Abstract
Hemoglobins are ubiquitous proteins found in bacteria, plants, and animals with diverse functions other than the classical transport/storage of oxygen. Different functions are expected to correspond to substantially different structures, such as the hexa- and penta-coordination of the iron atom. It is now widely believed that pentacoordinate hemoglobins evolved from the hexacoordinate ones, both in plants and in animals. Since plant hemoglobins evolved more recently than in animals, they represent a simpler and thus useful system to investigate protein sequence/structure features that specifically supported, guided by molecular evolution, the capacity for oxygen transport. In the present work, we selected a fully hexacoordinate globin, AHb2 from Arabidopsis thaliana and the pentacoordinate oxygen-transporting LegHb from yellow lupin, that share a high degree of sequence identity. Our aim is to identify the structural determinants for oxygen transport by analyzing the structural/dynamical differences of a hexacoordinate and a pentacoordinate globin using all-atom molecular dynamics simulations. Using comparative MD simulations, we were able to go beyond the simple sequence alignment, pointing out important differences between these two hemoglobins especially at the level of the CD region, whose dynamics was found, in turn, to be strongly correlated with that of the distal region.








Similar content being viewed by others
References
Hunt PW, Watts RA, Trevaskis B, Llewelyn DJ, Burnell J et al (2001) Expression and evolution of functionally distinct haemoglobin genes in plants. Plant Mol Biol 47:677–692
de Sanctis D, Pesce A, Nardini M, Bolognesi M, Bocedi A et al (2004) Structure-function relationships in the growing hexa-coordinate hemoglobin sub-family. TBMB 56:643–651. doi:10.1080/15216540500059640
Burmester T, Weich B, Reinhardt S, Hankeln T (2000) A vertebrate globin expressed in the brain. Nature 407:520–523. doi:10.1038/35035093
Vinogradov SN, Hoogewijs D, Bailly X, Mizuguchi K, Dewilde S et al (2007) A model of globin evolution. Gene 398:132–142. doi:10.1016/j.gene.2007.02.041
Trevaskis B, Watts RA, Andersson CR, Llewellyn DJ, Hargrove MS et al (1997) Two hemoglobin genes in Arabidopsis thaliana: the evolutionary origins of leghemoglobins. Proc Natl Acad Sci USA 94:12230–12234
Hoy JA, Robinson H, Trent JT, Kakar S, Smagghe BJ et al (2007) Plant hemoglobins: a molecular fossil record for the evolution of oxygen transport. J Mol Biol 371:168–179. doi:10.1016/j.jmb.2007.05.029
Koder RL, Anderson JLR, Solomon LA, Reddy KS, Moser CC et al (2009) Design and engineering of an O(2) transport protein. Nature 458:305–309. doi:10.1038/nature07841
Martí MA, Capece L, Bikiel DE, Falcone B, Estrin DA (2007) Oxygen affinity controlled by dynamical distal conformations: the soybean leghemoglobin and the Paramecium caudatum hemoglobin cases. Proteins 68:480–487. doi:10.1002/prot.21454
Hargrove MS, Brucker EA, Stec B, Sarath G, Arredondo-Peter R et al (2000) Crystal structure of a nonsymbiotic plant hemoglobin. Structure 8:1005–1014
McCammon JA, Gelin BR, Karplus M (1977) Dynamics of folded proteins. Nature 267:585–590
Frauenfelder H, Chen G, Berendzen J, Fenimore PW, Jansson H et al (2009) A unified model of protein dynamics. Proc Natl Acad Sci 106:5129–5134. doi:10.1073/pnas.0900336106
Nishihara Y, Kato S, Hayashi S (2010) Protein collective motions coupled to ligand migration in myoglobin. Biophys J 98:1649–1657. doi:10.1016/j.bpj.2009.12.4318
Bruno S, Faggiano S, Spyrakis F, Mozzarelli A, Cacciatori E et al (2007) Different roles of protein dynamics and ligand migration in non-symbiotic hemoglobins AHb1 and AHb2 from Arabidopsis thaliana. Gene 398:224–233. doi:10.1016/j.gene.2007.02.042
Harutyunyan E, Safonova T, Kuranova I, Popov A, Teplyakov A et al (1995) The structure of deoxy-leghemoglobin and oxy-leghemoglobin from lupin. J Mol Biol 251:104–115
Fiser A, Sali A (2003) Modeller: generation and refinement of homology-based protein structure models. Meth Enzymol 374:461–491. doi:10.1016/S0076-6879(03)74020-8
Benkert P, Künzli M, Schwede T (2009) QMEAN server for protein model quality estimation. Nucleic Acids Res 37:W510–W514. doi:10.1093/nar/gkp322
Morrone JA, Zhou R, Berne BJ (2010) Molecular dynamics with multiple time scales: how to avoid pitfalls. J Chem Theory Comput 6:1798–1804. doi:10.1021/ct100054k
Lindorff-Larsen K, Piana S, Palmo K, Maragakis P, Klepeis JL et al (2010) Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins 78:1950–1958. doi:10.1002/prot.22711
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926. doi:10.1063/1.445869
Henry ER, Levitt M, Eaton WA (1985) Molecular dynamics simulation of photodissociation of carbon monoxide from hemoglobin. Proc Natl Acad Sci USA 82:2034–2038
Kleywegt GJ, Jones TA (1994) Detection, delineation, measurement and display of cavities in macromolecular structures. Acta Crystal D Biol Crystal 50:178–185. doi:10.1107/S0907444993011333
Scorciapino MA, Robertazzi A, Casu M, Ruggerone P, Ceccarelli M (2009) Breathing motions of a respiratory protein revealed by molecular dynamics simulations. J Am Chem Soc 131:11825–11832. doi:10.1021/ja9028473
Mezei M (2010) Simulaid: a simulation facilitator and analysis program. J Comput Chem 31:2658–2668. doi:10.1002/jcc.21551
Aleksiev T, Potestio R, Pontiggia F, Cozzini S, Micheletti C (2009) PiSQRD: a web server for decomposing proteins into quasi-rigid dynamical domains. Bioinformatics 25:2743–2744. doi:10.1093/bioinformatics/btp512
Bruno S, Faggiano S, Spyrakis F, Mozzarelli A, Abbruzzetti S et al (2007) The reactivity with CO of AHb1 and AHb2 from Arabidopsis thaliana is controlled by the distal HisE7 and internal hydrophobic cavities. J Am Chem Soc 129:2880–2889. doi:10.1021/ja066638d
Dantsker D, Samuni U, Friedman J, Agmon N (2005) A hierarchy of functionally important relaxations within myoglobin based on solvent effects, mutations and kinetic model. Bba-Proteins Proteom 1749:234–251. doi:10.1016/j.bbapap.2005.04.002
Schotte F, Lim M, Jackson TA, Smirnov AV, Soman J et al (2003) Watching a protein as it functions with 150-ps time-resolved X-ray crystallography. Science 300:1944–1947. doi:10.1126/science.1078797
Brunori M, Bourgeois D, Vallone B (2004) The structural dynamics of myoglobin. J Struct Biol 147:223–234. doi:10.1016/j.jsb.2004.04.008
Scorciapino MA, Robertazzi A, Casu M, Ruggerone P, Ceccarelli M (2010) Heme proteins: the role of solvent in the dynamics of gates and portals. J Am Chem Soc 132:5156–5163. doi:10.1021/ja909822d
Springer BA, Egeberg KD, Sligar SG, Rohlfs RJ, Mathews AJ et al (1989) Discrimination between oxygen and carbon monoxide and inhibition of autooxidation by myoglobin. Site-directed mutagenesis of the distal histidine. J Biol Chem 264:3057–3060
Huang X, Boxer SG (1994) Discovery of new ligand binding pathways in myoglobin by random mutagenesis. Nat Struct Biol 1:226–229
Olson J (2007) Ligand pathways in myoglobin: a review of trp cavity mutations. TBMB
Uchida T, Ishimori K, Morishima I (1997) The effects of heme pocket hydrophobicity on the ligand binding dynamics in myoglobin as studied with leucine 29 mutants. J Biol Chem 272:30108–30114
Cohen J, Arkhipov A, Braun R, Schulten K (2006) Imaging the migration pathways for O2, CO, NO, and Xe inside myoglobin. Biophys J 91:1844–1857. doi:10.1529/biophysj.106.085746
Gibson QH, Regan R, Elber R, Olson JS, Carver TE (1992) Distal pocket residues affect picosecond ligand recombination in myoglobin. An experimental and molecular dynamics study of position 29 mutants. J Biol Chem 267:22022–22034
Nienhaus K, Dominici P, Astegno A, Abbruzzetti S, Viappiani C et al (2010) Ligand migration and binding in nonsymbiotic hemoglobins of Arabidopsis thaliana. Biochemistry 49:7448–7458. doi:10.1021/bi100768g
Whitaker TL, Berry MB, Ho EL, Hargrove MS, Phillips GN et al (1995) The D-helix in myoglobin and in the beta subunit of hemoglobin is required for the retention of heme. Biochemistry 34:8221–8226
Elber R (2010) Ligand diffusion in globins: simulations versus experiment. Curr Opin Struct Biol 20:162–167. doi:10.1016/j.sbi.2010.01.002
Garrocho-Villegas V, Gopalasubramaniam SK, Arredondo-Peter R (2007) Plant hemoglobins: what we know six decades after their discovery. Gene 398:78–85. doi:10.1016/j.gene.2007.01.035
Acknowledgments
We acknowledge the CYBERSAR and CINECA award, the latter under the ISCRA initiative for the availability of high-performance computing resources and support. CW thanks the ERASMUS program for supporting her visit to Department of Physics, University of Cagliari.
Author information
Authors and Affiliations
Corresponding author
Additional information
Dedicated to Professor Akira Imamura on the occasion of his 77th birthday and published as part of the Imamura Festschrift Issue.
Electronic supplementary material
Below is the link to the electronic supplementary material.
214_2011_1041_MOESM1_ESM.tiff
Figure S1 Secondary structure analysis of the AHb2’s D region. The black horizontal lines represent the structures where the corresponding residue has been found to be involved in a helical conformation. Supplementary material 1 (TIFF 15752 kb)
214_2011_1041_MOESM2_ESM.tiff
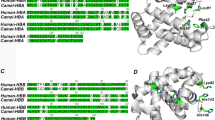
Figure S2 Residues RMSF calculated from Ca coordinates. Residue numbers correspond to AHb2 amino acids sequence, while LegHb has been aligned with respect to the latter on the basis of Fig. 1. RMSF have been calculated for the second half of the MD simulation in order to eliminate AHb2 relaxation contribution. The two data sets have been normalized. Supplementary material 2 (TIFF 15344 kb)
Rights and permissions
About this article
Cite this article
Scorciapino, M.A., Wallon, C. & Ceccarelli, M. MD simulations of plant hemoglobins: the hexa- to penta-coordinate structural transition. Theor Chem Acc 130, 1105–1114 (2011). https://doi.org/10.1007/s00214-011-1041-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00214-011-1041-6