliana. Goujon et al. [28] detected this compound by MS spectroscopy and recorded its fragmentation pattern. Liang et al. [25] identified five malate derivatives in turnip leaves (Brassica rapa) by NMR spectroscopy with the assistance of HPLC–MS, including hydroxyferuloylmalate as a new compound. Rochfort et al. [19] also detected two major compounds in pak choi with the molecular masses of [M − H]− at m/z 309 and 339 but were unable to identify them.
Compound 4 was identified as sinapine with m/z ion at 310.1628 and the choline ester of sinapic acid at the retention time of 5.76 min (Table 1). Sinapine is more stable in negative ESI mode which is why an ESI–MS/MS analysis was performed in positive mode to verify ion breakdown. In positive mode of ions, the breakdown into fragments of ethyl ester of sinapic acid (loss of trimethyl amine) [29] and into fragments of sinapic acid (loss of H2O and CH3OH) was observed at m/z 207.0647 and 175.0392. The molecular mass for sinapine calculated by MS is the same as reported by Harbaum et al. [22].
The presence of sinapoylcholine (sinapine) has been reported in rapeseed, which imparts bitter taste, astringency, and dark color to repressed-mustard products [13, 14]. Sinapine, choline ester of sinapic acid followed by sinapoyl glucose, is the main compound making up to 1–2 % of the seed dry matter in rapeseed and canola [12, 30].
Three acylated flavonol compounds 3, 5, and 6 (Table 1; Figs. 1, 2, 3) were identified as quercetin-sinapoyl-dihexose-pentoside (m/z 977.2541 − [M − H]−), kaempferol-sinapoyl-dihexoside-pentoside (m/z 961.2581 − [M − H]−), and kaempferol-feruloyl-dihexoside-pentoside (m/z 931.2497 − [M − H]−), respectively. The ESI mass spectrum of acylated flavonols showed fragment ions at [M − 146 (pentose)]− corresponding to m/z 831, 815, and 785, respectively; next fragment ions at [M-206 (sinapic acid) and M − 176 (ferulic acid)] − m/z 609, 625, and 609, respectively (Fig. 2). The last fragment ions at [M − 324 (dihexose)] were aglycons, that is, m/z 285 (kaempferol), m/z 301 (quercetin), and m/z 285 (kaempferol), respectively, as listed in Table 1.
This fragmentation pattern corresponds to data in the literature [31], namely that in all acylated flavonol dihexosides, which were mainly quercetin and kaempferol glycosides, the MS fragmentation follows the following pattern: first, the sugar moiety at the -7-O position is split off, followed by the acyl- at the -3-O position and finally a dihexosides at the -3-O position. The preferential fragmentation of the glycosyl derivatives is at the -7 position compared to the glycosylation at -3 position. Acylations are present. The cinnamoyls derived from compounds (quercetin-3-O-di-Glc-7-O-Glc) and (kaempferol-3-O-diGlc-7-O-Glc) are widely distributed in Brassicaceae. These compounds were described in leaves of pack choi [B. rapa L. ssp. chinensis L. (Hanelt)] [19], B. rapa L. ssp. sylvestris L., flower of B. rapa [32], and leaves, stems, and flower buds of B. rapa var. rapa L. [33].
Thus, all of these plants containing acylated compounds studied in the negative mode MS/MS showed the presence of the base peak at (m/z 162 [M − H]−) corresponding to the -hexosyl derivatives and also at (m/z 324 [M − H]−) corresponding to the –sophorosyl derivatives. The fragmentation of these compounds was similar to that indicated in our samples. The only difference concerned the base peak which was produced during MS/MS experiment which was 146 amu (atomic mass unit) loss instead of 162 amu loss. This led to glycosylation with pentose instead of a hexose. The qualitative study of flavonoids in watercress (Nasturtium officinale) leaves also showed a characteristic glycosylation pattern with rhamnose at the -7 position [34]. Acylated flavonol glycosides were identified not only on the basis of the increase in mass of the parent ions but also on the basis of the wavelength maxima (330–336 nm) of their UV spectra (Table 1). They are characterized by a maximum with a high absorption at 333–334 nm and show a little maximum that coincide with the flavonoid band II at 266 nm.
Compound 2 in the L. sativum sprouts analyzed by UPLC-DAD-MS/MS method that had retention time at 5.28 min was identified as nonacyled kaempferol-dihexose-pentose (Table 1; Figs. 1, 2, 3). As a result of deprotonation of ion m/z 755.2021 in the first step, the loss of 146 amu takes place which indicates that one pentose moiety is lost. Next, dihexoside moiety (2 × 162 amu = 324 amu) is lost which gives rise to a deprotonated kaempferol ion (m/z 285.0403) as a base peak. This kind of fragmentation indicates the occurrence of glycosylation in accordance with literature data which show that in Brassica species, only sugar moieties such as glucose and rhamnose are predominantly bound in positions -7 [19, 32, 33, 35, 36]. In the disaccharide moieties present in analyzed compounds [2, 3, 5, and 6 (Table 1)] after fragmentation, the loss of 324 amu and low abundance of 162 amu selon occur, which, according to Ferreres et al. [31], corresponds to a diglucoside with a 1–6 linkage, that is, gentiobioside.
During the identification of flavonoids in L. sativum sprouts, this information has been used for allocating the position of the glucoside moieties and the interglucosidic linkage. We did not observe MS fragmentation that produces three characteristic losses of 180, 162, and 120 for sophorosides, that is, probably two glucose molecules with 1-2-interglucoside linkage.
Quantitative determination of the identified compounds was carried out in triplicate for two samples germinated in light and darkness. All flavonoid and hydroxycinnamic acid derivatives were quantified individually in the plant extracts and expressed in milligrams per gram of dry material (Table 1). The flavonoid contents were calculated on the basis of kaempferol and quercetin aglycone. The content of sinapic acid derivatives was calculated on the basis of sinapic acid standard. Previously, result described by Gil and MacLoed [37] and Glendening and Poulton [38] presented that L. sativum sprouts which grow in light during the first week of germination contain high concentration of benzylglucosinolate. Flavonoids were detected in sprouts that germinated in the presence of light in higher concentration (10.51 mg/g of dm) than in L. sativum sprouts that were cultivated in darkness (6.17 mg/g of dm), whereas sinapic acid derivatives were more concentrated in sample that germinated without light (49.79 mg/g of dm) than with the day light (45.24 mg/g of dm). Especially, two times more sinapine was present in sample cultivated in darkness. However, the total content of phenolics in both samples was almost the same: 54.00 mg/g of dm for samples that germinated in the presence of light and 54.99 mg/g of dm for those in darkness.
Rochfort et al. [19] determined the total contents of aglycones kaempferol, isorhamnetin, and quercetin to range between 0.7 and 1.2 mg/g of dm in B. rapa L. ssp. chinensis L. These results were obtained for aglycones after hydrolysis reactions. Martínez-Sánchez et al. [34] found that N. officinale R.Br. leaves showed the highest content with mean values of 263 mg of polyphenols per 100 g fw; while for mizuna, salad rocket, and wild rocket leaves, the content was the lowest (99.1, and 139.0 mg/100 g fw, respectively). This value was also higher than the content reported for other Brassicaceae vegetables such as different broccoli cultivars, cabbage, Chinese cabbage, and Brussels sprouts. Singh et al. [39] found different contents of polyphenols in Brassica vegetable and showed that polyphenols may be influenced by several factors, including genetic growing period and environmental influences, especially exposition to light.
After the analysis of L. sativum L. sprouts was performed by UPLC-PDA-Q/TOF-MS, the ChromaLynx™ software scanned the extracts for thirteen minutes. Good separation and resolution results are obtained by utilizing particles of only 1.7 μm and pressure-tolerant reversed-phase (≤800 bar).
Conclusion
As a result, a noticeably shorter chromatography time with defined peaks will be achieved in comparison with traditional HPLC systems. Aside from the time of analysis, the detectors that were used play an important role. The selectivity and sensitivity of TOF-MS detectors make it a good alternative to PDA for determination of low UV-sensitive substances which are difficult to analyze in small amounts. Since the hybrid Q/TOF-mass detector offers not only high selectivity and sensitivity but also great mass accuracy (ensured by permanent injection of leucine enkephalin), the assumed peaks can be assigned to individual substances. In spite of its selectivity and sensitivity, the hybrid Q/TOF-mass detector reached its limit with regard to accurate mass measurement. The determination and the evaluation of L. sativum extracts is much simpler using the UPLC-Q/TOF-MS. Even though they only occur in low concentrations, the combination of the retention time and the accurate molecular mass allows for a fast qualification of substances with the use of ChromaLynx™ software.
The major compounds from among sinapic acid derivatives to be found in L. sativum sprouts germinated both in light and in darkness were sinapic acid and sinapine, and from among flavonol derivatives: kaempferol-sinapoyldihexose-pentose.
References
Carbajal D, Casaco A, Arruzazabala L, Gonzalez R, Fuentes V (1991) J Ethnopharm 33:21–24
Hasegawa K, Mizutani J, Kosemura S, Yamamura S (1992) Plant Physiol (USA) 100(2):1059–1061
Sumangala S, Nagappa G, Malleshi G, Mingruo G (2005) Plant Foods Hum Nutr 59(3):105–111
Mathews S, Singhal RS, Kulkarni PR (1993) Nahrung 37:69–71
Gokavi SS, Malleshi NG, Guo M (2004) Plant Foods Hum Nutr 59:105–111
Al-Yahya MA, Mossa JS, Ageel AM, Rafatullah S (1994) Phytomedicine 1:155–159
Paranjape AN, Mehta AA (2006) Iranian J Pharm Ther 5(1):55–59
La Hamedan WA (2010) J Am Sci 6(11):873–879
Fahey JW, Zalcmann AT, Talalay P (2001) Phytochemistry 56:5–51
Chandel KPS, Shukla G, Sharma N (1996) Biodiversity in medicinal and aromatic plants in India: conservation and utilization. National Bureau of Plant Genetic Resources, New Delhi
Chalas J, Claise C, Edeas M, Messaoudi C, Vergnes L, Abella A, Lindenbaum A (2001) Pharmacotherapy 55:54–60
Shahidi F, Naczk M (1992) JAOCS 69:917–924
Larsen L, Olsen O, Ploeger A, Sorenson H (1983) In: Proceedings 6th international rapeseed congress, Paris, 1577–1582
Sosulski FW, Dąbrowski KJ (1984) J Agric Food Chem 32:131–133
Nayak PS, Upadhyay A, Dwivedi SK, Rao S (2012) EJEAFChe 11(3):156–162
Nayak PS, Upadhyaya SD, Upadhyaya A (2009) J Sci Res 1:121–127
Liorach R, Espin JC, Tomas-Barberan FA, Ferreres F (2003) J Agric Food Chem 51:2181–2187
Ferreres F, Valentao P, Llorach R, Pinheiro C, Cardoso U, Pereira JA, Sousa C, Seabra RM, Andrade PB (2005) J Agric Food Chem 53:2901–2907
Rochfort SJ, Imsic M, Jones R, Trenerry VC, Tomkins B (2006) J Agric Food Chem 54:4855–4860
Kim EH, Kim SH, Chung JI, Chi HY, Kim JA, Chung IM (2006) Eur Food Res Technol 222:201–208
Janicki B, Kupcewicz B, Napierala A, Madzielewska A (2005) Folia Biol 53:121–125
Harbaum B, Hubbermann EM, Wolff C, Herges R, Zhu Z, Schwarz K (2007) J Agric Food Chem 55:8251–8260
Thiyam U, Pickardt C, Ungewiss J, Baumert A (2009) Eur Food Res Technol 229:825–831
Ferreres F, Sousa C, Vrchovska V, Valentao P, Pereira JA, Seabra RM (2006) Eur Food Res Technol 222:88–98
Liang YS, Kim HK, Lefeber AWM, Erkelens C, Choi YH, Verpoorte R (2006) J Chromatogr A 1112:148–155
Brandl W, Herrmann K, Grotjahn L (1984) Z Naturforsch 39C:515–520
Nielsen JK, Olsen O, Petersen LH, Sorensen H (1984) Phytochemistry 23:1741–1743
Goujon T, Sibout R, Pollet B, Maba B, Nussaume L, Bechthold N, Lu F, Ralph J, Mila I, Barriere Y, Lapierre C, Jouanin L (2003) Plant Mol Biol 51:973–989
Marnoch R, Diosady LL (2006) JAOCS 83(1):65–69
Wanasundara U, Amarowicz R, Shahidi F (1994) J Agric Food Chem 42:1285–1290
Ferreres F, Llorach R, Gil-Izquierdo A (2004) J Mass Spectrom 39(3):312–321
Sasaki K, Takahashi TA (2002) Phytochemistry 61:339–343
Fernandes F, Valentao P, Sousa C, Pereira JA, Seabra RM, Andrade PB (2007) Food Chem 105:1003–1010
Martínez-Sánchez A, Gil-Izquierdo A, Gil MI, Ferreres F (2008) J Agric Food Chem 56:2330–2340
Baumert A, Milkowski C, Schmidt J, Nimtz M, Wray V, Strack D (2005) Phytochemistry 66:1334–1345
Ferreres F, Sousa C, Valentao P, Seabra RM, Pereira JA, Andrade PB (2007) Food Chem 101:549–558
Gil V, Macloed AJ (1980) Phytochemistry 19:1365–1368
Glendening TM, Poulton JE (1988) Glucosinolate biosynthesis. Plant Physiol 86:319–321
Singh J, Upadhyay AK, Prasad K, Bahadur A, Rai M (2007) J Food Compos Anal 20:106–112
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Oszmiański, J., Kolniak-Ostek, J. & Wojdyło, A. Application of ultra performance liquid chromatography-photodiode detector-quadrupole/time of flight-mass spectrometry (UPLC-PDA-Q/TOF-MS) method for the characterization of phenolic compounds of Lepidium sativum L. sprouts. Eur Food Res Technol 236, 699–706 (2013). https://doi.org/10.1007/s00217-013-1925-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-013-1925-x