Abstract
DNA acquisition promotes the spread of resistance to antibiotics and virulence among bacteria. It is also linked to several natural phenomena including recombination, genome dynamics, adaptation and speciation. Horizontal DNA transfer between bacteria occurs via conjugation, transduction or competence for natural transformation by DNA uptake. Among these, competence is the only mechanism of transformation initiated and entirely controlled by the chromosome of the recipient bacteria. While the molecular mechanisms allowing the uptake of extracellular DNA are increasingly characterized, the function of competence for natural transformation by DNA uptake, the selective advantage maintaining it and the reasons why bacteria take up DNA in the first place are still debated. In this synthesis, I review some of the literature and discuss the four hypotheses on how and why do bacteria take up DNA. I argue that DNA uptake by bacteria is an accidental by-product of bacterial adhesion and twitching motility. Adhesion and motility are generally increased in stressful conditions, which may explain why bacteria increase DNA uptake in these conditions. In addition to its fundamental scientific relevance, the new hypothesis suggested here has significant clinical implications and finds further support from the fact that antibiotics sometimes fail to eliminate the targeted bacterium while inevitably causing stress to others. The widespread misuse of antibiotics may thus not only be selecting for resistant strains, but may also be causing bacteria to take up more DNA with the consequent increase in the chances of acquiring drug resistance and virulence—a scenario in full concordance with the previously reported induction of competence genes by antibiotics in Streptococcus pneumoniae and Legionella pneumophila.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Acquisition of new genetic material by bacteria occurs by transduction, conjugation or competence for natural transformation by uptake of extracellular DNA (hereafter referred to simply as competence). These mechanisms of horizontal gene transfer (HGT) are sources of genetic diversity in the otherwise mostly clonal world of bacteria and can even interfere with speciation by blurring species boundaries [see (Fraser et al. 2007)]. HGT is central to the spread of virulence and resistance to antibiotics among human and livestock pathogens (Vazquez et al. 2001; Courvalin 2005; Sabia et al. 2005; Bartlett 2006; Diep et al. 2006; Messi et al. 2006) (e.g. Methicillin-resistant Staphylococcus aureus, Vancomycin-resistant enterococci, including Clostridium difficile, and more virulent/resistant Neisseria meningitides and Neisseria gonorrhoeae). Indeed, the relatively recent deaths caused by a chimeric, virulent and multidrug-resistant Escherichia coli strain in Europe are just a small reminder of the serious implications of such natural phenomenon. Neither conjugation nor transduction are initiated by the recipient cell, and HGT through those DNA transfer mechanisms is a side effect of plasmid transmission (which are symbiotic, accessory or selfish elements) and infection by bacteriophages (which are parasites). Contrarily, the chromosome of the recipient cell initiates and entirely controls competence, making this the only mechanism through which acquisition of foreign DNA molecules is not perceived as side effect (Redfield 2001).
Discovered by Frederick Griffith (Griffith 1928) and best defined by David Dubnau as ‘a genetically programmed physiological state permitting the efficient uptake of macromolecular DNA’ (Dubnau 1999), competence is a complex heterogeneous phenomenon. Indeed (1) most bacteria were never reported as competent, whereas some bacteria are known to develop such state. (2) Competence can be constitutive [e.g. Neisseria meningitidis and N. gonorrhoeae (Catlin 1960; Biswas et al. 1977)] or occasional [e.g. Bacillus subtilis, Haemophilus influenzae and Streptococcus pneumoniae (Singh and Pitale 1968; Herriott et al. 1970; Havarstein et al. 1995)]. (3) Its inducers are species specific [e.g. temperature and media alterations (Herriott et al. 1970; Ranhand 1976; Lopez et al. 1983; Auzat et al. 1999; MacFadyen et al. 2001)]. (4) Depending on the species, the fraction of competent cells varies from ≤25 % [e.g. Pseudomonas stutzeri, Streptomyces virginiae, Streptomyces kasugaensis, B. subtilis and Acinetobacter calcoaceticus (Roelants et al. 1976; Smith et al. 1981; Lorenz et al. 1992; Palmen et al. 1993)], to ~100 % [e.g. Azotobacter vinelandii, H. influenzae and S. pneumoniae (Smith et al. 1981; Glick et al. 1985)]. (5) Most competent species take up DNA indiscriminately, but some ‘prefer’ con-specific DNA [see (Lorenz and Wackernagel 1994; Dubnau 1999)]. (6) In competent Pasteurellaceae and Neisseriaceae species, the ‘preference’ toward the bacterium’s own species DNA requires that the incoming DNA contains short specific sequences called DNA uptake enhancing sequences (DUES) (Sisco and Smith 1979; Graves et al. 1982; Bakkali 2007). In A. vinelandii, Campylobacter coli and P. stutzeri, however, the preferential uptake of con-specific DNA requires no DUES (Doran and Page 1983; Wang and Taylor 1990).
Except from few differences relating to the presence/absence of an outer membrane, development of competence is similar between Gram-positive and Gram-negative bacteria and consists of (1) DNA binding, (2) exonuclease degradation of its 3′ strand simultaneous to (3) the interiorization of the 5′ strand, and finally, (4) digestion of the incoming single stranded DNA in the cytoplasm or, occasionally, its recombination with the cell’s chromosome (Lorenz and Wackernagel 1994; Dubnau 1999; Chen and Dubnau 2003; Chen et al. 2005). Research on the genetic control of the competent state shows that it involves a complex network of genes [see (Dubnau 1999; Chen and Dubnau 2003; Chen et al. 2005)] controlling a variety of processes including oxidative, nucleic acids and sugar metabolism (Macfadyen et al. 1996; Auzat et al. 1999; MacFadyen et al. 2001), DNA recombination and repair (Raymond-Denise and Guillen 1992; Kruger et al. 1997; Mortier-Barriere et al. 1998), quorum sensing (Li et al. 2002), heat shock (Turgay et al. 1997) and secretion systems (Chen and Dubnau 2003; Averhoff 2004).
In contrast with the scientific advances and consensus on the nature of the molecular basis and series of events that take place during competence, the origin, reasons of being, maintenance and evolution of this phenomenon are still subjects of ongoing debate. The phylogenetic origin and distribution of competence are hitherto undefined. Lorenz and Wackernagel reported ~40 naturally competent species (Lorenz and Wackernagel 1994), and since then, ~20 more were added, and this number will certainly keep increasing. Competence could therefore be more common than we currently know. Its scattered distribution across taxa ostensibly suggests independent evolution. However, scattering could result from bias in research and from the small number of species reported as competent compared to the number of phylogenetic branches. In fact, the likelihood of such a complex phenomenon independently evolving in several lineages is negligible. Competence is probably an ancestral feature of at least most lineages—a possibility supported by findings in (Redfield et al. 2006). Species not known as competent could therefore be competent under unknown conditions or may have lost this ability. For instance, Pseudomonas fluorescens and Agrobacterium tumefaciens seem competent in nature but not under laboratory conditions (Demaneche et al. 2001), whereas Pasteurella multocida was never reported as competent although it belongs to a family of competent species and has the genes involved in competence and the DUES ‘preferred’ by H. influenzae (Bakkali et al. 2004).
To explain the evolution and maintenance of competence, four hypotheses were suggested. Each of these was the result of neat logic and great intellectual formulations and was supported by elegantly designed experiments (see references herein). Nonetheless, the matter is still subject of controversy as none of these hypotheses can explain all the cases of competent species. A new hypothesis is therefore needed to provide a unifying explanation and account for the weaknesses on which I will focus my following discussion of each of the four existing hypotheses.
Competence for chromosome repair (Michod et al. 1988) relies on the up-regulation of DNA repair genes in competent B. subtilis (Love et al. 1985) and the increase in its transformation and survival when cells are UV treated before adding DNA to the medium, but not after (Michod et al. 1988). However, it is argued that DNA repair cannot explain the increase in survival, that extracellular DNA is often damaged and useless for chromosome repair, and that DNA damaging agents do not induce competence neither in H. influenzae nor in B. subtilis (Mongold 1992; Redfield 1993a). Yet, there is a plethora of situations where most extracellular DNA will not be damaged (e.g. in host tissues or in soil). Nonetheless, to me, the repair hypothesis has at least three major weaknesses: First, most of the DNA taken up is eliminated/degraded (Lorenz and Wackernagel 1994; Dubnau 1999; Chen and Dubnau 2003; Chen et al. 2005). Although no energy cost is too high if the final outcome is survival, eliminating one DNA strand at the cell surface means spending energy and a 50 % reduction in the chances of successful chromosome repair. Subsequent cytoplasm-localized degradation of the DNA means further energy cost and reduction in the repair material that DNA might be—why should bacteria ‘waste’ that repair material as well the energy required for degrading it if what they want is chromosome repair? It is true that S. pneumoniae, L. pneumophila and probably other competent bacteria seem to lack the standard SOS DNA repair system, so that any contribution to the repair system might be selected for, many competent bacteria already possess the standard DNA repair mechanisms and, in principle, would not need competence for repair. Second, it is DNA uptake and its entry into the cytoplasm what induces the DNA repair machinery and not the reverse—up-regulation of the DNA repair genes seems to be an SOS response to the presence of single stranded DNA inside the cell. Third, and probably more importantly, uptake of DNA that corresponds to the damaged chromosomal region is so unlikely for it to explain the evolution of competence, especially since most competent bacteria take up DNA indiscriminately and not only from their own species. One may argue that restriction–modification and cytoplasmic nucleases would eliminate the non-homologous DNA. The benefit of competence as repair mechanism would then depend on the prevalence of the homologous DNA compared to the non-homologous one. In high population densities, one could expect the DNA from the same species to be frequent but, in a non-homogenous medium and in hosts, most of the DNA would come from other species (for instance, most of the DNA in human lung would be human and not from the pathogen). Furthermore, even if the homologous were frequent enough, it still must contain the damaged region of the chromosome that the bacterium seeks to repair and this would depend on the length of the DNA fragment. It is true that bacteria can take up large DNA fragments but, outside the cell, DNA tends to be degraded and fragmented due to the action of the many agents, chemical and physical, as well as to the different enzymes secreted by other living beings. Indeed, even if the DNA taken up was always from the own species and as big as 20 kb and that the bacterial genome were small as 1 Mb, in 98 % of the times, the competent bacterium will take up DNA that would be useless for repairing its chromosome. In a way, one may make the analogy between taking up random DNA fragments for chromosome repair and buying random mechanical pieces with the hope that one might repair a damaged car engine.
Competence for recombination is a widely accepted idea recognizing competence as a mechanism for genetic diversification comparable to eukaryotic sex [see (Narra and Ochman 2006)]. DUESs could therefore be mate recognition tags (Redfield 1991) preventing interspecific HGT [e.g. DUESs would prevent H. influenzae from taking N. gonorrhoeae DNA and vice versa (Sisco and Smith 1979; Graves et al. 1982)]. However, to our current knowledge, DUESs are limited to some species of the Pasteurellaceae and Neisseriaceae families (Sisco and Smith 1979; Graves et al. 1982; Bakkali et al. 2004; Redfield et al. 2006; Bakkali 2007). They are not species specific, hence unable to prevent interspecific DNA uptake between species as distant as P. multocida, H. influenzae, Actinobacillus actinomycetemcomitans and Mannheimia succiniciproducens—these and other species have the same DUES (Bakkali et al. 2004; Redfield et al. 2006). Furthermore, it is argued that the extracellular DNA derives from dead cells and is therefore more likely to reduce fitness than confer advantage (Redfield 1988; Redfield et al. 1997). It is hard to see the point of bacteria taking up DNA for recombination to face challenging conditions if the DNA comes from dead cells that could not cope with those conditions (consider antibiotic treatment situations). One may argue that some of the extracellular DNA does not come from dead cells but is secreted by living ones or comes from cells actively killed by the competent bacteria. But DNA secretion and/or cell killing does not seem to be general characteristics of all competent bacteria, and the secreted DNA, or DNA from killed cells, is not as abundant as the one from dead cells. In addition, recombination of the strong (the killer) bacterium with DNA from the weak (the killed) bacterium is selectively counterproductive, and, for the secreted DNA to be of any selective use, ‘generous’ bacteria must ‘kindly’ secrete DNA that contains the advantage-conferring gene/mutation, so that others bacteria can take it up and use it to survive multiply and compete. This must be one of the utmost altruistic group selection situations! Another situation where extracellular DNA is not primarily from dead cells is biofilms. However, there, the DNA plays more a role of an aggregating matrix rather than a source of information. In addition, as with the chromosome repair hypothesis, I do not see why bacteria should degrade most of the DNA they bind to if what they want is recombination—why waste DNA and energy? Also, bacteria have no mate choice or sexual selection, and while unlikely to integrate non-homologous DNA, most competent bacteria take up DNA indiscriminately [see (Lorenz and Wackernagel 1994; Dubnau 1999)], even from eukaryotes! [e.g. Acinetobacter sp., P. stutzeri and Ralstonia solanacearum take up plant DNA (Bertolla et al. 2000; de Vries et al. 2001)]. What about the cost of taking up free bacteriophage DNA? And the ‘suicide plasmid’ (Li et al. 2001)? Competence allows uptake of DNA from dead cells, even of other species, it can thus hardly be seen as bacterial sex.
The Competence for nutrition hypothesis (Redfield 1993b) suggests that bacteria take up DNA for its nucleotides and has been adopted by different works [e.g. (Macfadyen et al. 1996; MacFadyen et al. 2001; Palchevskiy and Finkel 2006)]. Competence induction by starving H. influenzae (Herriott et al. 1970) and its reversion by adding nucleic acids precursors to the medium (MacFadyen et al. 2001) support this hypothesis. But, why should some species, including H. influenzae, ‘prefer’ con-specific DNA (Sisco and Smith 1979; Graves et al. 1982)? Why do they ignore DNA of other species? In fact, this ‘diet’ restriction discounts a ‘safer’ DNA, as con-specific DNA is more likely to recombine with the chromosome, while it could be damaged and/or coming from dead (i.e. unfit and aged) cells. Also, why should bacteria discard one DNA strand (i.e. 50 % of the potentially nutritive nucleotides)? Why does competence develop even in complex media [e.g. A. calcoaceticus, N. meningitidis and N. gonorrhoeae (Catlin 1960; Biswas et al. 1977; Palmen et al. 1993)]? Furthermore, A. calcoaceticus, B. subtilis, N. gonorrhoeae and S. pneumoniae release DNA to the medium while being competent (Lorenz et al. 1991; Moscoso and Claverys 2004; Hamilton et al. 2005). Similarly, anaerobic and heat shifts induce competence in B. subtilis, S. pneumoniae and even H. influenzae (Goodgal and Herriott 1961; Espinosa et al. 1980; Chapuy-Regaud et al. 2001) could stressed bacteria become hungry because of the stress? Competence involves DNA uptake from dead conspecifics and discharge of >50 % of the DNA bound. If it is a nutritional mechanism, it must be rather abnormal and evoking nutritional anomalies—namely cannibalism and bulimia.
Recently, competence was suggested to be sustained in B. subtilis by episodic selection in favor of non-growing cells in conditions where dividing cells are killed (Johnsen et al. 2009). At first, maintenance based on episodic situations where competence may confer advantage seems a priori a sensible possibility that may potentially work for whatever function competence may have. However, Johnsen et al.’s work (Johnsen et al. 2009) is based on B. subtilis, which is only occasionally (episodically) competent. Furthermore, when this species is competent, only ~20 % of its cells take up DNA (the environmental strains are even less competent than that [see (Young et al. 1963; Nijland et al. 2010) and references therein]. It is still to be seen whether episodic selection could apply to other species—especially the ones constitutively competent and those where ~100 % of the cells take up DNA. Similarly, this hypothesis is still to be tested with selective forces other than strong selections against dividing cells. Indeed, competence inducers such as heat, nutritional and oxidative stresses are expected to affect the cells ability to divide but would not kill the dividing cells. In any case, Johnsen et al.’s work (Johnsen et al. 2009) relies on deterministic modeling supplemented by tightly controlled (see unrealistic) experiments and even for the species tested (B. subtilis) the selective advantage it defends seems more in favor of non-growth in anti-growth conditions than of DNA uptake. The latter is suggested to enhance the advantage of non-growth (hence, the episodic selection hypothesis is a recombinational one), but the authors do not take into account that the DNA taken up from the environment could also be neutral, deleterious or parasitic (bacteriophage). They assume that the DNA taken up is always positively selected (80 % fitness increase) and abundant (for each cell, there is a good DNA molecule)—a clearly unrealistic situation. If a bacterium stops dividing in an antibiotic-containing environment, it has a chance to resist and survive and the last thing it should do is to gamble and take up DNA from that environment as most likely that DNA is from a dead antibiotic sensitive cell.
In my opinion, for competence to be a nutritional or a recombination/sex mechanism, at least two conditions must be met: (1) other cross-membrane mechanisms for metabolites transport or HGT must be insufficient for conferring adequate food supply or genetic diversification to bacteria (otherwise competence will not be needed and selection would not favor it), and (2) DNA uptake must not be accidental. In minimal media, competence may, at first, seem needed to acquire nucleotides from DNA. However, B. subtilis and S. pneumoniae develop competence although they secrete non-specific DNases that should degrade extracellular DNA and make its nucleotides available for cross-membrane transport (Burke 1982; Moscoso and Claverys 2004). As to genetic diversification, although competence seems ancestral and commoner than we currently know, it is less frequently reported than conjugation and transduction; meaning that its contribution to HGT might be lower. Currently, no single hypothesis can by itself explain a function for competence, nor how natural selection can favor it, without generalization of findings that might be specific to a single or group of species. As if we believed that every natural phenomenon needs to have a function for it to exist, we strive to pinpoint one for competence. Surprisingly, the subjacently assumed non-accidental nature of DNA uptake was never challenged. Understanding why bacteria bind and take up DNA in the first place could therefore be helpful.
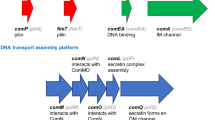
DNA uptake involves type IV pili (T4P) [e.g. (Rudel et al. 1995a, b; Wolfgang et al. 1998; Chen and Dubnau 2003; Averhoff 2004; Varga et al. 2006)] and the less characterized but related structures called ‘pseudo-pili’ (Chen and Dubnau 2003; Chen et al. 2005). The well-characterized T4P, on which I will focus, are adhesion and motility structures that function by (1) polymerization of pilin subunits [pilA (Wu and Kaiser 1997), pilE in Neisseria (Fyfe et al. 1995)] assisted by the ATPase pilB [pilF in Neisseria (Lauer et al. 1993)], (2) secretion of the polymerized rod through the secretin pilQ (Collins et al. 2001), (3) adhesion of the T4P to a surface—normally via its tip, pilC (Rudel et al. 1995a, b; Kirchner and Meyer 2005)—and (4) its retraction by depolymerization involving the ATPase pilT (Merz et al. 2000). This sequence of events allows adhesion to surfaces, including host tissues [e.g. (Kubiet et al. 2000; Kirchner and Meyer 2005)], and its repetition using different T4P, normally located at alternate cell poles [e.g. (Mignot et al. 2005, 2007; Margolin 2006)], allows movement [see (Mattick 2002; Nudleman and Kaiser 2004; Christie et al. 2005)].
T4P is ~3–4 μm long and ~60 Å wide and has no central canal (Forest and Tainer 1997) (Fig. 1); so DNA cannot pass through it, and to enter the cell, it needs to cross the membranes through the ~6.5 ηm diameter pilQ pore (Collins et al. 2001). An agreed-upon mechanism describing DNA uptake was suggested in (Dubnau 1999; Chen and Dubnau 2003, 2004; Chen et al. 2005). But why and how do bacteria bind DNA in the first place? Although never directly stated, from the different hypotheses on the function and maintenance of competence, one can infer that there seems to be a de facto assumption that bacteria may actively seek DNA. But there is no evidence or indication to support such possibility. In fact, there may be evidence to supporting the more logical passive DNA binding possibility as there is at least one bacterium, A. vinelandii, that seems to bind DNA even when not competent (Doran and Page 1983). Therefore, the null (by-default) hypothesis (i.e. bacteria bind DNA just by ‘accident’) might still stand. In such case, with T4P being adhesion and motility structures, DNA uptake may as well be a by-product of these motility and adhesion functions.
Obeying the laws of physics, bacteria move twitchingly if they adhere to surfaces that have higher inertia than the bacterial cell itself (Fig. 2). However, if T4P adheres to a DNA fragment that is smaller and has less mass and electrostatic energy than the bacterial cell, its retraction should pull the DNA toward the cell rather than the cell toward the DNA (Fig. 2). Once at the pilQ pore, the DNA [~2 ηm diameter and ~50 ηm bend-persistence length (Manning 2006)] would kink and form a loop ~43.5 ηm wider than the pore. The T4P and the pore would thus be blocked. This resembles an imaginary 12 cm round arm pulling a 4 cm diameter 1 m bend-persistence length tube through a 13 cm diameter window. Obstruction of the T4P and/or the membrane pore could be a selecting force for presence of the membrane-localized nucleases that reduce DNA stiffness by eliminating one of its strands [e.g. endA, nucA and nucT (Provvedi et al. 2001; Berge et al. 2002; O’Rourke et al. 2004)]. While the nuclease is located at the only membrane, Gram-positive bacteria have and they necessarily degrade and release the DNA strand into the medium; for Gram-negative bacteria, only inner membrane nucleases are thus far experimentally detected (and it releases the degraded DNA into the periplasm). Outer membrane nucleases in Gram-negative bacteria are thus far just a theoretical possibility that is neither experimentally proven nor excluded. Still, whether the impasse happens at the inner or at the outer/only membrane, the outcome is the same; a blocked pilus and pore structures. Bacteria might hence be accidentally binding and taking up DNA as they try to move (e.g. when escaping stressful conditions [Kozlovsky et al. 1999; Lacasta et al. 1999; Tang et al. 2004; Nachin et al. 2005; Wang et al. 2006; Gomez-Gomez et al. 2007)] (Fig. 3). Indeed, bacterial pili are known to bind and introduce ‘unwanted things’ into the cell (e.g. plasmids and bacteriophage DNA [Bradley 1975; Karaolis et al. 1999)], so DNA binding and uptake during competence could be just another accidental by-product of the pili extension and retraction. One could thus speculate that bacteria not known as competent could be competent less frequently or under strict conditions, might not use T4P or similar structures for adhesion and twitching motility, their adhesins’ specificity might exclude DNA or might have membrane-localized, periplasmic and/or cytoplasmic 5′-3′ and 3′-5′ nucleases that degrade any DNA bound or taken up—so that the DNA end up with no possibility of entering the cell and/or surviving in the protoplasm to recombine with the chromosome and, thus, transform the bacterium. Indeed, the reducing effect of the nucleases on bacteria′s transformation by DNA uptake capacity is demonstrated by the increase in transformability of the cyanobacterium Synechocystis sp. after elimination of its cytoplasmic exonuclease, RecJ (Kufryk et al. 2002).
Schematic explanation of the accidental nature of DNA binding by type IV pili of the motile/adhering bacteria and the possible outcomes with regard to DNA uptake and bacterial transformability. The interrogation marks signal options that while possible remain speculative at this point as they are still not experimentally demonstrated
When facing unfavorable conditions, bacteria would be stressed and could increase their motility [e.g. (Kozlovsky et al. 1999; Lacasta et al. 1999; Jelsbak and Sogaard-Andersen 2000; Tang et al. 2004; Nachin et al. 2005; Wang et al. 2006; Gomez-Gomez et al. 2007)]; meaning that T4P extension and retraction cycles increase and so should be the likelihood of DNA binding and uptake. Accordingly, T4P respond to factors like quorum sensing (Kendall and Sperandio 2007), pH (Brener and DeVoe 1983; Manetti et al. 2010), and temperature (Liles et al. 1998), and T4P-dependent motility is light regulated and linked to transformation in Synechocystis (Bhaya et al. 2000; Yoshihara et al. 2002). Indeed, competence was suspected to be a general SOS stress response (Li et al. 2002; Claverys et al. 2006; Charpentier et al. 2011). It is even deadly to a sub-fraction of S. pneumoniae (Steinmoen et al. 2002)—probably because this species produces a competence-stimulating peptide that is also antibacterial (Oggioni et al. 2004). Antibiotics misuse—which does not always eliminate the target while stressing non-targeted bacteria—might therefore not only be selecting for resistant strains, but also increasing DNA uptake and the likelihood of acquiring drug resistance and virulence. This seems true at least for S. pneumoniae and L. pneumophila where antibiotics treatment induces competence (Prudhomme et al. 2006; Charpentier et al. 2011).
The hypothesis suggested here tackles the weaknesses of the previous ones and explains experimental observations including (1) instead of taking full advantage of the DNA, bacteria get rid of as much of it as they probably can (they therefore seem not to want it), (2) A. vinelandii seem to bind DNA even when not competent (Doran and Page 1983) (a clear case of accidental DNA binding), (3) extracellular DNA is important for biofilm formation (Whitchurch et al. 2002) (the case of biofilms could be relevant for the current hypothesis as it shows that bacteria can stick to DNA not only for the purpose of taking it up. Hence, at least in this occasion, DNA seems an adhesion substrate rather than a genetic information or nutritional material), (4) DNA is a major constituent of the neutrophil extracellular bacteria traps (Brinkmann et al. 2004) (that probably exploit the accidental binding of DNA by bacteria to entrap and phagocyte them), and (5) all competence inducers cause stress (e.g. starvation, heat, antibiotics) that increases twitching motility and, thus, pili activity (polymerization and retraction) hence the consequent possibility of DNA binding, uptake, and the following degradation or occasional integration into the chromosome [e.g. (Kozlovsky et al. 1999; Lacasta et al. 1999; Jelsbak and Sogaard-Andersen 2000; Tang et al. 2004; Nachin et al. 2005; Wang et al. 2006; Gomez-Gomez et al. 2007)]. (6) Transformation and adhesion are linked in H. pylori (Lin et al. 2006).
The inhibition of competence in H. influenzae by adding nucleotide precursors to the medium (Macfadyen et al. 1996, 2001), the phase variation in DNA uptake in some species, the up-regulation of several cytoplasmic proteins—especially nucleases and DNA repair ones—and the overrepresentation of DUESs in the genomes of some species with preferential uptake of con-specific DNA (Sisco and Smith 1979; Graves et al. 1982; Bakkali et al. 2004) seem to contradict the hypothesis suggested here. However, addition of nucleotide precursors relieves the starvation stress that induces competence in H. influenzae, which may reduce cell motility and accidental DNA binding and uptake. On the other hand, phase variation in DNA uptake is due to variation in T4P production and states of adhesion and motility [e.g. (Villar et al. 2001)]. Similarly, nucleases, DNA repair proteins and other membrane-localized and cytoplasmic proteins are not exclusive to the competence state [e.g. EndA (Claverys et al. 2006) and SsbB (Thanassi et al. 2002)] or to competent species [e.g. DprA (Smeets et al. 2006) and RecA (Sung and Klein 2006)]. They may, thus, not have evolved as competence proteins and might be better seen as SOS response systems for dealing with DNA binding or interiorization into the cell as part of their role in secretion, transport, enzymatic, chromosome repair and/or defense against ‘parasitic’ DNA systems. On the other hand, proteins such as B. subtilis ComEC and ComEA have thus far no other demonstrated function but in competence. However, these proteins were detected in only few species and are not fully studied as to discard their involvement in other processes. In fact, at least ComEC is orthologous to H. influenzae Rec2, which is also involved in phage recombination (Kupfer and McCarthy 1992), S. pneumoniae CelB, which is also involved both in biofilm formation and virulence (Wu et al. 2012) and in the use of and growth on cellobiose (Zeng and Burne 2009; Shafeeq et al. 2011), and to N. gonorrhoeae ComA gene, which is known to be involved in pilin variation (Facius and Meyer 1993). All in all, while the membrane-localized proteins (such as the Com and Nup proteins are constitutive parts of these structures, the cytoplasmic ones [for a review see (Claverys et al. 2009)] would be up-regulated upon detection of foreign and damaged DNA in the cytoplasm no matter what its origin could be, even phage DNA and DNA artificially introduced by electroporation. Once the DNA has been accidentally bound, the cell uses different proteins to deal with the situation. This would explain why in B. subtilis entry of exogenous DNA occurs at the cell poles (Hahn et al. 2005; Chen et al. 2006; Hahn et al. 2009), where the pseudo-pili as well as the competence-related proteins are located (Kaiser 1979; Henrichsen 1983; Kaiser 2000; Mattick 2002; Hahn et al. 2005, 2009), and where the polymerization–depolymerization cycles of the pili allow motility (McBride 2001; Mignot et al. 2005). Some of the competence-related proteins (such as the Com proteins) are expressed even by non-competent bacteria [such as Streptococcus sanguinis strain 133–179 (Zhu et al. 2011)]. As to DUESs, they are hitherto limited to some Pasteurellaceae and Neisseriaceae species. Their evolution seems therefore subsequent to that of competence, so their evolution must be related to specificities in the DNA binding by these species. Given that T4P, especially of pathogens, show differences in their adhesion specificities [e.g. (Kubiet et al. 2000; Kirchner and Meyer 2005)], DUESs may have been gradually driven to overrepresentation in unconstrained regions of these species’ genomes (all pathogenic) as a possible consequence of this adhesion specificity [Bakkali in preparation]. This would inevitably result in preferential uptake of con-specific DNA. This dynamic is what the molecular drive hypothesis on DUESs evolution suggests [see (Bakkali et al. 2004; Bakkali 2007)]. At the end, the accidental binding and uptake of DNA nicely fits the suggested model on DUES evolution (Bakkali et al. 2004; Bakkali 2007), as well as the facts that DUESs must have evolved after competence evolution, they are limited to pathogens, the latter have specific adhesion to their host tissues, and this specificity relates to characteristics of the T4P (which binds the DNA).
In conclusion, DNA uptake could be an ‘unintended’ by-product of bacterial adhesion and twitching motility. However, accidental binding and uptake of DNA does not completely rule out any of the available hypotheses on the potential benefit of competence, and cells may occasionally benefit from different contributions of competence depending on the species and/or situation. For instance, bacteria with indiscriminate DNA uptake may benefit from potential occasional nutritional contribution of DNA uptake (if DNA uptake coincides with a nucleotide starvation situation), and an occasional advantage of recombination may help species ‘preferring’ con-specific DNA and those frequently exposed to antibiotics/stress (if DNA uptake coincides with a challenging environment and the DNA taken up happens to be an advantageous piece from a cell that died for reasons unrelated to the challenging situation). However, the coincidence between DNA uptake and the cell’s need for nucleotides is not always guaranteed and may not be as frequent as to explain the evolution and maintenance of competence. Less likely is the coincidence between uptake of an advantageous DNA and the cell’s need for that particular piece of DNA. Hence, competence could also be selectively neutral and drifting throughout the bacterial evolutionary history. It may have been lost in some bacteria while still drifting in others. This could explain its sporadic distribution across taxa. Whatever the case, antibiotics misuse is most likely favoring the increase and spread of DNA uptake and competence [e.g. (Prudhomme et al. 2006)] while selecting for bacteria capable of acquiring new genetic material that potentially confers drug resistance. The consequent increase in multidrug resistance and virulence adds to increased bacterial dispersal assisted by extensive worldwide displacement of humans, livestock and other goods.
Finally, it is worth mentioning that, if demonstrating the existence of existing things is hard, demonstrating the non-existence of non-existing things is much harder. Some of the crucial tests on the validity of my hypothesis include examining whether bacteria sense the presence of extracellular DNA, testing whether bacteria actively search for DNA and analyzing DNA binding to cells in the presence of competing non-nutritive substances that are devoid of genetic information.
References
Auzat I et al (1999) The NADH oxidase of Streptococcus pneumoniae: its involvement in competence and virulence. Mol Microbiol 34:1018–1028
Averhoff B (2004) DNA transport and natural transformation in mesophilic and thermophilic bacteria. J Bioenerg Biomembr 36:25–33
Bakkali M (2007) Genome dynamics of short oligonucleotides: the example of bacterial DNA uptake enhancing sequences. PLoS ONE 2:e741
Bakkali M, Chen TY, Lee HC, Redfield RJ (2004) Evolutionary stability of DNA uptake signal sequences in the Pasteurellaceae. Proc Natl Acad Sci USA 101:4513–4518
Bartlett JG (2006) Narrative review: the new epidemic of Clostridium difficile-associated enteric disease. Ann Intern Med 145:758–764
Berge M, Moscoso M, Prudhomme M, Martin B, Claverys JP (2002) Uptake of transforming DNA in Gram-positive bacteria: a view from Streptococcus pneumoniae. Mol Microbiol 45:411–421
Bertolla F et al (2000) Plant genome complexity may be a factor limiting in situ the transfer of transgenic plant genes to the phytopathogen Ralstonia solanacearum. Appl Environ Microbiol 66:4161–4167
Bhaya D, Bianco NR, Bryant D, Grossman A (2000) Type IV pilus biogenesis and motility in the cyanobacterium Synechocystis sp. PCC6803. Mol Microbiol 37:941–951
Biswas GD, Sox T, Blackman E, Sparling PF (1977) Factors affecting genetic transformation of Neisseria gonorrhoeae. J Bacteriol 129:983–992
Bradley DE (1975) The occurrence of pili associated with a plasmid of the W compatibility group. Biochem Biophys Res Commun 64:918–925
Brener D, DeVoe IW (1983) Effects of culture pH on the expression of meningococcal pili. Curr Microbiol 8:57–61
Brinkmann V et al (2004) Neutrophil extracellular traps kill bacteria. Science 303:1532–1535
Burke WF Jr (1982) Bacillus subtilis extracellular nuclease production associated with the spoOH sporulation locus. J Gen Microbiol 128:1591–1597
Catlin BW (1960) Transformation of Neisseria meningitidis by deoxyribonucleates from cells and from culture slime. J Bacteriol 79:579–590
Chapuy-Regaud S, Duthoit F, Malfroy-Mastrorillo L, Gourdon P, Lindley ND, Trombe MC (2001) Competence regulation by oxygen availability and by Nox is not related to specific adjustment of central metabolism in Streptococcus pneumoniae. J Bacteriol 183:2957–2962
Charpentier X, Kay E, Schneider D, Shuman HA (2011) Antibiotics and UV radiation induce competence for natural transformation in Legionella pneumophila. J Bacteriol 193:1114–1121
Chen I, Dubnau D (2003) DNA transport during transformation. Front Biosci 8:s544–s556
Chen I, Dubnau D (2004) DNA uptake during bacterial transformation. Nat Rev Microbiol 2:241–249
Chen I, Christie PJ, Dubnau D (2005) The ins and outs of DNA transfer in bacteria. Science 310:1456–1460
Chen I, Provvedi R, Dubnau D (2006) A macromolecular complex formed by a pilin-like protein in competent Bacillus subtilis. J Biol Chem 281:21720–21727
Christie PJ, Atmakuri K, Krishnamoorthy V, Jakubowski S, Cascales E (2005) Biogenesis, architecture, and function of bacterial type IV secretion systems. Annu Rev Microbiol 59:451–485
Claverys JP, Prudhomme M, Martin B (2006) Induction of competence regulons as a general response to stress in Gram-positive bacteria. Annu Rev Microbiol 60:451–475
Claverys JP, Martin B, Polard P (2009) The genetic transformation machinery: composition, localization, and mechanism. FEMS Microbiol Rev 33:643–656
Collins RF, Davidsen L, Derrick JP, Ford RC, Tonjum T (2001) Analysis of the PilQ secretin from Neisseria meningitidis by transmission electron microscopy reveals a dodecameric quaternary structure. J Bacteriol 183:3825–3832
Courvalin P (2005) Antimicrobial drug resistance: “prediction is very difficult, especially about the future”. Emerg Infect Dis 11:1503–1506
de Vries J, Meier P, Wackernagel W (2001) The natural transformation of the soil bacteria Pseudomonas stutzeri and Acinetobacter sp. by transgenic plant DNA strictly depends on homologous sequences in the recipient cells. FEMS Microbiol Lett 195:211–215
Demaneche S, Kay E, Gourbiere F, Simonet P (2001) Natural transformation of Pseudomonas fluorescens and Agrobacterium tumefaciens in soil. Appl Environ Microbiol 67:2617–2621
Diep BA, Carleton HA, Chang RF, Sensabaugh GF, Perdreau-Remington F (2006) Roles of 34 virulence genes in the evolution of hospital- and community-associated strains of methicillin-resistant Staphylococcus aureus. J Infect Dis 193:1495–1503
Doran JL, Page WJ (1983) Heat sensitivity of Azotobacter vinelandii genetic transformation. J Bacteriol 155:159–168
Dubnau D (1999) DNA uptake in bacteria. Annu Rev Microbiol 53:217–244
Espinosa M, Joenje H, Venema G (1980) DNA binding and deoxyribonuclease activity in Bacillus subtilis during temperature-induced competence development. J Gen Microbiol 121:79–84
Facius D, Meyer TF (1993) A novel determinant (comA) essential for natural transformation competence in Neisseria gonorrhoeae and the effect of a comA defect on pilin variation. Mol Microbiol 10:699–712
Forest KT, Tainer JA (1997) Type-4 pilus-structure: outside to inside and top to bottom—a minireview. Gene 192:165–169
Fraser C, Hanage WP, Spratt BG (2007) Recombination and the nature of bacterial speciation. Science 315:476–480
Fyfe JA, Carrick CS, Davies JK (1995) The pilE gene of Neisseria gonorrhoeae MS11 is transcribed from a sigma 70 promoter during growth in vitro. J Bacteriol 177:3781–3787
Glick BR, Brooks HE, Pasternak JJ (1985) Transformation of Azotobacter vinelandii with plasmid DNA. J Bacteriol 162:276–279
Gomez–Gomez JM, Manfredi C, Alonso JC, Blazquez J (2007) A novel role for RecA under non-stress: promotion of swarming motility in Escherichia coli K-12. BMC Biol 5:14
Goodgal SH, Herriott RM (1961) Studies on transformations of Hemophilus influenzae. I. Competence. J Gen Physiol 44:1201–1227
Graves JF, Biswas GD, Sparling PF (1982) Sequence-specific DNA uptake in transformation of Neisseria gonorrhoeae. J Bacteriol 152:1071–1077
Griffith F (1928) The significance of pneumococcal types. J Hyg 27:113–159
Hahn J, Maier B, Haijema BJ, Sheetz M, Dubnau D (2005) Transformation proteins and DNA uptake localize to the cell poles in Bacillus subtilis. Cell 122:59–71
Hahn J, Kramer N, Briley K Jr, Dubnau D (2009) McsA and B mediate the delocalization of competence proteins from the cell poles of Bacillus subtilis. Mol Microbiol 72:202–215
Hamilton HL, Dominguez NM, Schwartz KJ, Hackett KT, Dillard JP (2005) Neisseria gonorrhoeae secretes chromosomal DNA via a novel type IV secretion system. Mol Microbiol 55:1704–1721
Havarstein LS, Coomaraswamy G, Morrison DA (1995) An unmodified heptadecapeptide pheromone induces competence for genetic transformation in Streptococcus pneumoniae. Proc Natl Acad Sci USA 92:11140–11144
Henrichsen J (1983) Twitching motility. Annu Rev Microbiol 37:81–93
Herriott RM, Meyer EM, Vogt M (1970) Defined nongrowth media for stage II development of competence in Haemophilus influenzae. **J Bacteriol 101:517–524
Jelsbak L, Sogaard-Andersen L (2000) Pattern formation: fruiting body morphogenesis in Myxococcus xanthus. Curr Opin Microbiol 3:637–642
Johnsen PJ, Dubnau D, Levin BR (2009) Episodic selection and the maintenance of competence and natural transformation in Bacillus subtilis. Genetics 181:1521–1533
Kaiser D (1979) Social gliding is correlated with the presence of pili in Myxococcus xanthus. Proc Natl Acad Sci USA 76:5952–5956
Kaiser D (2000) Bacterial motility: how do pili pull? Curr Biol 10:R777–R780
Karaolis DK, Somara S, Maneval DR Jr, Johnson JA, Kaper JB (1999) A bacteriophage encoding a pathogenicity island, a type-IV pilus and a phage receptor in cholera bacteria. Nature 399:375–379
Kendall MM, Sperandio V (2007) Quorum sensing by enteric pathogens. Curr Opin Gastroenterol 23:10–15
Kirchner M, Meyer TF (2005) The PilC adhesin of the Neisseria type IV pilus-binding specificities and new insights into the nature of the host cell receptor. Mol Microbiol 56:945–957
Kozlovsky Y, Cohen I, Golding I, Ben-Jacob E (1999) Lubricating bacteria model for branching growth of bacterial colonies. Phys Rev E Stat Phys Plasmas Fluids Relat Interdiscip Topics 59:7025–7035
Kruger E, Msadek T, Ohlmeier S, Hecker M (1997) The Bacillus subtilis clpC operon encodes DNA repair and competence proteins. Microbiology 143(Pt 4):1309–1316
Kubiet M, Ramphal R, Weber A, Smith A (2000) Pilus-mediated adherence of Haemophilus influenzae to human respiratory mucins. Infect Immun 68:3362–3367
Kufryk GI, Sachet M, Schmetterer G, Vermaas WF (2002) Transformation of the cyanobacterium Synechocystis sp. PCC 6803 as a tool for genetic mapping: optimization of efficiency. FEMS Microbiol Lett 206:215–219
Kupfer DM, McCarthy D (1992) rec-2-dependent phage recombination in Haemophilus influenzae. J Bacteriol 174:4960–4966
Lacasta AM, Cantalapiedra IR, Auguet CE, Penaranda A, Ramirez-Piscina L (1999) Modeling of spatiotemporal patterns in bacterial colonies. Phys Rev E Stat Phys Plasmas Fluids Relat Interdiscip Topics 59:7036–7041
Lauer P, Albertson NH, Koomey M (1993) Conservation of genes encoding components of a type IV pilus assembly/two-step protein export pathway in Neisseria gonorrhoeae. Mol Microbiol 8:357–368
Li YH, Lau PC, Lee JH, Ellen RP, Cvitkovitch DG (2001) Natural genetic transformation of Streptococcus mutans growing in biofilms. J Bacteriol 183:897–908
Li YH et al (2002) A quorum-sensing signaling system essential for genetic competence in Streptococcus mutans is involved in biofilm formation. J Bacteriol 184:2699–2708
Liles MR, Viswanathan VK, Cianciotto NP (1998) Identification and temperature regulation of Legionella pneumophila genes involved in type IV pilus biogenesis and type II protein secretion. Infect Immun 66:1776–1782
Lin TL, Shun CT, Chang KC, Wang JT (2006) Isolation and characterization of a competence operon associated with transformation and adhesion in Helicobacter pylori. Microbes Infect 8:2756–2765
Lopez P, Perez Urena MT, Espinosa M (1983) Influence of temperature-induced competence in the genetic transformation of Bacillus subtilis. Microbios 38:205–216
Lorenz MG, Wackernagel W (1994) Bacterial gene transfer by natural genetic transformation in the environment. Microbiol Rev 58:563–602
Lorenz MG, Gerjets D, Wackernagel W (1991) Release of transforming plasmid and chromosomal DNA from two cultured soil bacteria. Arch Microbiol 156:319–326
Lorenz MG, Reipschlager K, Wackernagel W (1992) Plasmid transformation of naturally competent Acinetobacter calcoaceticus in non-sterile soil extract and groundwater. Arch Microbiol 157:355–360
Love PE, Lyle MJ, Yasbin RE (1985) DNA-damage-inducible (din) loci are transcriptionally activated in competent Bacillus subtilis. Proc Natl Acad Sci USA 82:6201–6205
Macfadyen LP, Dorocicz IR, Reizer J, Saier MH Jr, Redfield RJ (1996) Regulation of competence development and sugar utilization in Haemophilus influenzae Rd by a phosphoenolpyruvate:fructose phosphotransferase system. Mol Microbiol 21:941–952
MacFadyen LP, Chen D, Vo HC, Liao D, Sinotte R, Redfield RJ (2001) Competence development by Haemophilus influenzae is regulated by the availability of nucleic acid precursors. Mol Microbiol 40:700–707
Manetti AG et al (2010) Environmental acidification drives S. pyogenes pilus expression and microcolony formation on epithelial cells in a FCT-dependent manner. PLoS ONE 5:e13864
Manning GS (2006) The persistence length of DNA is reached from the persistence length of its null isomer through an internal electrostatic stretching force. Biophys J 91:3607–3616
Margolin W (2006) Gliding motility: anticipating the next move with a molecular clock. Curr Biol 16:R85–R87
Mattick JS (2002) Type IV pili and twitching motility. Annu Rev Microbiol 56:289–314
McBride MJ (2001) Bacterial gliding motility: multiple mechanisms for cell movement over surfaces. Annu Rev Microbiol 55:49–75
Merz AJ, So M, Sheetz MP (2000) Pilus retraction powers bacterial twitching motility. Nature 407:98–102
Messi P, Guerrieri E, de Niederhausern S, Sabia C, Bondi M (2006) Vancomycin-resistant enterococci (VRE) in meat and environmental samples. Int J Food Microbiol 107:218–222
Michod RE, Wojciechowski MF, Hoelzer MA (1988) DNA repair and the evolution of transformation in the bacterium Bacillus subtilis. Genetics 118:31–39
Mignot T, Merlie JP Jr, Zusman DR (2005) Regulated pole-to-pole oscillations of a bacterial gliding motility protein. Science 310:855–857
Mignot T, Shaevitz JW, Hartzell PL, Zusman DR (2007) Evidence that focal adhesion complexes power bacterial gliding motility. Science 315:853–856
Mongold JA (1992) DNA repair and the evolution of transformation in Haemophilus influenzae. Genetics 132:893–898
Mortier-Barriere I, de Saizieu A, Claverys JP, Martin B (1998) Competence-specific induction of recA is required for full recombination proficiency during transformation in Streptococcus pneumoniae. Mol Microbiol 27:159–170
Moscoso M, Claverys JP (2004) Release of DNA into the medium by competent Streptococcus pneumoniae: kinetics, mechanism and stability of the liberated DNA. Mol Microbiol 54:783–794
Nachin L, Nannmark U, Nystrom T (2005) Differential roles of the universal stress proteins of Escherichia coli in oxidative stress resistance, adhesion, and motility. J Bacteriol 187:6265–6272
Narra HP, Ochman H (2006) Of what use is sex to bacteria? Curr Biol 16:R705–R710
Nijland R, Burgess JG, Errington J, Veening JW (2010) Transformation of environmental Bacillus subtilis isolates by transiently inducing genetic competence. PLoS ONE 5:e9724
Nudleman E, Kaiser D (2004) Pulling together with type IV pili. J Mol Microbiol Biotechnol 7:52–62
Oggioni MR et al (2004) Antibacterial activity of a competence-stimulating peptide in experimental sepsis caused by Streptococcus pneumoniae. Antimicrob Agents Chemother 48:4725–4732
O’Rourke EJ, Pinto AV, Petroni EA, Tolmasky ME, Ielpi L (2004) Evidence for the active role of a novel nuclease from Helicobacter pylori in the horizontal transfer of genetic information. J Bacteriol 186:2586–2593
Palchevskiy V, Finkel SE (2006) Escherichia coli competence gene homologs are essential for competitive fitness and the use of DNA as a nutrient. J Bacteriol 188:3902–3910
Palmen R, Vosman B, Buijsman P, Breek CK, Hellingwerf KJ (1993) Physiological characterization of natural transformation in Acinetobacter calcoaceticus. J Gen Microbiol 139:295–305
Provvedi R, Chen I, Dubnau D (2001) NucA is required for DNA cleavage during transformation of Bacillus subtilis. Mol Microbiol 40:634–644
Prudhomme M, Attaiech L, Sanchez G, Martin B, Claverys JP (2006) Antibiotic stress induces genetic transformability in the human pathogen Streptococcus pneumoniae. Science 313:89–92
Ranhand JM (1976) Effect of pH on competence development and deoxyribonucleic acid uptake in Streptococcus sanguis (Wicky). J Bacteriol 126:205–212
Raymond-Denise A, Guillen N (1992) Expression of the Bacillus subtilis dinR and recA genes after DNA damage and during competence. J Bacteriol 174:3171–3176
Redfield RJ (1988) Evolution of bacterial transformation: is sex with dead cells ever better than no sex at all? Genetics 119:213–221
Redfield RJ (1991) Bacteria mating preferences. Nature 352:25–26
Redfield RJ (1993a) Evolution of natural transformation: testing the DNA repair hypothesis in Bacillus subtilis and Haemophilus influenzae. Genetics 133:755–761
Redfield RJ (1993b) Genes for breakfast: the have-your-cake-and-eat-it-too of bacterial transformation. J Hered 84:400–404
Redfield RJ (2001) Do bacteria have sex? Nat Rev Genet 2:634–639
Redfield RJ, Schrag MR, Dean AM (1997) The evolution of bacterial transformation: sex with poor relations. Genetics 146:27–38
Redfield RJ, Findlay WA, Bosse J, Kroll JS, Cameron AD, Nash JH (2006) Evolution of competence and DNA uptake specificity in the Pasteurellaceae. BMC Evol Biol 6:82
Roelants P, Konvalinkova V, Mergeay M, Lurquin PF, Ledoux L (1976) DNA uptake by Streptomyces species. Biochim Biophys Acta 442:117–122
Rudel T, Scheurerpflug I, Meyer TF (1995a) Neisseria PilC protein identified as type-4 pilus tip-located adhesin. Nature 373:357–359
Rudel T, Facius D, Barten R, Scheuerpflug I, Nonnenmacher E, Meyer TF (1995b) Role of pili and the phase-variable PilC protein in natural competence for transformation of Neisseria gonorrhoeae. Proc Natl Acad Sci USA 92:7986–7990
Sabia C, Bondi M, Messi P, De Niederhausern S, Manicardi G (2005) Study of five penicillinase producing Neisseria gonorrhoeae isolated in Italy. New Microbiol 28:223–229
Shafeeq S, Kloosterman TG, Kuipers OP (2011) CelR-mediated activation of the cellobiose-utilization gene cluster in Streptococcus pneumoniae. Microbiology 157:2854–2861
Singh RN, Pitale MP (1968) Competence and deoxyribonucleic acid uptake in Bacillus subtilis. J Bacteriol 95:864–866
Sisco KL, Smith HO (1979) Sequence-specific DNA uptake in Haemophilus transformation. Proc Natl Acad Sci USA 76:972–976
Smeets LC, Becker SC, Barcak GJ, Vandenbroucke-Grauls CM, Bitter W, Goosen N (2006) Functional characterization of the competence protein DprA/Smf in Escherichia coli. FEMS Microbiol Lett 263:223–228
Smith HO, Danner DB, Deich RA (1981) Genetic transformation. Annu Rev Biochem 50:41–68
Steinmoen H, Knutsen E, Havarstein LS (2002) Induction of natural competence in Streptococcus pneumoniae triggers lysis and DNA release from a subfraction of the cell population. Proc Natl Acad Sci USA 99:7681–7686
Sung P, Klein H (2006) Mechanism of homologous recombination: mediators and helicases take on regulatory functions. Nat Rev Mol Cell Biol 7:739–750
Tang Y, Guest JR, Artymiuk PJ, Read RC, Green J (2004) Post-transcriptional regulation of bacterial motility by aconitase proteins. Mol Microbiol 51:1817–1826
Thanassi JA, Hartman-Neumann SL, Dougherty TJ, Dougherty BA, Pucci MJ (2002) Identification of 113 conserved essential genes using a high-throughput gene disruption system in Streptococcus pneumoniae. Nucleic Acids Res 30:3152–3162
Turgay K, Hamoen LW, Venema G, Dubnau D (1997) Biochemical characterization of a molecular switch involving the heat shock protein ClpC, which controls the activity of ComK, the competence transcription factor of Bacillus subtilis. Genes Dev 11:119–128
Varga JJ, Nguyen V, O’Brien DK, Rodgers K, Walker RA, Melville SB (2006) Type IV pili-dependent gliding motility in the Gram-positive pathogen Clostridium perfringens and other Clostridia. Mol Microbiol 62:680–694
Vazquez JA, Berron S, Gimenez MJ, de la Fuente L, Aguilar L (2001) In vitro susceptibility of Neisseria meningitidis isolates to gemifloxacin and ten other antimicrobial agents. Eur J Clin Microbiol Infect Dis 20:150–151
Villar MT, Hirschberg RL, Schaefer MR (2001) Role of the Eikenella corrodens pilA locus in pilus function and phase variation. J Bacteriol 183:55–62
Wang Y, Taylor DE (1990) Natural transformation in Campylobacter species. J Bacteriol 172:949–955
Wang WB, Lai HC, Hsueh PR, Chiou RY, Lin SB, Liaw SJ (2006) Inhibition of swarming and virulence factor expression in Proteus mirabilis by resveratrol. J Med Microbiol 55:1313–1321
Whitchurch CB, Tolker-Nielsen T, Ragas PC, Mattick JS (2002) Extracellular DNA required for bacterial biofilm formation. Science 295:1487
Wolfgang M, Lauer P, Park HS, Brossay L, Hebert J, Koomey M (1998) PilT mutations lead to simultaneous defects in competence for natural transformation and twitching motility in piliated Neisseria gonorrhoeae. Mol Microbiol 29:321–330
Wu SS, Kaiser D (1997) Regulation of expression of the pilA gene in Myxococcus xanthus. J Bacteriol 179:7748–7758
Wu MC, Chen YC, Lin TL, Hsieh PF, Wang JT (2012) Cellobiose-specific phosphotransferase system of Klebsiella pneumoniae and its importance in biofilm formation and virulence. Infect Immun 80:2464–2472
Yoshihara S, Geng X, Ikeuchi M (2002) pilG Gene cluster and split pilL genes involved in pilus biogenesis, motility and genetic transformation in the cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol 43:513–521
Young FE, Spizizen J, Crawford IP (1963) Biochemical aspects of competence in the Bacillus subtilis transformation system. I. Chemical composition of cell walls. J Biol Chem 238:3119–3125
Zeng L, Burne RA (2009) Transcriptional regulation of the cellobiose operon of Streptococcus mutans. J Bacteriol 191:2153–2162
Zhu L, Zhang Y, Fan J, Herzberg MC, Kreth J (2011) Characterization of competence and biofilm development of a Streptococcus sanguinis endocarditis isolate. Mol Oral Microbiol 26:117–126
Acknowledgments
I am very thankful to my wife Pernille Lavgesen for her continuous support. I am also thankful to all the friends and colleagues who commented on this manuscript, especially Prof. Rosemary Redfield who was the reason why I got introduced into the fascinating world of microbiology and whose influence on me goes beyond the way of thinking—Rosie you are and will always be one of my greatest sources of inspiration: THANK YOU. I also wish to thank Dr. Andrew Cameron and the anonymous reviewer for their helpful comments on this manuscript. To Nadia L. Bakkali and Elias L. Bakkali, and in memory of my beloved father, Prof. Abdeslam Bakkali 1945–2011.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Djamel Drider.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Bakkali, M. Could DNA uptake be a side effect of bacterial adhesion and twitching motility?. Arch Microbiol 195, 279–289 (2013). https://doi.org/10.1007/s00203-013-0870-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-013-0870-1