Abstract
Long-term and continuous monitoring of the microRNA (miRNA) expression in living cells is essential in biomedical research, but it is currently limited by fast consumption and easy digestion of probes in the intracellular environment. Herein, we report polydopamine-modified gold nanoparticles (AuNPs@PDA) as protective and efficient nanocarriers for DNA hairpin probes (hpDNA), achieving long-term monitoring (48 h) of the miRNA (let-7a) levels in living cells after drug treatments. This method enabled excellent sensitivity and high selectivity toward let-7a with a limit of detection of 0.51 nM (n = 3) and a linear range from 1 to 100 nM. More importantly, AuNPs@PDA can not only efficiently improve the loading of hpDNA on each nanoparticle, but also effectively protect hpDNA from hydrolysis in the cell microenvironment, finally realizing the continuous monitoring of let-7a in living cells for 48 h. This simple method would be of great significance for drug screening and precision medicine.
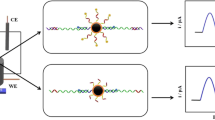
Graphical abstract





Similar content being viewed by others
References
Stefani G, Slack FJ. Small non-coding RNAs in animal development. Nat Rev Mol Cell Biol. 2008;9(3):219–30. https://doi.org/10.1038/nrm2347.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;7(5):522–31. https://doi.org/10.1038/nrg1379.
Esquela-Kerscher A, Slack FJ. Oncomirs — microRNAs with a role in cancer. Nat Rev Cancer. 2006;6(4):259–69. https://doi.org/10.1038/nrc1840.
Zhao X, Li Y, Sun R, Fan Y, Mu X, Wang Y. Electrical potential-assisted DNA-RNA hybridization for rapid microRNA extraction. Anal Bioanal Chem. 2022. https://doi.org/10.1007/s00216-022-03979-8.
Tian J, Chu H, Zhu L, Xu W. Duplex-specific nuclease-resistant triple-helix DNA nanoswitch for single-base differentiation of miRNA in lung cancer cells. Anal Bioanal Chem. 2020;412(19):4477–82. https://doi.org/10.1007/s00216-020-02713-6.
Wang X, Cao L, Wang Y, Wang X, Liu N. Regulation of let-7 and its target oncogenes (review). Oncol Lett. 2012;3(5):955–60. https://doi.org/10.3892/ol.2012.609.
van Schooneveld E, Wildiers H, Vergote I, Vermeulen PB, Dirix LY, Van Laere SJ. Dysregulation of microRNAs in breast cancer and their potential role as prognostic and predictive biomarkers in patient management. Breast Cancer Res. 2015;17(1):21. https://doi.org/10.1186/s13058-015-0526-y.
Bombonati A, Sgroi DC. The molecular pathology of breast cancer progression. J Pathol. 2011;223(2):308–18. https://doi.org/10.1002/path.2808.
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 2005;65(16):7065. https://doi.org/10.1158/0008-5472.CAN-05-1783.
Yu F, Yao H, Zhu P, Zhang X, Pan Q, Gong C, Huang Y, Hu X, Su F, Lieberman J, Song E. Let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell. 2007;131(6):1109–23. https://doi.org/10.1016/j.cell.2007.10.054.
Liu X, Liu L, Xu Q, Wu P, Zuo X, Li A. MicroRNA as a novel drug target for cancer therapy. Expert Opin Biol Ther. 2012;12(5):573–80. https://doi.org/10.1517/14712598.2012.671293.
Alimova IN, Liu B, Fan Z, Edgerton SM, Dillon T, Lind SE. Metformin inhibits breast cancer cell growth, colony formation and induces cell cycle arrest in vitro. Cell Cycle. 2009;8(6):909–15. https://doi.org/10.4161/cc.8.6.7933.
Yin BC, Liu YQ, Ye BC. One-step, multiplexed fluorescence detection of microRNAs based on duplex-specific nuclease signal amplification. J Am Chem Soc. 2012;134(11):5064–7. https://doi.org/10.1021/ja300721s.
Qiu X, Wang P, Cao Z. Hybridization chain reaction modulated DNA-hosted silver nanoclusters for fluorescent identification of single nucleotide polymorphisms in the let-7 miRNA family. Biosens Bioelectron. 2014;60:351–7. https://doi.org/10.1016/j.bios.2014.04.040.
Zhu D, Huang J, Lu B, Zhu Y, Wei Y, Zhang Q. Intracellular microRNA imaging with MoS2-supported nonenzymatic catassembly of DNA hairpins. ACS Appl Mater Interfaces. 2019;11(23):20725–33. https://doi.org/10.1021/acsami.9b04883.
Zhao P, Li N, Astruc D. State of the art in gold nanoparticle synthesis. Coord Chem Rev. 2013;257(3–4):638–65. https://doi.org/10.1016/j.ccr.2012.09.002.
Qian R-C, Cao Y, Zhao L-J, Gu Z, Long Y-T. A two-stage dissociation system for multilayer imaging of cancer biomarker-synergic networks in single cells. Angew Chem Int Ed. 2017;56(17):4802–5. https://doi.org/10.1002/anie.201702415.
Choi C, Li J, Wei K, Xu YJ, Ho L, Zhu M. A gold@polydopamine core–shell nanoprobe for long-term intracellular detection of microRNAs in differentiating stem cells. J Am Chem Soc. 2015;137(23):7337–46. https://doi.org/10.1021/jacs.5b01457.
Zhu C, Zeng Z, Hai L, Li F, Fan C, Zhang H. Single-layer MoS2-based nanoprobes for homogeneous detection of biomolecules. J Am Chem Soc. 2013;135(16):5998–6001. https://doi.org/10.1021/ja4019572.
Baptista P, Pereira E, Eaton P, Doria G, Miranda A, Gomes I. Gold nanoparticles for the development of clinical diagnosis methods Anal. Bioanal Chem. 2008;391(3):943–50. https://doi.org/10.1007/s00216-007-1768-z.
Dai G, Choi CKK, Zhou Y, Bai Q, Xiao Y, Yang C, Choi CHJ, Ng DKP. Immobilising hairpin DNA-conjugated distyryl boron dipyrromethene on gold@polydopamine core–shell nanorods for microRNA detection and microRNA-mediated photodynamic therapy. Nanoscale. 2021;13:6499. https://doi.org/10.1039/D0NR09135A.
Sy KHS, Ho LWC, Lau WCY, Ko H, Choi CHJ. Morphological diversity, protein adsorption, and cellular uptake of polydopamine-coated gold nanoparticles. Langmuir. 2018;34:14033–45. https://doi.org/10.1021/acs.langmuir.8b02572.
Jiang J, Zhu L, Zhu L, Zhu B, Xu Y. Surface characteristics of a self-polymerized dopamine coating deposited on hydrophobic polymer films. Langmuir. 2011;27(23):14180–7. https://doi.org/10.1021/la202877k.
Qiang W, Li W, Li X, Chen X, Xu D. Bioinspired polydopamine nanospheres: a superquencher for fluorescence sensing of biomolecules. Chem Sci. 2014;5(8):3018–24. https://doi.org/10.1039/c4sc00085d.
Xu S, Zhang G, Fang B, Xiong Q, Duan H, Lai W. Lateral flow immunoassay based on polydopamine-coated gold nanoparticles for the sensitive detection of zearalenone in maize. ACS Appl Mater Interfaces. 2019;11:31283–90. https://doi.org/10.1021/acsami.9b08789.
Dai W, Lu H, Yang F, Dong H, Zhang X. Accurate detection of intracellular microRNAs using functional Mo2C quantum dots nanoprobe. Chem Commun. 2019;55:10615–8. https://doi.org/10.1039/C9CC04261J.
Zhang H, Wang Y, Zhao D, Zeng D, Xia J, Aldalbahi A. Universal fluorescence biosensor platform based on graphene quantum dots and pyrene-functionalized molecular beacons for detection of microRNAs. ACS Appl Mater Interfaces. 2015;7(30):16152–6. https://doi.org/10.1021/acsami.5b04773.
Mahani M, Mousapour Z, Divsar F, Nomani A, Ju H. A carbon dot and molecular beacon based fluorometric sensor for the cancer marker microRNA-21. Microchim Acta. 2019;186(3):132. https://doi.org/10.1007/s00604-019-3233-z.
Bian F, Sun L, Cai L, Wang Y, Zhao Y, Wang S. Molybdenum disulfide-integrated photonic barcodes for tumor markers screening. Biosens Bioelectron. 2019;133:199–204. https://doi.org/10.1016/j.bios.2019.02.066.
Yu X, Hu L, Zhang F, Wang M, Xia Z, Wei W. MoS2 quantum dots modified with a labeled molecular beacon as a ratiometric fluorescent gene probe for FRET based detection and imaging of microRNA. Microchim Acta. 2018;185:239. https://doi.org/10.1007/s00604-018-2773-y.
Han Y, Zou R, Wang L, Chen C, Gong H, Cai C. An amine-functionalized metal-organic framework and triple-helix molecular beacons as a sensing platform for miRNA ratiometric detection. Talanta. 2021;228(1):122199. https://doi.org/10.1016/j.talanta.2021.122199.
Zhao H, Qu Y, Yuan F, Quan X. A visible and label-free colorimetric sensor for miRNA-21 detection based on peroxidase-like activity of graphene/gold-nanoparticle hybrids. Anal Methods. 2016;8:2005–12. https://doi.org/10.1039/C5AY03296B.
Ghosh R, Singh LC, Shohet JM, Gunaratne PH. A gold nanoparticle platform for the delivery of functional microRNAs into cancer cells. Biomaterials. 2013;34(3):807–16. https://doi.org/10.1016/j.biomaterials.2012.10.023.
Wang N, Song L, Qiu Y, Xing H, Li J. Hybridization-activated spherical DNAzyme for cascading two-photon fluorescence emission: applied for intracellular miRNA measurement by two-photon microscopy. Sensors Actuatorors B Chem. 2019;286:250–7. https://doi.org/10.1016/j.snb.2019.01.135.
Heneghan HM, Miller N, Lowery AJ, Sweeney KJ, Newell J, Kerin MJ. Circulating microRNAs as novel minimally invasive biomarkers for breast cancer. Ann Surg. 2010;251(3):499–505. https://doi.org/10.1097/SLA.0b013e3181cc939f.
Oliveras-Ferraros C, Cufí S, Vazquez-Martin A, Torees-Garcia VZ, Barco SD, Martin-Castillo BA. Micro(mi)RNA expression profile of breast cancer epithelial cells treated with the anti-diabetic drug metformin: induction of the tumor suppressor miRNA let-7a and suppression of the TGFβ-induced oncomiR miRNA-181a. Cell Cycle. 2011;10(7):1144–51. https://doi.org/10.4161/cc.10.7.15210.
Acknowledgements
This work was supported by the Scientific Technology Project of Shenzhen City (JCYJ20200109142410170 and JCYJ20210324120601004), the National Natural Science Foundation of China (21801259 and 21974153), the Guangdong Natural Science Foundation (2022A1515011318), the Scientific Technology Project of Guangzhou City (202103000003), and the Guangdong Science and Technology Plan Project Grant (2020B1212060077).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liang, Y., Yang, H., Yin, W. et al. Long-term continuous monitoring of microRNA in living cells using modified gold nanoprobe. Anal Bioanal Chem 414, 6157–6166 (2022). https://doi.org/10.1007/s00216-022-04182-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-04182-5