Abstract
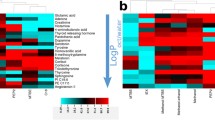
In this study, the efficiency of five peptide-extraction methods—acetonitrile (ACN) precipitation, ultrafiltration, C18 solid-phase extraction (SPE), dispersed SPE with mesoporous carbon CMK-3, and mesoporous silica MCM-41—was quantitatively investigated. With 28 tryptic peptides as target analytes, these methods were evaluated on the basis of recovery and reproducibility by using high-performance liquid chromatography–triple-quad tandem mass spectrometry in selected-reaction-monitoring mode. Because of the distinct extraction mechanisms of the methods, their preferences for extracting peptides of different properties were revealed to be quite different, usually depending on the pI values or hydrophobicity of peptides. When target peptides were spiked in bovine serum albumin (BSA) solution, the extraction efficiency of all the methods except ACN precipitation changed significantly. The binding of BSA with target peptides and nonspecific adsorption on adsorbents were believed to be the ways through which BSA affected the extraction behavior. When spiked in plasma, the performance of all five methods deteriorated substantially, with the number of peptides having recoveries exceeding 70 % being 15 for ACN precipitation, and none for the other methods. Finally, the methods were evaluated in terms of the number of identified peptides for extraction of endogenous plasma peptides. Only ultrafiltration and CMK-3 dispersed SPE performed differently from the quantitative results with target peptides, and the wider distribution of the properties of endogenous peptides was believed to be the main reason.

Distribution of the recoveries of target peptides after treatment by the five peptide-extraction methods






Similar content being viewed by others
Abbreviations
- DSPE:
-
Dispersed solid-phase extraction
- FA:
-
Formic acid
- MW:
-
Molecular weight
- pI:
-
Isoelectric point
- QQQ:
-
Triple quadrupole
References
Anderson NL (2002) The Human Plasma Proteome: History, Character, and Diagnostic Prospects. Mol Cell Proteomics 1(11):845–867
Liotta LA, Ferrari M, Petricoin E (2003) Written in blood. Nature 425(6961):905–905
Petricoin EF, Belluco C, Araujo RP, Liotta LA (2006) The blood peptidome: a higher dimension of information content for cancer biomarker discovery. Nat Rev Cancer 6(12):961–967
Diamandis EP (2006) Peptidomics for cancer diagnosis: present and future. J Proteome Res 5(9):2079–2082
Aebersold R, Mann M (2003) Mass spectrometry-based proteomics. Nature 422(6928):198–207
Petricoin EF, Ardekani AM, Hitt BA, Levine PJ, Fusaro VA, Steinberg SM, Mills GB, Simone C, Fishman DA, Kohn EC, Liotta LA (2002) Use of proteomic patterns in serum to identify ovarian cancer. Lancet 359(9306):572–577
Sadek PC, Carr PW, Bowers LD, Haddad LC (1986) A radiochemical study of irreversible adsorption of proteins on reversed-phase chromatographic packing materials. Anal Biochem 153(2):359–371
Gillette MA, Mani DR, Carr SA (2005) Place of pattern in proteomic biomarker discovery. J Proteome Res 4(4):1143–1154
Chen J, Anderson M, Misek DE, Simeone DM, Lubman DM (2007) Characterization of apolipoprotein and apolipoprotein precursors in pancreatic cancer serum samples via two-dimensional liquid chromatography and mass spectrometry. J Chromatogr A 1162(2):117–125
Chertov O, Biragyn A, Kwak LW, Simpson JT, Boronina T, Hoang VM, Prieto DA, Conrads TP, Veenstra TD, Fisher RJ (2004) Organic solvent extraction of proteins and peptides from serum as an effective sample preparation for detection and identification of biomarkers by mass spectrometry. Proteomics 4(4):1195–1203
Kay R, Barton C, Ratcliffe L, Matharoo-Ball B, Brown P, Roberts J, Teale P, Creaser C (2008) Enrichment of low molecular weight serum proteins using acetonitrile precipitation for mass spectrometry based proteomic analysis. Rapid Commun Mass Spectrom 22(20):3255–3260
Kawashima Y, Fukutomi T, Tomonaga T, Takahashi H, Nomura F, Maeda T, Kodera Y (2010) High-yield peptide-extraction method for the discovery of subnanomolar biomarkers from small serum samples. J Proteome Res 9(4):1694–1705
Hu LH, Ye ML, Zou HF (2009) Recent advances in mass spectrometry-based peptidome analysis. Expert Rev Proteomic 6(4):433–447
Tirumalai RS (2003) Characterization of the Low molecular weight human serum proteome. Mol Cell Proteomics 2(10):1096–1103
Tammen H, Schulte I, Hess R, Menzel C, Kellmann M, Mohring T, Schulz-Knappe P (2005) Peptidomic analysis of human blood specimens: Comparison between plasma specimens and serum by differential peptide display. Proteomics 5(13):3414–3422
Orvisky E, Drake SK, Martin BM, Abdel-Hamid M, Ressom HW, Varghese RS, An Y, Saha D, Hortin GL, Loffredo CA, Goldman R (2006) Enrichment of low molecular weight fraction of serum for MS analysis of peptides associated with hepatocellular carcinoma. Proteomics 6(9):2895–2902
Zheng X, Baker H, Hancock WS (2006) Analysis of the low molecular weight serum peptidome using ultrafiltration and a hybrid ion trap-Fourier transform mass spectrometer. J Chromatogr A 1120(1–2):173–184
Greening DW, Simpson RJ (2010) A centrifugal ultrafiltration strategy for isolating the low-molecular weight (≤25 K) component of human plasma proteome. J Proteomics 73(3):637–648
Aristoteli LP, Molloy MP, Baker MS (2007) Evaluation of endogenous plasma peptide extraction methods for mass spectrometric biomarker discovery. J Proteome Res 6(2):571–581
Johnson KL, Mason CJ, Muddiman DC, Eckel JE (2004) Analysis of the low molecular weight fraction of serum by LC-dual ESI-FT-ICR mass spectrometry: precision of retention time, mass, and ion abundance. Anal Chem 76(17):5097–5103
Koomen JM, Li DH, Xiao LC, Liu TC, Coombes KR, Abbruzzese J, Kobayashi R (2005) Direct tandem mass spectrometry reveals limitations in protein profiling experiments for plasma biomarker discovery. J Proteome Res 4(3):972–981
Mouton-Barbosa E, Roux-Dalvai F, Bouyssie D, Berger F, Schmidt E, Righetti PG, Guerrier L, Boschetti E, Burlet-Schiltz O, Monsarrat B, de Peredo AG (2010) In-depth exploration of cerebrospinal fluid by combining peptide ligand library treatment and label-free protein quantification. Mol Cell Proteomics 9(5):1006–1021
Tian R, Zhang H, Ye M, Jiang X, Hu L, Li X, Bao X, Zou H (2007) Selective Extraction of Peptides from Human Plasma by Highly Ordered Mesoporous Silica Particles for Peptidome Analysis. Angew Chem 119(6):980–983
Qin H, Gao P, Wang F, Zhao L, Zhu J, Wang A, Zhang T, Ra W, Zou H (2011) Highly efficient extraction of serum peptides by ordered mesoporous carbon. Angew Chem Int Ed 50(51):12218–12221
Capriotti AL, Caruso G, Cavaliere C, Piovesana S, Samperi R, Laganà A (2012) Comparison of three different enrichment strategies for serum low molecular weight protein identification using shotgun proteomics approach. Anal Chim Acta 740:58–65
De Bock M, de Seny D, Meuwis M-A, Servais A-C, Minh TQ, Closset J, Chapelle J-P, Louis E, Malaise M, Merville M-P (2010) Comparison of three methods for fractionation and enrichment of low molecular weight proteins for SELDI-TOF-MS differential analysis. Talanta 82(1):245–254
Tucholska M, Scozzaro S, Williams D, Ackloo S, Lock C, Siu KWM, Evans KR, Marshall JG (2007) Endogenous peptides from biophysical and biochemical fractionation of serum analyzed by matrix-assisted laser desorption/ionization and electrospray ionization hybrid quadrupole time-of-flight. Anal Biochem 370(2):228–245
Biosa G, Addis MF, Tanca A, Pisanu S, Roggio T, Uzzau S, Pagnozzi D (2011) Comparison of blood serum peptide enrichment methods by Tricine SDS-PAGE and mass spectrometry. J Proteomics 75(1):93–99
Potier DN, Griffiths JR, Unwin RD, Walker MJ, Carrick E, Willamson AJK, Whetton AD (2012) An assessment of peptide enrichment methods employing mTRAQ quantification approaches. Anal Chem 84(13):5604–5610
Picotti P, Aebersold R, Domon B (2007) The implications of proteolytic background for shotgun proteomics. Mol Cell Proteomics 6(9):1589–1598
Fan R, Huh S, Yan R, Arnold J, Yang P (2008) Gated proton transport in aligned mesoporous silica films. NatMater 7(4):303–307
Hartmann M (2005) Ordered mesoporous materials for bioadsorption and biocatalysis. Chem Mater 17(18):4577–4593
Tenzer S, Docter D, Kuharev J, Musyanovych A, Fetz V, Hecht R, Schlenk F, Fischer D, Kiouptsi K, Reinhardt C, Landfester K, Schild H, Maskos M, Knauer SK, Stauber RH (2013) Rapid formation of plasma protein corona critically affects nanoparticle pathophysiology. Nat Nanotechnol 8(10):772–781
Acknowledgments
The financial support by the National Nature Science Foundation of P. R. China (grant Nos. 21005080, 21275144, 21475131, 91317313, 21235005, 21321064) is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 3.46 MB)
Rights and permissions
About this article
Cite this article
Du, Y., Wu, D., Wu, Q. et al. Quantitative evaluation of peptide-extraction methods by HPLC–triple-quad MS–MS. Anal Bioanal Chem 407, 1595–1605 (2015). https://doi.org/10.1007/s00216-014-8389-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-014-8389-0