Abstract
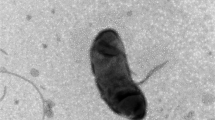
A novel bacterium, designated CAU 1574T, was isolated from marine sand. Cells were Gram stain negative, aerobic, gliding and rod shaped. Growth was observed at 20–37 °C (optimum, 30 °C), a pH of 5.5–10.0 (optimum, 8.0), and 0–3.0% (w/v) NaCl concentrations (optimum, 1%). Based on the results of 16S rRNA gene sequence analyses, strain CAU 1574T belonged to the genus Echinicola, and showed the highest similarity to Echinicola shivajiensis JCM 17847T (97.5%). Phylogenomic analysis based on consisting of 92 core genes extracted from the genome sequences showed that strain CAU 1574T was affiliated with species in the genus Echinicola. The average nucleotide identity (ANI), average amino acid identity (AAI), and digital DNA–DNA hybridization (dDDH) values between strain CAU 1574T and the closely related species were below the cut-off values of 95–96, 90, and 70%, respectively used for species demarcation. The chemotaxonomic data of CAU 1574T were as follows: major isoprenoid quinone, MK-7; predominant polar lipids, phosphatidylethanolamine, two unidentified aminophospholipids and two unidentified lipids; major fatty acids, iso-C15:0, summed feature 3 (C16:1 ω6c/C16:1 ω7c). The 5.4 Mb genome included 20 contigs and 4237 protein-coding genes with a 39.8 mol% G + C content. Based on the phylogenetic, phenotypic, and physiological result, strain CAU 1574T represents a novel species of this genus Echinicola, for which the name Echinicola arenosa sp. nov. is proposed. The type strain is CAU 1574T (= KCTC 82410T = MCCC 1K05669T).


Similar content being viewed by others
Data availability
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene and the complete genome sequences of strain CAU 1574T are MN540912 and JACYTQ000000000, respectively.
References
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T et al (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:75
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N et al (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87
Bowman JP (2000) Description of Cellulophaga algicola sp. nov., isolated from the surfaces of Antarctic algae, and reclassification of Cytophaga uliginosa (ZoBell and Upham 1944) Reichenbach 1989 as Cellulophaga uliginosa comb. nov. Int J Syst Evol Microbiol 50:1861–1868
Cappuccino JG, Sherman N (2010) Microbiology: a laboratory manual, 9th edn. Benjamin Cummings, San Francisco, USA
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Embley TM, Wait R (1994) Structural Lipids of Eubacteria. In: Goodfellow M, O'Donnell (eds) Chemical methods in prokaryotic systematics. Chichester, John Wiley and Sons, New York, USA, pp 121–161
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evol 39:783–791
Gomori G (1955) Preparation of buffers for use in enzyme studies. Methods Enzymol 1:138–146
Hiraishi A, Ueda Y, Ishihara J, Mori T (1996) Comparative lipoquinone analysis of influent sewage and activated sludge by high-performance liquid chromatography and photodiode array detection. J Gen Appl Microbiol 42:457–469
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HH (ed) Mammalian protein metabolism. Academic Press, New York, pp 21–132
Jung YJ, Yang SH, Kwon KK, Bae SS (2017) Echinicola strongylocentroti sp. nov., isolated from a sea urchin Strongylocentrotus intermedius. Int J Syst Evol Microbiol 67:670–675
Kim H, Joung Y, Ahn TS, Joh K (2011) Echinicola jeungdonensis sp. nov., isolated from a solar saltern. Int J Syst Evol Microbiol 61:2065–2068
Kim JH, Kanjanasuntree R, Kim DH, Lee JS, Sukhoom A, Kantachote D et al (2019) Arenibacillus arenosus gen. nov., sp. nov., a member of the family Rhodobacteraceae isolated from sea sand. Int J Syst Evol Microbiol 69:153–158
Kim JH, Ward AC, Kim W (2015) Kangiella chungangensis sp. nov. isolated from a marine sand. Anton Van Leeuwenhoek 107:1291–1298
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Konstantinidis KT, Tiedje JM (2005) Genomic insights that advance the species definition for prokaryotes. Proc Natl Acad Sci USA 102:2567–2572
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Larkin MA, Blackshields G, Brown NP, Chenn R, McGettigan PA, McWilliam H et al (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Lee DW, Lee AH, Lee H, Kim JJ, Khim JS, Yim UH et al (2017) Echinicola sediminis sp. nov., a marine bacterium isolated from coastal sediment. Int J Syst Evol Microbiol 67:3351–3357
Liang P, Sun J, Li H, Liu M, Xue Z, Zhang Y (2016) Echinicola rosea sp. nov., a marine bacterium isolated from surface seawater. Int J Syst Evol Microbiol 66:3299–3304
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A et al (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Na S-I, Kim YO, Yoon S-H, Ha S-M, Baek I, Chun J (2018) UBCG: up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:281–285
Nam SW, Kim W, Chun J, Goodfellow M (2004) Tsukamurella pseudospumae sp. nov., a novel actinomycete isolated from activated sludge foam. Int J Syst Evol Microbiol 54:1209–1212
Nedashkovskaya OI, Kim SB, Vancanneyt M, Lysenko AM, Shin DS, Park MS et al (2006) Echinicola pacifica gen. nov., sp. nov., a novel flexibacterium isolated from the sea urchin Strongylocentrotus intermedius. Int J Syst Evol Microbiol 56:953–958
Nedashkovskaya OI, Kim SB, Hoste B, Shin DS, Beleneva IA, Vancanneyt M et al (2007) Echinicola vietnamensis sp. nov., a member of the phylum Bacteroidetes isolated from seawater. Int J Syst Evol Microbiol 57:761–763
Parte AC, Carbasse JS, Meier-Kolthoff JP, Reimer LC, Göker M (2020) List of prokaryotic names with standing in nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol 70:5607
Rodriguez-R LM, Konstantinidis KT (2016) The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Prep 4:e1900v1
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (2006) Bacterial identification by gas chromatographic analysis of fatty acids methyl esters (GC-FAME). Technical Note 101, Microbial ID Inc., Newark, DE, USA
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RG, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 607–654
Srinivas TNR, Tryambak BK, Kumar PA (2012) Echinicola shivajiensis sp. nov., a novel bacterium of the family “Cyclobacteriaceae” isolated from brackish water pond. Anton Van Leeuwenhoek 101:641–647
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Xing YT, Xu L, Wang HT, Huang XX, Wang S, Sun JQ (2020) Echinicola soli sp. nov., isolated from alkaline saline soil. Int J Syst Evol Microbiol 70:4139–4144
Funding
This work was supported by a grant from the National Institute of Biological Resources (NIBR), funded by the Ministry of Environment (MOE) of the Republic of Korea (NIBR201902203), and the Chung-Ang University Research Grants in 2020.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: WK. Conducted the experiments: JB, VW, J-SL, analyzed the data: J-HK, J-HY, AS. Contributed reagents, materials, and analysis tools: WK. Wrote the paper: JB, VW, WK.
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared that no competing interests exist.
Ethics approval
This article does not contain any studies with human participants and/or animals performed by any of the authors. The formal consent is not required in this study.
Consent to participate
All authors have seen a copy of the manuscript and have approved its submission.
Consent to participate and/or Consent to publish
This research does not involve in human subjects, so the informed consent to participate and consent to publish are not obtained.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Baek, J., Weerawongwiwat, V., Kim, JH. et al. Echinicola arenosa sp. nov., isolated from marine sand. Arch Microbiol 203, 5675–5681 (2021). https://doi.org/10.1007/s00203-021-02553-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02553-7