Abstract
A Gram-positive staining, aerobic, endospore-forming bacterial strain, isolated from the rhizosphere of Zea mays was studied for its detailed taxonomic allocation. Based on the 16S rRNA gene sequence similarity comparisons, strain JJ-42 T was shown to be a member of the genus Paenibacillus, most closely related to the type strain of Paenibacillus pectinilyticus (98.8%). The 16S rRNA gene sequence similarity to all other Paenibacillus species was below 98.5%. The pairwise average nucleotide identity (ANI) and digital DNA−DNA hybridization (dDDH) values of the JJ-42 T genome assembly against publicly available Paenibacillus type strain genomes were below 92% and 47%, respectively. The quinone system of strain JJ-42 T consisted exclusively of menaquinone MK-7. The polar lipid profile consisted of the major components diphosphatidylglycerol, phosphatidylethanolamine, phosphatidylglycerol, three aminophospholipids (APL), and one unidentified lipid. The major fatty acids were iso- and anteiso-branched with the major compound anteiso C15:0. Physiological and biochemical characteristics allowed a further phenotypic differentiation of strain JJ-42 T from the most closely related species. Thus, JJ-42 T represents a novel species of the genus Paenibacillus, for which the name Paenibacillus allorhizoplanae sp. nov. is proposed, with JJ-42 T (= LMG 32089 T = CCM 9085 T = DSM 111786 T = CIP 111891 T) as the type strain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Paenibacillus, initially proposed by Ash et al. (1993) is now accommodating more than 280 species (https://lpsn.dsmz.de/genus/paenibacillus) isolated from many different sources. It is obvious that species of Paenibacillus have been isolated often as endophytes (Carro et al. 2013; Lai et al. 2015; Kittiwongwattana and Thawai 2015; Gao et al. 2015) and from other plant-associated environments as the rhizosphere (Elo et al. 2001; Ma et al. 2007; Kim et al. 2009a, b; Hong et al. 2009; Beneduzi et al. 2010; Zhang et al. 2013; 2015; Wang et al. 2014; Son et al. 2014; Han et al. 2015; Kämpfer et al. 2016, 2017a,b, 2021), seeds (Liu et al. 2015) or the phyllosphere (Rivas et al. 2005, 2006). In general, endospore-forming bacilli, including members of the genus Paenibacillus, are of particular interest for their capacity to promote plant growth and form endospores (Grady et al. 2016). The purpose of this study was to analyze strain JJ-42 T in detail for its taxonomic allocation and for the presence of genes associated with plant−growth promotion.
Materials and methods
Isolation and culture conditions
Field-grown maize plants (Zea mays) grown in Dunbar, Nebraska were manually uprooted a few weeks after planting. Roots, approximately 15 cm in length, were separated from the surrounding soil by vigorous shaking such that only the most tightly adhering soil remained. Bacteria were collected from the root surface by immersing the root in sterile water followed by plating dilutions on Nutrient Agar (Sigma-Aldrich). Strain JJ-42 T was initially isolated at Auburn University in this manner. Subsequent cultivation of JJ-42 T was performed on tryptone soy agar, TSA (Oxoid) at 28 °C for 24 h.
Molecular characterization
16S rRNA gene phylogeny
For a first phylogenetic placement the 16S rRNA gene of strain JJ-42 T was PCR amplified with primer system Eub9f (5´-GAGTTTGATCMTGGCTCAG-3´) and Eub1492R (5´-ACGGYTACCTTGTTACGACTT-3´) (Lane 1991) and sequenced with the Sanger dideoxy sequencing technology using primers Eub9f and E786F (5´-GATTAGATACCCTGGTAG-3´), respectively, as described previously (Schauss et al. 2015). The sequence was manually processed and corrected based on the electropherograms using MEGA11 (Tamura et al. 2021).
A first phylogenetic assignment was performed by BLAST analysis against the EzBioCloud 16S rRNA gene sequence database, including all type strain 16S rRNA gene sequences and BLASTN BLAST + 2.13.0 (reduced to sequences of type material) of the NCBI (https://blast.ncbi.nlm.nih.gov/). The 16S rRNA gene sequences of strain JJ-42 T and nonincluded-related type strains were subsequently added to LTP_2020 database of the “All-Species Living Tree'' Project (LTP; Yarza et al. 2008) using ARB release 5.2 (Ludwig et al. 2004). The updated database version LTP_04_2021 (released in September 2021) was used. The imported 16S rRNA gene sequences were aligned in the alignment explorer using the pt server generated for the respective database. The alignment was checked manually before subsequent analyses. The sequences were added to the database tree tree_Ltp_all_04_21 containing all type strain 16S rRNA gene sequences of species described until May 2021 using the parsimony quick ad marked tool of ARB and the gap95_q0_to_q5 filter.
All strains of the resulting cluster and the type strain of the type species Paenibacillus polymyxa and of two additional species, Paenibacillus peoriae and Paenibacillus kribbensis, which clustered with the type species, were included in the analysis. Type strains of two Cohnella species, Cohnella thermotolerans and Cohnella xylanilytica, were used as outgroup. The genus Cohnella is a monophyletic genus related to the paraphyletic genus Paenibacillus. A maximum-likelihood tree was calculated with RAxML v7.04 (Stamatakis 2006), GTR-GAMMA and rapid bootstrap analysis and a maximum-parsimony tree with DNAPARS v 3.6 (Felsenstein 2005). Both trees were calculated with 100 re-samplings (bootstrap analysis; Felsenstein 1985) and based on the 16S rRNA gene sequences between gene termini 95 to 1475 (Escherichia coli numbering, Brosius et al. 1978).
Genomic features
The whole-genome sequencing of strains JJ-42 T was carried out with a NextSeq500 instrument using the Nextera XT DNA library preparation kit (Illumina) and a 2 × 150 bp paired-end protocol, yielding 3,876,943 read pairs (140 × average sequencing depth, 356 bp average insert size). Read processing, genome assembly and scaffolding, and gene prediction and annotation were performed using fq2dna v21.06 (https://gitlab.pasteur.fr/GIPhy/fq2dna).
Different plant−beneficial function contributing (PBFC) genes (as listed in Cherif-Silini et al. 2019) were searched using tblastn against the draft genome of JJ-42 T, assessing the presence of different putative PBFC genes related to different features.
Pairwise average nucleotide and amino acid identity (ANI and AAI, respectively) values were computed using OGRI_B (https://gitlab.pasteur.fr/GIPhy/OGRI) between the draft genome of JJ-42 T and every Paenibacillus type strain genome selected for the 16S rRNA gene sequence based phylogenetic analysis (when publicly available).
Physiology and chemotaxonomy
Detailed phenotypic characterization of strain JJ-42 T was performed in comparison to P. pectinilyticus KCTC 13222 T. Cell morphology and motility were observed under a Zeiss light microscope at a magnification of 1000, using cells that had been grown for 3 days at 28 °C on TSA (Oxoid). Gram staining was performed by the modified Hucker method according to Gerhardt et al. (1994). Oxidase activity was tested with an oxidase reagent text kit following the instructions of the manufacturer (bioMérieux, France). The KOH test for the determination of spores was carried out according to Moaledj (1986). The growth was tested on different agar media, including R2A (Oxoid), nutrient broth agar (NB, Oxoid), tryptic soy agar (TSA, Becton Dickinson), malt agar (Merck), PYE [0.3% (w/v) yeast extract, and 0.3% (w/v) casein peptone, respectively, 15 g agar L−1, pH 7.2], CASO agar (Carl Roth), K7 [0.1% (w/v) of yeast extract, peptone, and glucose, 15 g L−1 agar, pH 6.8], medium 65 (M65, according to DSMZ), DEV agar (DEV, Merck), Nutrient agar (NA, Becton Dickinson), Luria–Bertani agar (LB, Sigma-Aldrich), Marine agar 2216 (MA, Becton Dickinson), Columbia agar with sheep blood (Oxoid), and MacConkey agar (Oxoid). The growth was evaluated after 48 h incubation at 28 °C. Temperature-dependent growth was determined on Columbia agar with sheep blood at 4, 10, 15, 20, 25, 28, 30, 36, 45, 50, and 55 °C. pH and salinity-dependent growth was tested in R2A broth incubated at 28 °C. The pH was adjusted to pH 4.5 to 10.5 (1 pH unit intervals incrementing) using HCl and NaOH. For salinity-dependent growth, 1 to 8% (w/v) NaCl was added (in 1% intervals incrementing). The growth was monitored after 72 h of incubation.
Further physiological characterization of the strains was performed with the API20NE and APIZYM test systems according to the instruction of the manufacturer (bioMérieux) and with the methods described by Kämpfer et al. (1991) and Kämpfer (1990). All tests were incubated at 28 °C.
For analyses of quinones and polar lipids cells were grown in half-concentrated nutrient broth at 25 °C for 7 days. Quinones were extracted as described by Minnikin et al. (1984) and by Wiertz et al. (2013), and analyzed with an Agilent 1260 infinity HPLC system. Polar lipids were extracted and analyzed by thin-layer chromatography (TLC) according to Minnikin et al. (1984). Molybdophosphoric acid was used for visualization of all polar lipids. Aminolipids were stained by ninhydrin, molybdenum blue reagent for the detection of phospholipids, and α-naphthol reagent for the detection of sugar-containing lipids.
Fatty acid analysis was performed in parallel with JJ-42 T and P. pectinilyticus KCTC 13222 T. The fatty acids were extracted and analyzed as described by Kämpfer & Kroppenstedt (1996). Strains were grown under identical conditions (TS-medium after 72 h incubation at 28 °C) and the cells for extractions were taken from colonies of the same size. Fatty acids were identified with the Sherlock version 2.11, TSBA40 Rev. 4.1.
Results and discussion
Molecular and genome characteristics
The final corrected 16S rRNA gene sequence of strain JJ-42 T (OP023150) had a size of 1,466 nt and spanned gene termini 8 to 1,475 (numbering according to the Escherichia coli rrnB sequence published by Brosius et al. 1978).
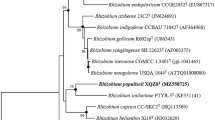
Strain JJ-42 T was placed within the cluster of Paenibacillus type strains by BLAST analyses. Strain JJ-42 T showed the highest 16S rRNA gene sequence similarity to the type strain of Paenibacillus pectinilyticus (98.8%), followed by the type strains of Paenibacillus qinlingensis and Paenibacillus oryzisoli (both 98.4%). Sequence similarities to all other type strains were below 98%. Independent of the applied treeing method, strain JJ-42 T formed a stable cluster (100% bootstrap support) with the type strains of P. pectinilyticus, P. qinlingensis, P. oryzisoli, and P. plantarum (Fig. 1).
Maximum-likelihood tree showing the phylogenetic position of strain JJ-42 T among the closest related Paenibacillus species. The tree was generated in ARB using RAxML (GTR-GAMMA, rapid bootstrap analysis) and based on the 16S rRNA gene sequences between positions 95–1475 according to E. coli numbering (Brosius et al. 1978). GenBank/EMBL/DDBJ accession numbers are given in parentheses. Numbers at branch nodes refer to bootstrap values > 70% (100 replicates). Circle marks nodes that were also present in the maximum-parsimony tree. Larger circles were supported by high bootstrap values in the maximum-parsimony tree. Type strains of Cohnella species were used as outgroup. Bar, 0.1 substitutions per nucleotide position
The resulting draft genome was made of 8,162,391 bps on 78 contigs (N50, 232,911) with G + C content of 45.07 mol%. A total of 7,004 coding sequences and 103 tRNA was inferred. Genome sequence authenticity was assessed by aligning the 16S rRNA segment derived from Sanger sequencing (OP023150) against the de novo assembly (CAKMMW00000000) using blastn, leading to > 99.4% pairwise sequence similarity.
ANI and AAI values are reported in Table S1, together with the associated digital DNA−DNA hybridization (dDDH) values (formula 2; https://ggdc.dsmz.de/). All these estimated pairwise similarity values are far below the commonly admitted species delineation cutoffs (ANI, 95%; AAI, 95%; dDDH, 70%). A phylogenomic classification of these genome sequences was also inferred using JolyTree v2.0 (Criscuolo 2019, 2020), such a tree confirming that strain JJ-42 T formed a stable cluster with the type strains of P. plantarum, P. oryzisoli, and P. pectinilyticus (Fig. 2).
Whole-genome-based tree showing the phylogenetic placement of strain JJ-42 T among type strains of closely related Paenibacillus species. This minimum evolution tree was inferred using JolyTree (https://gitlab.pasteur.fr/GIPhy/JolyTree). Two publicly available Cohnella type strain genomes were used as an outgroup. The genome sequence accession is specified between parentheses next after each taxon name. Branch supports were assessed by the rate of elementary quartets, as estimated by JolyTree (only supports > 0.5 were specified). Bar, 0.025 nucleotide substitutions per site
Different plant−beneficial function contributing (PBFC) genes (as listed in Cherif-Silini et al. 2019) were searched using tblastn against the draft genome of JJ-42 T, assessing the presence of different putative PBFC genes related to plant−root colonization (Table S2), nutrient acquisition (Table S3), growth-promoting traits (Table S4), oxidative stress protection (Table S5), drug and heavy metal resistance (Table S6), disease resistance (Table S7), and degradation of aromatic compounds (Table S8). Finally, seven biosynthetic gene clusters for secondary metabolites were found using antiSMASH (Blin et al. 2021) bacterial version (Table S9).
Phenotype
The results of the physiological characterization, performed using methods described previously (Kämpfer 1990; Kämpfer et al. 1991), are given in Table 1 and in the species description.
The polar lipid profile consisted of diphosphatidylglycerol, phosphatidylethanolamine, phosphatidylglycerol, three unidentified aminophospholipids, and one unidentified polar lipid (Fig. S1). The quinone system of strain JJ-42T consisted exclusively of menaquinone MK-7. The presence of menaquinone MK-7 as well as the polar lipid profile are in agreement with the characteristics of other species of the genus Paenibacillus.
The fatty acids comprised mainly iso- and anteiso-branched fatty acids and the fatty acid profile was very similar to the most closely related Paenibacillus species. The detailed fatty acid profile obtained from cells grown is shown in Table S10.
Conclusion
Based on the summary of genotypic and phenotypic, we describe a novel species of the genus Paenibacillus, for which the name Paenibacillus allorhizoplanae is proposed with JJ-42 T as proposed type strain.
Description of Paenibacillus allorhizoplanae sp. nov.
Paenibacillus allorhizoplanae (Gr. masc. adj. allos, other; N.L. gen. n. rhizoplanae, a specific epithet ('of the rhizoplane'); N.L. gen. n. allorhizoplanae, related, but distinct from Paenibacillus rhizoplanae).
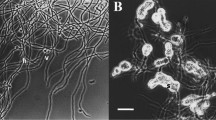
Cells (with rounded ends) stain Gram-positive. No chains or filaments could be observed after growth on TSA at 28 °C for 48 h. Cells were 2.0–3.0 µm in length and 0.8–1.0 µm in width) and showed no motility. Oval endospores are formed in a subterminal position in swollen sporangia. No other cell inclusions could be detected. Colonies grown on tryptone soy agar (after 48 h of incubation on TSA) are circular, convex, and beige with a shiny appearance and an average diameter of 2–3 mm. The growth at 28 °C on R2A (Oxoid), NB (Oxoid), TSA, Malt agar (Merck), PYE, CASO agar (Carl Roth), K7, M65, DEV agar (DEV, Merck), NA (Becton Dickinson), LB (Sigma-Aldrich), and Columbia agar with sheep blood (Oxoid). No growth on MA (Becton Dickinson) and MacConkey agar (Oxoid).
Good growth on blood agar between 25 and 45 °C (optimal growth at 36 °C), growth occurred between 15 and 45 °C, but not at 4, 10, 50 °C, and above. Optimal pH for growth in R2A broth at 28 °C is pH 7–8; growth occurs between pH 4.5 and 10.5. Optimal growth in R2A broth at 28 °C without and in the presence of 1% NaCl, growth occurs between 0 and 3% NaCl, but not at 4% or above. Both, pH- and salinity-dependent growth tested in R2A broth at 28 °C. Tests for catalase and oxidase activities are negative. According to API 20 NE, positive for the activity of ß-glucosidase (esculin hydrolysis), PNPG beta-galactosidase, and the assimilation of potassium gluconate, and negative for the activity of nitrate reduction, indol production, D-glucose fermentation, arginine dihydrolase, urease, gelatin hydrolysis, and the assimilation of D-glucose, L-arabinose, D-mannose, D-mannitol, N-acetylglucosamine, D-maltose, capric acid, adipic acid, malic acid, trisodium citrate, and phenylacetic acid.
According to API ZYM, positive for leucine arylamidase, acid phosphatase, naphtol-AS-BI-phosphohydrolase, and β-galactosidase, and negative for alkaline phosphatase, esterase (C4), esterase lipase (C8), lipase (C14), valine arylamidase, cystine arylamidase, trypsin, α-chymotrypsin, α-galactosidase, β-glucuronidase, α-glucosidase, β-glucosidase, N-acetyl-β-glucosaminidase, α-mannosidase, and α-fucosidase.
Some sugars or sugar-related compounds were utilized: L-arabinose (weak), D-galactose, D-gluconate, glucose (weak), D-melibiose, ribose, D-trehalose, and D-maltitol are utilized as sole sources of carbon.
Arbutin, D-cellobiose, D-fructose, i-inositol, D-mannose, D-maltose, salicin, sucrose, D-sorbitol, D-xylose, acetate, N-acetyl-D-glucosamine, cis-aconitate, trans-aconitate, adipate, D-adonitol, 4-aminobutyrate, azelate, citrate, itaconate, malate, mesaconate, 2-oxoglutarate, propionate, putrescine, pyruvate and L-rhamnose are not utilized as sole carbon source.
The quinone system contains only menaquinone MK-7. The polar lipid profile is composed of the major lipids diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine, three unidentified aminophospholipids (APL1-3) and one unidentified polar lipid (L).
The major fatty acids are anteiso C15:0 and iso C16:0. The genomic DNA G + C content is 45.07 mol% (based on the genome sequence).
The type stain JJ-42 T (= LMG 32089 T = CCM 9085 T = DSM 111786 T = CIP 111891 T) was isolated from the root surface of a field-grown corn plant in Dunbar, Nebraska USA.
The genome sequence of the type strain is available under accession number CAKMMW00000000 and the 16S rRNA gene sequence under OP023150.
Availability of data and material
The new generated sequences were uploaded to the GenBank database at the National Center for Biotechnology Information (NCBI) and are available. The complete genome sequence of strain JJ-42 T has been deposited under the GenBank/EMBL/DDBJ accession numbers CAKMMW000000000 and the 16S rRNA gene sequence under OP023150.
References
Ash C, Priest FG, Collins MD (1993) Molecular identification of rRNA group 3 bacilli (Ash, Farrow, Wallbanks and Collins) using a PCR probe test. Antonie Van Leeuwenhoek 64:253–260. https://doi.org/10.1007/bf00873085
Beneduzi A, Costa PB, Parma M et al (2010) Paenibacillus riograndensis sp. nov., a nitrogen-fixing species isolated from the rhizosphere of Triticum aestivum. Int J Syst Evol Microbiol 60:128–133. https://doi.org/10.1099/ijs.0.011973-0
Blin K, Shaw S, Kloosterman AM et al (2021) antiSMASH 6.0: improving cluster detection and comparison capabilities. Nucleic Acids Res 49(W1,2):W29–W35. https://doi.org/10.1093/nar/gkab335
Brosius J, Palmer ML, Kennedy PJ et al (1978) Complete nucleotide-sequence of a 16S ribosomal-RNA gene from Escherichia coli. PNAS 75:4801–4805. https://doi.org/10.1073/pnas.75.10.4801
Carro L, Flores-Félix JD, Cerda-Castillo E et al (2013) Paenibacillus endophyticus sp. nov., isolated from nodules of Cicer arietinum. Int J Syst Evol Microbiol 63:4433–4438. https://doi.org/10.1099/ijs.0.050310-0
Cherif-Silini H, Thissera B, Chenari Bouket A et al (2019) Durum wheat stress tolerance induced by endophyte pantoea agglomerans with genes contributing to plant functions and secondary metabolite arsenal. Int J Mol Sci 20:3989. https://doi.org/10.3390/ijms20163989
Criscuolo A (2019) A fast alignment-free bioinformatics procedure to infer accurate distance-based phylogenetic trees from genome assemblies. Res Ideas Outcomes 5:e36178. https://doi.org/10.3897/rio.5.e36178
Criscuolo A (2020) On the transformation of MinHash-based uncorrected distances into proper evolutionary distances for phylogenetic inference. F1000Research 9:1309. https://doi.org/10.12688/f1000research.26930.1
Elo S, Suominen I, Kämpfer P et al (2001) Paenibacillus borealis sp. nov., a nitrogen-fixing species isolated from spruce forest humus in Finland. Int J Syst Evol Microbiol 51:535–545
Felsenstein J (1985) Confidence limits of phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Felsenstein J (2005) PHYLIP (Phylogeny Inference Package) version 3.6. Distributed by the author. Department of Genome Sciences, University of Washington, Seattle
Gao JL, Lv FY, Wang XM et al (2015) Paenibacillus wenxiniae sp. nov., a nifH gene-harbouring endophytic bacterium isolated from maize. Antonie Van Leeuwenhoek 108:1015–1022. https://doi.org/10.1007/s10482-015-0554-8
Gerhardt P, Murray RGE, Wood W et al (1994) Methods for General and Molecular Bacteriology. American Society for Microbiology, Washington
Grady EN, MacDonald J, Liu L et al (2016) Current knowledge and perspectives of Paenibacillus: a review. Microb Cell Fact 15:203. https://doi.org/10.1186/s12934-016-0603-7
Han T-Y, Tong X-M, Wang Y-W et al (2015) Paenibacillus populi sp. nov., a novel bacterium isolated from the rhizosphere of Populus alba. Antonie Van Leeuwenhoek 108:659–666. https://doi.org/10.1007/s10482-015-0521-4
Hong YY, Ma YC, Zhou YG et al (2009) Paenibacillus sonchi sp. nov., a nitrogen-fixing species isolated from the rhizosphere of Sonchus oleraceus. Int J Syst Evol Microbiol 59:2656–2661. https://doi.org/10.1099/ijs.0.009308-0
Kämpfer P (1990) Evaluation of the Titertek-Enterobac-Automated system (TTE-AS) for identification of Enterobacteriaceae. Zentbl Bakteriol 273:164–172. https://doi.org/10.1016/s0934-8840(11)80244-6
Kämpfer P, Kroppenstedt RM (1996) Numerical analysis of fatty acid patterns of coryneform bacteria and related taxa. Can J Microbiol 42:989–1005. https://doi.org/10.1139/m96-128
Kämpfer P, Steiof M, Dott W (1991) Microbiological characterisation of a fuel-oil contaminated site including numerical identification of heterotrophic water and soil bacteria. Microbiol Ecol 21:227–243. https://doi.org/10.1007/bf02539156
Kämpfer P, Busse HJ, Kloepper JW et al (2016) Paenibacillus cucumis sp. nov., isolated from a cucumber plant. Int J Syst Evol Microbiol 66:2599–2603
Kämpfer P, Busse HJ, McInroy JA et al (2017a) Paenibacillus nebraskensis sp. nov., isolated from the root surface of field-grown maize. Int J Syst Evol Microbiol 67:4956–4961. https://doi.org/10.1099/ijsem.0.002357
Kämpfer P, Busse HJ, McInroy JA et al (2017b) Paenibacillus rhizoplanae sp. nov., isolated from the rhizosphere of Zea mays. Int J Syst Evol Microbiol 67:1058–1063. https://doi.org/10.1099/ijsem.0.001779
Kämpfer P, Busse HJ, McInroy JA et al (2021) Paenibacillus allorhizosphaerae sp. Nov., from soil of the rhizosphere of Zea mays. Int J Syst Evol Microbiol. https://doi.org/10.1099/ijsem.0.005051
Kim BC, Lee KH, Kim MN et al (2009a) Paenibacillus pini sp. nov., a cellulolytic bacterium isolated from the rhizosphere of pine tree. J Microbiol 47:699–704. https://doi.org/10.1007/s12275-009-0343-z
Kim BC, Lee KH, Kim MN et al (2009b) Paenibacillus pinihumi sp. nov., a cellulolytic bacterium isolated from the rhizosphere of Pinus densiflora. J Microbiol 47:530–535. https://doi.org/10.1007/s12275-009-0270-z
Kittiwongwattana C, Thawai C (2015) Paenibacillus lemnae sp. nov., an endophytic bacterium of duckweed (Lemna aequinoctialis). Int J Syst Evol Microbiol 65:107–112. https://doi.org/10.1099/ijs.0.067876-0
Lai WA, Hameed A, Lin SY et al (2015) Paenibacillus medicaginis sp. nov. a chitinolytic endophyte isolated from the root nodule of alfalfa (Medicago sativa L.). Int J Syst Evol Microbiol 65:3853–3860. https://doi.org/10.1099/ijsem.0.000505
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, United Kingdom, pp 115–175
Liu Y, Zhai L, Wang R et al (2015) Paenibacillus zeae sp. nov., isolated from maize (Zea mays L.) seeds. Int J Syst Evol Microbiol 65:4533–4538. https://doi.org/10.1099/ijsem.0.000608
Ludwig W, Strunk O, Westram R et al (2004) ARB: a software environment for sequence data. Nucleic Acid Res 32:1363–1371. https://doi.org/10.1093/nar/gkh293
Ma Y, Xia Z, Liu X et al (2007) Paenibacillus sabinae sp. nov., a nitrogen-fixing species isolated from the rhizosphere soils of shrubs. Int J Syst Evol Microbiol 57:6–11. https://doi.org/10.1099/ijs.0.64519-0
Minnikin DE, O’Donnell AG, Goodfellow M et al (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Method 2:233–241. https://doi.org/10.1016/0167-7012(84)90018-6
Moaledj K (1986) Comparison of Gram-staining and alternate methods, KOH test and aminopeptidase activity in aquatic bacteria: their application to numerical taxonomy. J Microbiol Methods 5:303–310. https://doi.org/10.1016/0167-7012(86)90056-4
Rivas R, Mateos PF, Martínez-Molina E et al (2005) Paenibacillus phyllosphaerae sp. nov., a xylanolytic bacterium isolated from the phyllosphere of Phoenix dactylifera. Int J Syst Evol Microbiol 55:743–746. https://doi.org/10.1099/ijs.0.63323-0
Rivas R, García-Fraile P, Mateos PF et al (2006) Paenibacillus cellulosilyticus sp. nov., a cellulolytic and xylanolytic bacterium isolated from the bract phyllosphere of Phoenix dactylifera. Int J Syst Evol Microbiol 56:2777–2781. https://doi.org/10.1099/ijs.0.64480-0
Schauss T, Busse HJ, Golke J et al (2015) Empedobacter stercoris sp. nov., isolated from an input sample of a biogas plant. Int J Syst Evol Microbiol 65:3746–3753. https://doi.org/10.1099/ijsem.0.000486
Son J-S, Kang H-U, Ghim S-Y (2014) Paenibacillus dongdonensis sp. nov., isolated from rhizospheric soil of Elymus tsukushiensis. Int J Syst Evol Microbiol 64:2865–2870. https://doi.org/10.1099/ijs.0.061077-0
Stamatakis A (2006) RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690. https://doi.org/10.1093/bioinformatics/btl446
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis Version 11. Mol Biol Evol 38(7):3022–3027. https://doi.org/10.1093/molbev/msab120
Wang DS, Jiang YY, Wie XM et al (2014) Paenibacillus quercus sp. nov., isolated from rhizosphere of Quercus aliena var. acuteserrata. Antonie Van Leeuwenhoek 105:1173–1178. https://doi.org/10.1007/s10482-014-0178-4
Wiertz R, Schulz SC, Müller U et al (2013) Corynebacterium frankenforstense sp. nov. and Corynebacterium lactis sp. nov., isolated from raw cow milk. Int J Syst Evol Microbiol 63:4495–4501. https://doi.org/10.1099/ijs.0.050757-0
Xin K, Li M, Chen C et al (2017) Paenibacillus qinlingensis sp. nov., an indole-3-acetic acid-producing bacterium isolated from roots of Sinopodophyllum hexandrum (Royle) Ying. Int J Syst Evol Microbiol 67:589–595. https://doi.org/10.1099/ijsem.0.001666
Yarza P, Richter M, Peplies J et al (2008) The all-species living tree project: A 16S rRNA-based phylogenetic tree of all sequenced type strains. Syst Appl Microbiol 31:241–250. https://doi.org/10.1016/j.syapm.2008.07.001
Zhang J, Wang ZT, Yu HM et al (2013) Paenibacillus catalpae sp. nov., isolated from the rhizosphere soil of Catalpa speciosa. Int J Syst Evol Microbiol 63:1776–1781. https://doi.org/10.1099/ijs.0.040659-0
Zhang L, Gao JS, Zhang S et al (2015) Paenibacillus rhizoryzae sp. nov., isolated from rice rhizosphere. Int J Syst Evol Microbiol 65:3053–3059. https://doi.org/10.1099/ijs.0.000376
Acknowledgements
We thank Gundula Will, Maria Sowinsky and Katja Grebing for excellent technical assistance and Prof. Aharon Oren for his help with the etymology of the specific epithet. We also acknowledge the help of the HPC Core Facility of the Institut Pasteur for this work.
Funding
Open Access funding enabled and organized by Projekt DEAL. No funding was received.
Author information
Authors and Affiliations
Contributions
P.K. conceived and designed the study and provided fatty acid data. P.K. S.G. A.C. J.M. A.L. and D.C. wrote the manuscript. S.G. A.L. and P.K. conducted the experiments. A.L. analyzed the lipids and quinones. A.C. and L. L. performed the genomic and phylogenomic analyses.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose. The authors declare no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
The manuscript is submitted with the consent of all authors.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kämpfer, P., Lipski, A., Lamothe, L. et al. Paenibacillus allorhizoplanae sp. nov. from the rhizoplane of a Zea mays root. Arch Microbiol 204, 630 (2022). https://doi.org/10.1007/s00203-022-03225-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-022-03225-w