Abstract
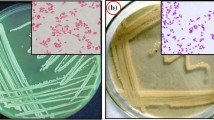
A gram-stain positive, aerobic, motile, rod-shaped bacterium, designated strain LAM7117T, was isolated from a sulfonylurea herbicides degrading consortium enriched with birch forest soil. The optimal temperature and pH for the growth of strain LAM7117T were 35 °C and 7.5, respectively. Strain LAM7117T could grow in the presence of NaCl with concentration up to 9% (w/v). Strain LAM7117T formed a distinct phylogenetic subclade within the genus Arthrobacter in the phylogenetic trees built with 16S rRNA gene sequences and shared the highest similarity with A. crystallopoietes JCM 2522T (97.7%). The values of digital DNA–DNA relatedness and Avery Nucleotide Identity based on the genome sequences between LAM7117T and A. crystallopoietes JCM 2522T were 21.4 and 77.4%, respectively. The genomic DNA G + C content was 65.9 mol%. The major cellular fatty acids were anteiso-C15:0, iso-C16:0 and anteiso-C17:0. The cell wall peptidoglycan contained the amino acids as glycine, lysine, alanine and glutamic acid. The major polar lipids present in strain LAM7117T were diphosphatidylglycerol, phosphatidylglycerol, phosphatidyl inositol, two unidentified glycolipids and one unidentified lipid. The predominant menaquinones of strain LAM7117T were MK-8 and MK-9. Based on the phenotypic characteristics, chemotaxonomic data and genotypic analyses, strain LAM7117T should be classified as a novel species of genus Arthrobacter, for which the name Arthrobacter sulfonylureivorans sp. nov. is proposed. The type strain is LAM7117T (= JCM 32824T = CGMCC 1.16681T).

Similar content being viewed by others
Abbreviations
- JCM:
-
Japan collection of microorganisms
- ANI:
-
Average nucleotide identity
References
Busse, HJ, Wieser M, Buczolits S (2012) Genus III Arthrobacter Conn & Dimmick 1947, 301AL emend. Koch, Schumann & Stackebrandt 1995, 838. In: Bergey’s manual of systematic bacteriology, 2nd edn, vol 5, pp 578–624
Chen YG, Tang SK, Zhang YQ, Li ZY, Yi LB (2009) Arthrobacter halodurans sp. nov., a new halotolerant bacterium isolated from sea water. Antonie Van Leeuwenhoek 96:63–70
Chen YG, Tang SK, Zhang YQ, Li ZY, Yi LB, Wang YX, Li WJ, Cui XL (2009) Arthrobacter halodurans sp. nov., a new halotolerant bacterium isolated from sea water. Antonie Van Leeuwenhoek 96:63–70
Cheng J, Zhang MY, Zhao JC, Xu H, Zhang Y (2017) Arthrobacter ginkgonis sp. nov., an actinomycete isolated from rhizosphere of Ginkgo biloba L. Int J Syst Evol Microbiol 67:319–324
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466
Collins MD, Shah HN (1984) Fatty acid, menaquinone and polar lipid composition of Rothia dentocariosa. Arch Microbiol 137:247–249
Conn HJ, Dimmick I (1947) Soil bacteria similar in morphology to Mycobacterium and Corynebacterium. J Bacteriol 54:291–303
Dong XZ, Cai MY (2001) Determinative manual for routine bacteriology. Scientific Press, Beijing
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Funke G, Hutson RA, Bernard KA, Pfyffer GE, Wauters G, Collins MD (1996) Isolation of Arthrobacter spp. from clinical specimens and description of Arthrobacter cumminsii sp. nov. Arthrobacter woluwensis sp. nov. J Clin Microbiol 34:2356–2363
Heyrman J, Verbeeren J, Schumann P, Swings J, De Vos P (2005) Six novel Arthrobacter species isolated from deteriorated mural paintings. Int J Syst Evol Microbiol 55:1457–1464
Kageyama A, Morisaki K, Omura S, Takahashi Y (2008) Arthrobacter oryzae sp. nov. Arthrobacter humicola sp. nov. Int J Syst Evol Microbiol 58:53–56
Kim KK, Lee KC, Oh HM, Kim MJ, Eom MK, Lee JS (2008) Arthrobacter defluvii sp. nov., 4-chlorophenol-degrading bacteria isolated from sewage. Int J Syst Evol Microbiol 58:1916–1921
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura MA (1980) Simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Lechevalier MP, Lechevalier HA (1980) The chemotaxonomy of actinomycetes. In: Actinomycete taxonomy (SIM special publication no. 6), pp 27–291
Li Y, Kawamura Y, Fujiwara N, Naka T, Liu H, Huang X, Kobayashi K, Ezaki T (2004) Rothia aeria sp. nov., Rhodococcus baikonurensis sp. nov. Arthrobacter russicus sp. nov., isolated from air in the Russian space laboratory Mir. Int J Syst Evol Microbiol 54:827–835
Ma JP, Wang Z, Lu P, Wang HJ, Waseem Ali S, Li SP, Huang X (2009) Biodegradation of the sulfonylurea herbicide Chlorimuron-ethyl by the strain Pseudomonas sp. LW3. FEMS Microbiol Lett 296:203–209
MacKenzie SL (1987) Gas chromatographic analysis of amino acids as the N-heptafluorobutyryl isobutyl esters. J Assoc Off Anal Chem 70:151–160
Minnikin DE, Odonnell AG, Goodfellow M, Alderson G, Athalye M (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Moraine RA, Rogovin P (1966) Kinetics of polysaccharide B-1459 fermentation. Biotechnol Bioeng 8:511–524
Pindi PK, Manorama R, Begum Z, Shivaji S (2010) Arthrobacter antarcticus sp. nov isolated from an Antarctic marine sediment. Int J Syst Evol Microbiol 60:2263–2266
Ruan ZY, Wang YW, Song JL, Jiang SH, Wang HM (2014) Kurthia huakuii sp. nov., isolated from biogas slurry, and emended description of the genus Kurthia. Int J Syst Evol Microbiol 64:518–521
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sakamoto M, Suzuki M, Umeda M, Ishikawa I, Benno Y (2002) Reclassification of Bacteroides forsythus as Tannerella forsythensis corrig., gen. nov., comb. nov. Int J Syst Evol Microbiol 52:841–849
SantaCruz-Calvo L, Gonzalez-Lopez J, Manzanera M (2013) Arthrobacter siccitolerans sp. nov., a highly desiccation-tolerant, xeroprotectant-producing strain isolated from dry soil. Int J Syst Evol Microbiol 63:4174–4180
Skerman VBD (1967) A guide to the identification of the genera of bacteria, vol 35, 2nd edn. Williams & Wilkins, Baltimore, pp 1–92
Skyring Quadling GWC (1970) Soil bacteria: a principal component analysis and guaninecytosine contents of some arthrobacter-coryneform soil isolates and of some named cultures. Can J Microbiol 16:95–106
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods forgeneral and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 607–654
Sun C, Fu GY, Zhang CY, Hu J, Xu L (2016) Isolation and complete genome sequence of Algibacter alginolytica sp. nov., a novel seaweed-degrading bacteroidetes bacterium with diverse putative polysaccharide utilization loci. Appl Environ Microbiol 82:2975–2987
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA7: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The clustal × windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tindall BJ, Sikorski J, Smibert RM, Krieg NR (2007) Phenotypiccharacterization and the principles of comparative systematics. In: Reddy CA, Beveridge TJ, Breznak JA, Marzluf G, Schmidt TM, Snyder LR (eds) Methods for general and molecular microbiology, 3rd edn. American Society for Microbiology, Washington, DC, pp 330–393
Wang HF, Li L, Zhang YG, Hozzein WN, Zhou XK, Liu WH, Duan YQ, Li WJ (2015) Arthrobacter endophyticus sp. nov., an endophytic actinobacterium isolated from root of Salsola affinis C. A. Mey. Int J Syst Evol Microbiol 65:2154–2160
Wayne LG, Moore WEC, Stackebrandt E, Kandler O, Colwell RR (1987) Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Evol Microbiol 37:463–464
Xiang W, Liu C, Wang X, Du J, Xi L (2011) Actinoalloteichus nanshanensis sp. nov., isolated from the rhizosphere of a fig tree (Ficus religiosa). Int J Syst Evol Microbiol 61:1165–1169
Xu XW, Huo YY, Wang CS, Oren A, Cui HL (2011) Pelagibacterium halotolerans gen. nov., sp. nov. Pelagibacterium luteolum sp. nov., novel members of the family Hyphomicrobiaceae. Int J Syst Evol Microbiol 61:1817–1818
Yassin AF, Sproër C, Siering C, Hupfer H, Schumann P (2011) Arthrobacter equi sp. nov., isolated from veterinary clinical material. Int J Syst Evol Microbiol 61:2089–2094
Acknowledgements
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain LAM7117T is MG009252. The GenBank accession number for the draft genome sequence of strain LAM7117T is RJAE01000000.
Funding
This work was supported by National Natural Science Foundation of China (NSFC, No. 31670006). This work was also supported by the National Key R&D Program of China (2017YFD0800702) and the Fundamental Research Funds for the Chinese Academy of Agricultural Sciences (Y2020GH14).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Han, X., Zhang, Q., Ma, Q. et al. Arthrobacter sulfonylureivorans sp. nov., isolated from a sulfonylurea herbicides degrading consortium enriched with birch forest soil. Arch Microbiol 203, 1039–1045 (2021). https://doi.org/10.1007/s00203-020-02097-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-02097-2