Abstract
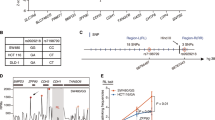
GWAS-identified 10q22.3 loci with lead SNP rs704017 are significantly associated with CRC risk in both Asian and European populations. However, the functional mechanism of this region is unclear. In this study, we performed a fine-mapping analysis to identify the causal SNPs. To identify potential functional SNPs in linkage disequilibrium with the lead SNP, we searched for the potential target genes using a Hi-C database and an RNA interfering-based on-chip approach. The results indicated that rs12263636 (r2 = 0.41) showed the highest potential to be functional. It resided in a region with enhancer markers and a topologically associating domain. We found that RPS24 was the only gene that significantly promoted the proliferation rate of CRC cells and might have promoter–enhancer interaction with rs12263636. Dual-luciferase reporter assays confirmed that the risk alleles of two variants (rs3740253 and rs7071351) in RPS24 promoter could increase the expression of luciferase. Case control study consisting of 1134 cases and 2039 health controls confirmed that both the two variants were associated with risk of CRC (rs3740253: P = 0.0079, OR = 1.15, 95% CI 1.04–1.28; rs7071351: P = 0.0085, OR = 1.15, 95% CI 1.04–1.28). And plasmid containing mutant haplotypes containing all the three mutations (rs12263636 or rs3740253 and rs7071351) could most significantly increase luciferase expression, compared with any haplotype of the three mutations. The study explained the functional mechanism for the 10q22.3 loci and provided new insights into the prevention and treatment of CRC.





Similar content being viewed by others
References
Badhai J, Frojmark AS, Davey JE, Schuster J, Dahl N (2009) Ribosomal protein S19 and S24 insufficiency cause distinct cell cycle defects in Diamond-Blackfan anemia. Biochim Biophys Acta 1792(10):1036–1042. https://doi.org/10.1016/j.bbadis.2009.08.002
Carithers LJ, Ardlie K, Barcus M et al (2015) A novel approach to high-quality postmortem tissue procurement: the GTEx project. Biopreserv Biobank 13(5):311–319. https://doi.org/10.1089/bio.2015.0032
Chang J, Tian J, Yang Y et al (2018) A rare missense variant in TCF7L2 associates with colorectal cancer risk by interacting with a GWAS-identified regulatory variant in the MYC enhancer. Cancer Res 78(17):5164–5172. https://doi.org/10.1158/0008-5472.CAN-18-0910
Chen W, Zheng R, Baade PD et al (2016) Cancer statistics in China, 2015. CA 66(2):115–132. https://doi.org/10.3322/caac.21338(A Cancer Journal for Clinicians)
Chen XF, Zhu DL, Yang M et al (2018) An osteoporosis risk SNP at 1p36.12 acts as an allele-specific enhancer to modulate LINC00339 expression via long-range loop formation. Am J Hum Genet 102(5):776–793. https://doi.org/10.1016/j.ajhg.2018.03.001
Choesmel V, Fribourg S, Aguissa-Toure AH et al (2008) Mutation of ribosomal protein RPS24 in Diamond-Blackfan anemia results in a ribosome biogenesis disorder. Hum Mol Genet 17(9):1253–1263. https://doi.org/10.1093/hmg/ddn015
Gong J, Tian J, Lou J et al (2018) A polymorphic MYC response element in KBTBD11 influences colorectal cancer risk, especially in interaction with an MYC-regulated SNP rs6983267. Ann Oncol 29(3):632–639. https://doi.org/10.1093/annonc/mdx789
Gundert M, Edelmann D, Benner A et al (2019) Genome-wide DNA methylation analysis reveals a prognostic classifier for non-metastatic colorectal cancer (ProMCol classifier). Gut 68(1):101–110. https://doi.org/10.1136/gutjnl-2017-314711
Inoue E, Watanabe Y, Egawa J et al (2015) Rare heterozygous truncating variations and risk of autism spectrum disorder: whole-exome sequencing of a multiplex family and follow-up study in a Japanese population. Psychiatry Clin Neurosci 69(8):472–476. https://doi.org/10.1111/pcn.12274
Jia WH, Zhang B, Matsuo K et al (2013) Genome-wide association analyses in East Asians identify new susceptibility loci for colorectal cancer. Nat Genet 45(2):191–196. https://doi.org/10.1038/ng.2505
Karaderi T, Drong AW, Lindgren CM (2015) Insights into the genetic susceptibility to type 2 diabetes from genome-wide association studies of obesity-related traits. Curr Diabetes Rep. https://doi.org/10.1007/s11892-015-0648-8
Li J, Chang J, Tian J et al (2018) A rare variant P507L in TPP1 interrupts TPP1-TIN2 interaction, influences telomere length, and confers colorectal cancer risk in Chinese population. Cancer Epidemiol Biomarkers Prev 27(9):1029–1035. https://doi.org/10.1158/1055-9965.EPI-18-0099(A Publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology)
Patel RS, Ye S (2011) Genetic determinants of coronary heart disease: new discoveries and insights from genome-wide association studies. Heart 97(18):1463–1473. https://doi.org/10.1136/hrt.2010.219675
Rao SSP, Huntley MH, Durand NC et al (2015) A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 162(3):687–688. https://doi.org/10.1016/j.cell.2015.07.024
Rao SSP, Huang SC, St Hilaire BG et al (2017) Cohesin loss eliminates all loop domains. Cell 171(2):305. https://doi.org/10.1016/j.cell.2017.09.026
Ron G, Globerson Y, Moran D, Kaplan T (2017) Promoter-enhancer interactions identified from Hi-C data using probabilistic models and hierarchical topological domains. Nat Commun. https://doi.org/10.1038/S41467-017-023863
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA 68(1):7–30. https://doi.org/10.3322/caac.21442(A Cancer Journal for Clinicians)
Tian J, Chang J, Gong J et al (2019) Systematic functional interrogation of genes in GWAS loci identified ATF1 as a key driver in colorectal cancer modulated by a promoter-enhancer interaction. Am J Hum Genet 105(1):29–47. https://doi.org/10.1016/j.ajhg.2019.05.004
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. CA 65(2):87–108. https://doi.org/10.3322/caac.21262(A Cancer Journal for Clinicians)
Visser M, Kayser M, Palstra RJ (2012) HERC2 rs12913832 modulates human pigmentation by attenuating chromatin-loop formation between a long-range enhancer and the OCA2 promoter. Genome Res 22(3):446–455. https://doi.org/10.1101/gr.128652.111
Wang Z, Chatterjee N (2017) Increasing mapping precision of genome-wide association studies: to genotype and impute, sequence, or both? Genome Biol 18(1):118. https://doi.org/10.1186/s13059-017-1255-6
Wang Y, Sui JK, Li X et al (2015) RPS24 knockdown inhibits colorectal cancer cell migration and proliferation in vitro. Gene 571(2):286–291. https://doi.org/10.1016/j.gene.2015.06.084
Zeng C, Matsuda K, Jia WH et al (2016) Identification of susceptibility loci and genes for colorectal cancer risk. Gastroenterology 150(7):1633–1645. https://doi.org/10.1053/j.gastro.2016.02.076
Zhang B, Jia WH, Matsuda K et al (2014a) Large-scale genetic study in East Asians identifies six new loci associated with colorectal cancer risk. Nat Genet 46(6):533–542. https://doi.org/10.1038/ng.2985
Zhang B, Jia WH, Matsuo K et al (2014b) Genome-wide association study identifies a new SMAD7 risk variant associated with colorectal cancer risk in East Asians. Int J Cancer 135(4):948–955. https://doi.org/10.1002/ijc.28733
Zhu Z, Zhang F, Hu H et al (2016) Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet 48(5):481–487. https://doi.org/10.1038/ng.3538
Zou D, Lou J, Ke J et al (2018) Integrative expression quantitative trait locus-based analysis of colorectal cancer identified a functional polymorphism regulating SLC22A5 expression. Eur J Cancer 93:1–9. https://doi.org/10.1016/j.ejca.2018.01.065
Acknowledgements
We gratefully acknowledge the members of the Miao lab for the suggestions and contributions to this work. This work was supported by the National Natural Science Foundation of China [81502875 to J.C.]; the National Key Research and Development Plan Program [2016YFC1302702, 2016YFC1302703 to X.M.]; National Program for Support of Top-notch Young Professionals, National Natural Science Foundation of China [81171878, 81222038 to X.M.]; the National Science Fund for Distinguished Young Scholars of China and the Fok Ying Tung Foundation for Young Teachers in the Higher Education Institutions of China [131038 to X.M.].
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zou, D., Zhang, H., Ke, J. et al. Three functional variants were identified to affect RPS24 expression and significantly associated with risk of colorectal cancer. Arch Toxicol 94, 295–303 (2020). https://doi.org/10.1007/s00204-019-02600-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-019-02600-9