Abstract
It has been a challenge to analyze minute amounts of proteomic samples in a facile and robust manner. Herein, we developed a quantitative proteomics workflow by integrating suspension trapping (S-Trap)–based sample preparation and label-free data-independent acquisition (DIA) mass spectrometry and then applied it for the analysis of microgram and even nanogram amounts of exosome samples. S-Trap–based sample preparation outperformed the traditional in-solution digestion-based approach and the commonly used filter-aided sample preparation (FASP)–based approach with regard to the number of proteins and peptides identified. Moreover, S-Trap–based sample preparation coupled with DIA mass spectrometry also showed the highest reproducibility for protein quantification. In addition, this approach allowed for identification and quantification of exosome proteins with low starting amounts (down to 50 ~ 200 ng). Finally, the proposed method was successfully applied to label-free quantification of exosomal proteins extracted from MDA-MB-231 breast cancer cells and MCF-10A non-tumorigenic epithelial breast cells. Prospectively, we envision the integrated S-Trap sample preparation coupled with DIA quantification strategy as a promising alternative for highly efficient and sensitive analysis of trace amounts of proteomic samples (e.g., exosomal samples).
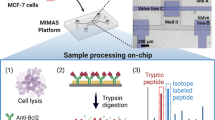
Graphical abstract






Similar content being viewed by others
References
Aebersold R, Mann M. Mass-spectrometric exploration of proteome structure and function. Nature. 2016;537(7620):347–55.
Zhang Y, Fonslow BR, Shan B, Baek M-C, Yates JR. Protein analysis by shotgun/bottom-up proteomics. Chem Rev. 2013;113(4):2343–94.
Ye X, Tang J, Mao Y, Lu X, Yang Y, Chen W, et al. Integrated proteomics sample preparation and fractionation: method development and applications. TrAC Trends Anal Chem. 2019;120:115667.
Ma J, Zhang L, Liang Z, Shan Y, Zhang Y. Immobilized enzyme reactors in proteomics. TrAC Trends Anal Chem. 2011;30(5):691–702.
Wiśniewski JR, Zougman A, Nagaraj N, Mann M. Universal sample preparation method for proteome analysis. Nat Methods. 2009;6(5):359–62.
Liebler DC, Ham A-JL. Spin filter–based sample preparation for shotgun proteomics. Nat Methods. 2009;6(11):785–785.
Kulak NA, Pichler G, Paron I, Nagaraj N, Mann M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat Methods. 2014;11(3):319–24.
Zhu Y, Piehowski PD, Zhao R, Chen J, Shen Y, Moore RJ, et al. Nanodroplet processing platform for deep and quantitative proteome profiling of 10–100 mammalian cells. Nat Commun. 2018;9(1):882.
Yi L, Piehowski PD, Shi T, Smith RD, Qian W-J. Advances in microscale separations towards nanoproteomics applications. J Chromatogr A. 2017;1523:40–8.
Zougman A, Selby PJ, Banks RE. Suspension trapping (STrap) sample preparation method for bottom-up proteomics analysis. Proteomics. 2014;14(9):1006–1000.
HaileMariam M, Eguez RV, Singh H, Bekele S, Ameni G, Pieper R, et al. S-Trap, an ultrafast sample-preparation approach for shotgun proteomics. J Proteome Res. 2018;17(9):2917–24.
Hughes CS, Moggridge S, Müller T, Sorensen PH, Morin GB, Krijgsveld J. Single-pot, solid-phase-enhanced sample preparation for proteomics experiments. Nat Protoc. 2019 Jan;14(1):68–85.
Chen W, Wang S, Adhikari S, Deng Z, Wang L, Chen L, et al. Simple and integrated spintip-based technology applied for deep proteome profiling. Anal Chem. 2016;88(9):4864–71.
Lange V, Picotti P, Domon B, Aebersold R. Selected reaction monitoring for quantitative proteomics: a tutorial. Mol Syst Biol. 2008;4(1):222.
Peterson AC, Russell JD, Bailey DJ, Westphall MS, Coon JJ. Parallel reaction monitoring for high resolution and high mass accuracy quantitative, targeted proteomics. Mol Cell Proteomics. 2012;11(11):1475–88.
Gillet LC, Navarro P, Tate S, Röst H, Selevsek N, Reiter L, et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteomics. 2012;11(6):O111.016717.
Ludwig C, Gillet L, Rosenberger G, Amon S, Collins BC, Aebersold R. Data-independent acquisition-based SWATH-MS for quantitative proteomics: a tutorial. Mol Syst Biol. 2018;14(8).
Li W, Chi H, Salovska B, Wu C, Sun L, Rosenberger G, et al. Assessing the relationship between mass window width and retention time scheduling on protein coverage for data-independent acquisition. J Am Soc Mass Spectrom. 2019;30(8):1396–405.
Mittelbrunn M, Sánchez-Madrid F. Intercellular communication: diverse structures for exchange of genetic information. Nat Rev Mol Cell Biol. 2012;13(5):328–35.
Simons M, Raposo G. Exosomes–vesicular carriers for intercellular communication. Curr Opin Cell Biol. 2009;21(4):575–81.
Théry C, Zitvogel L, Amigorena S. Exosomes: composition, biogenesis and function. Nat Rev Immunol. 2002;2(8):569–79.
Simpson RJ, Lim JW, Moritz RL, Mathivanan S. Exosomes: proteomic insights and diagnostic potential. Expert Rev Proteomics. 2009;6(3):267–83.
Hou R, Li Y, Sui Z, Yuan H, Yang K, Liang Z, et al. Advances in exosome isolation methods and their applications in proteomic analysis of biological samples. Anal Bioanal Chem. 2019;411(21):5351–61.
Fricke F, Lee J, Michalak M, Warnken U, Hausser I, Suarez-Carmona M, et al. TGFBR2-dependent alterations of exosomal cargo and functions in DNA mismatch repair-deficient HCT116 colorectal cancer cells. Cell Commun Signal. 2017;15(1):14.
Mitchell MI, Ben‐Dov IZ, Liu C, Ye K, Chow K, Kramer Y, et al. Extracellular vesicle capture by AnTibody of CHoice and Enzymatic Release (EV‐CATCHER): a customizable purification assay designed for small‐RNA biomarker identification and evaluation of circulating small‐EVs. J Extracell Vesicles. 2021;10(8).
Aldeghaither DS, Zahavi DJ, Murray JC, Fertig EJ, Graham GT, Zhang Y-W, et al. A mechanism of resistance to antibody-targeted immune attack. Cancer Immunol Res. 2019;7(2):230–43.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Methodol. 1995;57(1):289–300.
Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57.
Galperin MY, Cochrane GR. Nucleic acids research annual database issue and the NAR online Molecular Biology Database Collection in 2009. Nucleic Acids Res. 2009 Jan 1;37(Database):D1–4.
Ludwig KR, Schroll MM, Hummon AB. Comparison of in-solution, FASP, and S-Trap based digestion methods for bottom-up proteomic studies. J Proteome Res. 2018;17(7):2480–90.
Keerthikumar S, Chisanga D, Ariyaratne D, Saffar H, Anand S, Zhao K, Samuel M, Pathan M, Jois M, Chilamkurti N, Gangoda L, Mathivanan S. ExoCarta: a web-based compendium of exosomal cargo. J Mol Biol. 2016;428(4):688–92.
Risha Y, Minic Z, Ghobadloo SM, Berezovski MV. The proteomic analysis of breast cell line exosomes reveals disease patterns and potential biomarkers. Sci Rep. 2020 Dec;10(1):13572.
Cho K-C, Clark DJ, Schnaubelt M, Teo GC, Leprevost F da V, Bocik W, et al. Deep proteomics using two dimensional data independent acquisition mass spectrometry. Anal Chem. 2020;92(6):4217–25.
Yamaguchi H, Condeelis J. Regulation of the actin cytoskeleton in cancer cell migration and invasion. Biochim Biophys Acta BBA - Mol Cell Res. 2007;1773(5):642–52.
Kanojia D, Morshed RA, Zhang L, Miska JM, Qiao J, Kim JW, et al. βIII-Tubulin regulates breast cancer metastases to the brain. Mol Cancer Ther. 2015;14(5):1152–61.
Hoshino A, Costa-Silva B, Shen T-L, Rodrigues G, Hashimoto A, Tesic Mark M, et al. Tumour exosome integrins determine organotropic metastasis. Nature. 2015;527(7578):329–35.
Tang H, Wang Y, Zhang B, Xiong S, Liu L, Chen W, et al. High brain acid soluble protein 1(BASP1) is a poor prognostic factor for cervical cancer and promotes tumor growth. Cancer Cell Int. 2017;17(1):97.
Hsiao K-C, Shih N-Y, Fang H-L, Huang T-S, Kuo C-C, Chu P-Y, et al. Surface α-enolase promotes extracellular matrix degradation and tumor metastasis and represents a new therapeutic target. Karamanos NK, editor. PLoS ONE. 2013;8(7):e69354.
Tu S-H, Chang C-C, Chen C-S, Tam K-W, Wang Y-J, Lee C-H, et al. Increased expression of enolase α in human breast cancer confers tamoxifen resistance in human breast cancer cells. Breast Cancer Res Treat. 2010 Jun;121(3):539–53.
Li S, Li X, Yang S, Pi H, Li Z, Yao P, et al. Proteomic landscape of exosomes reveals the functional contributions of CD151 in triple-negative breast cancer. Mol Cell Proteomics. 2021;20:100121.
Cascone T, McKenzie JA, Mbofung RM, Punt S, Wang Z, Xu C, et al. Increased tumor glycolysis characterizes immune resistance to adoptive T cell therapy. Cell Metab. 2018;27(5):977-987.e4.
Wortzel I, Dror S, Kenific CM, Lyden D. Exosome-mediated metastasis: communication from a distance. Dev Cell. 2019;49(3):347–60.
Costa-Silva B, Aiello NM, Ocean AJ, Singh S, Zhang H, Thakur BK, et al. Pancreatic cancer exosomes initiate pre-metastatic niche formation in the liver. Nat Cell Biol. 2015;17(6):816–26.
Acknowledgements
This work is partially supported by NIH/NCI grant P30 CA051008 and GUMC institutional support. The Orbitrap Lumos Tribrid mass spectrometer is partially supported by Dekelbaum Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, C., Zhou, S., Mitchell, M.I. et al. Coupling suspension trapping–based sample preparation and data-independent acquisition mass spectrometry for sensitive exosomal proteomic analysis. Anal Bioanal Chem 414, 2585–2595 (2022). https://doi.org/10.1007/s00216-022-03920-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-03920-z