Abstract
Numerous empirical scoring schemes are arbitrary, yielding contradictory results for different protein inhibitor families. This study is aimed to validate the predictive potential of alternative nonempirical model of inhibitory activity based on first principle quantum chemical results for new class of small molecules targeting the menin-mixed lineage leukemia (MLL) interaction. Presented nonempirical interaction energy model comprising long-range electrostatic multipole and approximate dispersion terms is universal and computationally affordable, outperforming a number of commonly used empirical scoring functions, especially when combined with analysis of solvation energy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Computational methods used to estimate the inhibitory activity are mostly based on arbitrary empirical scoring functions or force fields that provide rather differing estimates [1, 2]. More accurate quantum chemical predictions are impractical in virtual high-throughput screening of drug candidates due to their significant computational cost, and therefore their applicability to protein–ligand interactions has been rather limited [3, 4]. Therefore, simplified nonempirical models for the description of inhibitory activity are needed. They can be derived from the systematical partitioning of the ab initio computed interaction energy into well-defined contributions, yielding approximate nonempirical yet affordable models of general applicability.
The specific binding of mixed lineage leukemia (MLL) or MLL fusion proteins to menin results in acute MLL leukemias [5] that are considered mostly incurable [6,7,8]. This protein–protein interaction (PPI) has been validated as a therapeutic target in MLL leukemias with both genetic [5] and pharmacological methods [9,10,11,12,13], and inhibition of the menin–MLL interaction seems to be very important for development of targeted therapy against this subtype of aggressive leukemias.
Recently [14], we have demonstrated that menin-MLL inhibitors can be successfully described with nonempirical model based on the long-range components of the interaction energy, namely multipole electrostatic and approximate dispersion. Not only our model was able to correctly rank novel compounds, but it was also revealed that the main force responsible for binding of these compounds to menin was the electrostatic interaction. These previously identified inhibitors (thienopyrimidines), with a lead compound MI-2-2, directly interacted with menin in the MLL binding site with low nanomolar affinity [9, 10]. As it has been subsequently shown, this class of menin-MLL inhibitors was inefficient in vivo [12]. As a result, the inhibitors of menin–MLL interaction were further developed by Grembecka et al., and another class of these inhibitors was introduced. This work aims at the analysis of the novel class of menin-MLL inhibitors described in Ref. [12], which are analogs of the lead compound MI-136. Both classes of menin-MLL inhibitors are compared in Methods Section.
Similar analysis of EphA2-ephrin A1 inhibitors [15], another class of compounds targeting protein–protein interaction, has revealed that nonempirical models based on enthalpic contributions to the binding energy are limited to the sets of inhibitors with similar values of the solvation free energy, \(\varDelta G_{\mathrm{solv}}\). The initial \(\varDelta G_{\mathrm{solv}}\) calculations have shown that our model cannot be applied to the full set of inhibitors, and therefore, we have finally selected only those compounds, whose solvation energy is comparable. The analysis of solvation energy of the analyzed compounds prior to interaction energy calculations could be performed to determine the applicability of our scoring model in given case.
2 Methods
2.1 Preparation of complexes
The analyzed inhibitors are shown in Table 1. The common scaffold encompasses a thieno[2,3-d]pyrimidine, piperidine and indole ring systems with different R1, R2 and R3 substituents on indole. Therefore, to limit the computational cost, only indol with \(\text {{--}CH}_{3}\) group on carbon C5 and R1, R2 and R3 substitution was taken into account. The part of the scaffold not included in the calculations is marked in gray in Table 1.
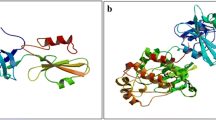
The crystal structures of menin in complex with MI-503 (PDB accession code 4X5Y [12]; 1.59 Å resolution) or MI-136 (PDB accession code 4X5Z [12]; 1.86 Å resolution) were selected as relevant for the analysis of binding of inhibitors listed in Table 1. The main difference between these complexes, as shown in Fig. 1, is related to the positioning of Glu363 and Glu366 residues, since the side chains of both glutamic acid residues need to shift in order to accommodate the MI-503 compound with rather large substitution at R3 position. Thus, all compounds possessing R3 substituents (compounds 18–26) were modeled on the basis of compound 27 (MI-503). The remaining inhibitors (11–17) were modeled on the basis of compound 9 (MI-136).
Both hydrogen and the missing heavy atoms of all compounds considered herein were added with Schrödinger Maestro program [16], and their positions were minimized with OPLS 2005 force field [17]. Building and optimization of the protein hydrogen atoms were performed following the Protein Preparation Wizard [18] protocol, as applied to both protein structures, i.e., 4X5Z and 4X5Y. Determination of the optimal hydrogen bonding was carried out with Maestro-implemented PROPKA [19,20,21,22].
Subsequently, to obtain more reliable positions of amino acid residues surrounding the compounds 11–26, the corresponding complexes were solvated with TIP3 water model [23] and reoptimized in CHARMM program [24] (version c36b1). Both CHARMM General Force Field v. 2b7 [25] and CHARMM22 All-Hydrogen [26,27,28] force field parameters were used. Missing parameters for compounds 11–26 were generated with CGenFF program at http://cgenff.paramchem.org [25, 29,30,31] (interface version 1.0.0). Indole ring and all amino acid residues further than 8Å from each inhibitor were kept frozen throughout 1000 steps of steepest descent minimization followed by conjugate gradient optimization continued until RMS gradient of \(0.01\; {\text {kcal}}\cdot {\text {mol}}^{-1}\cdot \)Å was reached. As a result, an individual menin-inhibitor complex for each analyzed structure was obtained; that is, 18 distinct complexes (including original 4X5Z and 4X5Y structures) were analyzed in what follows.
Amino acid residues representing menin binding site (Fig. 2) were selected based on the distance from inhibitors. Except for Glu363 residue, all the first-shell amino acid residues surrounding the inhibitor fragments were included in the binding site model. Due to flexibility of this particular residue, its solvent exposition and the undertaken optimization protocol, Glu363 was not optimized correctly as the proper orientation toward inhibitors was not obtained in case of each complex. The proposed binding site representation consists of 8 amino acid residues, namely nonpolar Met322, Ala325, Gly326, Trp341, Val367, Val371 residues, polar Tyr323 residue and negatively charged Glu366 residue.
Peptide bonds between the following pairs of residue: Glu366-Val367, Ala325-Gly326 and Met322-Tyr323 were not cut to conduct interaction energy calculations. The remaining residues (Trp341 and Val371) were included separately. Dangling bonds resulting from cutting the amino acid residues out of the protein structure were filled with hydrogen atoms minimized in Maestro [16] using OPLS 2005 force field [17].
2.2 Ab initio binding energy calculations
Binding energy between menin and inhibitors was calculated within the Hybrid Variation-Perturbation Theory (HVPT) [32, 33]. This method allows to analyze the physical nature of interactions within relatively large systems at a reasonable computational cost [34,35,36,37,38,39]. The HVPT scheme of the interaction energy decomposition defines partitioning of the second-order Møller–Plesset binding energy (\(E_{\mathrm{MP2}}\)) into the electrostatic multipole (\(E_{\mathrm{EL,MTP}}^{(10)}\)), penetration (\(E_{\mathrm{EL,PEN}}^{(10)}\)), exchange (\(E_{\mathrm{EX}}^{(10)}\)), delocalization (\(E_{\mathrm{DEL}}^{(R0)}\)) and correlation (\(E_{\mathrm{CORR}}^{(2)}\)) terms, as shown in Eq. (1). The zero value of the second superscript represents uncorrelated interaction energy contributions, and the \(E_{\mathrm{CORR}}^{(2)}\) term denotes the inter- and intramolecular correlation contribution.
\(E_{\mathrm{EL,MTP}}^{(10)}\) term in Eq. (1) represents the electrostatic long-range multipole binding energy calculated from atomic multipole expansion [40]. The total electrostatic interaction energy, \(E_{\mathrm{EL}}^{(10)}\), accounts for both \(E_{\mathrm{EL,MTP}}^{(10)}\) and the short-range penetration term, \(E_{\mathrm{EL,PEN}}^{(10)}\). The first-order repulsive exchange term, \(E_{\mathrm{EX}}^{(10)}\), is obtained as the difference between the first-order Heitler–London energy, \(E^{(10)}\), and the total electrostatic interaction energy, \(E_{\mathrm{EL}}^{(10)}\). The higher order delocalization energy, \(E_{\mathrm{DEL}}^{(R0)}\), represents the classical induction and charge transfer terms. It is calculated as \(E_{\mathrm{DEL}}^{(R0)} = E_{\mathrm{SCF}} - E^{(10)}\), where \(E_{\mathrm{SCF}}\) stands for the counterpoise-corrected self-consistent field (SCF) variational energy. The correlation term, \(E_{\mathrm{CORR}}^{(2)}\), is the difference between the second-order Møller–Plesset interaction energy, \(E_{\mathrm{MP2}}\), and the converged SCF energy, \(E_{\mathrm{SCF}}\): \(E_{\mathrm{CORR}}^{(2)} = E_{\mathrm{MP2}} - E_{\mathrm{SCF}}\). It accounts for dispersion and exchange–dispersion interactions as well as intramolecular correlation contribution. More details can be found in our previous papers (e.g., see Refs. [14, 41]). All of the subsequent corrections to \(E_{\mathrm{MP2}}\) interaction energy [Eq. (1)] could be categorized into long- and short-range interactions varying with the intermolecular distance, R, as \(R^{-n}\) and \(e^{-\gamma R}\), respectively.
The interaction energy between each residue (Trp341 and Val371 monomers or Glu366-Val367, Ala325-Gly326 and Met322-Tyr323 dimers) and each inhibitor fragment was calculated with a modified version [33] of the GAMESS program [42] using the 6-31G(d) [43, 44] basis set. The counterpoise correction was applied to avoid basis set superposition errors [45]. Multipole electrostatic energy terms were calculated using the Cumulative Atomic Multipole Moments (CAMM) approach [46] (implemented in GAMESS) based on the correlated wave function, with the multipole expansion truncated at the \(R^{-4}\) term.
Since the dispersion contribution within the HVPT decomposition scheme is included in the computationally demanding \(E_{\mathrm{CORR}}^{(2)}\) correlation energy term, the recently derived atom-atom potential function, \(E_{\mathrm{Das}}\)[47, 48] was chosen to describe this type of interactions in our approximate nonempirical model for the description of inhibitory activity (\(E_{\mathrm{EL,MTP}}^{(10)}+E_{Das}\)). \(E_{\mathrm{Das}}\) is a far more affordable alternative to the costly computations of \(E_{\mathrm{CORR}}^{(2)}\), as it scales with the square number of atoms \( O(A^{2})\), in contrast to ab initio \(E_{\mathrm{CORR}}^{(2)}\) calculations scaling at least with the fifth power of number of orbitals, \( O(N^{5})\).
2.3 Solvation energy calculation
\(\varDelta G_{\mathrm{solv}}\) was calculated in Gaussian09 with Polarizable Continuum Model (PCM) using the integral equation formalism variant (IEFPCM) [49,50,51]. MP2/6-31G(d) level of theory was applied. ExternalIteration [52, 53], DoVacuum and SMD [54] options were employed in the course of single point calculations performed for geometries obtained at the preparation stage (Sect. 2.1).
2.4 Empirical scoring
To serve as a comparison to the nonempirical approach, empirical scoring was performed with a variety of scoring functions including ChemPLP from PLANTS docking program [55] and LigScore, LS [56], Picewise Linear Potential, PLP [57, 58], Jain [59], Potential of Mean Force, PMF [60, 61] and Ludi [62, 63] implemented in Discovery Studio 2017 [64]. In all these cases, the scoring was performed with a 8 Å radius sphere centered on the ligand. In addition, scoring with AutoDock Vina was performed. It involved PyMOL [65] and PyMOL AutoDock/Vina plugin [66] for preparation of the receptors and inhibitors and was carried out with 10 Å cubic grid centered on the ligand. All these calculations involved only scoring of the compounds’ poses without any re-docking. In contrast to the nonempirical models applied herein, all mentioned docking and scoring programs required the usage of full protein structures.
The performance of a particular scoring method, either nonempirical or empirical, was evaluated by means of the Pearson’s correlation coefficient calculated with respect to the experimentally determined inhibitory activity [12]. The results were also compared in terms of the success rate of prediction, \(N_{\mathrm{pred}}\) (predictability), which refers to the percentage of concordant pairs among all possible pairs of complexes within a given set (nonparametric statistics). A concordant pair denotes two complexes with computationally evaluated relative stability being of the same sign as the relative experimental binding affinity (see Ref. [67] for details).
3 Results and discussion
3.1 Comparison of menin-MLL inhibitor classes
In this work, a novel class of menin-MLL inhibitors, developed on the basis of previously examined thienopyrimidines [14], is analyzed. A significant modification of the scaffold of these compounds involved introducing a cyano-indole ring, which is connected to the thienopyrimidine moiety with a piperidine linker. On the basis of the lead compound MI-136 (Table 1), the substitutions on the indole ring were explored [12]. Comparison of the most potent inhibitors representing each class is given in Fig. 3.
a Inhibitor MI-503 (compound 27) representing a distinct class of inhibitors analyzed herein. The part of the scaffold not included in the nonempirical calculations is marked in gray; b Inhibitor MI-2-2 representing the thienopyrimidine class of inhibitors analyzed in Ref. [14]
As already discussed, the inhibitors in this study were truncated to the substituted indole part of the scaffold. It seemed vital for the efficiency of ab initio calculations, since the size of novel inhibitors nearly doubled compared to the previously studied class of compounds.
Importantly, the binding sites of both classes of menin-MLL inhibitors overlap only partially (Fig. 4). The common part involves the thienopyrimidine and piperidine moieties, which are omitted in current study to decrease the computational cost. The substituted indole fragment considered herein is positioned beyond the binding site analyzed previously [14]. In fact, after exclusion of incorrectly modeled Glu363 residue (marked in yellow in Fig. 4) from the model examined, only Tyr323 residue is present in both binding site representations. Moreover, in contrast to previous model [14], in which charged or polar amino acid residues were abundant, this time the binding site representation comprises mostly hydrophobic amino acid residues (Val367, Val371, Met322, Ala325, Trp341) and only one charged Glu366 residue is present. This suggests a different nature of interactions between these two inhibitor classes and their respective menin binding sites.
MI-2-2 and MI-503 (blue and red, respectively) in their corresponding menin binding sites; **Glu363 residue shown in yellow was excluded from the model analyzed herein since it was not positioned correctly. It was present in the previous study [14] (the corresponding Glu363 conformation is given in blue); *Tyr323 is the only amino acid residue present in both studies
3.2 Solvation energy of inhibitors
As an initial assessment of the applicability of nonempirical models to the chosen inhibitors’ set, solvation free energy, \(\varDelta G_{\mathrm{solv}}\), was computed (Fig. 5). Since theoretical models applied herein are restricted to the enthalpic contribution to the binding free energy, their performance is limited to the analysis of compounds with similar solvation energy. According to our recent results obtained for protein-protein inhibitors targeting EphA2-ephrin A1 interaction [15], large differences in the ligand solvation energy limit the applicability of the nonempirical models tested herein. On the other hand, roughly comparable values of solvation free energy of inhibitors, as demonstrated by relatively low standard deviation of \(\varDelta G_{\mathrm{solv}}\) values, appear to ensure that the nonempirical models operating on the basis of enthalpic contribution provide reasonable inhibitory activity estimates. Therefore, monitoring the differences in solvation free energy within the analyzed group of inhibitors prior to the interaction energy calculations could indicate whether the nonempirical model could be applied to the entire set of compounds. Nevertheless, this estimate should be treated with caution as the results provided by various solvation models do not necessarily give exact solutions [68,69,70].
Standard deviation of the solvation free energy values demonstrated in Fig. 5 is equal to \(4.3\; {\text {kcal}}\cdot {\text {mol}}^{-1}\), indicating the presence of differences that might affect the performance of the nonempirical models based on the interaction energy values. However, for a reduced set of 11 inhibitors obtained by omitting the compounds with the extreme differences in \(\varDelta G_{\mathrm{solv}}\) values (compounds 9, 13, 14, 15, 18, 24 and 25; see Fig. 5), the value of standard deviation of solvation free energy is decreased to \(2.1\; {\text {kcal}}\cdot {\text {mol}}^{-1}\). Accordingly, nonempirical models employed in what follows are more likely to be predictive when applied to the reduced set of inhibitors. Admittedly, such a selection of ligands based on \(\varDelta G_{\mathrm{solv}}\) differences is rather an arbitrary approach, where inhibitors could be excluded iteratively to reach even lower standard deviation values and, probably, better predictive abilities of the nonempirical approach. However, more extensive elimination of the compounds to lower the value of the \(\varDelta G_{\mathrm{solv}}\) standard deviation does not necessarily result in a significant improvement of the performance of the nonempirical models of inhibitory activity, as determined by monitoring of the changes in the correlation coefficient between the given interaction energy model and the experimental binding potency (see below). While the latter will be discussed in the subsequent section in more details, it should be noted here that further limiting of the size of the reduced set of inhibitors (i.e., excluding more that 7 compounds marked in Fig. 5) results in only minor improvement of the correlation coefficient values, despite the significant drop in the \(\varDelta G_{\mathrm{solv}}\) standard deviation. Presumably, the nonempirical models employed herein are characterized by some inherent level of precision, arising from the structural accuracy of the complexes undergoing analysis. Due to certain approximations and/or assumptions undertaken while modeling the geometries of the receptor-ligand complexes, it would be unreasonable to expect that the computational model could provide perfect agreement with the experimental data. Our results seem to demonstrate that sensitivity of the selection of the inhibitors based on the \(\varDelta G_{\mathrm{solv}}\) differences is limited, as some variability of the latter is still allowed without the loss of the performance of the nonempirical approach.
3.3 Interaction energy derived with nonempirical methods
Nonempirical results of interaction energy computed for all inhibitors are given in Table 2. To verify the performance of the particular level of theory in terms of predicting the experimental binding potency, experimentally established inhibitory activity [12] was employed to determine the Pearson correlation coefficient (R) and percentage of successful predictions (\(N_{\mathrm{pred}}\)). As assumed initially, the correlation between the experimental and computational results was observed only for the reduced (\(R_{\mathrm{(r)}}\)) set of compounds featuring the solvation free energy comparable throughout that group. This finding is further supported by the increased values of the success rate of prediction derived for the reduced set of compounds, \(N_{\mathrm{pred (r)}}\) (Table 2).
As indicated by significant correlation coefficient (\(R=-\,0.78\)) and the highest \(N_\mathrm{pred}\) value (81.8%), obtained for both the \(E_\mathrm{SCF}\) and the most robust MP2 level of theory, the binding of this class of menin-MLL inhibitors appears to be governed rather by delocalization and dispersion contributions than by the electrostatic interactions only. It can be seen in Table 2 that \(E_\mathrm{EL}^{(10)}\) term within the reduced set of inhibitors is associated with \(R_\mathrm{(r)}\) value of \(-0.61\), suggesting a weaker relationship between binding energy at this particular level of theory and the experimental binding potency. This is in contrast to the thienopyrimidine class of menin-MLL inhibitors studied before [14], in which case it was the electrostatic interaction that was dominant. The different physical nature underlying the binding within these two sets of menin-MLL inhibitors seems to stem from the differences in the binding site composition (mostly nonpolar amino acid residues versus charged or polar binding site model in a previous work [14], see Fig. 3). Despite the significant contribution of the correlation energy that manifests itself in the large difference between the \(E_\mathrm{MP2}\) and \(E_\mathrm{SCF}\) levels of theory (Table 2), the dispersive interactions do not seem to influence the relative binding strength; that is, their contribution is essentially similar within the reduced set of menin-MLL inhibitors.
Another difference concerning the molecular basis of binding of the two inhibitor classes is that including the repulsive exchange term, \(E_\mathrm{EX}^{(10)}\), actually improves the inhibitory activity predictions, as the correlation coefficient associated with \(E^{(10)}\) energy exceeds the value characterizing the \(E_\mathrm{EL}^{(10)}\) term (\(R_\mathrm{(r)}\) values for \(E_\mathrm{EL}^{(10)}\) and \(E^{(10)}\) levels of theory are equal to \(-\,0.68\) and \(-\,0.80\), respectively; see Table 2). This is in sharp contrast with the results obtained for thienopyrimidine menin-MLL inhibitors that featured the complete loss of the predictive abilities upon including the exchange term [14]. In that previous case, the short-range components of the interaction energy appeared to be exaggerated, probably due to the presence of the excessively short intermolecular contacts. While the positive values of \(E^{(10)}\) interaction energy observed herein (Table 2) might also suggest suboptimal positioning of the inhibitors relative to the binding site residues, including the short-range exchange term seems to be important for proper modeling of the inhibitory activity.
The approximate model of interaction energy, \(E_\mathrm{EL,MTP}^{(10)}+E_{Das}\), comprising multipole electrostatic and approximate dispersion, is described by a rather moderate correlation coefficient (\(R=-\,0.62\)) or the percentage of correct predictions (\(N_\mathrm{pred}=70.9\%\)), which are similar to the values obtained for the electrostatic interaction energy. Nevertheless, due to low computational cost, comparable to that of empirical scoring functions, and the lack of arbitrary empirical parameters, this particular model could still provide qualitative estimates of the binding potency.
3.4 Empirical scoring
The results of scoring with empirical functions are compared in Table 3 with the \(E_\mathrm{SCF}\) and \(E_\mathrm{EL,MTP}^{(10)}+E_{Das}\) energy values. It is clear that for the full set of inhibitors, all of the employed scoring functions fail to properly reproduce the experimental ranking of the inhibitory activity, either in terms of the correlation (\(R_\mathrm{(f)}\)) or predictability (\(N_\mathrm{pred (f)}\)). In the reduced set of ligands encompassing the inhibitors with similar solvation energy, only three empirical scoring functions appear to possess the predictive capabilities, as the \(R_\mathrm{(r)}\) values of PLP2, PLP1 and AutoDock Vina exceed \(-\,0.6\) (Table 3). The set of inhibitors studied herein appears to be particularly challenging in terms of obtaining the proper ligand ranking, as most of the empirical scoring functions tested currently were capable of providing at least the qualitative estimates of the inhibitory activity of thienopyrimidine class of menin-MLL inhibitors [14]. Despite the difficulties in predicting the binding potency of the current series of menin-MLL inhibitors with empirical scoring functions, the ab initio self-consistent field energy, \(E_\mathrm{SCF}\), yields the best estimate of the inhibitory activity from all of the proposed models. The performance of the \(E_\mathrm{EL,MTP}^{(10)}+E_{Das}\) model is comparable to the best predictions obtained with the empirical scoring functions. Noticeably, the computational cost associated with the nonempirical \(E_\mathrm{EL,MTP}^{(10)}+E_{Das}\)model is as low as that characterizing the empirical scoring approaches without the requirement of development or employment of any arbitrary parametrization.
4 Conclusions
In this study, the nonempirical model of inhibitory activity is validated for the new complexes of menin-MLL inhibitors, which were modeled on the basis of previously examined thienopyrimidines [14]. The modification accounted for an introduction of a cyano-indole ring, connected to the thienopyrimidine moiety with a piperidine linker. The substitutions on indole proposed in Ref. [12] were analyzed herein. A systematic series of quantum chemical calculations of the interaction energy at the consecutive levels of theory and subsequent comparison of the binding energy with the experimental activity of the menin-MLL inhibitors has revealed a significant correlation for the most robust \(E_\mathrm{MP2}\) interaction energy. The same performance was obtained by more affordable \(E_\mathrm{SCF}\) binding energy. Satisfactory correlation was achieved for the electrostatic level of theory (\(E_\mathrm{EL}^{(10)}\)). These findings suggest that in case of the analyzed class of menin-MLL inhibitors both delocalization and dispersion interactions are important for correct representation of the binding potency. This is in contrast to our findings from previous study on thienopyrimidine class of menin-MLL inhibitors [14], in which the electrostatic interactions were vital for the description of the relative binding strength. Our results reflect the nature of different binding site representations: Previously studied model of menin binding site [14] could be assumed polar (with numerous charged and polar amino acid residues), and the model presented herein was rather nonpolar (abundant hydrophobic residues).
In order to achieve significant correlation between the theoretical models and experimental inhibitory activity, the solvation free energy should be comparable across the series of analyzed compounds. After limiting the analyzed set of inhibitors to those exhibiting similar \(\varDelta G_\mathrm{solv}\) values, the correlation between the inhibitory activity and the interaction energy on each level of theory has been obtained. Importantly, reasonable estimates of the binding potency within the reduced set of ligands were achieved only with a number of empirical functions, while most of the empirical approaches tested herein failed to reproduce the experimental inhibitory activity. Noticeably, that currently available solvation energy estimates are not always reliable [68, 69] and further progress in this area could facilitate our methodology.
Although the predictability of \(E_\mathrm{EL,MTP}^{(10)}+E_{Das}\) model is moderate in the case of this particular set of menin-MLL inhibitors, it still outperforms most of the empirical scoring functions used for comparison. Therefore, it could be useful in structure-based development of novel inhibitors, including PPI inhibitors, for which it is challenging to provide satisfactory predictions of ligand binding affinity with currently available scoring functions.
References
Leach AR, Shoichet BK, Peishoff CE (2006) Prediction of protein-ligand interactions. docking and scoring: successes and gaps. J Med Chem 49:5851–5855
Plewczyński D, Łaźniewski M, Augustyniak R, Ginalski K (2011) Can we trust docking results? Evaluation of seven commonly used programs on PDBbind database. J Comput Chem 32:742–755
Yilmazer ND, Korth M (2013) Comparison of molecular mechanics, semi-empirical quantum mechanical and density functional theory methods for scoring protein-ligand interactions. J Phys Chem B 117:8075–8084
Ryde U, Söderhjelm P (2016) Ligand-binding affinity estimates supported by quantum-mechanical methods. Chem Rev 116:5520–5566
Yokoyama A, Somervaille TCP, Smith KS, Rozenblatt-Rosen O, Meyerson M, Cleary ML (2005) The menin tumor suppressor protein is an essential oncogenic cofactor for MLL-associated leukemogenesis. Cell 123(2):207–218
Marschalek R (2011) Mechanisms of leukemogenesis by mll fusion proteins. Br J Haematol 152(2):141–154
Slany RK (2005) When epigenetics kills: MLL fusion proteins in leukemia. Hematol Oncol 23(1):1–9
Tomizawa D, Koh K, Sato T, Kinukawa N, Morimoto A, Isoyama K, Kosaka Y, Oda T, Oda M, Hayashi Y, Eguchi M, Horibe K, Nakahata T, Mizutani S, Ishii E (2007) Outcome of risk-based therapy for infant acute lymphoblastic leukemia with or without an MLL gene rearrangement, with emphasis on late effects: a final report of two consecutive studies, MLL96 and MLL98, of the japan infant leukemia study group. Leukemia 21(11):2258–2263
Grembecka J, He S, Shi A, Purohit T, Muntean AG, Sorenson RJ, Showalter HD, Murai MJ, Belcher AM, Hartley T, Hess JL, Cierpicki T (2012) Menin-MLL inhibitors reverse oncogenic activity of MLL fusion proteins in leukemia. Nat Chem Biol 8(3):277–284
Shi A, Murai M, S H, Lund G, Hartley T, Purhoit T, Reddy G, Chruszcz M, Grembecka J, Cierpicki T (2012) Structural insights into inhibition of the bivalent menin-MLL interaction by small molecules in leukemia. Blood 120:4461–4469
He S, Senter TJ, Pollock J, Han C, Upadhyay SK, Purohit T, Gogliotti RD, Lindsley CW, Cierpicki T, Stauffer SR, Grembecka J (2014) High-affinity small-molecule inhibitors of the menin-mixed lineage leukemia (MLL) interaction closely mimic a natural protein-protein interaction. J Med Chem 57(4):1543–1556
Borkin D, He S, Miao H, Kempinska K, Pollock J, Chase J, Purohit T, Malik B, Zhao T, Wang J, Wen B, Zong H, Jones M, Danet-Desnoyers G, Guzman ML, Talpaz M, Bixby DL, Sun D, Hess JL, Muntean AG, Maillard I, Cierpicki T, Grembecka J (2015) Pharmacologic inhibition of the menin-MLL interaction blocks progression of MLL leukemia in vivo. Cancer Cell 27(4):589–602
Borkin D, Pollock J, Kempinska K, Purohit T, Li X, Wen B, Zhao T, Miao H, Shukla S, He M, Sun D, Cierpicki T, Grembecka J (2016) Property focused structure-based optimization of small molecule inhibitors of the protein-protein interaction between menin and mixed lineage leukemia (MLL). J Med Chem 59(3):892–913
Jedwabny W, Kłossowski S, Purohit T, Cierpicki T, Grembecka J, Dyguda-Kazimierowicz E (2017a) Theoretical models of inhibitory activity for inhibitors of protein-protein interactions: targeting menin-mixed lineage leukemia with small molecules. Med Chem Commun 8:2216–2227
Jedwabny W, Lodola A, Dyguda-Kazimierowicz E, Jedwabny W, Lodola A, Dyguda-Kazimierowicz E (2018) Theoretical model of EphA2-Ephrin A1 inhibition. Molecules 23(7):1688
Schrödinger LLC, New York, NY (2012) Schrödinger Suite 2012, Maestro version 9.3
Banks JL, Beard HS, Cao Y, Cho AE, Damm W, Farid R, Felts AK, Halgren TA, Mainz DT, Maple JR, Murphy R, Philipp DM, Repasky MP, Zhang LY, Berne BJ, Friesner RA, Gallicchio E, Levy RM (2005) Integrated modeling program, applied chemical theory (IMPACT). J Comput Chem 26:1752–1780
Schrödinger LLC, New York, NY (2015) Schrödinger Suite 2015-2, Protein Preparation Wizard, Epik version 3.2, Impact version 6.7, Prime version 4.0
Li H, Robertson AD, Jensen JH (2005) Very fast empirical prediction and interpretation of protein pKa values. Proteins 61:704–721
Bas DC, Rogers DM, Jensen JH (2008) Very fast prediction and rationalization of pKa values for protein-ligand complexes. Proteins 73:765–783
Olsson MHM, Søndergard CR, Rostkowski M, Jensen JH (2011) PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J Chem Theory Comput 7:525–537
Søndergard CR, Olsson MHM, Rostkowski M, Jensen JH (2011) Improved treatment of ligands and coupling effects in empirical calculation and rationalization of pKa values. J Chem Theory Comput 7:2284–2295
Mahoney MW, Jorgensen WL (2000) A five-site model for liquid water and the reproduction of the density anomaly by rigid, nonpolarizable potential functions. J Chem Phys 112(20):8910–8922
Brooks BR, Bruccoleri RE, Olafson BD, States DJ, Swaminathan S, Karplus M (1983) CHARMM–a program for macromolecular energy, minimization, and dynamics calculations. J Comput Chem 4(2):187–217
Vanommeslaeghe K, Hatcher E, Acharya C, Kundu S, Zhong S, Shim J, Darian E, Guvench O, Lopes P, Vorobyov I, MacKerell AD Jr (2010) CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem 31(4):671–690
MacKerell AD, Bashford D, Bellott M, Dunbrack RL, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph-McCarthy D, Kuchnir L, Kuczera K, Lau FTK, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, Reiher WE, Roux B, Schlenkrich M, Smith JC, Stote R, Straub J, Watanabe M, Wiorkiewicz-Kuczera J, Yin D, Karplus M (1998) All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B 102(18):3586–3616
Mackerell AD, Feig M, Brooks CL (2004) Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J Comput Chem 25(11):1400–1415
Stote RH, Karplus M (1995) Zinc-binding in proteins and solution–a simple but accurate nonbonded representation. Proteins-Struct Funct Genet 23(1):12–31
Vanommeslaeghe K, MacKerell AD Jr (2012) Automation of the CHARMM General Force Field (CGenFF) I: bond perception and atom typing. J Chem Inf Model 52(12):3144–3154
Vanommeslaeghe K, Raman EP, MacKerell AD Jr (2012) Automation of the CHARMM General Force Field (CGenFF) II: assignment of bonded parameters and partial atomic charges. J Chem Inf Model 52(12):3155–3168
Yu W, He X, Vanommeslaeghe K, MacKerell AD Jr (2012) Extension of the CHARMM general force field to sulfonyl-containing compounds and its utility in biomolecular simulations. J Comput Chem 33(31):2451–2468
Sokalski WA, Roszak S, Pecul K (1988) An efficient procedure for decomposition of the SCF interaction energy into components with reduced basis set dependence. Chem Phys Lett 153:153–159
Góra RW, Sokalski WA, Leszczyński J, Pett V (2005) The nature of interactions in the ionic crystal of 3-pentenenitrile, 2-nitro-5-oxo, ion(-1) sodium. J Phys Chem B 109:2027–2033
Grembecka J, Kędzierski P, Sokalski WA (1999) Non-empirical analysis of the nature of the inhibitor-active site interactions in leucine aminopeptidase. Chem Phys Lett 313:385–392
Kędzierski P, Sokalski WA, Krauss M (2000) Nonempirical analysis of nature of catalytic effects in ribonuclease a active site. J Comput Chem 21:432–445
Sokalski WA, Kędzierski P, Grembecka J (2001) Ab initio study of physical nature of interactions between enzyme active site fragments in vacuo. Phys Chem Chem Phys 3:657–663
Dyguda E, Grembecka J, Sokalski WA, Leszczyński J (2005) Origins of the activity of PAL and LAP enzyme inhibitors: toward ab initio binding affinity prediction. J Am Chem Soc 127:1658–1659
Szefczyk B, Mulholland AJ, Ranaghan KE, Sokalski WA (2004) Differential transition-state stabilization in enzyme catalysis: quantum chemical analysis of interactions in the chorismate mutase reaction and prediction of the optimal catalytic field. J Am Chem Soc 126:16148–16159
Langner KM, Kędzierski P, Sokalski WA, Leszczyński J (2006) Physical nature of ethidium and proflavine interactions with nucleic acid bases in the interaction plane. J Phys Chem B 110:9720–9727
Sokalski WA, Sawaryn A (1987) Correlated molecular and cumulative atomic multipole moments. J Chem Phys 87:526–534
Jedwabny W, Panecka-Hofman J, Dyguda-Kazimierowicz E, Wade RC, Sokalski WA (2017b) Application of a simple quantum chemical approach to ligand fragment scoring for Trypanosoma brucei pteridine reductase 1 inhibition. J Comput Aided Mol Des 31:715–728
Schmidt MW, Baldridge KK, Boatz JA, Elbert ST, Gordon MS, Jensen JH, Koseki S, Matsunaga N, Nguyen KA, Su SJ, Windus TL, Dupuis M, Montgomery JA (1993) General atomic and molecular electronic structure system. J Comput Chem 14(11):1347–1363
Hariharan PC, Pople JA (1973) The influence of polarization functions on molecular-orbital hydrogenation energies. Theoret Chimica Acta 28(3):213–222
Francl MM, Pietro WJ, Hehre WJ, Binkley JS, DeFrees DJ, Pople JA, Gordon MS (1982) Self-consistent molecular orbital methods. XXIII. A polarization-type basis set for second row elements. J Chem Phys 77(7):3654–3665
Boys S, Bernardi F (1970) The calculation of small molecular interactions by the differences of separate total energies. Some procedures with reduced errors. Mol Phys 19:553–566
Sokalski WA, Poirier RA (1983) Cumulative atomic multipole representation of the molecular charge distribution and its basis set dependence. Chem Phys Lett 98(1):86–92
Pernal K, Podeszwa R, Patkowski K, Szalewicz K (2009) Dispersionless density functional theory. Phys Rev Lett 103:263201–263204
Podeszwa R, Pernal K, Patkowski K, Szalewicz K (2010) Extension of the Hartree-Fock plus dispersion method by first-order correlation effects. J Phys Chem Lett 1(2):550–555, 27
Tomasi J, Mennucci B, Cances E (1999) The IEF version of the PCM solvation method: an overview of a new method addressed to study molecular solutes at the QM ab initio level. J Mol Struct Theochem 464(1–3):211–226
Pascualahuir JL, Silla E, Tunon I (1994) GEPOL–an improved description of molecular-surfaces.3. A new algorithm for the computation of a solvent-excluding surface. J Comput Chem 15(10):1127–1138
Tomasi J, Mennucci B, Cammi R (2005) Quantum mechanical continuum solvation models. Chem Rev 105(8):2999–3093
Improta R, Scalmani G, Frisch MJ, Barone V (2007) Toward effective and reliable fluorescence energies in solution by a new state specific polarizable continuum model time dependent density functional theory approach. J Chem Phys 127(7):074504
Improta R, Barone V, Scalmani G, Frisch MJ (2006) A state-specific polarizable continuum model time dependent density functional theory method for excited state calculations in solution. J Chem Phys 125(5):054103
Marenich AV, Cramer CJ, Truhlar DG (2009) Universal solvation model based on solute electron density and on a continuum model of the solvent defined by the bulk dielectric constant and atomic surface tensions. J Phys Chem B 113(18):6378–6396
Korb O, Stutzle T, Exner TE (2009) Empirical scoring functions for advanced protein-ligand docking with PLANTS. J Chem Inf Model 49(1):84–96
Krammer A, Kirchhoff PD, Jiang X, Venkatachalam CM, Waldman M (2005) LigScore: a novel scoring function for predicting binding affinities. J Mol Gr Model 23:395–407
Gehlhaar DK, Verkhivker GM, Rejto PA, Sherman CJ, Fogel DB, Fogel LJ, Freer ST (1995) Molecular recognition of the inhibitor AG-1343 by HIV-1 protease: conformationally flexible docking by evolutionary programming. Chem Biol 2:317–324
Gehlhaar DK, Bouzida D, Rejto PA (1999) Rational drug design: novel methodology and practical applications, vol 719. American Chemical Society, Washington, DC
Jain AN (1996) Scoring noncovalent protein-ligand interactions: a continuous differentiable function tuned to compute binding affinities. J Comput Aid Mol Des 10:427–440
Muegge I, Martin YC (1999) A general and fast scoring function for protein-ligand interactions: a simplified potential approach. J Med Chem 42:791–804
Muegge I (2006) PMF scoring revisited. J Med Chem 49:5895–5902
Böhm HJ (1994) The development of a simple empirical scoring function to estimate the binding constant for a protein-ligand complex of known three-dimensional structure. J Comput Aid Mol Des 8:243–256
Böhm HJ (1998) Prediction of binding constants of protein ligands: a fast method for the prioritization of hits obtained from the de novo design or 3D database search programs. J Comput Aid Mol Des 12:309–323
BIOVIA (2017) Discovery Studio modeling environment. Release 2017. Dassault Systèmes, San Diego
(2012) PyMOL(TM) Molecular Graphics System, Version 1.5.0.1. Schrödinger, New York
Seeliger D, de Groot BL (2010) Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J Comput Aid Mol Des 24:417–422
Langner KM, Beker W, Sokalski WA (2012) Robust predictive power of the electrostatic term at shortened intermolecular distances. J Phys Chem Lett 3:2785–2789
Yokogawa D (2018) Toward accurate solvation free energy calculation with the reference interaction site model self-consistent field: introduction of a new bridge function. J Chem Theory Comput. https://doi.org/10.1021/acs.jctc.8b00314
Ho JM, Klamt A, Coote ML (2010) Comment on the correct use of continuum solvent models. J Phys Chem A 114(51):13442–13444
Ángyán JG (1992) Common theoretical framework for quantum chemical solvent effect theor ies. J Math Chem 10(1):93–137
Acknowledgements
We thank Professor W. Andrzej Sokalski for thoughtful reading and suggestions. The project was financed by the National Science Centre Poland, Grant Number 2016/21/N/ST4/00516. Calculations were carried out using resources provided by Wrocław Centre for Networking and Supercomputing, Poznań Supercomputing and Networking Center, Warsaw Interdisciplinary Centre for Mathematical and Computational Modelling, and Biovia Polish Academic Country Wide license. We also thank Wrocław University of Technology and University of Michigan for support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published as part of the special collection of articles In Memoriam of János Ángyán.
This paper is dedicated to the memory of János G. Ángyán.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Jedwabny, W., Cierpicki, T., Grembecka, J. et al. Validation of approximate nonempirical scoring model for menin-mixed lineage leukemia inhibitors. Theor Chem Acc 137, 148 (2018). https://doi.org/10.1007/s00214-018-2350-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00214-018-2350-9