Abstract
Understanding the gut microbiota characteristics of endangered species such as the Eurasian otter (Lutra lutra), especially in their early stages of life, could be essential for improving their management and ex situ conservation strategies. Here, we analyzed the gut microbiota diversity, composition, and function of captive Eurasian otters at different ages using high-throughput 16S rRNA gene sequencing. We found that: (1) Clostridiaceae was abundant in all age stages; (2) Lactococcus in cubs is thought to predominate for digesting milk; (3) bacteria associated with amino acid metabolism increase with age, while bacteria associated with carbohydrate metabolism decrease with age, which is likely due to decrease in dietary carbohydrate content (e.g., milk) and increase in dietary protein contents (e.g., fishes) with age; and (4) fish-related bacteria were detected in feces of healthy adults and juveniles. Overall, the gut microbiota of captive Eurasian otters was taxonomically and functionally different by age, which is thought to be attributed to the difference in the diet in their life stages. This study provided baseline information regarding the gut microbiota of Eurasian otters for the first time and contributes to improvement in their management in captivity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Eurasian otter (Lutra lutra) belongs to the subfamily Lutrinae and is a top predator in freshwater ecosystems (Blanco-Garrido et al. 2008). Although the Eurasian otter was originally found throughout most of Europe, Asia, and North Africa, its population has declined in recent decades due to habitat destruction, pollution, and urbanization. Therefore, the Eurasian otter is currently designated as a near threatened species on the International Union for Conservation of Nature (IUCN) Red List and is listed in Appendix I of the Convention on International Trade in endangered species (CITES) to protect this species. In East Asia, it was declared in 2012 that the Eurasian otter has already become extinct in Japan (Nakanishi and Izawa 2019). The distribution area of the Eurasian otter has dramatically declined in China over the last half-century, and this species has not been recorded on Taiwan Island for over two decades (Li and Chan 2018). Moreover, the population of the Eurasian otter in Korea has also declined rapidly during the last several decades, even though it had once been abundant throughout the country (Jo and Won 2019).
Ex situ conservation (in captivity) is also very important in addition to in-situ conservation (in the wild) for the management of endangered species (Che-Castaldo et al. 2018). Among several approaches in ex situ conservation, studies on gut microbiota are essential because gut microbiota play an important role in maintaining the host’s health (Karmacharya et al. 2019). While it has been reported that hand-reared otter cubs have successfully grown up after being fed puppy milk replacers with low protein and lactose, several cubs have died as a result of weakness from diarrhea during the process (Ito et al. 2020). In humans, it is known that gut microbiota development influences infants’ health and subsequent host physiology (Moore and Townsend 2019). Therefore, gaining an understanding of the gut microbiota characteristics of the Eurasian otter in an early age may lead to improved husbandry skills of this endangered species. However, to the best of our knowledge, there is no valid information on the microbiota of the Eurasian otter during infancy, although several studies have been conducted using adult individuals (An et al. 2017; Finlayson-Trick et al. 2017) and other otter species, including the North American river otter (Lontra canadensis) (Finlayson-Trick et al. 2017; Guo et al. 2020a), neotropical otter (Lontra longicaudis) (Rodriguez-Rey and Santamaria-Vanegas 2020), and sea otter (Enhydra lutris) (Miller et al. 2010).
In this study, the gut microbiota diversity, composition, and function of Eurasian otters at different age stages were analyzed and compared using high-throughput 16S rRNA gene sequencing to provide baseline information regarding the gut microbiota of this species. As mentioned above, the Eurasian otter is extremely rare and ex situ conservation is essential. Research on physiology and ecology, including gut microbiota conditions, in captivity is vital for ex situ conservation, and this time, by clarifying the baseline characteristics of the gut microbiota of Eurasian otters, we aimed to improve husbandry management in captivity. To the best of our knowledge, this was the first observation of the gut microbiota characteristics of Eurasian otters in their early stages of life.
Materials and methods
Studied animals
A total of seven Eurasian otters kept at the Korean Otter Research Center were assessed in this study (Table 1; Fig. S1). Among them, three otters were born in captivity, and four otters were rescued in the wild and artificially raised. The age of rescued otter cubs was estimated based on their weight and length by a previously reported method (Reuther 1999). For comparison of the characteristics of gut microbiota by age, all individuals were categorized into four growth stages: cubs at the age of 1–2 M; cubs at the age of 3 M; cubs at the age of 4–5 M; and juveniles (8 M–2 Y) and adults (≥ 2 Y) (M = months, Y = years). The juvenile and adult otters were mainly fed 1 kg of Channel catfish (Ictalurus punctatus) once per day with seasonal feeding of landlocked salmon (Oncorhynchus masou masou) during winter. Conversely, the feeding of the cubs varied based on their developmental stage: (1) 50–80 ml of diluted Kitten Milk Replacer (KMR; Pet-Ag, Inc., Hampshire, IL, USA) was fed four times per day during 1–2 M; (2) 30 g of weaning food, including diluted Kitten Milk Replacer, defrosted Alaska pollock (Gadus chalcogrammus) (Frozen Alaska Pollack fillet; Youngwoo Corp., Busan, South Korea) and cans of kitten (Today’s recipe: Tuna pate, [Topper, Pocheon, South Korea] or Fancy Feast: Tender ocean whitefish feast kitten [Purina, Saint Louis, MO, USA]) or puppy food (Cesar Puppy beef and eggs, Mars Petcare, Wodonga, VIC, Australia), was fed four times per day during 3 M; and (3) 100–120 g of weaning food, including defrosted Alaska pollock and cans of kitten or puppy food, was fed three times per day during 4–5 M. No supplemental food was fed to any of the otters. Unfortunately, cub No. 5 died at 3 M, and cubs No. 6 and 7 died at 5 M due to unknown causes.
Fecal sample collection
In total, 40 fecal samples were collected noninvasively after natural voiding from September 2018 to June 2019. Fecal samples were collected immediately after excretion and stored frozen at − 20 °C until DNA extraction. The feces excreted on the dry floor for the marking purpose were collected to minimize contamination by water. All methods of sample collection were performed in accordance with a previously reported method (National Research Council 2011). Fecal samples were collected noninvasively by the animal keeper as part of normal rearing operations. Therefore, the approval by Institutional Animal Care and Use Committee was not required for this study.
DNA extraction
The genomic DNA was extracted from about 0.5 g of each fecal sample by using the DNeasy PowerSoil Kit (QIAGEN, Venlo, The Netherlands) with modification (Hospodsky et al. 2010; An et al. 2017; Kumari et al. 2019). In addition to the DNeasy Powersoil Kit’s beads, 0.1-mm diameter glass beads (300 mg) and 0.5-mm diameter glass beads (100 mg) were supplemented for the 3-min physical disruption step by the mini beadbeater-24 (Biospec Products, Inc., Bartlesville, OK, USA). After bead beating, DNA was extracted, purified, and eluted to 50 μl of TE (10 mM Tris–HCl, 1 mM EDTA, pH = 8.0) based on the kit’s protocol. The extracted DNA samples were stored at − 80 °C until subsequent DNA sequencing analysis.
DNA sequencing
The extracted DNA was amplified using universal bacterial primers targeting the V3 and V4 regions of the 16S rRNA gene (Herlemann et al. 2011) with MiSeq adapter sequences. The PCR was carried out in a 50 μl mixture composed of 2 × PCR Solution Premix Taq™ DNA polymerase (Takara Bio Inc., Otsu, Shiga, Japan), 1 μM of each primer, and 1 μl of each extract. The PCR conditions were as follows: 5 min at 95 °C for the denaturing step, followed by 35 cycles consisting of 30 s at 95 °C, 30 s at 55 °C, and 60 s at 72 °C. The final elongation step was performed for 10 min at 72 °C on a T100™ thermal cycler (Bio-Rad Laboratories, Inc., Hercules, CA, USA). The amplicons were purified using AMPure XP beads (Beckman Coulter, Inc., Brea, CA, USA) and indexed with a Nextera XT Index Kit v2 (Illumina, Inc., San Diego, CA, USA) in a 50 μl mixture consisting of 2 × PCR Solution Premix Taq™ DNA polymerase (Takara Bio Inc.), 5 μl of each index primer, and 5 μl of each purified DNA. The index PCR conditions were as follows: 3 min at 95 °C for the denaturing step, followed by 8 cycles consisting of 30 s at 95 °C, 30 s at 55 °C, and 30 s at 72 °C. The final elongation step was 5 min at 72 °C. The indexed PCR products were purified using AMPure XP beads (Beckman Coulter) and normalized to 4 nM with 10 mM Tris–HCl (pH = 8.5). The subsequent products were pooled with an internal control PhiX (30%) and loaded on a v3 600 cycle-kit reagent cartridge (Illumina, Inc.). The 2 × 300 bp paired-end sequencing was performed using an Illumina MiSeq (Illumina, Inc.). Raw sequence data are available in the Sequence Read Archive of the National Center for Biotechnology Information under project number PRJNA613985. The raw sequence reads were demultiplexed using MiSeq Reporter v2.5 (Illumina, Inc.) for subsequent bacterial taxonomical and functional analyses.
Taxonomical analyses
The demultiplexed reads were processed using USEARCH version 11.0.667 (Edgar 2010). Forward and reverse reads were joined and low-quality reads with lengths less than 200 bp and/or > 1.0 expected errors were removed using the -fastq_filter command. Unique sequences were identified among joined and filtered reads using the -fastx_uniques command. Using the UNOISE3 algorithm (Edgar and Flyvbjerg 2015; Edgar 2016), the reads with sequencing errors and chimeric sequences were removed. Simultaneously, the resultant reads were clustered into zero-radius operational taxonomic units (ZOTUs) at a 100% sequence identity level (Edgar 2018b). Using the SINTAX algorithm (Edgar 2016, 2018a), each ZOTU was taxonomically assigned against the RDP training set v16 (rdp_16s_v16.fa.gz) with a cutoff value of 0.8. Additionally, α and β diversity analyses based on ZOTUs were performed by the “phyloseq” package (McMurdie and Holmes 2013) on R version 4.0.4. The Jaccard similarity coefficient and Bray–Curtis similarity coefficient were computed to analyze the similarity of bacterial assemblage memberships and structures across samples, respectively. Permutational multivariate analysis of variance (PERMANOVA) was performed to test the statistical difference in the gut microbiota across the age groups. The post hoc pairwise comparisons were performed by the “pairwiseAdonis” package (Martinez Arbizu 2020).
Functional analyses
Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt) (Langille et al. 2013) was used for functional analyses. The functional analyses were performed separately from the abovementioned taxonomic analyses from raw sequence reads according to the method of PICRUSt. Briefly, poly(N) tails of raw sequence reads were trimmed by the Trimmomatic-0.39 (Bolger et al. 2014). The forward and reverse reads were joined by the join_paired_ends.py script with a minimum allowed overlap of 10 bp (-j 10) on Quantitative Insights Into Microbial Ecology (Caporaso et al. 2010). The operational taxonomic units (OTUs) at a 97% sequence similarity threshold level were identified by the pick_closed_reference_otus.py script against the Greengenes database v13_5 (DeSantis et al. 2006; McDonald et al. 2012). The resultant OTUs were normalized by the normalize_by_copy_number.py script with the Greengenes database v13_5. The functions were predicted by the predict_metagenomes.py script with the Greengenes database v13_5 and Kyoto Encyclopedia of Genes and Genomes (KEGG) Orthologs (Kanehisa and Goto 2000; Kanehisa 2019; Kanehisa et al. 2021). Finally, the predicted functions were classified by the categorize_by_function.py script. The categorized functions were further analyzed by Statistical Analysis of Metagenomic Profiles version 2.1.3 (Parks et al. 2014) to compare the gut microbiota across the age groups.
Results
Sequencing statistics
After a series of quality filter steps, a total of 1,084,107 (average length of 448 bp) chimera-free high-quality sequences were recovered, with an average of 27,102 sequences per sample, ranging from 9749 to 45,373. These sequences were assigned to a total of 2548 ZOTUs at a 100% sequence identity level, which were sorted from 40 fecal samples. Each sample had 617 ± 259 ZOTUs on average, ranging from 234 to 1079.
Diversity
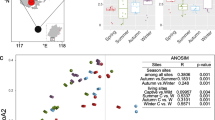
For diversity analyses, each library was rarefied to 9700 sequence reads. The rarefaction curves are shown in Fig. 1a. The α diversity analyses revealed increasing trends in the Chao1 estimator (species richness) (Fig. 1b) and the Shannon index (species diversity) (Fig. 1c) as the otters aged from birth to adulthood. However, the tendencies were not statistically significant (p > 0.05; Kruskal–Wallis test). The results of α diversity metrics for all samples are listed in Table S1. The β diversity analyses based on the Jaccard similarity coefficients (community membership) (Fig. 1d) and Bray–Curtis similarity coefficients (community structure) (Fig. 1e) revealed significant differences in the gut microbiota across the age categories (p < 0.05; PERMANOVA). The pairwise comparisons revealed differences in gut microbiota, both in terms of community structures and memberships, across the age categories, except between the groups of cubs at the ages of 3 M and 4–5 M.
Diversity of gut microbiota of captive Eurasian otters across the age categories of 1–2 months, 3 months, 4–5 months, and adults/juveniles. Rarefaction curves based on zero-radius operational taxonomic units (ZOTUs) at a 100% sequence identity level (a). Alpha diversity metrics based on the Chao1 estimator (b) and Shannon index (c). The bottom and top lines of the boxes indicate the 25th and 75th percentiles, respectively, and the lines in the boxes indicates the medians. The bottom and top whiskers indicate the 10th and 90th percentiles, respectively. Beta diversity based on the Jaccard similarity coefficients (d) and Bray–Curtis similarity coefficient (e) on non-metric multidimensional scaling (NMDS) plots, with different letters (a, b, and c) representing significant differences (p < 0.05) by pair-wise comparisons
Taxonomic compositions
The relative abundances of gut microbiota at the family level are shown in Fig. 2. There was a tendency towards greater Peptostreptococcaceae abundance as the otters aged, while Streptococcaceae became less abundant with age. On average, relative abundances of Streptococcaceae were 44%, 40%, 23%, and 0.1% for cubs during 1–2 M, 3 M, and 4–5 M, and adults/juveniles, respectively. Conversely, Peptostreptococcaceae was more abundant in adults/juveniles than in cubs, i.e., on average 6.5%, 13%, 20%, and 41% for cubs during 1–2 M, 3 M, 4–5 M, and adults/juveniles, respectively. Furthermore, Enterobacteriaceae was particularly abundant in cubs during 1–2 M, i.e., on average 16%, while it was less abundant in other age groups, i.e., 0.2%, 2.0%, and 0.1% for cubs during 3 M and 4–5 M, and adults/juveniles, respectively. Clostridiaceae was abundant in all age groups, i.e., on average 9.5%, 35%, 29%, and 22% for cubs during 1–2 M, 3 M, and 4–5 M, and adults/juveniles, respectively.
Relative abundance of bacterial families in gut microbiota of captive Eurasian otters. The alphabet character before the first hyphen in each sample ID represents “C” for cubs, “J” for juveniles, and “A” for adults. The number before the first hyphen indicates the otter's individual ID. For cubs, the number after the first hyphen represents the age in the month of the otter. The number after the last hyphen represents the sample ID within each sampling month for each otter
The relative abundances of gut microbiota at the genus rank are shown in Fig. 3. In the family Streptococcaceae, Streptococcus and Lactococcus were identified, with mean relative abundances of 27% and 0.10%, respectively. They were the most abundant in cubs during 1–2 M. In the family Peptostreptococcaceae, Terrisporobacter was identified, with a mean relative abundance of 2.7% and the highest abundance observed in the group of adults and juveniles. In the family Enterobacteriaceae, Escherichia/Shigella and Plesiomonas were identified, with respective mean relative abundances of 4.2% and 0.51%. Escherichia/Shigella were the most abundant in cubs during 1–2 M, while Plesiomonas was abundant in cubs in 4–5 M and adults/juveniles. Four phylogenetic clusters of Clostridium were identified, i.e., Clostridium sensu stricto, Clostridium XI, Clostridium XlVa, and Clostridium XVIII with respective mean relative abundances of 23%, 5.6%, 0.67%, and 0.24%. Clostridium sensu stricto and Clostridium XI were detected in both cubs and adults/juveniles. However, Clostridium XlVa and Clostridium XVIII were detected only in cubs, with the highest relative abundances observed in the cub No. 2 during 1–2 M. The similar tendency was observed for Bacteroides and Fusobacterium. Conversely, some bacterial genera were more abundant in older cubs (4–5 M) and adults/juveniles. The examples include Psychrobacter, Cetobacterium, Plesiomonas, Vagococcus, Dietzia, and Sporosarcina.
Relative abundance of identified bacterial genera or taxa in gut microbiota of captive Eurasian otters. The data of unidentified sequence reads are not included. The data shown represent 84.6% of all high-quality sequence reads. The alphabet character before the first hyphen in each sample ID represents “C” for cubs, “J” for juveniles, and “A” for adults. The number before the first hyphen indicates the otter’s individual ID. For cubs, the number after the first hyphen represents the age in the month of the otter. The number after the last hyphen represents the sample ID within each sampling month for each otter. The tree represents the Euclidean distance-based similarities of log-transformed bacterial compositions
Functional tendencies
PICRUSt predicted four of the KEGG level 2 functions that were significantly different across the age categories, i.e., amino acid metabolism, carbohydrate metabolism, lipid metabolism, and metabolism of cofactors and vitamins (p < 0.05; Kruskal–Wallis test) (Fig. 4). The relative abundance of genes associated with amino acid metabolism and metabolism of cofactors and vitamins were found to increase with age. Conversely, the relative abundance of genes associated with carbohydrate metabolism and lipid metabolism were found to decrease with age. Pairwise comparisons based on the post hoc Tukey–Kramer test confirmed statistical significance between the groups of cubs during 1–2 M and adults/juveniles (p < 0.05), except for lipid metabolism. All of the KEGG level 2 and 3 functions significantly different across the age categories are listed in Table S2.
Relative abundance of genes associated with a amino acid metabolism, b carbohydrate metabolism, c lipid metabolism, and d metabolism of cofactors and vitamins in a metagenome predicted by Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt). The results of significantly different predicted functions at the Kyoto Encyclopedia of Genes and Genomes (KEGG) level 2 functions are shown. The error bars represent standard deviations. The different letters (a, b, and c) representing significant differences (p < 0.05) by pair-wise comparisons based on the post hoc Tukey–Kramer test
Discussion
Understanding gut microbiota is important for health management in captivity and ex situ conservation of endangered species because mammalian gut microbiota provides a range of essential functions for the host from the digestion of complex food to signaling the host immune system (McFall-Ngai et al. 2013). In this study, the first comparative investigation of the gut microbiota of captive Eurasian otters was conducted to characterize fecal microbial diversity, composition, and function with age.
Although it is not statistically significant, we found that the α diversity metrics of Eurasian otters increased with age (Fig. 1b, c), as previously reported in the forest musk deer (Moschus berezovskii) (Zhao et al. 2019a) and the Asian elephant (Elephas maximus) (Kambe et al. 2020), and that their gut microbiota structures became similar to those of adults as they aged. For carnivorous animals, previous studies of gut microbiota compared by gender, age, subspecies, or diet in the tiger (Panthera tigris) (Jiang et al. 2020) and spotted hyena (Crocuta crocuta) (Chen et al. 2020) showed that diet might be the most significant factor to shape the gut microbiota. In this study, significant differences in the β diversity and changes in the gut microbiota were observed in the four age groups with different dietary contents (Figs. 1d, e, 2, 3). The dietary contents are considered to be a factor that caused the observed differences.
Differences in relative abundance in gut microbiota were compared based on age stages, and Clostridiaceae was abundant in all age stages (Fig. 2), which was consistent with a study that revealed that the relative abundance of Clostridiaceae was 52% for Eurasian otters (An et al. 2017). Meanwhile, Enterococcaceae abundance appeared to be high in the gut microbiota of otters from 5 M to adults, which was supported by previous studies that showed that Enterococcus spp. belonging to the family Enterococcaceae were abundantly detected in wild Eurasian otters (Oliveira et al. 2008). The shift in the predominant family from Streptococcaceae to Peptostreptococcaceae with age was thought to have corresponded to their life stages. The bacterial genus Streptococcus belonging to the family Streptococcaceae is known to be abundantly detected in human newborn infants (Chong et al. 2018), and Peptostreptococcaceae is the representative bacterial family of carnivore species (Nishida and Ochman 2018).
At the genus rank, Lactococcus was dominant in the 1- to 2-month-old cubs, and Terrisporobacter was dominant in the adults and juveniles (Fig. 3). A study of the same Mustelidae American mink (Neovison vison) by Compo et al. (2018) showed that Lactococcus was abundantly detected in freshly weaned kits than adult females, which supports that the genus Lactococcus is thought to predominate for digesting milk. Terrisporobacter has been reported to be the most predominant in the gut microbiota of human infants fed only powder milk (Cai et al. 2019). In animals, Terrisporobacter was found in the gut microbiota of giant pandas (Zhao et al. 2019b) and pigs (McCormack et al. 2019). Notably, Wang et al. (2019) showed that pigs fed low-protein supplemented with casein hydrolysate (LPC) diet increased the abundance of Lactobacillus and decreased the abundance of Terrisporobacter compared with low-protein supplemented with free all amino acids (LPA) diet. Therefore, Terrisporobacter might be involved in the digestion of free all amino acids, but its roles in adult otters need further investigation.
Our functional analysis revealed that bacteria associated with amino acid metabolism and metabolism of cofactors and vitamins were found to increase with age, while bacteria associated with carbohydrate metabolism and lipid metabolism were found to decrease with age (Fig. 4). In the study for the tigers reported by Jiang et al. (2020), carbohydrate metabolism, amino acid metabolism, energy metabolism, and metabolism of cofactors and vitamins were reported as the main functions of their gut microbiota. Jiang et al. (2020) also reported the absence of Candida, a yeast, in the gut microbiota of meat-fed adult tigers and argued that it was attributable to a high protein content of the tiger diet. Candida, which is found in the gut microbiota of dogs and humans, is known to be related to digestion of carbohydrate-rich foods (Hoffmann et al. 2013). Additionally, Jiang et al. (2020) reported that energy metabolism was more abundant in meat-fed adult tigers than milk-fed tiger cubs, indicating that the gut of meat-fed tigers harbors more bacteria that assist energy metabolism. Milk contains large amounts of carbohydrates. For instance, lactose, a type of carbohydrates, is known to account for about 40% of the energy contained in human breast milk (Grenov et al. 2016). In the Eurasian otters investigated in this study, we expect that increase in amino acid metabolism and a decrease in carbohydrate metabolism with age was due to a decrease in dietary carbohydrate content (e.g., milk) and an increase in dietary protein contents (e.g., fishes) with age.
Fusobacterium and Bacteroides, which are possible human pathogens, were detected in the gut microbiota of the cub No. 1 at the age of 1 month (Fig. 3). Fusobacterium is known to be associated with diseases such as colorectal cancer and acute appendicitis in humans (Zhong et al. 2014; Brennan and Garrett 2016). However, Fusobacteria is known to be detected frequently and abundantly in the gut of predatory animals, such as seals (Nelson et al. 2013), dolphins (Nishida and Ochman 2018), cheetahs (Ley et al. 2008), leopards (Han et al. 2019), polar bears (Ley et al. 2008), blue foxes, silver foxes, raccoon dogs (Peng et al. 2018; Liu et al. 2020), fringe-lipped bats (Nishida and Ochman 2018), vultures (Roggenbuck et al. 2014), and alligators (Keenan et al. 2013). Moreover, Jiang et al. (2020) reported that abundances of Fusobacterium increased in meat-fed tigers compared with mix-fed and milk-fed tigers, implying that the high abundance of Fusobacterium is associated with the digestion of meat. Similarly, Bacteroides was reported to be dominant in the gut microbiota of adult and juvenile spottedd hyenas (Chen et al. 2020), Amur tigers (He et al. 2018), and dholes, implying that Bacteroides is known as protein and fat degraders (Arumugam et al. 2011). These findings indicated that Fusobacterium and Bacteroides are unlikely to act as serious pathogens in otter cubs, too.
Several bacterial genera were abundantly detected only in feces of adult and juvenile otters (Fig. 3). Of them, some genera are known to be associated with fishes, such as Psychrobacter, Cetobacterium, Plesiomonas, Vagococcus, Dietzia, and Sporosarcina. For instance, the first species of Psychrobacter was reported to be isolated from fishes (Juni and Heym 1986). Cetobacterium and Plesiomonas were identified from the gut of freshwater fishes (Sugita et al. 1993; Larsen et al. 2014). Vagococcus sp. was isolated from rainbow trout as a fish pathogen (Ruiz-Zarzuela et al. 2005). Dietzia was reported to be isolated from a drain of a fish-egg-processing plant (Yumoto et al. 2002), while Sporosarcina was reported to be isolated from minced fish meat (Tsuda et al. 2015). Moreover, Sporosarcina was isolated from seawater in Korea (Yoon et al. 2001). The detection of fish-related bacteria in healthy adults and juveniles indicates that they may participate as part of the normal gut microbiota of adult Eurasian otters.
Unfortunately, all of the three cubs died due to unknown reasons. These cubs were rescued in the wild and artificially raised at the Korean Otter Research Center. Although the cause of cubs’ mortality is unknown, no particular abnormalities were observed in the health condition until shortly before death. In addition, there was no distinct difference between samples collected immediately before death in each individual (C1-4M-3, C2-5M-2, and C3-5M-4) and the samples collected before that. However, it is difficult to assert the relationship between changes in the gut microbiota and mortality because there were no surviving cubs to compare with the cubs that died at this time. We look forward to further analysis using surviving individuals as controls in the future. Meanwhile, in the first reported case of successful hand-rearing of a Eurasian otter cub abandoned by the parent otters in a captive environment in Japan, probiotic bacteria, such as probiotic strains of Enterococcus faecium and Lactobacillus acidophilus, were administered from 33 days-old (Ito et al. 2020). We do not know how probiotic bacteria if administered, could have improved the survivability of our rescued Eurasian otter cubs. Our study revealed that Lactococcus was dominant in the gut microbiota of 1- to 2-month-old Eurasian otter cubs, rather than Lactobacillus, which is reported to be predominant in giant panda weaning cubs (Guo et al. 2020b). Further research is warranted to investigate the roles of probiotic bacteria, including lactic acid bacteria such as Lactococcus and Lactobacillus, in the health status of Eurasian otters in their early stages of life.
As a part of ex situ conservation activity, understanding the characteristics of the gut microbiota of endangered species in captivity is important. However, it should be noted that animals in captivity experience a range of changes that may influence the gut bacteria, such as diet changes, treatments, and reduced contact with other individuals, species, and variable environmental substrates that act as sources of bacterial diversity (McKenzie et al. 2017). These effects have been confirmed in many animals, such as nonhuman primates (Clayton et al. 2016, 2018), carnivores (Chong et al. 2019; Guo et al. 2019), and herbivores (Gibson et al. 2019; Zhao et al. 2019a). In otters, there is only one report in the neotropical otter (Lontra longicaudis) (Rodriguez-Rey and Santamaria-Vanegas 2020); therefore, further research is necessary to reveal the captive effects on the gut microbiota of the Eurasian otter and on the resultant possible health consequences.
It should also be noted that seasons can have a certain impact on the gut microbiota of animals. For instance, the significant seasonal differences in gut microbiota were reported by studies in giant pandas (Guo et al. 2020b) and Canadian mink of the same Mustelidae (Compo et al. 2018). In our study, we found the statistically significant differences in the gut microbiota collected between summer and fall, and between summer and winter (p < 0.05; PERMANOVA) (Table S3). We expect that the difference is due to the fact that all of juvenile/adult fecal samples were collected in the summer. The season effects appear to be much smaller than the age effects since the seasonal differences were observed only for summer. Moreover, if we look only at the results of cubs, significant differences were observed by age (Fig. 1d, e), but not by season (Table S3). Specifically, we observed statistically significant differences between 1–2 and 3–5 M (Fig. 1d, e), but no significant seasonal differences between the autumn, winter, and spring in which all of cubs’ feces were collected (Table S3). Furthermore, we observed the age-sequential increases in the α diversity measures, i.e., Chao1 estimator and Shannon index, although it was not statistically significant (Fig. 1b, c). Similarly, our functional analysis revealed that relative abundances of genes related to each metabolism increase or decrease sequentially by age (Fig. 4). These results imply that the effect of age was greater than the effect of the season since these age-sequential tendencies may have been more obscured if other confounding factors such as seasons predominated. However, future research should completely eliminate confounding factors such as season for better discerning the age effect.
In conclusion, we provided the first report of the dominant bacteria in the gut microbiota of Eurasian otters by age. The distinct differences in the gut microbiota and functions by age were observed, which is thought to be attributed to the difference in the diet in their life stages. However, the roles of these bacteria in otter’s health and diseases remain unknown. Characterization of bacteria that increase the survivability of the rescued cubs, which may include known human probiotics as well as fish-related bacteria commonly found in healthy adult otters, is necessary. This study investigated gut microbiota of Eurasian otters by age for the first time. We expect that the accumulation of knowledge of their gut microbiota will help elucidating the beneficial microbes. Studies on gut microbiota may be of particular relevance to ex situ conservation of wildlife in captive environments, such as zoos, which require preventative health care as well as veterinary treatment to increase the survivability of animals rescued in the wild or abandoned by the parents.
Data availability
Raw sequence data are available in the Sequence Read Archive of the National Center for Biotechnology Information under project number PRJNA613985.
References
An C, Okamoto Y, Xu S, Eo KY, Kimura J, Yamamoto N (2017) Comparison of fecal microbiota of three captive carnivore species inhabiting Korea. J Vet Med Sci 79:542–546
Arumugam M, Raes J, Pelletier E, Le Paslier D, Yamada T, Mende DR, Fernandes GR, Tap J, Bruls T, Batto J-M, Bertalan M, Borruel N, Casellas F, Fernandez L, Gautier L, Hansen T, Hattori M, Hayashi T, Kleerebezem M, Kurokawa K, Leclerc M, Levenez F, Manichanh C, Nielsen HB, Nielsen T, Pons N, Poulain J, Qin J, Sicheritz-Ponten T, Tims S, Torrents D, Ugarte E, Zoetendal EG, Wang J, Guarner F, Pedersen O, de Vos WM, Brunak S, Doré J, Antolín M, Artiguenave F, Blottiere HM, Almeida M, Brechot C, Cara C, Chervaux C, Cultrone A, Delorme C, Denariaz G, Dervyn R, Foerstner KU, Friss C, van de Guchte M, Guedon E, Haimet F, Huber W, van Hylckama-Vlieg J, Jamet A, Juste C, Kaci G, Knol J, Kristiansen K, Lakhdari O, Layec S, Le Roux K, Maguin E, Mérieux A, Melo Minardi R, M’Rini C, Muller J, Oozeer R, Parkhill J, Renault P, Rescigno M, Sanchez N, Sunagawa S, Torrejon A, Turner K, Vandemeulebrouck G, Varela E, Winogradsky Y, Zeller G, Weissenbach J, Ehrlich SD, Bork P, Meta HITC (2011) Enterotypes of the human gut microbiome. Nature 473:174–180
Blanco-Garrido F, Prenda J, Narvaez M (2008) Eurasian otter (Lutra lutra) diet and prey selection in Mediterranean streams invaded by centrarchid fishes. Biol Invasions 10:641–648
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120
Brennan CA, Garrett WS (2016) Gut microbiota, inflammation, and colorectal cancer. Annu Rev Microbiol 70:395–411
Cai C, Zhang Z, Morales M, Wang Y, Khafipour E, Friel J (2019) Feeding practice influences gut microbiome composition in very low birth weight preterm infants and the association with oxidative stress: A prospective cohort study. Free Radic Biol Med 142:146–154
Caporaso J, Kuczynski J, Stombaugh J, Bittinger K, Bushman F, Costello E, Fierer N, Pena A, Goodrich J, Gordon J, Huttley G, Kelley S, Knights D, Koenig J, Ley R, Lozupone C, McDonald D, Muegge B, Pirrung M, Reeder J, Sevinsky J, Turnbaugh P, Walters W, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Che-Castaldo JP, Grow SA, Faust LJ (2018) Evaluating the contribution of North American zoos and aquariums to endangered species recovery. Sci Rep 8:9789
Chen L, Liu M, Zhu J, Gao Y, Sha W, Ding H, Jiang W, Wu S (2020) Age, gender, and feeding environment influence fecal microbial diversity in spotted hyenas (Crocuta crocuta). Curr Microbiol 77:1139–1149
Chong CYL, Bloomfield FH, O’Sullivan JM (2018) Factors affecting gastrointestinal microbiome development in neonates. Nutrients 10:274
Chong R, Grueber CE, Fox S, Wise P, Barrs VR, Hogg CJ, Belov K (2019) Looking like the locals - gut microbiome changes post-release in an endangered species. Anim Microbiome 1:8
Clayton JB, Vangay P, Huang H, Ward T, Hillmann BM, Al-Ghalith GA, Travis DA, Long HT, Tuan BV, Minh VV, Cabana F, Nadler T, Toddes B, Murphy T, Glander KE, Johnson TJ, Knights D (2016) Captivity humanizes the primate microbiome. Proc Natl Acad Sci USA 113:10376–10381
Clayton JB, Al-Ghalith GA, Long HT, Tuan BV, Cabana F, Huang H, Vangay P, Ward T, Minh VV, Tam NA, Dat NT, Travis DA, Murtaugh MP, Covert H, Glander KE, Nadler T, Toddes B, Sha JCM, Singer R, Knights D, Johnson TJ (2018) Associations between nutrition, gut microbiome, and health in a novel nonhuman primate model. Sci Rep 8:11159
Compo NR, Gomez DE, Tapscott B, Weese JS, Turner PV (2018) Fecal bacterial microbiota of Canadian commercial mink (Neovison vison): yearly, life stage, and seasonal comparisons. PLoS One 13:e020711
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Edgar RC (2016) SINTAX: a simple non-Bayesian taxonomy classifier for 16S and ITS sequences. bioRxiv 074161
Edgar RC (2018a) Accuracy of taxonomy prediction for 16S rRNA and fungal ITS sequences. PeerJ 6:e4652
Edgar RC (2018b) Updating the 97% identity threshold for 16S ribosomal RNA OTUs. Bioinformatics 34:2371–2375
Edgar RC, Flyvbjerg H (2015) Error filtering, pair assembly and error correction for next-generation sequencing reads. Bioinformatics 31:3476–3482
Finlayson-Trick ECL, Getz LJ, Slaine PD, Thornbury M, Lamoureux E, Cook J, Langille MGI, Murray LE, McCormick C, Rohde JR, Cheng Z (2017) Taxonomic differences of gut microbiomes drive cellulolytic enzymatic potential within hind-gut fermenting mammals. PLoS One 12:e0189404
Gibson KM, Nguyen BN, Neumann LM, Miller M, Buss P, Daniels S, Ahn MJ, Crandall KA, Pukazhenthi B (2019) Gut microbiome differences between wild and captive black rhinoceros – implications for rhino health. Sci Rep 9:7570
Grenov B, Briend A, Sangild PT, Thymann T, Rytter MH, Hother A-L, Mølgaard C, Michaelsen KF (2016) Undernourished children and milk lactose. Food Nutr Bull 37:85–99
Guo W, Mishra S, Wang C, Zhang H, Ning R, Kong F, Zeng B, Zhao J, Li Y (2019) Comparative study of gut microbiota in wild and captive giant pandas (Ailuropoda melanoleuca). Genes 10:827
Guo G, Eccles KM, McMillan M, Thomas PJ, Chan HM, Poulain AJ (2020a) The gut microbial community structure of the North American river otter (Lontra canadensis) in the Alberta oil sands region in Canada: relationship with local environmental variables and metal body burden. Environ Toxicol Chem 39:2516–2526
Guo W, Chen Y, Wang C, Ning R, Zeng B, Tang J, Li C, Zhang M, Li Y, Ni Q, Ni X, Zhang H, Li D, Zhao J, Li Y (2020b) The carnivorous digestive system and bamboo diet of giant pandas may shape their low gut bacterial diversity. Conserv Physiol 8:coz104
Han S, Guan Y, Dou H, Yang H, Yao M, Ge J, Feng L (2019) Comparison of the fecal microbiota of two free-ranging Chinese subspecies of the leopard (Panthera pardus) using high-throughput sequencing. PeerJ 7:e6684
He F, Liu D, Zhai J, Zhang L, Ma Y, Xu Y, Rong K, Ma J (2018) Metagenomic analysis revealed the effects of goat milk feeding and breast feeding on the gut microbiome of Amur tiger cubs. Biochem Biophys Res Commun 503:2590–2596
Herlemann DPR, Labrenz M, Jürgens K, Bertilsson S, Waniek JJ, Andersson AF (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5:1571–1579
Hoffmann C, Dollive S, Grunberg S, Chen J, Li H, Wu GD, Lewis JD, Bushman FD (2013) Archaea and fungi of the human gut microbiome: Correlations with diet and bacterial residents. PLoS One 8:e66019
Hospodsky D, Yamamoto N, Peccia J (2010) Accuracy, precision, and method detection limits of quantitative PCR for airborne bacteria and fungi. Appl Environ Microbiol 76:7004–7012
Ito S, Kurihara T, Ohta M, Yamada A, Seino S, Ueda M, Murata K (2020) The first successful cases of hand-rearing of Eurasian otter (Lutra lutra) in Japan. J Jpn Assoc Zool Gdns Aquar 61:67–76
Jiang H, Chen W, Su L, Huang M, Lin L, Su Q, Li G, Ahmad HI, Li L, Zhang X, Li H, Chen J (2020) Impact of host intraspecies genetic variation, diet, and age on bacterial and fungal intestinal microbiota in tigers. MicrobiologyOpen 9:e1050
Jo Y-S, Won C (2019) The sprainting behavior and habitat preference of the Eurasian otter Lutra lutra along a montane stream in South Korea. Mammal Study 45:3–11
Juni E, Heym GA (1986) Psychrobacter immobilis gen. nov., sp. nov.: genospecies composed of Gram-negative, aerobic, oxidase-positive coccobacilli. Int J Syst Evol Microbiol 36:388–391
Kambe J, Sasaki Y, Inoue R, Tomonaga S, Kinjo T, Watanabe G, Jin W, Nagaoka K (2020) Analysis of infant microbiota composition and the relationship with breast milk components in the Asian elephant (Elephas maximus) at the zoo. J Vet Med Sci 82:983–989
Kanehisa M (2019) Toward understanding the origin and evolution of cellular organisms. Protein Sci 28:1947–1951
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kanehisa M, Furumichi M, Sato Y, Ishiguro-Watanabe M, Tanabe M (2021) KEGG: integrating viruses and cellular organisms. Nucleic Acids Res 49:D545–D551
Karmacharya D, Manandhar P, Manandhar S, Sherchan AM, Sharma AN, Joshi J, Bista M, Bajracharya S, Awasthi NP, Sharma N, Llewellyn B, Waits LP, Thapa K, Kelly MJ, Vuyisich M, Starkenburg SR, Hero J-M, Hughes J, Wultsch C, Bertola L, Fountain-Jones NM, Sinha AK (2019) Gut microbiota and their putative metabolic functions in fragmented Bengal tiger population of Nepal. PLoS One 14:e0221868
Keenan SW, Engel AS, Elsey RM (2013) The alligator gut microbiome and implications for archosaur symbioses. Sci Rep 3:2877
Kumari P, Dong K, Eo KY, Lee W-S, Kimura J, Yamamoto N (2019) DNA metabarcoding-based diet survey for the Eurasian otter (Lutra lutra): development of a Eurasian otter-specific blocking oligonucleotide for 12S rRNA gene sequencing for vertebrates. PLoS One 14:e0226253
Langille MGI, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R, Beiko RG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821
Larsen AM, Mohammed HH, Arias CR (2014) Characterization of the gut microbiota of three commercially valuable warmwater fish species. J Appl Microbiol 116:1396–1404
Ley RE, Hamady M, Lozupone C, Turnbaugh PJ, Ramey RR, Bircher JS, Schlegel ML, Tucker TA, Schrenzel MD, Knight R, Gordon JI (2008) Evolution of mammals and their gut microbes. Science 320:1647–1651
Li F, Chan BPL (2018) Past and present: the status and distribution of otters (Carnivora: Lutrinae) in China. Oryx 52:619–626
Liu H, Li Z, Si H, Zhong W, Fan Z, Li G (2020) Comparative analysis of the gut microbiota of the blue fox (Alopex lagopus) and raccoon dog (Nyctereutes procyonoides). Arch Microbiol 202:135–142
Martinez Arbizu P (2020) pairwiseAdonis: Pairwise multilevel comparison using adonis. In, R package version 0.4
McCormack UM, Curião T, Metzler-Zebeli BU, Wilkinson T, Reyer H, Crispie F, Cotter PD, Creevey CJ, Gardiner GE, Lawlor PG (2019) Improvement of feed efficiency in pigs through microbial modulation via fecal microbiota transplantation in sows and dietary supplementation of inulin in offspring. Appl Environ Microbiol 85:e01255-e11219
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, Andersen GL, Knight R, Hugenholtz P (2012) An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6:610–618
McFall-Ngai M, Hadfield MG, Bosch TCG, Carey HV, Domazet-Lošo T, Douglas AE, Dubilier N, Eberl G, Fukami T, Gilbert SF, Hentschel U, King N, Kjelleberg S, Knoll AH, Kremer N, Mazmanian SK, Metcalf JL, Nealson K, Pierce NE, Rawls JF, Reid A, Ruby EG, Rumpho M, Sanders JG, Tautz D, Wernegreen JJ (2013) Animals in a bacterial world, a new imperative for the life sciences. Proc Natl Acad Sci USA 110:3229–3236
McKenzie VJ, Song SJ, Delsuc F, Prest TL, Oliverio AM, Korpita TM, Alexiev A, Amato KR, Metcalf JL, Kowalewski M, Avenant NL, Link A, Di Fiore A, Seguin-Orlando A, Feh C, Orlando L, Mendelson JR, Sanders J, Knight R (2017) The effects of captivity on the mammalian gut microbiome. Integr Comp Biol 57:690–704
McMurdie PJ, Holmes S (2013) phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217
Miller MA, Byrne BA, Jang SS, Dodd EM, Dorfmeier E, Harris MD, Ames J, Paradies D, Worcester K, Jessup DA, Miller WA (2010) Enteric bacterial pathogen detection in southern sea otters (Enhydra lutris nereis) is associated with coastal urbanization and freshwater runoff. Vet Res 41:01
Moore RE, Townsend SD (2019) Temporal development of the infant gut microbiome. Open Biol 9:190128
Nakanishi N, Izawa M (2019) Rediscovery of otters on the Tsushima islands, Japan by trail cameras. Mammal Study 44:215–220
National Research Council (2011) Guide for the care and use of laboratory animals, 8th edn. The National Academies Press, Washington
Nelson TM, Rogers TL, Carlini AR, Brown MV (2013) Diet and phylogeny shape the gut microbiota of Antarctic seals: a comparison of wild and captive animals. Environ Microbiol 15:1132–1145
Nishida AH, Ochman H (2018) Rates of gut microbiome divergence in mammals. Mol Ecol 27:1884–1897
Oliveira M, Sales-Luís T, Duarte A, Nunes SF, Carneiro C, Tenreiro T, Tenreiro R, Santos-Reis M, Tavares L, Vilela CL (2008) First assessment of microbial diversity in faecal microflora of Eurasian otter (Lutra lutra Linnaeus, 1758) in Portugal. Eur J Wildl Res 54:245–252
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30:3123–3124
Peng Y, Zhang Z, Li H, Li S, Shi Q, Zhang J (2018) Assessment of fecal microbiota in farmed silver fox (Vulpes vulpes fulva) and raccoon dog (Nyctereutes procyonoides). Acta Agric Scand A Anim Sci 68:142–151
Reuther C (1999) Development of weight and length of Eurasian otter (Lutra Lutra) cubs. IUCN SCC Otter Spec Group Bull 16:11–26
Rodriguez-Rey LC, Santamaria-Vanegas J (2020) Gut bacteria comparison between wild and captive neotropical otters. Univ Sci 25:359–384
Roggenbuck M, Bærholm Schnell I, Blom N, Bælum J, Bertelsen MF, Sicheritz-Pontén T, Sørensen SJ, Gilbert MTP, Graves GR, Hansen LH (2014) The microbiome of New World vultures. Nat Commun 5:5498
Ruiz-Zarzuela I, de Bias I, Gironés O, Ghittino C, MúAzquiz JL (2005) Isolation of Vagococcus salmoninarum in rainbow trout, Oncorhynchus mykiss (Walbaum), Broodstocks: characterization of the pathogen. Vet Res Commun 29:553–562
Sugita H, Nakamura T, Deguchi Y (1993) J Food Prot 56:949–953
Tsuda K, Nagano H, Ando A, Shima J, Ogawa J (2015) Isolation and characterization of psychrotolerant endospore-forming Sporosarcina species associated with minced fish meat (surimi). Int J Food Microbiol 199:15–22
Wang H, Shen J, Pi Y, Gao K, Zhu W (2019) Low-protein diets supplemented with casein hydrolysate favor the microbiota and enhance the mucosal humoral immunity in the colon of pigs. J Anim Sci Biotechnol 10:79
Yoon JH, Lee KC, Weiss N, Kho YH, Kang KH, Park YH (2001) Sporosarcina aquimarina sp. nov., a bacterium isolated from seawater in Korea, and transfer of Bacillus globisporus (Larkin and Stokes 1967), Bacillus psychrophilus (Nakamura 1984) and Bacillus pasteurii (Chester 1898) to the genus Sporosarcina as Sporosarcina globispora comb. nov., Sporosarcina psychrophila comb. nov. and Sporosarcina pasteurii comb. nov., and emended description of the genus Sporosarcina. Int J Syst Evol Microbiol 51:1079–1086
Yumoto I, Nakamura A, Iwata H, Kojima K, Kusumoto K, Nodasaka Y, Matsuyama H (2002) Dietzia psychralcaliphila sp. nov., a novel, facultatively psychrophilic alkaliphile that grows on hydrocarbons. Int J Syst Evol Microbiol 52:85–90
Zhao G, Ma T, Tang W, Li D, Mishra SK, Xu Z, Wang Q, Jie H (2019a) Gut microbiome of Chinese forest musk deer examined across gender and age. Biomed Res Int 2019:9291216
Zhao S, Li C, Li G, Yang S, Zhou Y, He Y, Wu D, Zhou Y, Zeng W, Li T, Qu Y, Li B, Deng W, Jin L, Yu X, Huang Y, Zhang H, Zou L (2019b) Comparative analysis of gut microbiota among the male, female and pregnant giant pandas (Ailuropoda melanoleuca). Open Life Sci 14:288–298
Zhong D, Brower-Sinning R, Firek B, Morowitz MJ (2014) Acute appendicitis in children is associated with an abundance of bacteria from the phylum Fusobacteria. J Pediatr Surg 49:441–446
Funding
The cost of DNA sequencing in this study was supported by Seoul National University.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest associated with this manuscript.
Ethical approval
Fecal samples of Eurasian otters were collected noninvasively by the animal keeper as part of normal rearing operations. Therefore, the approval by Institutional Animal Care and Use Committee was not required for this study.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Okamoto, Y., Ichinohe, N., Woo, C. et al. Contrasting gut microbiota in captive Eurasian otters (Lutra lutra) by age. Arch Microbiol 203, 5405–5416 (2021). https://doi.org/10.1007/s00203-021-02526-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02526-w