Abstract
Receiver Operating Characteristic (ROC) curve analysis and area under the ROC curve (AUC) are commonly used performance measures in diagnostic systems. In this work, we assume a setting, where a classifier is inferred from multivariate data to predict the diagnostic outcome for new cases. Cross-validation is a resampling method for estimating the prediction performance of a classifier on data not used for inferring it. Tournament leave-pair-out (TLPO) cross-validation has been shown to be better than other resampling methods at producing a ranking of data that can be used for estimating the ROC curves and areas under them. However, the time complexity of TLPOCV, \(O\left( n^2\right)\), means that it is impractical in many applications. In this article, a method called quicksort leave-pair-out cross-validation (QLPOCV) is presented in order to decrease the time complexity of obtaining a reliable ranking of data to \(O\left( n\log n\right)\). The proposed method is compared with existing ones in an experimental study, demonstrating that in terms of ROC curves and AUC values QLPOCV produces as accurate performance estimation as TLPOCV, outperforming both k-fold and leave-one-out cross-validation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Diagnostic tests are used to support decision-making in medicine, and their quality is commonly measured by Receiver Operating Characteristic (ROC) curve analysis. An ROC curve describes the ordered sequence of trade-offs between the diagnostic test correctly/incorrectly detecting a disease (Pepe 2003). In other words, ROC curve analysis describes disease detectability independently of the threshold effects. In addition, it is independent of disease prevalence (Metz 1978). ROC curve analysis is particularly useful when the data distributions are skewed or there are different benefits/costs for correctly/incorrectly detecting a disease (Fawcett 2006).
In this work, we assume that the diagnostic test is carried out by a classifier inferred from data using a machine learning algorithm. The classifier can be a non-linear function of multiple input variables. The prediction performance of the classifier cannot be reliably estimated on the data it was inferred from. For example, a 1-nearest neighbor classifier has always perfect fit to the data it was inferred from, but this estimate provides no information about its actual prediction performance on new data. Rather, the performance estimate should be done on independent test data (i.e., data not used in the training of the classifier). Consequently, the available data is split into training and test sets.
In medical research, the available data for an ROC curve analysis is usually scarce, as data sets tend to be small in sample size (Berisha et al. 2021), varying from some dozens to a few hundreds of observations. For this reason, splitting the available data into training and test set may be a challenge when evaluating the performance of a classifier. Hence, cross-validation methods provide means to make the most of the data in the evaluation process. Furthermore, cross-validation is suitable for all types of classifiers (i.e., linear, and non-linear) since a complex classifier cannot obtain high performance simply due to overfitting, as the evaluation of the classifier is always performed on data not used in the training phase.
Several cross-validation methods produce a ranking of data and hence could be used when ROC curve analysis is wanted. The most common cross-validation methods, leave-one-out cross-validation (LOOCV) and k-fold cross-validation, have been shown to produce biased estimates of the area under the ROC curve (AUC) (Airola et al. 2011; Forman and Scholz 2010; Montoya Perez et al. 2019; Parker et al. 2007; Smith et al. 2014). LOOCV and pooled k-fold cross-validation (PkFCV) produce rankings of data, unlike averaged k-fold cross-validation, but the biased estimates of AUC imply biased estimates of the ROC curves as well. Leave-pair-out cross-validation (LPOCV) has been shown to reduce the bias in AUC estimation in comparison to LOOCV and PkFCV (Airola et al. 2011), but does not provide a ranking of data and thus cannot be used when ROC curve analysis is wanted. Tournament leave-pair-out cross-validation (TLPOCV) has then been introduced as a method that provides a ranking of data while also producing an almost unbiased estimate of the AUC (Montoya Perez et al. 2019).
Computation time is something that must be taken into account when using methods that require many repetitions, e.g. simulation study, permutation test (Golland et al. 2005) or nested cross-validation (Varma and Simon 2006). All cross-validation methods may be time-consuming if the learning process is not efficient because in cross-validation a model is trained as many times as how many hold-out sets there are. In TLPOCV for a sample of size n, there are \(\frac{n\left( n-1\right) }{2}\) pairs that will be one by one left out as a hold-out set and the model is learned on the rest of the data. In other words, \(\frac{n\left( n-1\right) }{2}\) prediction functions are learned in total when executing TLPOCV. Tournament leave-pair-out CV is the least biased cross-validation method that also produces a ranking of data, but is very time-consuming as the sample size is increased.
Quicksort is a classical sorting algorithm, which can be used to create a ranking of data in a time-efficient manner (Ailon and Mohri 2008). In this article, we combine quicksort with LPOCV and show that the resulting ranking is as reliable for the ROC curve analysis as the ranking produced by TLPOCV. This means that quicksort leave-pair-out cross-validation (QLPOCV) is faster than TLPOCV but equally good.
The structure of the article is as follows: In Sect. 2 preliminary information is presented and previously developed methods are described in more detail. After that the quicksort algorithm is described, and our modification to add leave-pair-out cross-validation to it is presented in Sect. 3. The quality of the presented method is estimated with an experimental study by using both generated data and a real data set. The setup and the results of the experimental study are presented in Sect. 4. Finally, the article is concluded and future research is discussed in Sect. 5.
2 Preliminaries
2.1 ROC curve analysis
Receiver Operating Characteristic curve analysis is a statistical tool for describing the performance of a test that returns continuous results instead of directly returning binary labels (Bradley 1997; Fawcett 2006; Hanley and McNeil 1982; Pepe 2003). In our case, the continuous results are predictions made by a function \(f:{\mathcal {I}}\rightarrow {\mathbb {R}}\) which is returned by a machine learning algorithm. The labels are then obtained by a binary classifier
where t is a decision threshold for a positive label, i is the index of a datum whose label is being predicted and \(f_{{\mathcal {D}}_{\text {training}}}\) is a prediction function that is learned on data \({\mathcal {D}}_{\text {training}}\). For convenience, the subscript notation of training data is used only when necessary. In this article the prediction functions are inferred from a fixed training set. Sometimes the estimates of this kind are said to be conditional on the training data (Airola et al. 2011; Hastie et al. 2017). As we focus on binary classifiers, a data set is assumed to consist of data points from two classes, which are referred to as positives and negatives. Usually, the positives are associated with label 1 and negatives with 0 or -1. The feature representation of data is not relevant while explaining the methodology. Thus, we refer to the data points only by their indices. The index set of a sample of size n is \({\mathcal {I}}= \left\{ 1,2, ... ,n\right\} = {\mathcal {I}}_{+}\cup {\mathcal {I}}_{-}\), where \({\mathcal {I}}_{+}\) and \({\mathcal {I}}_{-}\) are subsets of the indices of positive and negative data points, respectively.
The quality of a classifier is often measured by several conditional probabilities. The probability of correctly predicting a positive sample unit as positive \({\mathbb {P}}\left( C_{t}\left( i\right) = 1 |i\in {\mathcal {I}}_{+}\right)\) is called sensitivity or true positive rate (TPR). Similarly for the negatives: \({\mathbb {P}}\left( C_{t}\left( i\right) = 0 |i\in {\mathcal {I}}_{-}\right)\) is called true negative rate or specificity. Also, the type I and II errors can be denoted as conditional probabilities \({\mathbb {P}}\left( C_{t}\left( i\right) = 1 |i\in {\mathcal {I}}_{-}\right)\) and \({\mathbb {P}}\left( C_{t}\left( i\right) = 0 |i\in {\mathcal {I}}_{+}\right)\), respectively. Type I error is also called false positive rate (FPR), and type II error false negative rate (Shapiro 1999). Later in this article, terms TPR and FPR are mostly used but sometimes sensitivity and specificity are used instead, so please note the connections between the terms: TPR is exactly the same as sensitivity, and FPR is the complement of specificity: \({\mathbb {P}}\left( C_{t}\left( i\right) = 1 |i\in {\mathcal {I}}_{-}\right) = 1 - {\mathbb {P}}\left( C_{t}\left( i\right) = 0 |i\in {\mathcal {I}}_{-}\right)\).
In ROC curve analysis, the conditional probabilities are calculated for every possible \(t\in \left( -\infty , \infty \right)\) as the predicted labels vary as a function of t. An ROC curve visualizes the sequence of all possible pairs \(\left( \text {FPR}\left( t\right) ,\text {TPR}\left( t\right) \right)\) of sensitivity and specificity values ordered by the values of t. The data can be ranked from the smallest to the greatest according to the predicted values: \(f\left( i_{1}\right) \le f\left( i_{2}\right) \le \dots \le f\left( i_{n}\right)\). If \(t < f\left( i_{1}\right)\), every datum is predicted to belong to the positive class, which implies that \(\text {TPR} = 1\) and \(\text {FPR} = 1\). If \(f\left( i_{j}\right) \le t < f\left( i_{j+1}\right)\) for any \(j\in \left\{ 1,\dots ,n-1\right\}\), data points \(i_{j+1},\dots ,i_{n}\) are predicted to belong to the positive class and data points \(i_{1},\dots ,i_{j}\) to the negative class, and \(\text {TPR} \in \left[ 0,1\right]\) and \(\text {FPR} \in \left[ 0,1\right]\) depending on the real labels. If \(f\left( i_{n}\right) < t\), every datum is predicted to belong to the negative class implying that \(\text {TPR} = 0\) and \(\text {FPR} = 0\).
The qualities of classifiers with different prediction functions can be compared by their ROC curves. An ROC curve goes from (0,0) to (1,1) in a unit square. The closer it is to the top left corner of the square, i.e. point (0,1), the better the classifier is. If the curve reaches that point, there exists a threshold for the classifier that perfectly separates the negatives from the positives. The ROC curve is a straight line on the diagonal from (0,0) to (1,1) if the classifier predicts the labels at random. There are several ways to estimate an ROC curve (Gonçalves et al. 2014). In this article, we use an empirical estimator in a way that is described in Sect. 4.1.
In addition to comparing the ROC curves, also the areas under them can be used to compare the qualities of the classifiers. Area under the ROC curve is a summary statistic of the ROC curve and its value is the probability that the classifier ranks a randomly chosen positive unit higher than it ranks a randomly chosen negative unit \({\mathbb {P}}\left( f\left( j\right) < f\left( i\right) |j\in {\mathcal {I}}_{-}, i \in {\mathcal {I}}_{+}\right)\).
True ROC curve and the corresponding AUC value are not usually known. If f was tested on a large \(\left( n\rightarrow \infty \right)\), representative sample, the TPR and FPR could be considered as population level probabilities. Then the resulting ROC curve is an approximation of the true ROC curve and its AUC approximates the true AUC.
2.2 Cross-validation for ROC curve analysis
If there was access to a large enough data set, it could be divided into separate training and test data sets so that f could be learned from the training data and the evaluation of C could be done on the independent test data as was described in Subsect. 2.1. However, in medical data analysis the available data set is usually so small that dividing it into training data and independent test data would lead to a situation where there are not enough data to learn a reliable model nor enough data to validate it. Cross-validation (CV) is then commonly used.
In cross-validation, the data set is repeatedly split into separate training and test sets. For each split, the training set is used to train f and build C, and the test set is then used to measure the performance of the obtained classifier. In k-fold CV, the data set is divided into k disjoint equally sized subsets, i.e. folds. Then the learning and performance evaluation are repeated k times so that each fold is exactly once used as test set and the other \(k-1\) times included in the training data. In order to perform ROC curve analysis, a pooled version of k-fold CV is needed. In PkFCV all predictions are gathered together and then an ROC curve is drawn or an AUC value is calculated. If only AUC value is wanted, then also averaged k-fold CV could be used. In that, an AUC value is calculated for each test fold and the final estimate of the AUC is the average of them (Bradley 1997). A special case of k-fold CV is leave-one-out CV, where \(k = n\). Then pooling is the only possible approach even for AUC estimation.
However, it may be risky to use pooled CV for AUC estimation, because the estimates may be biased (Airola et al. 2011; Forman and Scholz 2010; Montoya Perez et al. 2019; Parker et al. 2007; Smith et al. 2014). The bias is a consequence of comparing predictions from different CV rounds. Then the compared predictions are made by different prediction functions \(f_{{\mathcal {I}}\setminus l}\), where \(l\in \left\{ 1,\dots ,k\right\}\) refers to the data points in the \(l^{\text {th}}\) fold, which may differ a lot from one another if the learning algorithm is unstable. A learning algorithm is unstable if a small change in the training data changes the output of the algorithm greatly (Shalev-Shwartz and Ben-David 2014).
An extreme case of this pooling bias in CV can be demonstrated by the following example (Montoya Perez et al. 2019): Let us assume that the prediction function returns the probability of a positive label in the training data, which means that the prediction for an independent test set is a constant and AUC = 0.5. However, for LOOCV \(\widehat{\text {AUC}}_{\text{LOO}} = 0\), since the negative left out data points will always have higher predicted probability than the positive ones. The opposite effect of falsely getting \(\widehat{\text {AUC}}_{\text{LOO}} = 1\) can also take place with standard software implementations, for example, with the balanced support vector machine implementation in scikit-learn (Pedregosa et al. 2011), that automatically adjusts weights inversely proportional to class frequencies. Also, with PkFCV, such negative bias for AUC may appear if the left out fold has different class proportions than the training folds.
Leave-pair-out CV is an averaging based CV method, that has been shown to produce an almost unbiased estimate of the AUC (Airola et al. 2011)
where H is the Heaviside step function
The idea of LPOCV is to hold out each pair consisting of one positive and one negative unit as a test set and then use averaging when calculating the AUC value. Thus, only the predictions made with one prediction function are compared while also maximizing the size of training data. LPOCV is good for the AUC estimation, but cannot be used for ROC curve analysis, because it does not provide all the information that is needed.
As was explained in the Sect. 2.1, a ranking of data is required in order to perform ROC curve analysis. This can be obtained, while producing the almost unbiased estimate of AUC, by expanding the idea of LPOCV: in tournament leave-pair-out CV (Montoya Perez et al. 2019) each possible pair, not only the positive-negative pairs, is once held out as test set. The ranking is generated by sorting tournament scores of the sample units. A tournament score
describes how many times data point i was predicted to have a higher value than the other sample units. The reliability of TLPOCV has been validated experimentally (Montoya Perez et al. 2019). In TLPOCV, a prediction function is learned \(\left( {\begin{array}{c}n\\ 2\end{array}}\right)\) times. Therefore, it is time-consuming if there is even slightly larger data set.
3 Quicksort leave-pair-out cross-validation
Balcan et al. (2008) and Ailon and Mohri (2008) have studied the reduction from ranking to classification. Ailon and Mohri have shown that quicksort can be used to decrease the time complexity of the preference-based ranking setting. Quicksort is a sorting algorithm originally presented by Hoare (1962). It is based on the divide-and-conquer paradigm, which means that the problem is divided into sub-problems which are recursively solved, and finally the results of the sub-problems are combined to form the result of the original problem (Cormen et al. 1990). Ailon and Mohri (2008) have shown that their "algorithm produces high-quality global rankings in a time-efficient manner" and proved bounds for the ranking quality. This approach serves as the basis for the fast CV algorithm proposed in our study.
The algorithm presented by Ailon and Mohri (2008) uses a pairwise preference function, that takes as input two data points and predicts which one is more likely to belong to the positive class. The preference function is learned from a training data set using a machine learning algorithm. The simplest way to construct a ranking of data points for a new data set with the pairwise preference function is to compute a tournament graph where all data points are compared, and then sort the data according to number of "wins" in the tournament. This however requires \(O\left( n^2\right)\) comparisons. Ailon and Mohri propose a method that can compute an accurate ranking for data with only \(O\left( n\log n\right)\) comparisons as follows. First, a pivot element p is picked uniformly at random from the set of data points. Then, the pre-learned preference function is used to define the order of the pivot element and the other data points. This way two sub-problems are created and then the same process is run on the smaller sets of data points. One of the subsets consists of the data points that are preferred to the pivot element and the other subset consists of data points for which the pivot element is preferred to them. The algorithm returns the data points in order from the most preferred to the least preferred.
Quicksort leave-pair-out CV consists of the same steps as described above. What is different now is that instead of using a pre-learned preference function, the comparisons are done by using LPO prediction functions \(f_{{\mathcal {I}}\setminus \left\{ i,p\right\} }\) for data points i and pivot element p. In our calculations the order of preference is reversed. In QLPOCV, the hold-out pairs are formed so that the pivot element is paired with every other data point in the current set. This procedure is described for an index set \(I\subseteq {\mathcal {I}}\) in supplemental materials in Algorithm 1. In addition, a demonstration of executing the algorithm is presented in the supplemental materials together with discussion related to a graph presentation of the comparisons in TLPO and QLPO CV and the effect of possibly existing cycles on the order returned by QLPO. If there is at least one cycle in the corresponding graph, the order returned by QLPOCV depends on the randomly selected pivot elements.
The time complexity of quicksort is \(O\left( n\log n\right)\) in terms of number of calls to a preference function (Ailon and Mohri 2008). As there is randomness in the algorithm, the exact number of comparisons cannot be calculated. Hoare (1962) states that the average number of comparisons required by quicksort is \(2\ln {2}\times n\log _{2}n\). Another formula for the expected value of the number of comparisons together with a formula for variance of the number of comparisons has been discussed by (Iliopoulos 2013). It can be seen in Fig. 1 how remarkable the difference between the number of comparisons of quicksort and TLPOCV is as the sample size is increased. Since the time complexity of QLPOCV is the same as that of quicksort, it can be seen that QLPOCV requires a lot smaller number of comparisons than TLPOCV in order to produce a ranking of the data.
4 Experimental study
An experimental study was conducted in order to estimate the quality of the ranking produced by QLPOCV. Those results were compared with the results of TLPO, LOO and pooled 10-fold (P10F) CVs. The quality of the results was measured by ROC curves and AUC values. Three classification methods and both simulated and real data were used in the study.
4.1 Methods
The study was conducted as a Monte Carlo study, where the calculations were repeated and the results are described by averages and credible intervals (Edwards et al. 1963). The empirical ROC curves were vertically averaged at fixed FPRs. The fixed points were \(\left\{ 0, \frac{1}{|{\mathcal {I}}_{-}|}, \frac{2}{|{\mathcal {I}}_{-}|},\dots ,\right. \left. \frac{|{\mathcal {I}}_{-}|- 1}{|{\mathcal {I}}_{-}|}, 1 \right\}\). As the empirical ROC curve is a step function, there may be two different TPR values at one FPR value. The averages were calculated separately for lower and higher TPR values at the given FPR. A credible interval of an ROC curve is a range where the ROC curve is with a given probability. We used symmetric 95 % credible intervals, which means that they are formed by excluding equal amount (2.5 %) of the smallest and the largest TPR values at the fixed FPRs. True ROC curves and their AUC values were approximated so that the whole sample \({\mathcal {I}}\) was used for training the model and an independent test set was used for evaluating the model.
A classification algorithm determines the form of the prediction function f. The classification methods used in the study were regularized least-squares (RLS), logistic regression (LR) and random forest (RF). Even though RLS and LR are similar methods, they are both included in the study because LR is commonly used and RLS is better optimized for this type of problem. In RLS the learning task is to minimize a penalized version of the least-squares function (Hoerl and Kennard 1970; Rifkin et al. 2003). Also the version of LR that was used, was penalized. The penalty that was used in both classifiers is called Tikhonov regularization (Rifkin and Lippert 2007) and it is used to control the model complexity and prevent overfitting. Random forest is different from the other two classification methods in that it is neither linear nor parametric. RF is an ensemble method and uses bagging (Breiman 2001).
Python 3.7.6 was used to generate the simulation data and compute the predictions. ROC curve analysis, calculations of AUC values and visualizations were done with R 4.1.3. For RLS method, including fast cross-validation algorithms, we used RLScore library (Pahikkala and Airola 2016) implementation with regularization parameter set to one. For logistic regression and RF we used functions from Scikit-learn library (Pedregosa et al. 2011). For logistic regression we used l2 penalization with C=1 and liblinear solver, and for RF one hundred decision trees in a forest.
4.2 Simulation
4.2.1 Setup
In the first part of the experimental study, we used a non-linear generative model presented by Luckett et al. (2021) as a baseline for the data generation process. In that process, the features \({\mathbf {X}}\sim N_{n_{\text {f}}}\left( \mu {\mathbf {Z}},\sigma ^{2}{\mathbf {I}}\right)\), where \(n_{\text {f}}\) is the number of features, \({\mathbf {Z}}\) is a vector of ones or negative ones determining the sign of the distributions mean values, and \({\mathbf {I}}\) is an \(n_{\text {f}} \times n_{\text {f}}\) identity matrix. The probability for positive mean values is q and for negative mean values \(1-q\). Variance of each feature is a constant \(\sigma ^{2}\) and there is no explicit covariance between the features. A datum is in the positive class with a probability that is obtained by scaling expression \({\mathbf {X}}^{\top }\beta + X_{1}^{2} + X_{2}^{2} + 4X_{1} X_{2}\) to range \(\left[ 0,1\right]\) by a sigmoid function.
We added another parameter to the above expression so that we could measure the effects of the linear and non-linear parts:
We generated data populations of size \(10^{6}\) with parameters
and \(\theta\) was varied from 0 to 1 in steps of 0.25. From every population, we sampled an independent test data of size \(10^{4}\) and 1 000 separate samples of sizes 30 and 100 with fractions of positives 10 % and 50 %. The sample sizes and fractions of positives are similar to the ones that have been used previously (see (Airola et al. 2011; Montoya Perez et al. 2019)) and are chosen so that the simulation study reflects real world situations with small and possibly highly unbalanced data sets.
The data distributions of the positive and the negative classes are more or less overlapping in every population as is demonstrated in Fig. 2 for the first two features. When \(\theta = 0\), only features \(X_{1}\) and \(X_{2}\) are used in expression (1). In Fig. 2 it can be seen that the distributions of the positives and negatives are then highly overlapping and hence there is not much of a signal that could be used for predicting the class label. As \(\theta\) is increased, the linear combination of the features starts to increase the signal in data. According to the Fig. 2, the distributions are the least overlapping when the linear generative model has been used for generating the population data.
4.2.2 Results
In Fig. 3 are presented the resulting ROC curves of four settings of the simulation study. The selected cases are the extremes in two aspects: 1) The cases with the smallest and the largest number of positives and 2) the non-linear and linear data generation processes. It can be seen that the ROC curves are different from one case to another. For the case where the sample size is 30 and fraction of positives is 10 % (upper row), the average ROC curves for QLPO and TLPO are approximately equal to the approximated true ROC curve, whereas for P10F and LOO CV there can be seen a negative bias, especially when the learning algorithm is the random forest. For balanced samples of size 100 (lower row) such bias is not observed. The estimates are accurate but not precise as the credible intervals demonstrate. For unbalanced samples of size 30, the credible intervals cover almost the entire possible area. The credible intervals narrow as the number of positives is increased.
Average ROC curves and 95 % credible intervals over 1 000 repetitions for quicksort leave-pair-out, tournament leave-pair-out, pooled 10-fold and leave-one-out cross-validation methods for logistic regression, random forest and regularized least-squares classification algorithms. The subplots are for different sample sizes, fractions of positives and weights of the linear term: a) sample size = 30, fraction of positives = 0.1, and \(\theta = 0.0\), b) sample size = 30, fraction of positives = 0.1, and \(\theta = 1.0\), c) sample size = 100, fraction of positives = 0.5, and \(\theta = 0.0\), and d) sample size = 100, fraction of positives = 0.5, and \(\theta = 1.0\)
The ROC curves are quite different also when comparing them in the second aspect. When the data are generated by the non-linear generative model, the ROC curves are on average along the diagonal, as can be seen in the left column of Fig. 3. This means that for this kind of data, the learning algorithms are not able to learn anything and the predictions are made at random. Then again, the ROC curves are closer to the top left corner when the data are generated by the linear generative model (see the right column of the figure). Then there is some signal, which can be used for predicting the labels.
The shape of the ROC curve is obviously directly related to the area under it. The AUC values of every case of the simulation study are presented in Fig. 4. In that, one can see even more easily that the closer the data generation process is to the linear model, the better the algorithms are able to distinguish the positive class from the negative, i.e. the higher the AUC values. Again, it is visible that increasing the number of positives has a decreasing effect on the width of the credible intervals. Nevertheless, there are no large differences in the performance between the machine learning algorithms. When comparing the cross-validation methods to one another while keeping other parameters constant, the average \(\widehat{\text {AUC}}_{\text{QLPO}}\) and \(\widehat{\text {AUC}}_{\text{TLPO}}\) values are equal to the approximated true AUC whereas average \(\widehat{\text {AUC}}_{\text{P10F}}\) and \(\widehat{\text {AUC}}_{\text{LOO}}\) values are slightly less. Meaning that, QLPO and TLPO CV are equally good, and P10F and LOO CV are slightly worse.
Average and 95 % credible intervals of the AUC value over 1 000 repetitions for quicksort leave-pair-out, tournament leave-pair-out, pooled 10-fold and leave-one-out cross-validation methods in the unbalanced and balanced data cases for logistic regression, random forest and regularized least-squares classification algorithms
4.3 Real data
Analogous calculations were computed as well on a medical data set in order to validate the results of the simulation study. The real data set is the same as has been used in a previous study by Montoya Perez et al. ( 2019). The data set consists of 85 876 voxels, of which 9 268 are labelled as positives and 76 608 as negatives, and six voxel-wise features extracted from Diffusion-weighted imaging (DWI) parametric maps of 20 patients with histologically confirmed prostate cancer in the peripheral zone. The features are the parameters derived using DWI decay modelling: ADCm, ADCk and K as described in Toivonen et al. (2015) and the corresponding Gabor texture for each parametric map (Gabor-ADCm, Gabor-ADCk, Gabor-K). In previous studies, these features have shown potential for discriminating voxels belonging to prostate cancer (Ginsburg et al. 2011; Langer et al. 2009; Merisaari and Jambor 2015; Montoya Perez et al. 2016; Toivonen et al. 2015).
The calculations were repeated as many times as how many disjoint training sets were possible to sample from the real data. Hence, the number of repetitions is different for different sample sizes. Balanced samples of sizes 30 and 100 were drawn from the data without replacement 617 and 185 times, respectively. Each sample was once used as training data and the remaining data were used as independent test set. Finally, the results were averaged over repetitions.
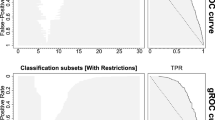
The average ROC curves and AUC values are presented in Fig. 5. These results are mainly consistent with the results of the simulation study. Now that the data are balanced, and there is some signal in the data, the ROC curves are above the diagonal and the credible intervals do not completely cover the possible area. The average \(\widehat{\text {AUC}}_{\text{QLPO}}\) and \(\widehat{\text {AUC}}_{\text{TLPO}}\) are closer than the average \(\widehat{\text {AUC}}_{\text{P10F}}\) and \(\widehat{\text {AUC}}_{\text{LOO}}\) to the approximated true AUC. The biggest difference in these and the simulation study results is the strong negative bias that is observed with logistic regression when LOO or P10F CV is used. A smaller negative bias is observed with RLS when LOO or pooled 10-fold CV is used. Sometimes this happens because of unstable classifiers and the bias is a consequence of pooling the results from different cross-validation rounds and thus from different classifiers.
Results of the real data experimental study for quicksort leave-pair-out, tournament leave-pair-out, pooled 10-fold and leave-one-out cross-validation methods for logistic regression, random forest and regularized least-squares classification algorithms. Upper row: Average ROC curves and their 95 % credible intervals. Lower row: Average AUC values and 95 % credible intervals
4.4 Runtime analysis
In Sect. 3 the time complexity of QLPO algorithm was compared to the time complexity of TLPOCV in terms of number of comparisons as a function of sample size. In order to compare the predicted values of two sample units i and j, a LPO prediction function \(f_{{\mathcal {I}}\setminus \left\{ i,j\right\} }\) needs to be learned. Thus, the number of prediction functions to be learned equals the number of comparisons for QLPO and TLPO cross-validations. Then again, for P10F the number of prediction functions needed is 10 for every sample size, and for LOOCV it is equal to the sample size.
The training time depends on the algorithm and how it is implemented. We noticed already when running the experimental study, that RLS is much faster than LR, and LR much faster than RF. Random forest is obviously slower because it is an ensemble method and every estimate is based on one hundred decision trees, meaning that for every prediction function, 100 decision trees need to be learned.
In order to highlight the importance of the choice of a learning algorithm and a cross-validation method on runtime, we executed a runtime analysis separately from the experimental study. The calculations were repeated 100 times, computed 10 in parallel on a computer equipped with Intel(R) Core(TM) i7-11800H (2,30 GHz). Average runtimes for sample sizes 30, 50 and 100 are presented in Table 1.
The results clearly demonstrate what was already discussed: RLS is a lot faster than LR, and RF is a lot slower than LR and RLS. What is little surprising is that TLPOCV actually is faster than QLPOCV when RLS is used. The implementation of RLS is based on utilizing Sherman-Morrison-Woodbury formula, which makes it possible to compute all predictions in time no greater than what it takes to learn the model once on the data (Pahikkala et al. 2008; Pahikkala and Airola 2016). Hence, it is faster to call it once with a large number of pairs than call it several times with a smaller number of pairs. Nevertheless, the difference is some milliseconds, so the choice between the methods should not affect the total runtime too much even when the calculations are repeated many times.
For LR and RF the runtimes are of completely different magnitude and vary more from one cross-validation method to another. For these algorithms, P10FCV is clearly the fastest, TLPOCV the slowest, and QLPOCV and LOOCV are about the same in the middle, as is expected according to the time complexities. Runtime is not the primary criterion, so even though the pooled CV methods are faster, they are not recommended over QLPOCV, because of the possible existence of the pooling bias.
5 Conclusion and future research
In this work, we presented a method called quicksort leave-pair-out cross-validation which combines leave-pair-out cross-validation with quicksort algorithm and produces a ranking of data that is needed for ROC curve analysis. The importance of this method compared to the previously presented TLPOCV is that in QLPOCV a remarkably smaller number of prediction functions needs to be learned in order to produce equally good results. The change from TLPOCV to QLPOCV decreases the time complexity from \(O\left( n^2\right)\) to \(O\left( n\log n\right)\) in terms of number of prediction functions to be learned as a function of sample size.
We compared this new method to previously existing methods by an experimental study consisting of two parts: one with generated data and the other with real data. The results demonstrated that QLPOCV produces as good estimates as TLPOCV of both the ROC curves and the AUC values. In addition, LOO and pooled 10-fold CV were used in the experimental study. As expected, the results of the latter CV methods were worse than the results of the former CV methods. Finally, we conducted a runtime analysis to highlight the importance of the choice of classifier and CV method. The runtime analysis showed that QLPOCV is significantly faster method than TLPOCV unless RLS classifier is used. Thus, based on the experimental study presented in this article, QLPOCV should be preferred to TLPOCV when other classifiers are used.
Several new questions arose while conducting this research, and they will be considered as future research. One of them is what is already briefly discussed in the supplemental material: the effect of possibly existing cycles in the corresponding tournament graph. The order returned by QLPOCV depends on the randomly chosen pivot elements if the learning algorithm is so unstable that there are cycles. This means that there is some probability distribution for the different orders, and the estimates of ROC curves and AUC values depend on the randomly chosen pivot elements, too. This kind of theoretical research would lead to achieving deeper understanding about the properties of the method.
References
Ailon N, Mohri M (2008) An efficient reduction of ranking to classification. In: 21st Annual Conference on Learning Theory, COLT (2008)
Airola A, Pahikkala T, Waegeman W, De Baets B, Salakoski T (2011) An experimental comparison of cross-validation techniques for estimating the area under the ROC curve. Comput Stat Data Anal 55(4):1828–1844
Balcan MF, Bansal N, Beygelzimer A, Coppersmith D, Langford J, Sorkin GB (2008) Robust reductions from ranking to classification. Mach Learn 72(1–2):139–153
Berisha V, Krantsevich C, Hahn PR, Hahn S, Dasarathy G, Turaga P, Liss J (2021) Digital medicine and the curse of dimensionality. NPJ Dig Med 4(1):1–8
Bradley AP (1997) The use of the area under the ROC curve in the evaluation of machine learning algorithms. Pattern Recogn 30(7):1145–1159
Breiman L (2001) Random forests. Mach Learn 45(1):5–32
Cormen T.H, Leiserson C.E, Rivest R.L (1990) Introduction to algorithms. MIT press
Edwards W, Lindman H, Savage LJ (1963) Bayesian statistical inference for psychological research. Psychol Rev 70(3):193
Fawcett T (2006) An introduction to ROC analysis. Pattern Recogn Lett 27(8):861–874
Forman G, Scholz M (2010) Apples-to-apples in cross-validation studies: pitfalls in classifier performance measurement. ACM SIGKDD Explor Newsl 12(1):49–57
Ginsburg S, Tiwari P, Kurhanewicz J, Madabhushi A (2011) Variable ranking with PCA: Finding multiparametric MR imaging markers for prostate cancer diagnosis and grading. In: International Workshop on Prostate Cancer Imaging. Springer, pp 146–157
Golland P, Liang F, Mukherjee S, Panchenko D (2005) Permutation tests for classification. In: International conference on computational learning theory, pp. 501–515. Springer
Gonçalves L, Subtil A, Oliveira MR, de Zea Bermudez P (2014) ROC curve estimation: an overview. REVSTAT-Statistical J 12(1):1–20
Hanley JA, McNeil BJ (1982) The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 143(1):29–36
Hastie T, Tibshirani R, Friedman J (2017) The elements of statistical learning, 2 edn. Springer series in statistics New York
Hoare CA (1962) Quicksort. Comput J 5(1):10–16
Hoerl AE, Kennard RW (1970) Ridge regression: biased estimation for nonorthogonal problems. Technometrics 12(1):55–67
Iliopoulos V (2013) The quicksort algorithm and related topics. Ph.D. thesis, Department of Mathematical Sciences, University of Essex
Langer DL, Van der Kwast TH, Evans AJ, Trachtenberg J, Wilson BC, Haider MA (2009) Prostate cancer detection with multi-parametric MRI: Logistic regression analysis of quantitative t2, diffusion-weighted imaging, and dynamic contrast-enhanced MRI. J Magnet Resonance Imag: An Official J Int Soc Magnet Resonance in Med 30(2):327–334
Luckett DJ, Laber EB, El-Kamary SS, Fan C, Jhaveri R, Perou CM, Shebl FM, Kosorok MR (2021) Receiver operating characteristic curves and confidence bands for support vector machines. Biometrics 77(4):1422–1430
Merisaari H, Jambor I (2015) Optimization of b-value distribution for four mathematical models of prostate cancer diffusion-weighted imaging using b values up to 2000 s/mm2: simulation and repeatability study. Magn Reson Med 73(5):1954–1969
Metz CE (1978) Basic principles of ROC analysis. Semin Nucl Med 8(4):283–298
Montoya Perez I, Airola A, Boström PJ, Jambor I, Pahikkala T (2019) Tournament leave-pair-out cross-validation for receiver operating characteristic analysis. Stat Methods Med Res 28(10–11):2975–2991
Montoya Perez I, Toivonen J, Movahedi P, Merisaari H, Pesola M, Taimen P, Boström P.J, Kiviniemi A, Aronen H.J, Pahikkala T, et al. Diffusion weighted imaging of prostate cancer: prediction of cancer using texture features from parametric maps of the monoexponential and kurtosis functions. In: 2016 Sixth International Conference on Image Processing Theory, Tools and Applications (IPTA), pp. 1–6. IEEE (2016)
Pahikkala T, Airola A (2016) RLScore: regularized least-squares learners. J Mach Learn Res 17(1):7803–7807
Pahikkala T, Airola A, Boberg J, Salakoski T (2008) Exact and efficient leave-pair-out cross-validation for ranking RLS. Proceedings of AKRR 2008:1–8
Parker BJ, Günter S, Bedo J (2007) Stratification bias in low signal microarray studies. BMC Bioinform 8(1):1–16
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, Blondel M, Prettenhofer P, Weiss R, Dubourg V, Vanderplas J, Passos A, Cournapeau D, Brucher M, Perrot M, Duchesnay E (2011) Scikit-learn: Machine learning in Python. J Mach Learn Res 12:2825–2830
Pepe MS (2003) The statistical evaluation of medical tests for classification and prediction. Oxford University Press, USA
Rifkin R, Yeo G, Poggio T et al (2003) Regularized least-squares classification. Nato Sci Series Sub Series III Comput Syst Sci 190:131–154
Rifkin RM, Lippert RA (2007) Notes on regularized least squares. Tech. rep, Massachusetts Institute of Technology, Cambridge
Shalev-Shwartz S, Ben-David S (2014)Understanding machine learning: From theory to algorithms. Cambridge university press
Shapiro DE (1999) The interpretation of diagnostic tests. Stat Methods Med Res 8(2):113–134
Smith GC, Seaman SR, Wood AM, Royston P, White IR (2014) Correcting for optimistic prediction in small data sets. Am J Epidemiol 180(3):318–324
Toivonen J, Merisaari H, Pesola M, Taimen P, Boström PJ, Pahikkala T, Aronen HJ, Jambor I (2015) Mathematical models for diffusion-weighted imaging of prostate cancer using b values up to 2000 s/mm2: Correlation with gleason score and repeatability of region of interest analysis. Magn Reson Med 74(4):1116–1124
Varma S, Simon R (2006) Bias in error estimation when using cross-validation for model selection. BMC Bioinform 7(1):1–8
Acknowledgements
The authors thank Peter J. Boström for his collaboration allowing the use of the real medical data for evaluating the proposed method. We would like to thank CSC, the Finnish IT Center for Science, for providing us with extensive computational resources.
Funding
Open Access funding provided by University of Turku (UTU) including Turku University Central Hospital. The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Academy of Finland (grants 311273, 313266).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author(s) declare(s) that there is no conflict of interest.
Availability of data and material
Data and results are available online: https://github.com/rimanu/QLPO.
Code availability
The code is available online: https://github.com/rimanu/QLPO.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Numminen, R., Montoya Perez, I., Jambor, I. et al. Quicksort leave-pair-out cross-validation for ROC curve analysis. Comput Stat 38, 1579–1595 (2023). https://doi.org/10.1007/s00180-022-01288-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00180-022-01288-3