Abstract
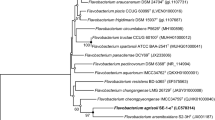
An aerobic, Gram-stain-negative, bright yellow-pigmented, oxidase and catalase-positive, non-motile, non-spore forming, rod-shaped strain designated DMN11T was isolated from the soil of crossroads of Jeju Island in South Korea. Colonies were circular, bright yellow-pigmented and smooth with regular edges and measured approximately 1–2 mm in diameter. Flexirubin-type pigments were absent. Phylogenetic tree analysis based on the 16SrRNA gene sequence revealed that the strain DMN11T formed a lineage within the family Flavobacteriaceae of the phylum Bacteroidetes, and it was the most closely related to Flavobacterium suzhouense XIN-1T and Flavobacterium hauense BX12T (98.6% and 98.2% similarity, respectively). The major isoprenoid quinone was MK-6. The major fatty acids were summed feature 3 (comprising C16:1ω7c and/or C16:1ω6c), iso-C15:0 and iso-C15:0 3OH. The polar lipid profile of the strain DMN11T showed the presence of phosphatidylethanolamine (PE) as major lipid. The DNA G+C content was 35.3 mol%, as determined by the thermal denaturation method. The mean levels of DNA–DNA relatedness of the strain DMN11T with F. suzhouense XIN-1T and F. hauense BX12T were 20.5% and 29.2%, respectively. Thus, the data accumulated in this study support the suggestion that the strain DMN11T is considered to represent a novel species of the genus Flavobacterieum, for which the name Flavobacterium edaphi sp. nov. is proposed. The type strain is DMN11T (= KCTC 62114T = JCM 32372T).

Similar content being viewed by others
References
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman J et al (eds) (1995) Short protocols in molecular biology: a compendium of methods from current protocols in molecular biology, 3rd edn. Wiley, New York
Bergey D, Harrison F, Breed R, Hammer B, Huntoon F et al (1957) Genus III. Flavobacterium. In: Breed RS (ed) Bergey’s manual of determinative bacteriology. Williams & Wilkins, Baltimore, pp 309–322
Bernardet JF, Bowman JP (2006) The genus Flavobacterium. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes: a handbook on the biology of bacteria, vol 7, 3rd edn. Springer, New York, pp 481–531
Bernardet JF, Bowman J (2011) Genus I. Flavobacterium. In: Whitman W (ed) Bergey’s manual of systematic bacteriology, vol 4, 2nd edn. Williams & Wilkins, Baltimore, pp 112–154
Bernardet J-F, Nakagawa Y, Holmes B (2002a) Proposed minimal standards for describing new taxa of the family Flavobacteriaceae and emended description of the family. Int J Syst Evol Microbiol 52:1049–1070
Bernardet JF, Nakagawa Y, Holmes B, Subcommittee on the taxonomy of Flavobacterium and Cytophaga-like bacteria of the International Committee on Systematics of Prokaryotes (2002b) Proposed minimal standards for describing new taxa of the family Flavobacteriaceae and emended description of the family. Int J Syst Evol Microbiol 52: 1049–1070
Breznak JA, Costilow RN (2007) Physicochemical factors in growth. In: Beveridge TJ, Breznak JA, Marzluf GA, Schmidt TM, Snyder LR (eds) Methods for general and molecular bacteriology, 3rd edn. American Society for Microbiology, Washington, DC, pp 309–329
Buck JD (1982) Nonstaining (KOH) method for determination of Gram reactions of marine bacteria. Appl Environ Microbiol 44:992–993
Collins MD, Jones D (1981) Distribution of isoprenoid quinone structural types in bacteria and their taxonomic implication. Microbial Rev 45:316–354
De Ley J, Cattoir JH, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Dong K, Xu B, Zhu F, Wang GJ (2013) Flavobacterium hauense sp. nov., isolated from soil, and emended descriptions of the genus Flavobacterium subsaxonicum, Flavobacterium beibuense and Flavobacterium rivuli. Int J Syst Evol Microbiol 63:3237–3242
Fautz E, Reichenbach H (1980) A simple test for flexirubin-type pigments. FEMS Microbiol Lett 8:87–91
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fu Y, Tang X, Lai Q, Zhang C, Zhong H, Li W, Liu Y, Chen L, Sun F, Shao Z (2011) Flavobacterium beibuense sp. nov., isolated from marine sediment. Int J Syst Evol Microbiol 61:205–209
Gillis M, De Ley J, Cleene M (1970) The determination of molecular weight of bacterial genome DNA from renaturation rates. Eur J Biochem 12:143–153
Gonzalez JM, Saiz-Jimenez C (2002) A fluorimetric method for the estimation of G+C mol% content in microorganisms by thermal denaturation temperature. Environ Microbiol 4:770–773
Hiraishi A, Ueda Y, Ishihara J, Mori T (1996) Comparative lipoquinone analysis of influent sewage and activated sludge by high performance liquid chromatography and photodiode array detection. J Gen Appl Microbiol 42:457–469
Huang L, Zhou J, Li X, Peng Q, Lu H et al (2013a) Charactrerization of new alginate lyase from newly isolated Flavobacterium sp. S20. J Ind Microbiol Biotechnol 40:113–122
Huang L, Zhou J, Li X, Peng Q, Lu H, Du Y (2013b) Characterization of a new alginate lyase from newly isolated Flavobacterium sp. S20. J Ind Microbiol Biotechnol 40:113–122
Kaur I, Kaur C, Khan F, Mayilraj S (2012) Flavobacterium rakeshii sp. nov., isolated from marine sediment, and emended description of Flavobacterium beibuense Fu et al. 2011. Int J Syst Evol Microbiol 62:2897–2902
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH et al (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbial 62:716–721
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Komagata K, Suzuki KI (1987) Lipid and cell-wall analysis in bacterial systematics. Methods Microbiol 19:161–205
Kumar S, Stecher G, Tamura K (2016) Mega 7: Molecular evolutionary genetics analysis in version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Kuykendall LD, Roy MA, O’Neill JJ, Devine TE (1988) Fatty acids, antibiotic resistance and deoxyribonucleic acid homology groups of Bradyrhizobium japonicum. Int J Syst Evol Microbiol 38:358–361
Li DD et al (2017) Flavobacterium arcticum sp. nov., isolated from Arctic seawater. Int J Syst Evol Microb 67:1070–1074
Loveland-Curtze J, Miteva VI, brenchley JE (2011) Evaluation of new flurometric DNA-DNA hybridization method. Can J Microbiol 57:250–255
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Miyashita M, Fujimura S, Nakagawa Y, Nishizawa M, Tomizuka N, Nakagawa T, Nakagawa J (2010) Flavobacterium algicola sp. nov., isolated from marine algae. Int J Syst Evol Microbiol 60:344–348
Rzhetsky A, Nei M (1992) A simple method for estimating and testing minimum-evolution trees. Mol Biol Evol 9:945–967
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, DC, pp 607–654
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tindall BJ, Sikorski J, Smibert RM, Kreig NR (2007) Phenotypic characterization and the principles of comparative systematics. In: Reddy CA, Beveridge TJ, Breznak JA, Marzluf G, Schmidt TM, Snyder LR (eds) Methods for general and molecular microbiology, 3rd edn. American Society for Microbiology, Washington, DC, pp 330–393
Touchon M, Barbier P, Bernardet JF, Loux V, Vacherie B, Barbe V, Rocha EPC, Duchaud E (2011) Complete genome sequence of the fish pathogen Flavobacterium branchiophilum. Appl Environ Microbiol 77:7656–7662
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Xu M, Xin Y, Tian J, Dong K, Yu Y, Zhang J, Liu H, Zhou Y (2011) Flavobacterium sinopsychrotolerans sp. nov., isolated from a glacier. Int J Syst Evol Microbiol 61:20–24
Zang H, Cheng MG, Sun B, Guo SH, Song M et al (2015) Flavobacterium suzhouense sp. nov., isolated from farmland river sludge. Int J Syst Evol Microbiol 65:370–374
Acknowledgements
We thank Prof Dr. Bernhard Schink (University of Konstanz, Konstanz, Germany) for the suggested genus and species names. This work was supported by the National Research Foundation of Korea (NRF) Grant funded by the Korea government (MSIT) (NRF-2017R1A2B4009448).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest.
Additional information
Communicated by Erko Stackebrandt.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chhetri, G., Yang, D., Choi, J. et al. Flavobacterium edaphi sp. nov., isolated from soil from Jeju Island, Korea. Arch Microbiol 201, 539–545 (2019). https://doi.org/10.1007/s00203-018-1593-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-018-1593-0