Abstract
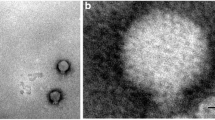
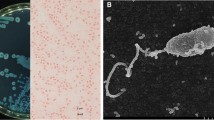
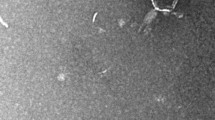
Cholera is a devastating diarrheal disease that accounts for more than 10% of children’s lives worldwide, but its treatment is hampered by a rise in antibiotic resistance. One promising alternative to antibiotic therapy is the use of bacteriophages to treat antibiotic-resistant cholera infections, and control Vibrio cholera in clinical cases and in the environment, respectively. Here, we report four novel, closely related environmental myoviruses, VP4, VP6, VP18, and VP24, which we isolated from two environmental toxigenic Vibrio cholerae strains from river Kuja and Usenge beach in Kenya. High-throughput sequencing followed by bioinformatics analysis indicated that the genomes of the four bacteriophages have closely related sequences, with sizes of 148,180 bp, 148,181 bp, 148,179 bp, and 148,179 bp, and a G + C content of 36.4%. The four genomes carry the phoH gene, which is overrepresented in marine cyanophages. The isolated phages displayed a lytic activity against 15 environmental, as well as one clinical, Vibrio cholerae strains. Thus, these novel lytic vibriophages represent potential biocontrol candidates for water decontamination against pathogenic Vibrio cholerae and ought to be considered for future studies of phage therapy.





Similar content being viewed by others
References
Abedon ST, Kuhl SJ, Blasdel BG, Kutter EM (2011) Phage treatment of human infections. Bacteriophage 1:66–85. https://doi.org/10.4161/bact.1.2.15845
Adriaenssens EM, Cowan DA (2014) Using signature genes as tools to assess environmental viral ecology and diversity. Appl Environ Microbiol 80:4470–4480. https://doi.org/10.1128/aem.00878-14
Almagro-Moreno S, Taylor RK (2013) Cholera: environmental reservoirs and impact on disease transmission. Microbiol Spectr. https://doi.org/10.1128/microbiolspec.OH-0003-2012
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/s0022-2836(05)80360-2
Aziz RK, Bartels D, Best AA et al (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:75. https://doi.org/10.1186/1471-2164-9-75
Aziz RK, Dwivedi B, Akhter S, Breitbart M, Edwards RA (2015) Multidimensional metrics for estimating phage abundance, distribution, gene density, and sequence coverage in metagenomes. Front Microbiol 6:381. https://doi.org/10.3389/fmicb.2015.00381
Aziz RK, Ackermann HW, Petty NK, Kropinski AM (2018) Essential steps in characterizing bacteriophages: biology, taxonomy, and genome analysis. Methods Mol Biol 1681:197–215. https://doi.org/10.1007/978-1-4939-7343-9_15
Baudoux AC, Hendrix RW, Lander GC et al (2012) Genomic and functional analysis of Vibrio phage SIO-2 reveals novel insights into ecology and evolution of marine siphoviruses. Environ Microbiol 14:2071–2086. https://doi.org/10.1111/j.1462-2920.2011.02685.x
Bhandare S, Colom J, Baig A et al (2019) Reviving phage therapy for the treatment of cholera. J Infect Dis 219:786–794. https://doi.org/10.1093/infdis/jiy563
Clokie MR, Millard AD, Mann NH (2010) T4 genes in the marine ecosystem: studies of the T4-like cyanophages and their role in marine ecology. Virol J 7:291. https://doi.org/10.1186/1743-422x-7-291
Cohen Y, Joseph Pollock F, Rosenberg E, Bourne DG (2013) Phage therapy treatment of the coral pathogen Vibrio coralliilyticus. Microbiol Open 2:64–74. https://doi.org/10.1002/mbo3.52
Comeau AM, Krisch HM (2008) The capsid of the T4 phage superfamily: the evolution, diversity, and structure of some of the most prevalent proteins in the biosphere. Mol Biol Evol 25:1321–1332. https://doi.org/10.1093/molbev/msn080
Davis JJ, Wattam AR, Aziz RK et al (2020) The PATRIC Bioinformatics Resource Center: expanding data and analysis capabilities. Nucleic Acids Res 48:D606-d612. https://doi.org/10.1093/nar/gkz943
Deen J, Mengel MA, Clemens JD (2020) Epidemiology of cholera. Vaccine 38(Suppl 1):A31-a40. https://doi.org/10.1016/j.vaccine.2019.07.078
El-Shibiny A, El-Sahhar S (2017) Bacteriophages: the possible solution to treat infections caused by pathogenic bacteria. Can J Microbiol 63:865–879. https://doi.org/10.1139/cjm-2017-0030
Faruque SM, Islam MJ, Ahmad QS et al (2005a) Self-limiting nature of seasonal cholera epidemics: role of host-mediated amplification of phage. Proc Natl Acad Sci USA 102:6119–6124. https://doi.org/10.1073/pnas.0502069102
Faruque SM, Naser IB, Islam MJ et al (2005b) Seasonal epidemics of cholera inversely correlate with the prevalence of environmental cholera phages. Proc Natl Acad Sci USA 102:1702–1707. https://doi.org/10.1073/pnas.0408992102
Gao EB, Huang Y, Ning D (2016) Metabolic genes within cyanophage genomes: implications for diversity and evolution. Genes (basel). https://doi.org/10.3390/genes7100080
Gao M, Qin Y, Fan H et al (2020) Characteristics and complete genome sequence of the virulent Vibrio alginolyticus phage VAP7, isolated in Hainan, China. Arch Virol 165:947–953. https://doi.org/10.1007/s00705-020-04535-4
Goldsmith DB, Crosti G, Dwivedi B et al (2011) Development of phoH as a novel signature gene for assessing marine phage diversity. Appl Environ Microbiol 77:7730–7739. https://doi.org/10.1128/aem.05531-11
Hendrix RW, Lawrence JG, Hatfull GF, Casjens S (2000) The origins and ongoing evolution of viruses. Trends Microbiol 8:504–508. https://doi.org/10.1016/s0966-842x(00)01863-1
Hendrix RW, Hatfull GF, Smith MC (2003) Bacteriophages with tails: chasing their origins and evolution. Res Microbiol 154:253–257. https://doi.org/10.1016/s0923-2508(03)00068-8
Hoque MM, Naser IB, Bari SM, Zhu J, Mekalanos JJ, Faruque SM (2016) Quorum regulated resistance of Vibrio cholerae against environmental bacteriophages. Sci Rep 6:37956. https://doi.org/10.1038/srep37956
Huang Y, Niu B, Gao Y, Fu L, Li W (2010) CD-HIT Suite: a web server for clustering and comparing biological sequences. Bioinformatics 26:680–682. https://doi.org/10.1093/bioinformatics/btq003
Jassim SA, Limoges RG (2014) Natural solution to antibiotic resistance: bacteriophages “The Living Drugs.” World J Microbiol Biotechnol 30:2153–2170. https://doi.org/10.1007/s11274-014-1655-7
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kauffman KM, Hussain FA, Yang J et al (2018) A major lineage of non-tailed dsDNA viruses as unrecognized killers of marine bacteria. Nature 554:118–122. https://doi.org/10.1038/nature25474
Kazakov AE, Vassieva O, Gelfand MS, Osterman A, Overbeek R (2003) Bioinformatics classification and functional analysis of PhoH homologs. Silico Biol 3:3–15
Kelly L, Ding H, Huang KH, Osburne MS, Chisholm SW (2013) Genetic diversity in cultured and wild marine cyanomyoviruses reveals phosphorus stress as a strong selective agent. ISME J 7:1827–1841. https://doi.org/10.1038/ismej.2013.58
Kieft K, Zhou Z, Anantharaman K (2020) VIBRANT: automated recovery, annotation and curation of microbial viruses, and evaluation of viral community function from genomic sequences. Microbiome 8:90. https://doi.org/10.1186/s40168-020-00867-0
Krzywinski M, Schein J, Birol I et al (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645. https://doi.org/10.1101/gr.092759.109
Kutter E (2009) Phage host range and efficiency of plating. Methods Mol Biol 501:141–149. https://doi.org/10.1007/978-1-60327-164-6_14
Kwan T, Liu J, DuBow M, Gros P, Pelletier J (2005) The complete genomes and proteomes of 27 Staphylococcus aureus bacteriophages. Proc Natl Acad Sci USA 102:5174–5179. https://doi.org/10.1073/pnas.0501140102
Letchumanan V, Chan KG, Pusparajah P et al (2016) Insights into bacteriophage application in controlling Vibrio species. Front Microbiol 7:1114. https://doi.org/10.3389/fmicb.2016.01114
Levy K, Woster AP, Goldstein RS, Carlton EJ (2016) Untangling the impacts of climate change on waterborne diseases: a systematic review of relationships between diarrheal diseases and temperature, rainfall, flooding, and drought. Environ Sci Technol 50:4905–4922. https://doi.org/10.1021/acs.est.5b06186
Li X, Sun Y, Liu J, Yao Q, Wang G (2019) Survey of the bacteriophage phoH gene in wetland sediments in northeast China. Sci Rep 9:911. https://doi.org/10.1038/s41598-018-37508-4
Lindell D, Jaffe JD, Coleman ML et al (2007) Genome-wide expression dynamics of a marine virus and host reveal features of co-evolution. Nature 449:83–86. https://doi.org/10.1038/nature06130
Loo K-Y, Letchumanan V, Law JW-F et al (2020) Incidence of antibiotic resistance in Vibrio spp. Rev Aquacult 12:2590–2608. https://doi.org/10.1111/raq.12460
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964. https://doi.org/10.1093/nar/25.5.955
Lutz C, Erken M, Noorian P, Sun S, McDougald D (2013) Environmental reservoirs and mechanisms of persistence of Vibrio cholerae. Front Microbiol 4:375. https://doi.org/10.3389/fmicb.2013.00375
Madhusudana Rao B, Lalitha KV (2015) Bacteriophages for aquaculture: are they beneficial or inimical. Aquaculture 437:146–154. https://doi.org/10.1016/j.aquaculture.2014.11.039
Maina AN, Mwaura FB, Oyugi J, Goulding D, Toribio AL, Kariuki S (2014) Characterization of Vibrio cholerae bacteriophages isolated from the environmental waters of the Lake Victoria region of Kenya. Curr Microbiol 68:64–70. https://doi.org/10.1007/s00284-013-0447-x
Maina A, Mwaura F, Jumba M, Kieft K (2021) Isolation, characterisation of four Gram-negative environmental bacteria and bacteriophage plaques formation on their lawns. Int J Pharm Bio Sci 11:121–129
Malik DJ, Sokolov IJ, Vinner GK et al (2017) Formulation, stabilisation and encapsulation of bacteriophage for phage therapy. Adv Colloid Interface Sci 249:100–133. https://doi.org/10.1016/j.cis.2017.05.014
Martiny AC, Coleman ML, Chisholm SW (2006) Phosphate acquisition genes in Prochlorococcus ecotypes: evidence for genome-wide adaptation. Proc Natl Acad Sci USA 103:12552–12557. https://doi.org/10.1073/pnas.0601301103
Mateus L, Costa L, Silva Y, Pereira C, Cunha A, Almeida A (2014) Efficiency of phage cocktails in the inactivation of Vibrio in aquaculture. Aquaculture 424:167–173. https://doi.org/10.1016/j.aquaculture.2014.01.001
McNair K, Aziz RK, Pusch GD, Overbeek R, Dutilh BE, Edwards R (2018) Phage genome annotation using the RAST pipeline. Methods Mol Biol 1681:231–238. https://doi.org/10.1007/978-1-4939-7343-9_17
Miller ES, Heidelberg JF, Eisen JA et al (2003) Complete genome sequence of the broad-host-range vibriophage KVP40: comparative genomics of a T4-related bacteriophage. J Bacteriol 185:5220–5233. https://doi.org/10.1128/jb.185.17.5220-5233.2003
Naser IB, Hoque MM, Abdullah A, Bari SMN, Ghosh AN, Faruque SM (2017) Environmental bacteriophages active on biofilms and planktonic forms of toxigenic Vibrio cholerae: potential relevance in cholera epidemiology. PLoS ONE 12:e0180838. https://doi.org/10.1371/journal.pone.0180838
Nelson EJ, Harris JB, Morris JG Jr, Calderwood SB, Camilli A (2009) Cholera transmission: the host, pathogen and bacteriophage dynamic. Nat Rev Microbiol 7:693–702. https://doi.org/10.1038/nrmicro2204
O’Flaherty S, Coffey A, Edwards R, Meaney W, Fitzgerald GF, Ross RP (2004) Genome of staphylococcal phage K: a new lineage of Myoviridae infecting Gram-positive bacteria with a low G+C content. J Bacteriol 186:2862–2871. https://doi.org/10.1128/jb.186.9.2862-2871.2004
Ou T, Gao XC, Li SH, Zhang QY (2015) Genome analysis and gene nblA identification of Microcystis aeruginosa myovirus (MaMV-DC) reveal the evidence for horizontal gene transfer events between cyanomyovirus and host. J Gen Virol 96:3681–3697. https://doi.org/10.1099/jgv.0.000290
Rambaut A (2007) FigTree, a graphical viewer of phylogenetic trees. Edinburgh, UK: Institute of Evolutionary Biology University of Edinburgh. http://tree.bio.ed.ac.uk/software/figtree/
Sambrook J, Russell DW, Russell DW (2001) Molecular cloning: a laboratory manual (3-volume set), 3rd edn. Cold Spring Harbor Laboratory Press, New York
Sen A, Ghosh AN (2005) New Vibrio cholerae O1 biotype ElTor bacteriophages. Virol J 2:28. https://doi.org/10.1186/1743-422x-2-28
Shah M (2014) An application of sequencing batch reactors in the identification of microbial community structure from an activated sludge. J Appl Env Microbiol 2:176–184. https://doi.org/10.12691/jaem-2-4-11
Silva-Valenzuela CA, Camilli A (2019) Niche adaptation limits bacteriophage predation of Vibrio cholerae in a nutrient-poor aquatic environment. Proc Natl Acad Sci USA 116:1627–1632. https://doi.org/10.1073/pnas.1810138116
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stenholm AR, Dalsgaard I, Middelboe M (2008) Isolation and characterization of bacteriophages infecting the fish pathogen Flavobacterium psychrophilum. Appl Environ Microbiol 74:4070–4078. https://doi.org/10.1128/aem.00428-08
Sulakvelidze A, Alavidze Z, Morris JG Jr (2001) Bacteriophage therapy. Antimicrob Agents Chemother 45:649–659. https://doi.org/10.1128/aac.45.3.649-659.2001
Sullivan MB, Coleman ML, Weigele P, Rohwer F, Chisholm SW (2005) Three Prochlorococcus cyanophage genomes: signature features and ecological interpretations. PLoS Biol 3:e144. https://doi.org/10.1371/journal.pbio.0030144
Sullivan MB, Huang KH, Ignacio-Espinoza JC et al (2010) Genomic analysis of oceanic cyanobacterial myoviruses compared with T4-like myoviruses from diverse hosts and environments. Environ Microbiol 12:3035–3056. https://doi.org/10.1111/j.1462-2920.2010.02280.x
Sullivan MJ, Petty NK, Beatson SA (2011) Easyfig: a genome comparison visualizer. Bioinformatics 27:1009–1010. https://doi.org/10.1093/bioinformatics/btr039
Talledo M, Rivera IN, Lipp EK et al (2003) Characterization of a Vibrio cholerae phage isolated from the coastal water of Peru. Environ Microbiol 5:350–354. https://doi.org/10.1046/j.1462-2920.2003.00411.x
Tetu SG, Brahamsha B, Johnson DA et al (2009) Microarray analysis of phosphate regulation in the marine cyanobacterium Synechococcus sp. WH8102. ISME J 3:835–849. https://doi.org/10.1038/ismej.2009.31
Turner D, Kropinski AM, Adriaenssens EM (2021) A roadmap for genome-based phage taxonomy. Viruses. https://doi.org/10.3390/v13030506
van Twest R, Kropinski AM (2009) Bacteriophage enrichment from water and soil. Methods Mol Biol 501:15–21. https://doi.org/10.1007/978-1-60327-164-6_2
Wang J, Jiang Y, Vincent M et al (2005) Complete genome sequence of bacteriophage T5. Virology 332:45–65. https://doi.org/10.1016/j.virol.2004.10.049
Wang X, Liu J, Yu Z, Jin J, Liu X, Wang G (2016) Novel groups and unique distribution of phage phoH genes in paddy waters in northeast China. Sci Rep 6:38428. https://doi.org/10.1038/srep38428
Weigele PR, Pope WH, Pedulla ML et al (2007) Genomic and structural analysis of Syn9, a cyanophage infecting marine Prochlorococcus and Synechococcus. Environ Microbiol 9:1675–1695. https://doi.org/10.1111/j.1462-2920.2007.01285.x
Yoshida T, Nagasaki K, Takashima Y et al (2008) Ma-LMM01 infecting toxic Microcystis aeruginosa illuminates diverse cyanophage genome strategies. J Bacteriol 190:1762–1772. https://doi.org/10.1128/jb.01534-07
Zeng Q, Chisholm SW (2012) Marine viruses exploit their host’s two-component regulatory system in response to resource limitation. Curr Biol 22:124–128. https://doi.org/10.1016/j.cub.2011.11.055
Acknowledgements
We wish to acknowledge David Goulding for help with TEM imaging. We are grateful for the generous financial assistance provided by Professor Elizabeth Kutter (The Evergreen State College, Olympia, WA, USA) and Tobi Nagel (Phages for Global Health) to phage researchers from Africa. The clinical Vibrio isolate was a kindly provided by the Department of Microbiology, University of Nairobi, Kenya.
Funding
RKA is funded by the Egyptian Academy for Scientific Research and Technology (ASRT) JESOR program (Project #3046). The funder has no interference with the scientific content of the article.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
None of the authors has a financial or personal conflict of interest to declare.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Maina, A.N., Mwaura, F.B., Jumba, M. et al. Novel PhoH-encoding vibriophages with lytic activity against environmental Vibrio strains. Arch Microbiol 203, 5321–5331 (2021). https://doi.org/10.1007/s00203-021-02511-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02511-3