Abstract
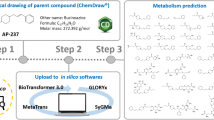
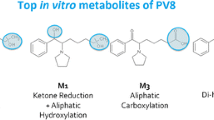
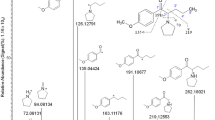
Synthetic cathinones constitute a family of new psychoactive substances, the consumption of which is increasingly worldwide. A lack of metabolic knowledge limits the detection of these compounds in cases of intoxication. Here, we used an innovative cross-disciplinary approach to study the metabolism of the newly emerging cathinone chloro-alpha-pyrrolidinovalerophenone (4-Cl-PVP). Three complementary approaches (in silico, in vitro, and in vivo) were used to identify putative 4-Cl-PVP metabolites that could be used as additional consumption markers. The in silico approach used predictive software packages. Molecular networking was used as an innovative bioinformatics approach for re-processing high-resolution tandem mass spectrometry data acquired with both in vitro and in vivo samples. In vitro experiments were performed by incubating 4-Cl-PVP (20 µM) for four different durations with a metabolically competent human hepatic cell model (differentiated HepaRG cells). In vivo samples (blood and urine) were obtained from a patient known to have consumed 4-Cl-PVP. The in silico software predicted 17 putative metabolites, and molecular networking identified 10 metabolites in vitro. On admission to the intensive care unit, the patient’s plasma and urine 4-Cl-PVP concentrations were, respectively, 34.4 and 1018.6 µg/L. An in vivo analysis identified the presence of five additional glucuronoconjugated 4-Cl-PVP derivatives in the urine. Our combination of a cross-disciplinary approach with molecular networking enabled the detection of 15 4-Cl-PVP metabolites, 10 of them had not previously been reported in the literature. Two metabolites appeared to be particular relevant candidate as 4-Cl-PVP consumption markers in cases of intoxication: hydroxy-4-Cl-PVP (m/z 282.1254) and dihydroxy-4-Cl-PVP (m/z 298.1204).





Similar content being viewed by others
References
Allard S, Allard P-M, Morel I, Gicquel T (2019) Application of a molecular networking approach for clinical and forensic toxicology exemplified in three cases involving 3-MeO-PCP, doxylamine, and chlormequat. Drug Test Anal 11:669–677. https://doi.org/10.1002/dta.2550
Allard S, Le Daré B, Allard P-M et al (2020) Comparative molecular networking analysis of a Rauwolfia plant powder and biological matrices in a fatal ingestion case. Forensic Toxicol 38:447–454. https://doi.org/10.1007/s11419-020-00531-0
Allibe N, Richeval C, Phanithavong M et al (2018) Fatality involving ocfentanil documented by identification of metabolites. Drug Test Anal 10:995–1000. https://doi.org/10.1002/dta.2326
Ameline A, Greney H, Monassier L et al (2019a) Metabolites to parent 3-MeO-PCP ratio in human urine collected in two fatal cases. J Anal Toxicol 43:321–324. https://doi.org/10.1093/jat/bky097
Ameline A, Richeval C, Gaulier J-M et al (2019b) Detection of the designer benzodiazepine flunitrazolam in urine and preliminary data on its metabolism. Drug Test Anal 11:223–229. https://doi.org/10.1002/dta.2480
Aninat C, Piton A, Glaise D et al (2006) Expression of cytochromes P450, conjugating enzymes and nuclear receptors in human hepatoma HepaRG cells. Drug Metab Dispos Biol Fate Chem 34:75–83. https://doi.org/10.1124/dmd.105.006759
Batisse A, Eiden C, Deheul S et al (2022) Chemsex practice in France: an update in addictovigilance data. Fundam Clin Pharmacol 36:397–404. https://doi.org/10.1111/fcp.12725
Carlier J, Diao X, Giorgetti R et al (2020) Pyrrolidinyl synthetic cathinones α-PHP and 4F-α-PVP metabolite profiling using human hepatocyte incubations. Int J Mol Sci 22:230. https://doi.org/10.3390/ijms22010230
Carlier J, Berardinelli D, Montanari E et al (2022) 3F-α-pyrrolydinovalerophenone (3F-α-PVP) in vitro human metabolism: multiple in silico predictions to assist in LC-HRMS/MS analysis and targeted/untargeted data mining. J Chromatogr B Analyt Technol Biomed Life Sci 1193:123162. https://doi.org/10.1016/j.jchromb.2022.123162
de Bruyn KC, Šícho M, Mazzolari A, Kirchmair J (2021) GLORYx: prediction of the metabolites resulting from phase 1 and phase 2 biotransformations of xenobiotics. Chem Res Toxicol 34:286–299. https://doi.org/10.1021/acs.chemrestox.0c00224
Di Trana A, Brunetti P, Giorgetti R et al (2021) In silico prediction, LC-HRMS/MS analysis, and targeted/untargeted data-mining workflow for the profiling of phenylfentanyl in vitro metabolites. Talanta 235:122740. https://doi.org/10.1016/j.talanta.2021.122740
Diao X, Huestis MA (2019) New synthetic cannabinoids metabolism and strategies to best identify optimal marker metabolites. Front Chem 7:109. https://doi.org/10.3389/fchem.2019.00109
Du Q-S, Huang R-B, Chou K-C (2008) Recent advances in QSAR and their applications in predicting the activities of chemical molecules, peptides and proteins for drug design. Curr Protein Pept Sci 9:248–259. https://doi.org/10.2174/138920308784534005
Dührkop K, Fleischauer M, Ludwig M et al (2019) SIRIUS 4: a rapid tool for turning tandem mass spectra into metabolite structure information. Nat Methods 16:299–302. https://doi.org/10.1038/s41592-019-0344-8
European Monitoring Centre for Drugs and Drug Addiction (2022) European drug report 2022: trends and developments. Publications Office, Luxembourg
Ferron P-J, Le Daré B, Bronsard J et al (2021) Molecular networking for drug toxicities studies: the case of hydroxychloroquine in COVID-19 patients. Int J Mol Sci 23:82. https://doi.org/10.3390/ijms23010082
Gertrudes JC, Maltarollo VG, Silva RA et al (2012) machine learning techniques and drug design. Curr Med Chem 19:4289–4297. https://doi.org/10.2174/092986712802884259
Gicquel T, Pelletier R, Richeval C et al (2021a) Metabolite elucidation of 2-fluoro-deschloroketamine (2F-DCK) using molecular networking across three complementary in vitro and in vivo models. Drug Test Anal. https://doi.org/10.1002/dta.3162
Gicquel T, Richeval C, Mesli V et al (2021b) Fatal intoxication related to two new arylcyclohexylamine derivatives (2F-DCK and 3-MeO-PCE). Forensic Sci Int 324:110852. https://doi.org/10.1016/j.forsciint.2021.110852
Hugbart C, Verres Y, Le Daré B et al (2020) Non-oxidative ethanol metabolism in human hepatic cells in vitro: Involvement of uridine diphospho-glucuronosyltransferase 1A9 in ethylglucuronide production. Toxicol in Vitro 66:104842. https://doi.org/10.1016/j.tiv.2020.104842
Kazmi SR, Jun R, Yu M-S et al (2019) In silico approaches and tools for the prediction of drug metabolism and fate: a review. Comput Biol Med 106:54–64. https://doi.org/10.1016/j.compbiomed.2019.01.008
Kelly JP (2011) Cathinone derivatives: a review of their chemistry, pharmacology and toxicology. Drug Test Anal 3:439–453. https://doi.org/10.1002/dta.313
Klingberg J, Keen B, Cawley A et al (2022) Developments in high-resolution mass spectrometric analyses of new psychoactive substances. Arch Toxicol 96:949–967. https://doi.org/10.1007/s00204-022-03224-2
La Maida N, Di Trana A, Giorgetti R et al (2021) A review of synthetic cathinone-related fatalities from 2017 to 2020. Ther Drug Monit 43:52–68. https://doi.org/10.1097/FTD.0000000000000808
Le Daré B, Allard S, Bouvet R et al (2020a) A case of fatal acebutolol poisoning: an illustration of the potential of molecular networking. Int J Legal Med 134:251–256. https://doi.org/10.1007/s00414-019-02062-9
Le Daré B, Ferron P-J, Allard P-M et al (2020b) New insights into quetiapine metabolism using molecular networking. Sci Rep 10:19921. https://doi.org/10.1038/s41598-020-77106-x
Le Daré B, Ferron P-J, Couette A et al (2021) In vivo and in vitro α-amanitin metabolism studies using molecular networking. Toxicol Lett 346:1–6. https://doi.org/10.1016/j.toxlet.2021.04.006
Lopes BT, Caldeira MJ, Gaspar H, Antunes AMM (2021) Metabolic profile of four selected cathinones in microsome incubations: identification of phase I and II metabolites by liquid chromatography high resolution mass spectrometry. Front Chem 8:609251. https://doi.org/10.3389/fchem.2020.609251
Majchrzak M, Celiński R, Kuś P et al (2018) The newest cathinone derivatives as designer drugs: an analytical and toxicological review. Forensic Toxicol 36:33–50. https://doi.org/10.1007/s11419-017-0385-6
Manier SK, Richter LHJ, Schäper J et al (2018) Different in vitro and in vivo tools for elucidating the human metabolism of alpha-cathinone-derived drugs of abuse. Drug Test Anal. https://doi.org/10.1002/dta.2355
Miotto K, Striebel J, Cho AK, Wang C (2013) Clinical and pharmacological aspects of bath salt use: a review of the literature and case reports. Drug Alcohol Depend 132:1–12. https://doi.org/10.1016/j.drugalcdep.2013.06.016
NIDA (2020) Synthetic cathinones (“bath salts”) drugfacts. In: Natl. Inst. Drug Abuse. https://nida.nih.gov/publications/drugfacts/synthetic-cathinones-bath-salts. Accessed 25 Jul 2022
Pelletier R, Le Daré B, Grandin L et al (2021) New psychoactive substance cocktail in an intensive care intoxication case elucidated by molecular networking. Clin Toxicol. https://doi.org/10.1080/15563650.2021.1931693
Quesnot N, Bucher S, Gade C et al (2018) Production of chlorzoxazone glucuronides via cytochrome P4502E1 dependent and independent pathways in human hepatocytes. Arch Toxicol 92:3077–3091. https://doi.org/10.1007/s00204-018-2300-2
Richeval C, Gicquel T, Hugbart C et al (2017) In vitro characterization of NPS metabolites produced by human liver microsomes and the HepaRG cell line using liquid chromatographyhigh resolution mass spectrometry (LC-HRMS) analysis: application to furanyl fentanyl. Curr Pharm Biotechnol 18:806–814. https://doi.org/10.2174/1389201018666171122124401
Schymanski EL, Jeon J, Gulde R et al (2014) Identifying small molecules via high resolution mass spectrometry: communicating confidence. Environ Sci Technol 48:2097–2098. https://doi.org/10.1021/es5002105
Seidl B, Schuhmacher R, Bueschl C (2022) CPExtract, a software tool for the automated tracer-based pathway specific screening of secondary metabolites in LC-HRMS data. Anal Chem 94:3543–3552. https://doi.org/10.1021/acs.analchem.1c04530
Tyzack JD, Kirchmair J (2019) Computational methods and tools to predict cytochrome P450 metabolism for drug discovery. Chem Biol Drug Des 93:377–386. https://doi.org/10.1111/cbdd.13445
Wagmann L, Frankenfeld F, Park YM et al (2020) How to study the metabolism of new psychoactive substances for the purpose of toxicological screenings-a follow-up study comparing pooled human liver S9, HepaRG cells, and zebrafish larvae. Front Chem 8:539. https://doi.org/10.3389/fchem.2020.00539
Wishart DS, Tian S, Allen D et al (2022) BioTransformer 3.0-a web server for accurately predicting metabolic transformation products. Nucleic Acids Res. https://doi.org/10.1093/nar/gkac313
Acknowledgements
This project was support by financial allowance “Défi Scientifique” Université Rennes 1. The authors thank David Fraser PhD (Biotech Communication SARL, Ploudalmezeau, France) for copy-editing assistance and Bernard Fromenty (NuMeCan Institute) for his helpful contribution.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Ethical approval
The study was conducted according to the guidelines of the Declaration of Helsinki, and approved by the Ethics Committee of University Hospital of Rennes.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pelletier, R., Le Daré, B., Ferron, PJ. et al. Use of innovative, cross-disciplinary in vitro, in silico and in vivo approaches to characterize the metabolism of chloro-alpha-pyrrolidinovalerophenone (4-Cl-PVP). Arch Toxicol 97, 671–683 (2023). https://doi.org/10.1007/s00204-022-03427-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-022-03427-7