Abstract
Summary
To scan novel candidate genes associated with osteoporosis, a two-stage transcriptome-wide association study (TWAS) of bone mineral density (BMD) was conducted. The BMD-associated genes identified by TWAS were then compared with the gene expression profiling of BMD in bone cells, B cells, and mesenchymal stem cells. We identified multiple candidate genes and gene ontology (GO) terms associated with BMD.
Introduction
Osteoporosis (OP) is a metabolic bone disease characterized by decrease in BMD. Our objective is to scan novel candidate genes associated with OP.
Methods
A transcriptome-wide association study (TWAS) was performed by integrating the genome-wide association study (GWAS) summary of bone mineral density (BMD) with two pre-computed mRNA expression weights of peripheral blood and muscle skeleton. Then, another independent GWAS data of BMD was used to verify the discovery results. The BMD-associated genes identified between discovery and replicate TWAS were further subjected to gene ontology (GO) analysis implemented by DAVID. Finally, the BMD-associated genes and GO terms were further compared with the mRNA expression profiling results of BMD to detect the common genes and GO terms shared by both DNA-level TWAS and mRNA expression profile analysis.
Results
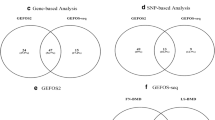
TWAS identified 95 common genes with permutation P value < 0.05 for peripheral blood and muscle skeleton, such as TMTC4 in muscle skeleton and DDX17 in peripheral blood. Further comparing the genes detected by discovery-replicate TWAS with the differentially expressed genes identified by mRNA expression profiling of OP patients found 18 overlapped genes, such as MUL1 in muscle skeleton and SPTBN1 in peripheral blood. GO analysis of the genes identified by discovery-replicate TWAS detected 12 BMD-associated GO terms, such as negative regulation of cell growth and regulation of glycogen catabolic process. Further comparing the GO results of discovery-replicate TWAS and mRNA expression profiles found 6 overlapped GO terms, such as membrane and cell adhesion.
Conclusion
Our study identified multiple candidate genes and GO terms for BMD, providing novel clues for understanding the genetic mechanism of OP.
Similar content being viewed by others
References
Ammann P, Rizzoli R (2003) Bone strength and its determinants. Osteoporos Int 14(Suppl 3):S13–S18
Cooper C, Campion G, Melton LJ 3rd (1992) Hip fractures in the elderly: a world-wide projection. Osteoporos Int 2(6):285–289
C WN et al (2014) The recent prevalence of osteoporosis and low bone mass in the United States based on bone mineral density at the femoral neck or lumbar spine. J Bone Miner Res 29(11):2520–2526
Coughlan T, Dockery F (2014) Osteoporosis and fracture risk in older people. Clin Med (Lond) 14(2):187–191
Ralston SH, de Crombrugghe B (2006) Genetic regulation of bone mass and susceptibility to osteoporosis. Genes Dev 20(18):2492–2506
McClung M, Baron R, Bouxsein M (2017) An update on osteoporosis pathogenesis, diagnosis, and treatment. Bone 98:37
Kemp JP, Morris JA, Medina-Gomez C, Forgetta V, Warrington NM, Youlten SE, Zheng J, Gregson CL, Grundberg E, Trajanoska K, Logan JG, Pollard AS, Sparkes PC, Ghirardello EJ, Allen R, Leitch VD, Butterfield NC, Komla-Ebri D, Adoum AT, Curry KF, White JK, Kussy F, Greenlaw KM, Xu C, Harvey NC, Cooper C, Adams DJ, Greenwood CMT, Maurano MT, Kaptoge S, Rivadeneira F, Tobias JH, Croucher PI, Ackert-Bicknell CL, Bassett JHD, Williams GR, Richards JB, Evans DM (2017) Identification of 153 new loci associated with heel bone mineral density and functional involvement of GPC6 in osteoporosis. Nat Genet 49(10):1468–1475
Estrada K, Styrkarsdottir U, Evangelou E, Hsu YH, Duncan EL, Ntzani EE, Oei L, Albagha OME, Amin N, Kemp JP, Koller DL, Li G, Liu CT, Minster RL, Moayyeri A, Vandenput L, Willner D, Xiao SM, Yerges-Armstrong LM, Zheng HF, Alonso N, Eriksson J, Kammerer CM, Kaptoge SK, Leo PJ, Thorleifsson G, Wilson SG, Wilson JF, Aalto V, Alen M, Aragaki AK, Aspelund T, Center JR, Dailiana Z, Duggan DJ, Garcia M, Garcia-Giralt N, Giroux S, Hallmans G, Hocking LJ, Husted LB, Jameson KA, Khusainova R, Kim GS, Kooperberg C, Koromila T, Kruk M, Laaksonen M, Lacroix AZ, Lee SH, Leung PC, Lewis JR, Masi L, Mencej-Bedrac S, Nguyen TV, Nogues X, Patel MS, Prezelj J, Rose LM, Scollen S, Siggeirsdottir K, Smith AV, Svensson O, Trompet S, Trummer O, van Schoor NM, Woo J, Zhu K, Balcells S, Brandi ML, Buckley BM, Cheng S, Christiansen C, Cooper C, Dedoussis G, Ford I, Frost M, Goltzman D, González-Macías J, Kähönen M, Karlsson M, Khusnutdinova E, Koh JM, Kollia P, Langdahl BL, Leslie WD, Lips P, Ljunggren Ö, Lorenc RS, Marc J, Mellström D, Obermayer-Pietsch B, Olmos JM, Pettersson-Kymmer U, Reid DM, Riancho JA, Ridker PM, Rousseau F, lagboom PES, Tang NLS, Urreizti R, van Hul W, Viikari J, Zarrabeitia MT, Aulchenko YS, Castano-Betancourt M, Grundberg E, Herrera L, Ingvarsson T, Johannsdottir H, Kwan T, Li R, Luben R, Medina-Gómez C, Th Palsson S, Reppe S, Rotter JI, Sigurdsson G, van Meurs JBJ, Verlaan D, Williams FMK, Wood AR, Zhou Y, Gautvik KM, Pastinen T, Raychaudhuri S, Cauley JA, Chasman DI, Clark GR, Cummings SR, Danoy P, Dennison EM, Eastell R, Eisman JA, Gudnason V, Hofman A, Jackson RD, Jones G, Jukema JW, Khaw KT, Lehtimäki T, Liu Y, Lorentzon M, McCloskey E, Mitchell BD, Nandakumar K, Nicholson GC, Oostra BA, Peacock M, Pols HAP, Prince RL, Raitakari O, Reid IR, Robbins J, Sambrook PN, Sham PC, Shuldiner AR, Tylavsky FA, van Duijn CM, Wareham NJ, Cupples LA, Econs MJ, Evans DM, Harris TB, Kung AWC, Psaty BM, Reeve J, Spector TD, Streeten EA, Zillikens MC, Thorsteinsdottir U, Ohlsson C, Karasik D, Richards JB, Brown MA, Stefansson K, Uitterlinden AG, Ralston SH, Ioannidis JPA, Kiel DP, Rivadeneira F (2012) Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture. Nat Genet 44(5):491–501
Massague J, Chen YG (2000) Controlling TGF-beta signaling. Genes Dev 14(6):627–644
Miyamoto T (2015) Mechanism underlying post-menopausal osteoporosis: HIF1alpha is required for osteoclast activation by estrogen deficiency. Keio J Med 64(3):44–47
Boudin E, Fijalkowski I, Hendrickx G, van Hul W (2016) Genetic control of bone mass. Mol Cell Endocrinol 432:3–13
Hindorff LA, Sethupathy P, Junkins HA, Ramos EM, Mehta JP, Collins FS, Manolio TA (2009) Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci U S A 106(23):9362–9367
Soldner F, Stelzer Y, Shivalila CS, Abraham BJ, Latourelle JC, Barrasa MI, Goldmann J, Myers RH, Young RA, Jaenisch R (2016) Parkinson-associated risk variant in distal enhancer of alpha-synuclein modulates target gene expression. Nature 533(7601):95–99
Tak YG, Farnham PJ (2015) Making sense of GWAS: using epigenomics and genome engineering to understand the functional relevance of SNPs in non-coding regions of the human genome. Epigenetics Chromatin 8:57
Grubert F, Zaugg JB, Kasowski M, Ursu O, Spacek DV, Martin AR, Greenside P, Srivas R, Phanstiel DH, Pekowska A, Heidari N, Euskirchen G, Huber W, Pritchard JK, Bustamante CD, Steinmetz LM, Kundaje A, Snyder M (2015) Genetic control of chromatin states in humans involves local and distal chromosomal interactions. Cell 162(5):1051–1065
Gusev A, Mancuso N, Won H et al (2018) Transcriptome-wide association study of schizophrenia and chromatin activity yields mechanistic disease insights. Nat Genet 50(4):538–548
Wu L et al (2018) A transcriptome-wide association study of 229,000 women identifies new candidate susceptibility genes for breast cancer. Nat Genet 50(7):968–978
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760
Zheng HF et al (2015) Whole-genome sequencing identifies EN1 as a determinant of bone density and fracture. Nature 526(7571):112–117
Kemp JP, Morris JA, Medina-Gomez C et al (2017) Identification of 153 new loci associated with heel bone mineral density and functional involvement of GPC6 in osteoporosis. Nat Genet 49(10):1468–1475
Loh PR, Tucker G, Bulik-Sullivan BK, Vilhjálmsson BJ, Finucane HK, Salem RM, Chasman DI, Ridker PM, Neale BM, Berger B, Patterson N, Price AL (2015) Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat Genet 47(3):284–290
Peng X et al (2008) In vivo genome-wide expression study on human circulating B cells suggests a novel ESR1 and MAPK3 network for postmenopausal osteoporosis. J Bone Miner Res 23(5):644–654
Benisch P, Schilling T, Klein-Hitpass L, Frey SP, Seefried L, Raaijmakers N, Krug M, Regensburger M, Zeck S, Schinke T, Amling M, Ebert R, Jakob F (2012) The transcriptional profile of mesenchymal stem cell populations in primary osteoporosis is distinct and shows overexpression of osteogenic inhibitors. PLoS One 7(9):e45142
Saito H, Shimokata K, Yamada Y, Nomura F, Yamori S (1992) Umbilical metastasis from small cell carcinoma of the lung. Chest 101(1):288–289
Smyth GK (2004) Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3:Article3
Yang YH et al (2002) Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res 30(4):e15
Berger JA, Hautaniemi S, Järvinen AK, Edgren H, Mitra SK, Astola J (2004) Optimized LOWESS normalization parameter selection for DNA microarray data. BMC Bioinformatics 5:194
Hopwood B, Tsykin A, Findlay DM, Fazzalari NL (2007) Microarray gene expression profiling of osteoarthritic bone suggests altered bone remodelling, WNT and transforming growth factor-β/bone morphogenic protein signalling. Arthritis Res Ther 9(5):R100
Gusev A, Ko A, Shi H, Bhatia G, Chung W, Penninx BWJH, Jansen R, de Geus EJC, Boomsma DI, Wright FA, Sullivan PF, Nikkola E, Alvarez M, Civelek M, Lusis AJ, Lehtimäki T, Raitoharju E, Kähönen M, Seppälä I, Raitakari OT, Kuusisto J, Laakso M, Price AL, Pajukanta P, Pasaniuc B (2016) Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet 48:245–252
Gusev A, Ko A, Shi H et al (2016) Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet 48(3):245–252
Lonsdale J, Thomas J, Salvatore M, Phillips R, Lo E, Shad S, Hasz R, Walters G, Garcia F, Young N, Foster B, Moser M, Karasik E, Gillard B, Ramsey K, Sullivan S, Bridge J, Magazine H, Syron J, Fleming J, Siminoff L, Traino H, Mosavel M, Barker L, Jewell S, Rohrer D, Maxim D, Filkins D, Harbach P, Cortadillo E, Berghuis B, Turner L, Hudson E, Feenstra K, Sobin L, Robb J, Branton P, Korzeniewski G, Shive C, Tabor D, Qi L, Groch K, Nampally S, Buia S, Zimmerman A, Smith A, Burges R, Robinson K, Valentino K, Bradbury D, Cosentino M, Diaz-Mayoral N, Kennedy M, Engel T, Williams P, Erickson K, Ardlie K, Winckler W, Getz G, DeLuca D, MacArthur D, Kellis M, Thomson A, Young T, Gelfand E, Donovan M, Meng Y, Grant G, Mash D, Marcus Y, Basile M, Liu J, Zhu J, Tu Z, Cox NJ, Nicolae DL, Gamazon ER, Im HK, Konkashbaev A, Pritchard J, Stevens M, Flutre T, Wen X, Dermitzakis ET, Lappalainen T, Guigo R, Monlong J, Sammeth M, Koller D, Battle A, Mostafavi S, McCarthy M, Rivas M, Maller J, Rusyn I, Nobel A, Wright F, Shabalin A, Feolo M, Sharopova N, Sturcke A, Paschal J, Anderson JM, Wilder EL, Derr LK, Green ED, Struewing JP, Temple G, Volpi S, Boyer JT, Thomson EJ, Guyer MS, Ng C, Abdallah A, Colantuoni D, Insel TR, Koester SE, Little AR, Bender PK, Lehner T, Yao Y, Compton CC, Vaught JB, Sawyer S, Lockhart NC, Demchok J, Moore HF (2013) The Genotype-Tissue Expression (GTEx) project. Nat Genet 45(6):580–585
Boomsma DI, Geus EJC, Vink JM, Stubbe JH, Distel MA, Hottenga JJ, Posthuma D, Beijsterveldt TCEM, Hudziak JJ, Bartels M, Willemsen G (2006) Netherlands Twin Register: from twins to twin families. Twin Res Hum Genet 9(6):849–857
Wright FA, Sullivan PF, Brooks AI, Zou F, Sun W, Xia K, Madar V, Jansen R, Chung W, Zhou YH, Abdellaoui A, Batista S, Butler C, Chen G, Chen TH, D’Ambrosio D, Gallins P, Ha MJ, Hottenga JJ, Huang S, Kattenberg M, Kochar J, Middeldorp CM, Qu A, Shabalin A, Tischfield J, Todd L, Tzeng JY, van Grootheest G, Vink JM, Wang Q, Wang W, Wang W, Willemsen G, Smit JH, de Geus EJ, Yin Z, Penninx BWJH, Boomsma DI (2014) Heritability and genomics of gene expression in peripheral blood. Nat Genet 46(5):430–437
Huang da W, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37(1):1–13
Huang da W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57
Calabrese GM, Mesner LD, Stains JP, Tommasini SM, Horowitz MC, Rosen CJ, Farber CR (2017) Integrating GWAS and co-expression network data identifies bone mineral density genes SPTBN1 and MARK3 and an osteoblast functional module. Cell Syst 4(1):46–59.e4
Zhang R, Zhang CY, Zhao Q, Li DH (2013) Spectrin: structure, function and disease. Sci China Life Sci 56(12):1076–1085
Wang C, Zhang Z, Zhang H, He JW, Gu JM, Hu WW, Hu YQ, Li M, Liu YJ, Fu WZ, Yue H, Ke YH, Zhang ZL (2012) Susceptibility genes for osteoporotic fracture in postmenopausal Chinese women. J Bone Miner Res 27(12):2582–2591
Vilhais-Neto GC, Maruhashi M, Smith KT, Vasseur-Cognet M, Peterson AS, Workman JL, Pourquié O (2010) Rere controls retinoic acid signalling and somite bilateral symmetry. Nature 463(7283):953–957
Zhang H, Sol-Church K, Rydbeck H, Stabley D, Spotila LD, Devoto M (2009) High resolution linkage and linkage disequilibrium analyses of chromosome 1p36 SNPs identify new positional candidate genes for low bone mineral density. Osteoporos Int 20(2):341–346
Li HY, Kung WC, Huang QY (2011) Bone mineral density is linked to 1p36 and 7p15-13 in a southern Chinese population. J Bone Miner Metab 29(1):80–87
Ma C, Shuai B, Shen L, Yang YP, Xu XJ, Li CG (2016) Serum carcinoembryonic antigen-related cell adhesion molecule 1 level in postmenopausal women: correlation with β-catenin and bone mineral density. Osteoporos Int 27(4):1529–1535
Michigami T et al (2000) Cell-cell contact between marrow stromal cells and myeloma cells via VCAM-1 and alpha(4)beta(1)-integrin enhances production of osteoclast-stimulating activity. Blood 96(5):1953–1960
Zeng Y, Zhang L, Zhu W, He H, Sheng H, Tian Q, Deng FY, Zhang LS, Hu HG, Deng HW (2017) Network based subcellular proteomics in monocyte membrane revealed novel candidate genes involved in osteoporosis. Osteoporos Int 28(10):3033–3042
Funding
This study was supported by the National Natural Scientific Foundation of China (81673112) and the Key Projects of International Cooperation among Governments in Scientific and Technological Innovation (2016YFE0119100).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ma, M., Huang, DG., Liang, X. et al. Integrating transcriptome-wide association study and mRNA expression profiling identifies novel genes associated with bone mineral density. Osteoporos Int 30, 1521–1528 (2019). https://doi.org/10.1007/s00198-019-04958-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00198-019-04958-z