Abstract
Obtaining structural information for lipids such as phosphatidylcholines, in particular the location of double bonds in their fatty acid constituents, is an ongoing challenge for mass spectrometry (MS) analysis. Here, we present a novel method utilizing the doping of liquid matrix-assisted laser desorption/ionization (MALDI) samples with divalent metal chloride salts, producing ions with the formula [L+M]2+ (L = lipid, M = divalent metal cation). Multiply charged lipid ions were not detected with the investigated trivalent metal cations. Collision-induced dissociation (CID) product ions from doubly charged metal-cationized lipids include the singly charged intact fatty acids [snx+M–H]+, where ‘x’ represents the position of the fatty acid on the glycerol backbone. The preference of the divalent metal cation to locate on the sn2 fatty acid during CID was found, enabling stereochemical assignment. Pseudo-MS3 experiments such as in-source decay (ISD)-CID and ion mobility-enabled time-aligned parallel (TAP) MS of [snx+M–H]+ provided diagnostic product ion spectra for determining the location of double bonds on the acyl chain and were applied to identify and characterize lipids extracted from soya milk. This novel method is applicable to lipid profiling in the positive ion mode, where structural information of lipids is often difficult to obtain.

MALDI of liquid lipid samples doped with divalent metal salt (e.g. BaCl2) produces doubly charged lipid-barium ions and enables structural elucidation via MS/MS and MS3 analysis
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Production of predominantly multiply charged ions of biological molecules is relatively novel in matrix-assisted laser desorption/ionization (MALDI) mass spectrometry (MS), compared with electrospray ionization (ESI) where they form readily. However, recent developments have introduced the combination of liquid MALDI samples and atmospheric pressure (AP)-MALDI sources specifically for this purpose [1,2,3]. For peptides and proteins, it is assumed in these cases that the processes leading to multiply charged ions are ’ESI-like’, with desolvation aided by the heated ion transfer tube common to these ion sources, and that the typical MALDI processes continue to favor the production of singly charged ions.
Multiply charged ions are advantageous for various MS analyzers for reasons ranging from increased detection sensitivity to more highly abundant collision-induced dissociation (CID) product ions, and access to electron-mediated MS/MS methods such as electron transfer dissociation (ETD). Typically, ETD and electron capture dissociation (ECD) rely on the reduction of the positive charge state for fragmentation to occur, so at least two charges are required on precursor ions for the MS detection of charged product ions [4, 5].
However, for lipid analysis the above attempts to produce multiply charged intact MALDI parent ions usually still fail. For survey scans or profiling work this is normally not a problem but when sample differences and further lipid characterization depend on the determination of the exact lipid structure and location of double bonds in the fatty acid side chains, singly charged lipids usually provide insufficient diagnostic fragment ions.
For instance, phosphatidylcholines (PCs) analyzed in positive ion mode with a soft ionization source typically form protonated [L+H]+ or sodiated [L+Na]+ ions [6,7,8]. PCs are zwitterionic and possess a permanent positive charge on the nitrogen atom of the phosphocholine head group. The phosphate anion’s negative charge is neutralized by a proton or other cation during ionization. CID of phospholipid cations revealed separate fragmentation mechanisms for [L+H]+ compared with [L+Alk]+ (where Alk = Li, Na) [9]. The product of [L+H]+ was almost exclusively the positively charged phosphocholine head group, with the bulk of the molecule remaining uncharged. Hence, little structural information was obtainable. Far more information was obtained with lithium adducts, where products of the neutral loss of one or the other fatty acid were detected. Lithium and the other Group 1 metals have been used in the literature for structural investigations [10,11,12,13,14], although their utility in obtaining double bond information from lipids typically requires multiple stages of MS/MS and is based on neutral loss rather than direct detection of product ions [14, 15].
Despite extensive investigations with additives, including metal (M) cations, [L+M]2+ ions have not been reported previously. In fact, it was once explicitly suggested that divalent cations did not complex with PCs because of their zwitterionic nature [16]. More recent studies have found that doubly charged lipid–divalent metal complexes can be produced as [Ln+M]2+ precursors by ESI and used for MS/MS analysis, relying on multiple lipid molecules bound to a metal center but thus producing little structural information for double bond assignment [17, 18]. Complexes with a high coordination number (>10) were produced with a biphasic ESI (BESI) source but again no detection of [L+M]2+ was reported in this study either [19]. Another study investigated ECD MS/MS of complexes [5]. While double bond location was again not possible, the stereochemistry of the fatty acids could be inferred through a preference for neutral loss at sn1.
Doubly sodiated and doubly charged lipid ions ([L+2Na]2+) produced with ESI were shown to be susceptible to ETD MS/MS analysis [4]. Ester bond cleavage was determined to be the dominant ETD process. Product ions directly corresponding to sodiated fatty acids were not detected although tentative assignment of double bond location was suggested from the fragmentation of other products.
For the structural characterization of double bond location in lipids, some advances have recently been made by using new techniques such as ozone-induced dissociation (OzID). It has been demonstrated that OzID is functional on chromatographic timescales, which is of importance for the separation of structural isomers by liquid chromatography (LC) and quantitative lipidomics [20, 21]. Ozone was introduced to a high-pressure region of the mass spectrometer in order to perform online ozonolysis at the fatty acid double bonds [22, 23]. Characteristic product ions, including radical cations, were produced from rapidly formed reaction products. Double bond location analysis by neutral loss scanning with a triple quadrupole mass spectrometer, and ozonolysis performed in the second quadrupole, were possible because of the rate and type of product ion formation [24]. CID prior to OzID revealed that a further level of information is achievable, e.g., the detection of sodiated free fatty acids [25,26,27]. Ozone ESI (OzESI), i.e., ESI in an ozone environment, has been used in a similar fashion to create C=C-cleaved reaction products [28]. Disadvantageously to what is an otherwise powerful method, ozone carries significant safety concerns and must be generated in situ.
As an alternative, PB-MS exploits the Paternò-Büchi photochemical reaction. In the presence of a carbonyl compound, a four-membered oxetane ring is formed across an alkene double bond on activation with UV radiation. In CID product ion spectra of lipids, a characteristic Δm/z of 26 per double bond indicates their position in the acyl chains [29]. Relative and absolute quantitation of constituent isomers has been reported based on the intensity of the diagnostic product ions [30]. Recently, this characteristic Δm/z of 26 has been applied to double bond determination in cholesteryl ester lithium adducts by MS3 analysis of the PB-reaction products [31]. The possibility of including the carbonyl reagent in the LC mobile phase makes PB-MS a good option for online lipidomic workflows.
Ultraviolet photodissociation (UVPD) is another promising approach for lipid characterization [32, 33]. UVPD has recently been implemented on commercial instrumentation, initially requiring significant non-trivial instrument modifications [34, 35]. It differs from collision- or electron-mediated dissociation methods in that the fragmentation energy is delivered by photon absorption. Typically, photons with sufficient energy are provided by a 193 nm pulsed laser. UVPD has just recently been applied to positive ion mode analysis of PCs and sphingolipids, which resulted in structurally diagnostic product ion detection from a single MS/MS stage [36, 37]. Overall, analytical performance was found to be comparable to other MS/MS methods, although the current sensitivity in product ion detection is lower. For lipid analysis, some UVPD product ions were found to be common to higher energy collisional dissociation (HCD) as recorded on Orbitrap mass spectrometers, although the most valuable diagnostic ions for double bond characterization were obtained by UVPD only.
Charge transfer dissociation (CTD) is a radical-induced dissociation method that has also been applied to phospholipids [38]. Interestingly, with CTD a charge-increased species [L+H]2+• was produced, as with metastable atom dissociation (MAD) [39]. Extensive information from the acyl chains was obtained through dissociation of [L+H]+ and [L+H]2+•, although at a relatively low fragment ion signal-to-noise ratio. Advantageously, this information was obtained through a single MS/MS stage. Free fatty acids were not detected. Thus, their identity was inferred from neutral losses and complex fragmentation spectra. CTD currently requires significant instrument modification compared with other methods included here, but yielded some completely different fragment ions for structural elucidation.
Differential mobility spectrometry-electron impact excitation of ions from organics (DMS-EIEIO) has been demonstrated to allow the determination of lipid cis and trans double bond isomers [40]. Characteristic fragment ions of singly charged trans isomers were generated on irradiation with electrons. Separation of the isomers with DMS enabled confident identification although such separation was not possible for all lipid species, such as sphingomyelins.
In the presented work, the addition of divalent metal salts (MCl2), in particular BaCl2, to liquid MALDI samples enabled double bond location for phosphatidylcholines to be assigned from MSn data of [L+M]2+ precursor ions (L = lipid, M = divalent metal cation). The metal salt additions were straightforward to implement into standard liquid MALDI sample preparation workflows for lipid profiling analysis, and no instrument modification beyond our in-house AP-MALDI source was necessary.
Experimental
HPLC-grade water, methanol, acetonitrile, and propan-2-ol were purchased from Fisher Scientific (Loughborough, UK). HPLC-grade hexane was purchased from Sigma-Aldrich (Gillingham, UK).
For MALDI MS analyses, a liquid support matrix (LSM) was prepared by dissolving 2,5-dihydroxybenzoic acid (DHB; >99%; Sigma-Aldrich) in water/acetonitrile (3:7; v:v) at a concentration of 25 mg/mL with glycerol added equal to 60% of the DHB solution volume.
Multivalent metal salts (MgCl2, CaCl2, SrCl2, BaCl2, MnCl2, CoCl2, ZnCl2, FeCl2, AlCl3, ScCl3, CrCl3, LaCl3) and glycerol were purchased from Sigma-Aldrich. Salts were dissolved in water at a concentration of 1 nmol/μL. The metal salts should be regarded as toxic and handled appropriately.
The lipid analyte standards PC (16:0/18:2(9Z,12Z)), PC (16:0/18:1(9Z)), PC (18:1(9Z)/16:0), and PE (16:0/16:0) were purchased from Sigma-Aldrich, whereas PC (16:0/18:0) was purchased from Avanti Polar Lipids (Alabaster, AL, USA). All PC and PE standards were dissolved in methanol. The sphingomyelin (SM) standard SM (d18:1/12:0) was purchased as an ethanolic solution from Avanti Polar Lipids through Stratech Scientific Ltd. (Newmarket, UK). Table 1 lists all standards used in this work.
Analysis was performed using a Synapt G2-Si HDMS mass spectrometer (Waters Corporation, Wilmslow, UK) equipped with an in-house developed AP-(MA)LDI source, featuring a Waters Research Enabled Software (WREnS)-controlled XY-stage. This setup has been previously reported [1]. In brief, an MNL100 nitrogen laser (LTB Lasertechnik Berlin, Berlin, Germany) with a wavelength of 337 nm, maximum pulse rate of 30 Hz, and a pulse width of approximately 3 ns was attenuated by a neutral density filter to 20–30 μJ/pulse for MALDI MS measurements. A home-made stainless steel ion transfer tube (70 mm length, 1 mm internal diameter) was added as the first ion extracting element to the ion block with a gap of approximately 3 mm between the MALDI target plate and inlet of the ion transfer tube and heated using a resistance wire with approximately 26 W delivered by a low-voltage DC power supply. A counter-flow gas (N2, 180 L/h, approximately 150 °C) was applied to the ion inlet while the MALDI target voltage was set to 3–4 kV and the cone voltage to 40 V. Experiments were run in the sensitivity mode (mass resolving power of approximately 10,000) with ion mobility (IM) measurements enabled. The time-of-flight (TOF) analyzer was calibrated on the [M+H]+ CID product ions of the [Glu1]-fibrinopeptide precursor [M+2H]2+ (Sigma-Aldrich). For IM analysis, data files were calibrated post-acquisition with polyalanine (Sigma-Aldrich) cluster ion peaks to allow drift time conversion to collision cross-section (CCS) if required. All calibrations were performed using the AP-MALDI source and LSM MALDI samples. Depending on the experiment, CID was enacted in the trap (pre-IM) and/or transfer (post-IM) cell of the TriWave device. Pseudo-MS3 experiments were performed through in-source decay (ISD) by raising the source cone voltage from the typical 40 V to 100–120 V and the trap collision potential to 40 V. Time-aligned parallel (TAP) experiments used a collision potential of 22 V (trap) and 40 V (transfer).
Liquid MALDI samples were prepared on a commercial stainless steel 96-well MALDI target plate (Waters) by combining (in order) 0.5 μL of each of the LSM, salt solution, and analyte solution.
Results and discussion
[L+M]2+ ion formation was initially noticed without any chromophore, such as DHB, in the liquid MALDI matrix under specific sampling conditions on the custom AP-MALDI source. A nuance of these AP-(MA)LDI MS analyses was that for any [L+M]2+ ion beam to be detected, the laser had to be focused at the edge of the liquid sample droplet. Firing the laser directly into the droplet center did not result in a detectable analyte ion beam. Additionally, and unlike liquid AP-MALDI MS with a matrix chromophore, higher laser pulse rates did not substantially enhance analyte ion signal intensity, arguably because of the greater ablation and associated changes in sample morphology and time required to ‘heal’ the droplet edge. The sample droplet depleted quickly at 30 Hz, and a pulse rate of 10 Hz was found to be optimal for ion signal and droplet longevity. After these initial discoveries, abundant [L+M]2+ ions formation was recorded with standard MALDI LSMs. Since MALDI samples prepared with LSMs exhibit superior ion current stability, these became the main type of samples for this work.
In general, in all measurements the base peak ion was the [L+H]+ ion, and there was some evidence to suggest that the formation of [L+H]+ and [L+M]2+ relies on different ionization processes. For instance, optimization experiments revealed separate trends in the intensity of these two ion types with varied BaCl2 amount (see Electronic Supplementary Material (ESM), Fig. S1). Doubly charged ions were only detected when divalent metal salts were added to the MALDI sample, forming [L+M]2+ ions.
TOF MS spectra varied noticeably with each different metal salt added to the MALDI sample. Typically, Mg2+as well as Mn2+ and Co2+ resulted in minor [L+M]2+ formation and significant spontaneous fragmentation. Fe2+ and Zn2+ were also tested and led to [L+M]2+ ions but were not further pursued as these metals have various naturally abundant isotopes and oxidation states. Interestingly, descending Group 2 of the Periodic Table revealed a trend of increased intensity, indicating a possible dependence on the metal ion radius. Figure 1 displays the relative intensities recorded for four different Group 2 metal chloride salts. With respect to [L+M]2+ ion signal intensity, Ba2+ adduct ion formation performed most optimally of all metal ions tested in a molar ratio of 500:1 (metal:lipid), resulting in spectra featuring [L+Ba]2+ and [2L+Ba]2+. [L+H]+ intensity was relatively consistent across the Group 2 metal ions attached to the standard PCs. Since Mg2+, Mn2+, and Co2+ have similar, smaller radii and smaller [L+M]2+ ion signal intensities than Ba2+, which has the largest radius of all metal cations investigated, there is further evidence for a dependency on the radius of the metal cation. Trivalent metal salts (Sc3+, Cr3+, La3+) did not produce detectable [L+M]3+, [L+M-H]2+, or other related ions. Thus, BaCl2 was used for all further experiments.
Liquid AP-MALDI ion signal intensities of [L+M]2+, [L+H]+, and [2L+M]2+ for PC (16:0/18:0) (1 pmol per sample droplet) with Group 2 elements (500 pmol per sample droplet) over 50 s of data acquisition. The MALDI sample droplet consisted of DHB LSM (0.5 μL), salt solution (0.5 μL, 1 nmol/μL), and analyte solution (0.5 μL, 2 pmol/μL). Intensity of 0 was reported if any peak was indistinguishable from the background noise signal. Error bars indicate the 95% confidence interval
For the PC and SM standards, [L+Ba]2+ was easily detected, although when compared with the [L+H]+ ion at a relatively higher intensity for the SM standard (see Fig. 2). Interestingly, both PCs and SMs possess a fixed, positively charged choline group, whereas PEs have a free amine group as head group. This head group is typically protonated under physiological conditions, and certainly under the acidic conditions of the LSM. A peak at m/z 414.72 for the PE standard could not be confirmed to be attributable to [L+Ba]2+ ions using MS and CID-MS/MS analysis. Dimers, previously reported by Ho et al. with the formula [L2-2H+2Ba]2+ were of low abundance [16]. Since the charge of the amine group is provided by a mobile proton, such proton should be able to leave easily when the divalent metal cation and PE form a complex; the result of which is the singly charged species [L–H+Ba]+. However, it appears that the ultimate formation of [L+H]+ rather than [L–H+Ba]+ as the predominant singly charged ion species is favored. The equivalent charge reduction is not possible for lipids containing choline. Should the formation of [L+M]2+ be specific to the lipids with a fixed positive charge, this offers an approach to the separation of isobaric PCs and PEs within a profile spectrum.
Liquid AP-MALDI-TOF MS spectra exhibiting [L+Ba]2+ (zoomed, underlined) and [L+H]+ of the phosphatidylcholines (a) PC (16:0/18:0) m/z 449.76, (b) PC (16:0/18:1) m/z 448.75, (c) PC (16:0/18:2) m/z 447.75, and sphingomyelin (d) SM (d18:1/12:0) m/z 392.22. The peak at m/z 414.72 in (e) could not be confirmed by CID to be the [L+Ba]2+ ion signal of the phosphatidylethanolamine PE (16:0/16:0)
CID fragment ion products of [L+Ba]2+ were further investigated (see Fig. 3). The product ions were classified into two major groups: intact fatty acids with the formula [snx+Ba-H]+ and protonated fragments resulting from the loss of a fatty acid [L+H–snx]+ (with x = 1 or 2). The former are highly abundant, a distinction from the CID product ions of singly charged, lipid-metal adducts where the fatty acids are detected as part of [L+H–snx]+ and inferred from neutral loss. Thus, further MS/MS analysis on individual [snx+Ba–H]+ is possible. Detection of [snx+Ba–H]+ suggests a putative alternative attachment of the divalent metal cation closer to the acyl chains or a relocation of the Ba2+ during CID. The ratios of [sn1+Ba–H]+/[sn2+Ba–H]+ within the same spectra suggest a mechanistic preference for product ion formation dependent on the sn1/sn2 position of the fatty acids. The dominant product ions were recorded for the fatty acids at the sn2 site, which is in agreement with previous research that indicates the sn2 bond as being more labile in CID of [L+H]+ and [L+Li]+ [41]. In contrast, ECD of PC (16:0/18:1) has been reported exhibiting almost equal signal intensities for ions resulting from the neutral loss of fatty acids from each site [5]. A preference for the most unsaturated fatty acid was not detected although this had been suggested with Ag+ adduct ions previously [15]. As doubly charged CID products were observed, it will be possible to perform ETD after CID in the future. CID of SM (d18:1/12:0) revealed a low abundance peak for the sn2 chain (m/z 336.1); other peaks were all head group-related, although with some additional peaks to those usually detected with CID.
Liquid AP-MALDI-CID MS/MS spectra of the precursor [L+Ba]2+ (*) for (a) PC (16:0/18:0), (b) PC (16:0/18:1), (c) PC (18:1/16:0), (d) PC (16:0/18:2), and (e) SM (d18:1/12:0). For the PCs, the dominant product ions are [sn1+Ba–H]+ and [sn2+Ba–H]+ (bold label), and [L+H–sn1]+ and [L+H–sn2]+ with other products including [L+H-sn1-N(CH3)3]+ and [L+H-sn2-N(CH3)3]+ (regular label), and [L+Ba–H-sn1]2+ and [L+Ba–H–sn2]2+ (underlined label). For the SM, many head group-related product ion peaks and a minor peak at m/z 336.10, presumably [amide+Ba–H]+ from the sn2 position, are detected
MS3 product ions from [C18:1+Ba–H]+ were detected in pseudo-MS3 experiments, by increasing the source cone voltage from 40 V to between 100 and 120 V, selecting the ISD product ion by the quadrupole, and then performing CID in the trap cell. ISD is often utilized for MALDI pseudo-MS3, but the high laser fluence typically used results in rapid depletion of the sample. However, AP-MALDI-ISD of peptides and proteins has previously been demonstrated using liquid samples, with ion signal stability for over 40 min [42].
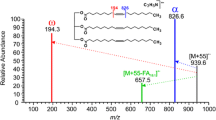
ISD product ions were selected by the quadrupole with a window of approximately m/z 2 around the [C18:n+Ba–H]+ fragment ion peak. A higher sample loading (100 pmol per droplet) was used to compensate for smaller ion current due to the additional fragmentation step. The BaCl2 amount was kept at 500 pmol as an increase into the nmol range proved detrimental to [L+Ba]2+ signal intensity. Data were collected for 5 min with consistent, stable ion current and low sample consumption. From the spectra shown in Fig. 4, evidence of charge-remote fragmentation of the acyl chain is evident, as peaks with a difference of m/z 14 are clearly visible, starting at m/z 209. Mechanisms for MS3 product ion formation are suggested in Fig. 5, including the radical ion at m/z 196 [43, 44]. The location of double bonds is easily revealed by the characteristic differences of m/z 40 between the acyl chain fragment ion signals. Surprisingly, the detection of m/z 333.1 in Fig. 4c might indicate that some impurities were present or the double bond moved in some cases, assuming sufficient energy was available. It can be assumed that it is less favorable for the double bond between C11 and C12 to be cleaved. However, Li et al. did present evidence of double bond cleavage occurring by CTD MS/MS [38]. Abundant ion signals of presumably Ba+• (m/z 137.9) and BaOH+ (m/z 154.9) were also detected. In the future, enhancement of the [snx+Ba–H]+ CID product ion signal might be possible by reducing the formation of these two ions, i.e., keeping Ba on the acyl chain. The data shows that the ions resulting from fragmentation further away from the carbonyl group are produced in lower abundance than those that were closer (see Fig. 4), which has also been observed for CID of OzID products [26].
Proposed fragmentation mechanisms for MS3 product ions, using [oleic acid+Ba–H]+ as an example. 1,4-Elimination (a) can occur where proximity to double bonds does not prohibit such process. Other mechanisms (b), (c) are likely multistep radical reactions, first involving C–C bond homolytic cleavage and subsequent H• loss. The radical ion with m/z 196 resulting from the cleavage of C2–C3 (d) is explained as there is no adjacent H to be lost
Since the pseudo-MS3 product ions were produced prior to the ion mobility cell, their drift time could be recorded. This recording shows that the smaller m/z product ions from all three standards match closely in drift time (see Fig. 6). The break in the trend between m/z 209 to 237 suggests that products of different chain length form different conformations, e.g., linear versus spiral conformation. Some conformations might form preferentially, which could also be the reason for the differences in MS3 product ion intensities. Unsaturated acyl chain product ions with greater m/z values exhibit different drift times compared with saturated acyl chain product ions.
Application to phospholipids extracted from soya milk
The addition of BaCl2 solution (1 nmol/μL) to a liquid MALDI sample of a soya milk lipid extract enabled the detection of [L+Ba]2+ ions related to many compounds detected as [L+H]+ using a liquid MALDI sample without the addition of BaCl2 (Fig. 7a). As an example, m/z 459.8 was the corresponding [L+Ba]2+ ion for the [L+H]+ ion at m/z 782.6. As a consequence, a single (minimally prepared) biosample can now be used for lipid profiling detecting singly charged lipids as [L+H]+ ions and at the same time achieving structural characterization by exploiting the fragmentation behavior of the doubly charged [L+Ba]2+ ions. Furthermore, the ion mobility setup of the Synapt G2-Si instrument can be used for another type of pseudo-MS3 experiment. TAP measurements enable two stages of CID to be performed (see ESM, Fig. S2 for a diagrammatic representation). First, doubly charged precursors can be selected with a quadrupole, which then undergo CID in the trap cell (22 V). Ion mobility separation of the MS2 products can then be followed by second-stage CID in the transfer cell (40 V), whilst keeping a record of the drift times of the MS2 product ions. The MS3 product ions can then be correlated to their precursor MS2 product ion since all exhibit the same drift time.
Liquid AP-MALDI pseudo-MS3 analysis of lipids from undiluted soya milk using TAP measurements and 1 pmol BaCl2 in the MALDI sample. (a) Positive ion mode MS profile. The inset details the spectral region around the [L+Ba]2+ ions. (b) CID MS/MS spectrum of the precursor ions within the selection window of m/z 459.5±1. (c) Pseudo-MS3 spectrum displaying the product ions that are drift time-aligned with the MS/MS product ion peak at m/z 417.16, which is putatively characterized to consist mainly of C18:2(Δ9, Δ12)
The spectra in Fig. 7b and c show the recorded product ion spectra of a soya milk lipid for the first and second stage CID, respectively. From these CID spectra and the MS precursor ion spectrum, the peak at m/z 459.8 can be assigned to phosphatidylcholine lipids, of which the majority contain isomers of the fatty acid C18:2 (m/z 417.1, [C18:2+Ba–H]+). This is further supported by the product ion peak at m/z 502.34, which corresponds to the loss of [C18:2+Ba–H]+. At both sn positions the acyl chains have to be C18:2 if the m/z value of 782.6 is assigned to the [L+H]+ ion. The TAP product ion peaks of [C18:2+Ba–H]+ shown in Fig. 7c reveal that the majority of the C18:2 isomers are C18:2(Δ9, Δ12). There is some evidence to suggest C18:2(Δ10, Δ12) is also present (m/z 293.01), another isomer of linoleic acid.
In summary, the identity of the most abundant soya milk lipid detected at m/z 782.6 and 459.8 should be PC[18:2(Δ9, Δ12)/18:2(Δ9, Δ12)]. This assignment is in good agreement with the literature stating that the fatty acid content of soybeans is around 50% C18:2(9Z,12Z) [45]. The stereochemistry could not be inferred, and there are likely less abundant isomers of linoleic acid present, which with higher IM resolution might soon be possible to resolve.
Conclusions
Structural information from PCs and SMs can be obtained through the generation of [L+Ba]2+ adduct ions and subsequent CID-MS/MS. The methodology is simple, just adding small amounts of inexpensive BaCl2 (or other divalent metal salts) to liquid MALDI samples. So far, these early results have been obtained on an in-house built AP-MALDI source optimized to produce multiply charged peptide ions. The complexity of lipidomic samples is not to be underestimated, and other ion sources (e.g., ESI) may allow for further improvements and analytical options, particularly through coupling with liquid chromatography (LC). There is also the potential to incorporate previously reported online and offline-coupled LC-MALDI for fractionation of complex samples [46, 47]. Equally important, gaining a further understanding of how the [L+M]2+ ions are produced might enable such optimizations in a faster and more tailored way.
The advantage of using a simple sample additive over instrumental modifications is one many laboratories should find easy to implement. This new method stands as a complementary technique to the aforementioned methods for lipid structure elucidation in the positive ion mode, with apparent specificity to species that contain a permanent positive charge. With regards to stereochemistry, the ongoing improvements to resolving power in the ion mobility-mass spectrometry community offer the chance to separate isobaric lipid isomers and MS2 products more effectively on future IM-MS instruments.
References
Ryumin P, Brown J, Morris M, Cramer R. Investigation and optimization of parameters affecting the multiply charged ion yield in AP-MALDI MS. Methods. 2016;104:11–20.
Koch A, Schnapp A, Soltwisch J, Dreisewerd K. Generation of multiply charged peptides and proteins from glycerol-based matrices using lasers with ultraviolet, visible and near-infrared wavelengths and an atmospheric pressure ion source. Int J Mass Spectrom. 2016;416:61–70.
Cramer R, Pirkl A, Hillenkamp F, Dreisewerd K. Liquid AP-UV-MALDI enables stable ion yields of multiply charged peptide and protein ions for sensitive analysis by mass spectrometry. Angew Chem Int Ed. 2013;2(8):2364–7.
Liang X, Liu J, LeBlanc Y, Covey T, Ptak AC, Brenna JT. Electron transfer dissociation of doubly sodiated glycerophosphocholine lipids. J Am Soc Mass Spectrom. 2007;18(10):1783–178.
James PF, Perugini MA, O'Hair RA. Electron capture dissociation of complexes of diacylglycerophosphocholine and divalent metal ions: competition between charge reduction and radical induced phospholipid fragmentation. J Am Soc Mass Spectrom. 2008;19(7):978–86.
Al-Saad KA, Zabrouskov V, Siems WF, Knowles NR, Hannan RM, Hill HH. Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry of lipids: ionization and prompt fragmentation patterns. Rapid Commun Mass Spectrom. 2003;17(1):87–96.
Schiller J, Suss R, Arnhold J, Fuchs B, Lessig J, Muller M. Matrix-assisted laser desorption and ionization time-of-flight (MALDI-TOF) mass spectrometry in lipid and phospholipid research. Prog Lipid Res. 2004;43(5):449–88.
Pulfer M, Murphy RC. Electrospray mass spectrometry of phospholipids. Mass Spectrom Rev. 2003;22(5):332–64.
Hsu F-F, Bohrer A, Turk J. Formation of lithiated adducts of glycerophosphocholine lipids facilitates their identification by electrospray ionization tandem mass spectrometry. J Am Soc Mass Spectrom. 1998;9(5):516–26.
Kliman M, May JC, McLean JA. Lipid analysis and lipidomics by structurally selective ion mobility-mass spectrometry. Biochim Biophys Acta. 2011;1811(11):935–45.
Sun G, Yang K, Zhao Z, Guan S, Han X, Gross RW. Matrix-assisted laser desorption/ionization time-of-flight mass spectrometric analysis of cellular glycerophospholipids enabled by multiplexed solvent dependent analyte−matrix interactions. Anal Chem. 2008;80(19):7576–85.
Hsu FF, Turk J. Elucidation of the double-bond position of long-chain unsaturated fatty acids by multiple-stage linear ion-trap mass spectrometry with electrospray ionization. J Am Soc Mass Spectrom. 2008;19(11):1673–80.
Trimpin S, Clemmer DE, McEwen CN. Charge-remote fragmentation of lithiated fatty acids on a TOF-TOF instrument using matrix-ionization. J Am Soc Mass Spectrom. 2007;18(11):1967–72.
Hsu FF, Turk J. Structural characterization of unsaturated glycerophospholipids by multiple-stage linear ion-trap mass spectrometry with electrospray ionization. J Am Soc Mass Spectrom. 2008;19(11):1681–91.
Yoo HJ, Haåkansson K. Determination of phospholipid regiochemistry by Ag(I) adduction and tandem mass spectrometry. Anal Chem. 2011;83(4):1275–83.
Ho YP, Huang PC, Deng KH. Metal ion complexes in the structural analysis of phospholipids by electrospray ionization tandem mass spectrometry. Rapid Commun Mass Spectrom. 2003;17(2):114–21.
James PF, Perugini MA, O'Hair RA. Size matters! Fragmentation chemistry of [Cu(L)n]2+ complexes of diacylglycerophosphocholines as a function of coordination number (n = 2–7). Rapid Commun Mass Spectrom. 2007;21(5):757–63.
Griffiths RL-K (2015) Additives for improved analysis of lipids by mass spectrometry: University of Birmingham
Prudent M, Mendez MA, Jana DF, Corminboeuf C, Girault HH. Formation and study of single metal ion–phospholipid complexes in biphasic electrospray ionization mass spectrometry. Metallomics. 2010;2(6):400–6.
Kozlowski RL, Campbell JL, Mitchell TW, Blanksby SJ. Combining liquid chromatography with ozone-induced dissociation for the separation and identification of phosphatidylcholine double bond isomers. Anal Bioanal Chem. 2015;407(17):5053–64.
Poad BL, Green MR, Kirk JM, Tomczyk N, Mitchell TW, Blanksby SJ. High-pressure ozone-induced dissociation for lipid structure elucidation on fast chromatographic timescales. Anal Chem. 2017;89(7):4223–9.
Thomas MC, Mitchell TW, Harman DG, Deeley JM, Nealon JR, Blanksby SJ. Ozone-induced dissociation: elucidation of double bond position within mass-selected lipid ions. Anal Chem. 2008;80(1):303–11.
Vu N, Brown J, Giles K, Zhang Q. Ozone-induced dissociation on a traveling wave high-resolution mass spectrometer for determination of double-bond position in lipids. Rapid Commun Mass Spectrom. 2017;31(17):1415–23.
Pham HT, Maccarone AT, Campbell JL, Mitchell TW, Blanksby SJ. Ozone-induced dissociation of conjugated lipids reveals significant reaction rate enhancements and characteristic odd-electron product ions. J Am Soc Mass Spectrom. 2013;4(2):286–96.
Pham HT, Maccarone AT, Thomas MC, Campbell JL, Mitchell TW, Blanksby SJ. Structural characterization of glycerophospholipids by combinations of ozone- and collision-induced dissociation mass spectrometry: the next step towards "top-down" lipidomics. Analyst. 2014;139(1):204–14.
Marshall DL, Pham HT, Bhujel M, Chin JS, Yew JY, Mori K. Sequential collision- and ozone-induced dissociation enables assignment of relative acyl chain position in triacylglycerols. Anal Chem. 2016;88(5):2685–6892.
Kozlowski RL, Mitchell TW, Blanksby SJ (2015) A rapid ambient ionization-mass spectrometry approach to monitoring the relative abundance of isomeric glycerophospholipids. Sci Rep 9243
Thomas MC, Mitchell TW, Blanksby SJ. On-line ozonolysis methods for the determination of double bond position in unsaturated lipids. In: Armstrong D, editor. Lipidomics: Vol. 1: Methods and Protocols. Totowa: Humana Press; 2009. p. 413–41.
Ma X, Xia Y. Pinpointing double bonds in lipids by Paterno-Buchi reactions and mass spectrometry. Angew Chem Int Ed. 2014;53(10):2592–6.
Ma X, Chong L, Tian R, Shi R, Hu TY, Ouyang Z. Identification and quantitation of lipid C=C location isomers: a shotgun lipidomics approach enabled by photochemical reaction. Proc Natl Acad Sci USA. 2016;113(10):2573–8.
Ren J, Franklin ET, Xia Y. Uncovering structural diversity of unsaturated fatty acyls in cholesteryl esters via photochemical reaction and tandem mass spectrometry. J Am Soc Mass Spectrom. 2017;28(7):1432–41.
Madsen JA, Cullen TW, Trent MS, Brodbelt JS. IR and UV photodissociation as analytical tools for characterizing lipid A structures. Anal Chem. 2011;83(13):5107–13.
O'Brien JP, Needham BD, Henderson JC, Nowicki EM, Trent MS, Brodbelt JS. One hundred ninety three nm ultraviolet photodissociation mass spectrometry for the structural elucidation of lipid A compounds in complex mixtures. Anal Chem. 2014;86(4):2138–45.
Fort KL, Dyachenko A, Potel CM, Corradini E, Marino F, Barendregt A. Implementation of ultraviolet photodissociation on a Benchtop Q Exactive mass spectrometer and its application to phosphoproteomics. Anal Chem. 2016;88(4):2303–010.
Theisen A, Yan B, Brown JM, Morris M, Bellina B, Barran PE. Use of ultraviolet photodissociation coupled with ion mobility mass spectrometry to determine structure and sequence from drift time selected peptides and proteins. Anal Chem. 2016;88(20):9964–71.
Klein DR, Brodbelt JS. Structural characterization of phosphatidylcholines using 193 nm ultraviolet photodissociation mass spectrometry. Anal Chem. 2017;89(3):1516–22.
Ryan E, Nguyen CQN, Shiea C, Reid GE. Detailed structural characterization of sphingolipids via 193 nm ultraviolet photodissociation and ultra high resolution tandem mass spectrometry. J Am Soc Mass Spectrom. 2017;28(7):1406–19.
Li P, Jackson GP. Charge transfer dissociation of phosphocholines: gas-phase ion/ion reactions between helium cations and phospholipid cations. J Mass Spectrom. 2017;52(5):271–82.
Deimler RE, Sander M, Jackson GP. Radical-induced fragmentation of phospholipid cations using metastable atom-activated dissociation mass spectrometry (MAD-MS). Int J Mass Spectrom. 2015;390:178–86.
Baba T, Campbell JL, Le Blanc JCY, Baker PRS. Distinguishing cis and trans isomers in intact complex lipids using electron impact excitation of ions from organics (EIEIO) mass spectrometry. Anal Chem. 2017; https://doi.org/10.1021/acs.analchem.6b04734.
Hsu F-F, Turk J. Electrospray ionization/tandem quadrupole mass spectrometric studies on phosphatidylcholines: the fragmentation processes. J Am Soc Mass Spectrom. 2003;14(4):352–63.
Ait-Belkacem R, Dilillo M, Pellegrini D, Yadav A, de Graaf EL, McDonnell LA. In-source decay and pseudo-MS3 of peptide and protein ions using liquid AP-MALDI. J Am Soc Mass Spectrom. 2016;27(12):2075–9.
Wysocki VH, Ross MM. Charge-remote fragmentation of gas-phase ions: mechanistic and energetic considerations in the dissociation of long-chain functionalized alkanes and alkenes. Int J Mass Spectrom Ion Processes. 1991;104(3):179–211.
Voinov VG, Claeys M. Charge-remote fragmentation characteristics of monounsaturated fatty acids in resonance electron capture: differentiation between cis and trans isomers. 11 Dedicated to Professor Aleksandar Stamatovic on the occasion of his 60th birthday. Int J Mass Spectrom. 2001;205(1):57–64.
Ivanov DS, Lević JD, Sredanović SA. Fatty acid composition of various soybean products. Food Feed Res. 2010;37(2):65–70.
Daniel JM, Laiko VV, Doroshenko VM, Zenobi R. Interfacing liquid chromatography with atmospheric pressure MALDI-MS. Anal Bioanal Chem. 2005;383(6):895–902.
Ryumin P, Brown J, Morris M, Cramer R. Protein identification using a nanoUHPLC-AP-MALDI MS/MS workflow with CID of multiply charged proteolytic peptides. Int J Mass Spectrom. 2017;416:20–8.
Acknowledgements
This research was conducted as part of a studentship funded by Waters Corporation and the Engineering and Physical Sciences Research Council (EPSRC) (DTG grant no. 1498422). Discussions with Dr. Andrew Russell (Department of Chemistry, University of Reading) are gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
All authors have given approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Data supporting the results reported in this paper are openly available from the University of Reading Research Data Archive at: http://dx.doi.org/10.17864/1947.130
Electronic supplementary material
ESM 1
(PDF 424 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Hale, O.J., Cramer, R. Collision-induced dissociation of doubly-charged barium-cationized lipids generated from liquid samples by atmospheric pressure matrix-assisted laser desorption/ionization provides structurally diagnostic product ions. Anal Bioanal Chem 410, 1435–1444 (2018). https://doi.org/10.1007/s00216-017-0788-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-017-0788-6