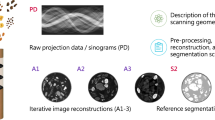
Abstract
Over recent years, many approaches have been proposed for the denoising or semantic segmentation of X-ray computed tomography (CT) scans. In most cases, high-quality CT reconstructions are used; however, such reconstructions are not always available. When the X-ray exposure time has to be limited, undersampled tomograms (in terms of their component projections) are attained. This low number of projections offers low-quality reconstructions that are difficult to segment. Here, we consider CT time-series (i.e. 4D data), where the limited time for capturing fast-occurring temporal events results in the time-series tomograms being necessarily undersampled. Fortunately, in these collections, it is common practice to obtain representative highly sampled tomograms before or after the time-critical portion of the experiment. In this paper, we propose an end-to-end network that can learn to denoise and segment the time-series’ undersampled CTs, by training with the earlier highly sampled representative CTs. Our single network can offer two desired outputs while only training once, with the denoised output improving the accuracy of the final segmentation. Our method is able to outperform state-of-the-art methods in the task of semantic segmentation and offer comparable results in regard to denoising. Additionally, we propose a knowledge transfer scheme using synthetic tomograms. This not only allows accurate segmentation and denoising using less real-world data, but also increases segmentation accuracy. Finally, we make our datasets, as well as the code, publicly available.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Since its invention by Hounsfield et al. [1], X-ray computed tomography (X-ray CT) has become increasingly popular amongst industrial users and researchers studying the hidden inner structure of many different biological and non-biological systems. In applications ranging from medicine and cell biology to material science and geology, X-ray microcomputed tomography is an essential imaging tool which provides researchers with detailed representations of what they are studying, usually in the form of a volumetric reconstruction. However, one of the most difficult tasks associated with almost all tomography studies is the semantic segmentation of the resulting volumetric datasets, which requires many human hours, in some cases weeks, if done manually. Fortunately, recent automatic techniques based upon deep learning [2,3,4,5,6] have shown an unprecedented improvement in segmentation accuracy, where sufficient annotated data are available. Deep learning approaches achieve this by learning an end-to-end mapping between the tomography volumes and the annotated data, and later apply it for the automatic annotation of new tomography volumes.
Nevertheless, there are situations in which only a limited time is available for the CT scanning of an object, which consequentially results in only a small number of projections being captured. This is the case in high-speed 4D CT datasets capturing metal corrosion, which comprise sets of consecutive tomograms depicting time-related events within the samples. Since only a small number of projections is attained for each component tomogram of a 4D dataset, higher numbers of tomograms may be captured, which increases the time-resolution of the dataset. Unfortunately, acquiring projections from a low number of angles makes the reconstruction of these undersampled tomograms an ill-posed problem. This results in low-quality reconstructions in which the level of noise is high and ray or streak artefacts are prevalent; see Fig. 1a–d. To address this issue, iterative reconstruction methods such as the Conjugate Gradient Least-Squares method (CGLS) [7] are used. Figure 1a, c shows a dataset reconstructed using the non-iterative Filter Back Projection method (FBP) [8] using a high number of projections. These methods are, however, designed to work analytically and regardless of the nature of the tomograms; when only a small number of projections are available, even with the use of the CGLS [7] method, there is still a high degree of noise, making further processing complex.
Cross section of our real-world dataset reconstructions and annotations. a FBP reconstruction from 3601 projections [9] (these are cropped and centred around the metallic pin of Label 1), b CGLS reconstruction from 91 projections [10] (these are cropped and centred around the metallic pin of Label 1), c, d are zoomed versions of a, b, respectively, and e are annotations for the same area [11]. Label 0 in b refers to the air outside the base material, Label 1 in e refers to the base material, Label 2 are the magnesium deposits and Label 3 are the air pockets
Motivated by the above, we present an end-to-end deep learning approach for the dual tasks of denoising and segmentation of reconstructions of undersampled tomograms, also referred to as low-dose or sparse-angle tomograms. As is common practice during 4D dataset collection, highly sampled tomograms with a large number of projections can be captured before or after the time-critical portion of the collection, “book ending” the lower resolution data with high fidelity, single time-point representations. It is therefore possible, using as prior the high-quality volume reconstructions of these representative highly sampled tomograms, to tackle both the denoising and segmentation of the low-quality reconstructions of the undersampled tomograms.
We achieve this by stacking two proposed DenseUSeg networks, described here. The stacked networks are inspired by Newell’s [12] approach of Stacked Hourglasses, with the first denoising the input and the second segmenting it. Stacking two networks creates an intermediate result that allows for an intermediate loss to be applied in order to obtain the denoised output. Additionally, stacking the networks enables the denoised output to be utilised by the follow-up network to achieve more accurate segmentation predictions. Training time required is also decreased, compared to training two networks separately. This is because there are software and hardware advantages due to parallelism, which allow more efficient and faster training when it is performed simultaneously rather than separately.
Finally, we propose a knowledge transfer scheme in order to achieve high levels of segmentation accuracy in situations where the available amount of annotated data is small. We achieve this by employing Kazantsev et al.’s [13] TomoPhantom software to construct synthetic tomograms, using random geometrical objects (e.g. cylinders, ellipsoids, etc.) to approximate the different classes present in the real-world data. The synthetic tomograms are then used to train networks. This way, key knowledge such as the attenuation levels of different classes and shape descriptive features can be transferred to the parameters of these networks’ convolutional layers, using procedurally synthesised tomograms that are by default already annotated. These networks are then used as a stepping stone, initialising the parameters of new networks before the latter are further trained/fine-tuned using a small amount real-world data. Provided that the synthetic tomograms approximate a broader family of different real-world tomograms with the same segmentation classes, these “pretrained” networks can be used multiple times for the fine-tuning of new networks targeted towards the segmentation or denoising of different real-world datasets. Therefore, the additional time cost of manufacturing synthetic tomograms and training “pretrained” networks is not repeated for every new real-world tomogram, the user would like to segment or denoise. As we further explore in Sects. 4 and 5, the knowledge learned from synthetic tomograms increases segmentation accuracy considerably. In fact, we find that the necessary amount of annotated data required for a highly accurate segmentation after training/fine-tuning, can be as low as 5% of the original training set of segmentation instances. This is essential for the application of our approach in industrial practice, where the absence of the sufficient annotated tomographic data is frequent.
In summary, in this paper:
-
1.
We propose our DenseUSeg architecture that extends upon the DenseSeg architecture [5] through the introduction of a “decoder” module similar to UNet [14].
-
2.
We propose an end-to-end deep learning approach that produces two different outputs, the denoised counterpart of the input and its segmentation map. This network, named Stacked-DenseUSeg, utilises the denoised output to further improve its segmentation accuracy compared to DenseUSeg and other state-of-the-art methods.
-
3.
We demonstrate that our approaches outperform the state-of-the-art over the Intersection over Union (IoU) metric for the semantic segmentation of low-quality reconstructions of undersampled tomograms.
-
4.
We show that our approaches are at least as good as state-of-the-art methods over the Peak Signal-to-Noise Ratio (PSNR) and Structural SIMilarity index (SSIM) for the denoising of the low-quality reconstructions. The reason the same methods are used for both the denoising and segmentation comparisons is that current published CT denoising approaches have not exploited the most recent advances in deep learning architectures that have proven to offer noteworthy results regardless of task.
-
5.
We propose a scheme that transfers knowledge from procedurally generated synthetic tomograms. This allows the use of less real-world data than before, higher segmentation accuracy and also reduced time required for training.
2 Related work
Over the recent years, deep learning approaches have offered remarkable results in the area of semantic segmentation. Generally, these approaches can be split into two categories: approaches using pixel-wise networks that label one pixel (or voxel) of the input and approaches using fully convolutional neural networks that label multiple pixels (or voxels).
In approaches that employ pixel-wise networks, the networks’ task is to predict the class of a single pixel at a time, using as input the pixel values of a neighbourhood around that pixel. One of the first attempts at pixel-wise segmentation came from Ciresan et al. [15] who used a convolutional network followed by fully connected layers to predict the class of individual pixels. Its performance was later improved by Pereira et al. [16], who deepened the architecture and demonstrated that by careful image preprocessing, segmentation accuracy can be improved further. Moeskops et al. [17] and Havaei et al. [18] introduced multiple pathways into the network to increase accuracy using multi-scale contextual information. In a recent publication, Kamnitsas et al. [19] introduced Conditional Random Fields (CFRs) to gain additional accuracy in segmentation. However, as shown by Salehi et al. [20], fully convolutional neural networks tend to have both better performance and faster training and testing times.
Fully convolutional approaches are designed to offer predictions for the whole (or a subsection) of the input image or volume. First proposed by Long et al. [21], fully convolutional networks replace the fully connected layers of pixel/voxel-wise networks with upscaling operations. It is then possible to attain accurate predictions for a more coarse version of the input, compared to a single pixel/voxel. Based on this principle, Badrinarayanan et al. [22] demonstrated that the classification architectures that were previously proposed in other publications can be easily modified for segmentation tasks. By partially using the same architecture (VGG-16 [23] in their case), it is possible to initiate the parameters of certain layers using the parameters of pretrained classification networks, leading to higher segmentation accuracy compared to using random parameter initialisation, due to the knowledge transferred from the classification networks. This fact, combined with recent publications [24, 25] which demonstrate how networks trained on synthetic datasets can be useful for real-world applications, is what inspired our proposed knowledge transfer scheme.
Later, Ronneberger et al. [14] introduced the UNet architecture, which uses a “decoder” module with upscaling operations, convolutions and skip connections to achieve new standards in segmentation accuracy. Similarly, Cicek et al. [3] modified this architecture for 3D volumes, proposing 3D-UNet. In 2016, an important year for deep learning, He et al. [26] presented their residual network architecture, ResNet, that combats the gradient degradation which appears in deep networks when they start to converge. Their proposed architecture, by summing feature maps with ones calculated in earlier layers, allowed the deep network to converge more easily by effectively learning just the residual information between the different feature maps. Their contribution inspired many deep learning approaches, some of them performing semantic segmentation [4, 4, 27]. One of the proposed approaches, introduced by Chen et al. [4], demonstrated that deep supervision [28] can offer an additional improvement of the segmentation accuracy. Recently, innovating towards the same direction, Huang et al. [29] proposed their DenseNet architecture offering better performance by further minimising the degradation problem. Using concatenation instead of summation, their proposed network is able to more flexibly utilise the feature maps produced by early layers, and converge even faster. Their architecture was also adapted for the task of segmentation, with Bui et al. [5] and Zhang et al. [6] demonstrating improved segmentation results. Lately, Li et al. [30] proposed such a dense network that includes a “decoder” module; however, their proposal for handling the network’s large memory demands is to sum the upscaled feature maps with the ones passed by skip connections. Nevertheless, as we will describe in Sect. 3 there is still a way to use concatenation instead of summation, which as demonstrated by Huang et al. [29] is a more flexible way to combine feature maps.
Deep learning methods for denoising the reconstructions of undersampled (low-dose or sparse-angle) tomograms have also been developed, though these are much fewer in number. Most of these [31,32,33,34,35,36] utilise simple deep learning architectures, as the recent advantages of deep learning have only been recently showcased. Because of that, the recent and deeper architectures like [3,4,5] have yet to be adopted for this task. However, based on the general trend in deep learning network architectures, deeper architectures tend to offer higher performances regardless of the task (classification, segmentation, etc.). For this reason, in Sect. 5 we compare our proposed architectures against the more recent and deeper [3,4,5] approaches.
3 The network architectures
3.1 Densely connected layers
In this section, we will showcase our DenseUSeg and Stacked-DenseUSeg networks (see [37] for the code) as shown in Fig. 2. Specifically, the Stacked-DenseUSeg network is composed of two of our proposed DenseUSeg networks stacked sequentially; the DenseUSeg architecture is formed by expanding upon the DenseSeg architecture proposed by Bui et al. [5] through the introduction of a “decoder” module similar to UNet [14]. This “decoder” using consecutive transpose convolutions and additional convolutional layers is proven to offer more accurate segmentation results. Similar to DenseSeg, our architecture is composed of the Dense Blocks proposed by Huang et al. [29]. Specifically, the output feature maps of the \(l\mathrm{th}\) Dense Module within a Dense Block are:
where \([x_0,x_1,\ldots ,x_{l-1}]\) is the concatenation of the feature maps formed in the earlier \(0,1,\ldots ,l-1\) layers and \(H_l\) the composite function of the \(l\mathrm{th}\) Dense Module. Based on the above and as shown in Fig. 2c, each Dense Module receives as input the concatenation of the all the previous Dense Modules’ feature maps. The number of feature maps after l Dense Modules is therefore \(k_0 + l \cdot g\) where \(k_0\) is the number of the initial feature maps provided to the Dense Block and g the growth rate, is the number of feature maps produced by each Dense Module.
Based on preliminary testing, we set the growth rate of DenseUSeg and Stacked-DenseUSeg to \(g=64\), in an effort to achieve good performance before further increases lead to diminishing returns.
3.2 DenseUSeg
In our architecture, before the use of the first Dense Block the input passes through 3, \(3 \times 3 \times 3\) convolutional layers that produce \(k_0=32\) feature maps followed by a \(2 \times 2 \times 2\) convolution with stride 2 to reduce their scale. The architecture then consists of 4 Dense Blocks each with 4 Dense Modules (see Fig. 2c) separated by 3 Transitional layers. Each of the Dense Modules contains a bottleneck \(1 \times 1 \times 1\) convolutional layer and a \(3 \times 3 \times 3\) layer. To mitigate the extensive increase in feature maps as the network becomes deeper, Transition Blocks are placed between each of the Dense Blocks. These include a \(1 \times 1 \times 1\) convolution that decreases the number of feature maps by a factor of \(\theta =0.5\), followed by a \(2 \times 2 \times 2\) convolution with stride 2 for scale reduction. This way the network condenses the information extracted from each scale of the input, before continuing with the extraction of information at smaller scales. It is also important to mention that, before each convolutional layer present either in the Dense and Transition Blocks, batch normalisation and the Rectified Linear Unit (ReLU) are employed to improve robustness [38] and introduce the necessary nonlinearity [39], respectively. Similarly, these layers are also present in the initial three convolutional layers, but are placed after the convolutions. Finally, a Dropout layer with rate \(p=0.2\) is placed at the end of each Dense Module, since this has been proven to help with over-fitting problems [40].
Moving forward to the architecture’s decoder, DenseUSeg uses a scheme similar to UNet [14]. The network upscales the feature maps of the last Dense Block using a \(4 \times 4 \times 4\) transpose convolutional layer with stride 2 and padding 1, similar to the transpose convolutions of U-Net [14]. These features are then concatenated with the feature maps of the earlier Dense Blocks with the same resolution (same number of voxels for the height, width and depth). The feature maps then pass through two consecutive convolutional layers, and the upscaling is repeated with similar transpose convolutions. Since the number of feature maps is very extensive after the Dense Blocks, both the transpose convolutions and the first of the two consecutive convolutions in the decoder reduce the number of convolutions by half. When the feature maps reach the same resolution as the input volume, they pass through a \(1 \times 1 \times 1\) convolution and a Softmax Layer which produces the probabilities for each of the voxels belonging in each of the classes. By assigning each voxel to the class which has the highest probability of belonging, the final segmentation map is formed.
3.3 Stacked-DenseUSeg
As Fig. 2b shows, when combining two DenseUSeg nets in our Stacked-DenseUSeg architecture the initial 3 convolutional layers (the ones between the feature maps A and B) are common. The denoising part of our architecture then produces the final feature maps of the denoised output (feature maps C), followed by the denoised prediction for the input (output D). These then pass through \(3 \times 3 \times 3\) convolutions (with batch normalisation and ReLU layers) and the resulting feature maps are summed with initial feature maps (B) to provide the input for the segmentation DenseUSeg. Since the input to the segmentation DenseUSeg receives information from the 3 convolutional layers (feature maps C) of the denoising DenseUSeg, the 3 convolutional layers are not repeated in the segmentation DenseUSeg. As shown in Sect. 5, the existence of a Dropout layer in DenseUSeg and DenseSeg [5] improves the performance only in the case where the input is noisy and has no effect when the input is noiseless. Therefore, while for the denoising DenseUSeg a Dropout layer with rate \(p=0.2\) is selected, in the respective segmentation network the rate is set to \(p=0\) as it receives a denoised input. The connection between the networks is inspired by Newell et al. [12], in which the authors stack multiple hourglass networks with intermediate supervision in order to improve the network’s final predictions. For our architecture, the goal is to produce a denoised output which assists in the accuracy of the final segmentation output. The stacked approach is necessary, as it allows for the creation of an intermediate output of the same size (number of voxel dimensions) as the input. This enables the application of an intermediate loss function to the network, in order to obtain the desired denoised output. Our Stacked-DenseUSeg architecture is therefore able to offer two different outputs within one forward pass. This is translated into a potential reduction of training time of two networks, since software can better schedule the required operations and may perform many of them in parallel. While training of the two networks could occur in parallel but separately, the segmentation output would not benefit from the denoised output due to the separation. Furthermore, in applications where only the denoising or the segmentation is desired, it is possible both the intermediate and the final output to be used either for segmentation or denoising. This means that the intermediate output would not be a secondary output, but just an entry point for additional supervision. As shown in [12, 28] additional supervision increases the accuracy of the final output.
Cross section of our synthetic tomogram reconstructions and annotations used during the knowledge transfer experiments. a, e FBP reconstruction from 3601 projections from synthetic tomograms, respectively, with [41] and without [42] rotating artefacts (these are cropped and centred around the metallic pin of Label 1), b, f CGLS reconstruction from 91 projections from synthetic tomograms, respectively, with [43] and without [44] rotating artefacts (these are cropped and centred around the metallic pin of Label 1), c, d, g, h are zoomed versions of a, b, e, f, respectively, and i are annotations for the same area [45]. Similar to the real-world data Label 0 in b refers to the air outside the base material, Label 1 in e refers to the base material, Label 2 are the magnesium deposits and Label 3 are the air pockets
4 Data and training
4.1 Representative real-world and synthetic tomograms
The real-world highly sampled tomogram used to train our networks was produced at the Diamond Light Source I13-2 beamline [46], and is used in [47]. The tomogram depicts a droplet of salt water on top of a 500 micron aluminium pin with magnesium deposits. It was captured at the end of a time-series of undersampled tomograms that forms a 4D dataset, itself part of a study to measure the corrosion of the metallic pin by the salt-water droplet over time [48]. Since the tomograms captured by Diamond Light Source are \(2160 \times 2560 \times 2560\) in size after reconstruction, even a single tomogram can in practice provide plenty of training instances after cropping. The use of a single tomogram (dataset) is a reflection of the limited availability of annotated data in this domain, where a set of multiple annotated tomograms cannot be a representative of future tomograms, due to the unique nature of the samples being imaged in every tomogram collection. This means that proposed networks in this domain would have to retrained on new representative highly sampled tomogram, that accompany the corresponding new time-series of undersampled tomograms. Manual annotation of these large \(2160 \times 2560 \times 2560\) highly sampled tomograms is a difficult and time-consuming process. As we note later, we incorporate the availability of only a very limited amount of training data into the core of our method, introducing synthetic data to help overcome this limitation.
The real-world, highly sampled tomogram, used in Sect. 5 is captured at the end of a time-series, and as such it has been possible to capture it with a high number of projections, namely 3601 instead of the 91 projections in the 4D dataset tomograms. Collection of these “book end” representative highly sampled tomograms is common practice for researchers dealing with 4D datasets. These tomograms are attained either before or after the capture of a 4D dataset, providing tomograms that can be used as reference, but without interfering with the time-critical part of the study. Even though not a part of the 4D dataset, they contain valuable information about the features of the samples under study which the researchers want to identify and label. Therefore, they can be used for training segmentation or denoising networks, which will later be applied on the reconstructions of the 4D dataset’s tomograms. In addition to such real-world, highly sampled datasets, synthetic, highly sampled tomograms are later utilised for the knowledge transfer experiments, as shown in Fig. 3. Such artificial datasets allow for better generalisation of the denoising and segmentation tasks, which elevates the accuracy of the networks. Furthermore, they are useful when the amount of annotated real-world tomograms is limited, which would otherwise not allow for the training of highly accurate networks.
For training, three highly sampled tomograms are used. Two of them are synthetically constructed and the other is a real-world tomogram. The real-world tomogram is reconstructed using the Savu Python package [49] and the synthetic tomograms using Kazantsev et al.’s [13] TomoPhantom software. For the synthetic tomograms, TomoPhantom constructs them using a file made by the user that lists a number of virtual objects placed in a virtual stage, in a similar way to how real-world objects are placed in a stage prior to them being CT scanned. The virtual objects can constructed to be of any shape formed from a predetermined list of geometrical shapes. The aforementioned file details the shape, size and orientation of the virtual objects as well as their virtual absorption rate, which will determine their brightness during the rendering of the synthetic reconstructions.
4.2 Tomogram reconstructions used for training
For each of the tomograms synthetic or real-world, two reconstructions are produced. The first are made using the Filter Back Projection method (FBP) [8] algorithm and all 3601 available projections. The resulting high-quality reconstructions [9, 41, 42] provide the ground truth used in the denoising experiments and in the denoising part of Stacked-DenseUSeg. The second reconstructions are made with the Conjugate Gradient Least-Squares method (CGLS) [7] reconstruction algorithm and uses only 91 projections (\(1\mathrm{st},41\mathrm{st},\ldots ,3601\mathrm{st}\) projections) ignoring some of the intermediate projections. These low-quality reconstructions [10, 43, 44], which are provided as input during training, have the same visual quality as the reconstructions from the undersampled tomograms in the 4D datasets, when only a small number of projections are used. The reason two different algorithms are used for the two reconstructions per tomogram is that the iterative CGLS method improves the reconstruction quality when applied to undersampled tomograms, while it offers poorer results than the more simple FBP method when applied to highly sampled tomograms. Figures 1, 3 provide cross sections of the two reconstructions for the real-world and synthetic tomograms, respectively, as well as their annotations, which depict the classes that the network aims to segment. The annotations [11] of the real-world tomogram are obtained using Luengo et al.’s SuRVoS [50] software tool using the high-quality reconstruction [9].
The synthetic tomograms’ annotations are predetermined [45]. Based on these the respective tomogram projections and reconstructions are created. In contrast to the real-world annotations, the synthetic annotations are not human estimations of the ground truth, rather they are the absolute ground truth. This is very significant since due to imaging artefacts, even in highly sampled tomograms, human annotators may unintentionally introduce bias to the data that will later be used for training, which in turn may reduce the accuracy of the final segmentation [51]. Synthetic datasets lack this potential bias, and by transferring knowledge from networks trained on synthetic data to networks that infer on real-world data, the latter may offer predictions in difficult cases which are potentially more accurate than human annotators. Two synthetic tomograms are used during experiments: one more realistic, with simulated rotating artefacts (Fig. 3a–d) that are also present in real-world data, and the other less realistic dataset, without such artefacts (Fig. 3e–h). This effectively gives us two levels of physical simulation quality to compare. For both tomograms, similarly to the real-world tomogram, there are two subsequent reconstructions: a high-quality reconstruction which provides the ground truth used in the denoising experiments and in the denoising part of Stacked-DenseUSeg (Fig. 3a,c,e,g), and a low-quality reconstruction used as input during training, because it has the same visual quality as the reconstructions from the real-world undersampled tomograms (Fig. 3b,d,f,h).
There are four classes to be segmented: the air outside the base material (Label 0 in Figs. 1c, 3i), the base material which is the water and aluminium (Label 1 in Figs. 1f, 3i), the magnesium deposits within the base material (Label 2 in Figs. 1f, 3i) and lastly the air pockets within the base material (Label 3 in Figs. 1f, 3i). In the synthetic tomograms, 5000 randomly oriented and sized virtual ellipsoids are randomly placed within a virtual cylinder for each class and assigned Labels 2 and 3. These simulate the magnesium deposits and air pockets, respectively, with the quantity chosen to roughly simulate the class balance present in the real-world data. The virtual cylinder in turn simulates the metallic pin found in the real-world data. Note that, due to the large number of ellipsoids introduced, there is occasional overlapping of ellipsoids which diversifies the shapes present in the tomograms (see Fig. 3c,g,i).
Since the training of the network is performed using a single tomogram at a time, they are normalised (if real-world data; synthetic tomograms are produced pre-normalised) and split into multiple non-overlapping \(64 \times 64 \times 64\) samples. Before entering the network each of the samples is either randomly rotated between 90, 180, 270 degrees, or horizontally or vertically mirrored. Of these samples, 70% are randomly chosen for training, 10% for validation and 20% for testing. This results in, for the real-world tomogram, 3723 samples used for training, 532 for validation and 1065 for testing. For the synthetic tomograms, 4646 samples are used for training, 663 for validation and 1329 for testing. The samples are cropped from the area of the tomograms that contain the desired classes to segment, because most voxels in the tomograms belong to Label 0 and sampling from the tomograms as a whole would create a great class imbalance. Thankfully, in most datasets (4D datasets or single tomograms) of this type [48] the region of interest which has to be segmented, is located in a single area of the tomogram. Using simple low-level imaging analysis techniques (edge detection, thresholding, etc.), it is possible to isolate this area, reducing the volume that the networks have to segment, and allowing for shorter inference times. During inference, the samples are chosen to be overlapping, and the final prediction is obtained by averaging predictions in areas of overlap. The predictions, that are the probabilities of which class each voxel belongs to, are used after averaging to infer the class of each voxel. Non-overlapping samples are used during training for clear separation between training, validation and testing samples. During inference, however, overlapping eliminates potential border artefacts in the output subvolumes and helps resolve potential uncertainties in the class of some voxels.
4.3 Hyperparameters and training settings
For backpropagation, the stochastic gradient-based optimisation method Adam [52] is used, with minibatches of 8. For DenseUSeg the learning rate is set to \(10^{-5}\), while for the Stacked-DenseUSeg the corresponding learning rates for both the segmentation part the denoising part of network are displayed in Fig. 4a. The reason for this duality of learning rates in the Stacked-DenseUSeg is that the denoising part has to train faster than the segmentation part, as the second is partially dependent on the output of the first. In all tested networks (ours and the state-of-the-art), a weight decay [53] of 0.0005 is used, that acts as a L2 regularisation which applies penalties when weights/parameters collectively get too large, which can lead to overfitting. Additionally, Fig. 4 displays the learning rate schedule regarding both the case of using a high amount of training data (Fig. 4a) and the case of low amount of training data (Fig. 4b).
The learning rate schedule of Stacked-DenseUSeg, both for its denoising part and its segmentation part. a is regarding the case where all or 50% of the training data is used for training (the later case probably for the case of knowledge transfer) and b is regarding the case where 10%, 5% or 1% of the training data is used for training or fine-tuning of a pretrained net to real-world applications following a knowledge transfer scheme
Each epoch lasts for 300 minibatches and the network passes through 100 epochs during training. In order to verify the accuracy of the network variations presented in Sect. 5, every training session is repeated 3 times in a threefold fashion. For each of these iterations the 70-10-20 split between training, validation and testing samples, respectively, is kept; however, the samples for validation and testing are different with no common samples between each iteration. The weighted cross-entropy loss criterion is used for the segmentation output and the Mean Square Error (MSE) for the denoising one. The weights for the cross-entropy are calculated as:
where P is a vector of the pixel counts of the different classes and S is a vector that contains the number of samples of each class is present. The combined criterion for Stacked-DenseUSeg is:
where \(\lambda \) balances the two losses and it is empirically set to 10. The \(L_{Cross\_Entropy}\) loss uses as ground truth the segmentation annotations while the \(L_{Mean\_Square\_Error}\) the high-quality reconstruction.
4.4 Metrics and execution time
As a segmentation metric, the Intersection over Union (IoU), also known as the Jaccard index, is used. Specifically, the IoU metric of class l is,
where \(v \in C_l\) are the voxels that belong to class l, are \(p \in C_l\) the predicted voxels that belong to class l. The IoU of class l can be calculated also from IoU’s second definition as seen in the second fraction of Eq. 4. \(TP_l\) is the total number of voxels predicted to belong to class l that actually do, \(FP_l\) is the total number of voxels predicted to belong to class l but do not, and \(FN_l\) the total number of voxels predicted to belong to any other class other than l, but they do in fact belong to class l. Therefore, the IoU of a class is penalised both by predicting voxels to belong to other classes when they belong to the class under consideration, and by predicting voxels to belong to this class when in fact they belong to others. This means that it is a good metric for segmentation, since it does achieve high values by overpredicting or underpredicting certain segmentation classes.
Furthermore, we also use as a segmentation metric, the F1 score, also known as the Sørensen–Dice coefficient (DSC). The F1 score of class l is,
The F1 score and the IoU are similar as they both penalise the same elements described earlier, in a similar but not identical manner. We include the F1 score as it is a popular metric for the measurement of segmentation accuracy [5, 30] in certain domains, which some readers may be more familiar with.
For denoising, the Peak Signal to Noise Ratio (PSNR) metric (which is a logarithmic representation of the mean square error, see Eq. 6), and the Structural Similarity Index (SSIM) [54] (see Eq. 7) are used to quantitatively evaluate the image restoration quality compared to the ground truth (in our case the high-quality reconstructions [9, 41, 42]). Namely PSNR is,
where \({\varvec{D}}\) is the denoised output of the network and \({\varvec{G}}\) the denoising ground truth. The PSNR metric represents the ratio between the maximum possible power of the signal, in our case is the low-quality reconstruction (produced with CGLS) of the high-sampled tomogram, and the power of noise that affects its fidelity and distance it from the ground truth, which in our case is the high-quality reconstruction (produced with FBP) of the high-sampled tomogram. PSNR is a metric of the signal-to-noise ratio and high PSNR values, expressed in the logarithmic decibel scale, signify higher restoration qualities. Additionally, the equation for the SSIM index is,
where \(\mu _{{\varvec{D}}}\) is the mean of \({\varvec{D}}\), \(\mu _{{\varvec{G}}}\) is the mean of \({\varvec{G}}\), \(\sigma _{{\varvec{D}}}\) is the variance of \({\varvec{D}}\), \(\sigma _{{\varvec{G}}}\) is the variance of \({\varvec{G}}\), \(\sigma _{{\varvec{DG}}}\) is the covariance of \({\varvec{D}}\) and \({\varvec{G}}\), L is the dynamic range of the voxel-values, and by default \(k_1=0.01\) and \(k_2=0.03\). The SSIM index opposite to PSNR, which estimates the signal-to-noise ratio and visual quality based on absolute errors, is a perception-based metric. It considers image degradation as a cause of quality decrease, but ignores illumination and contrast alterations that do not cause structural changes on what it is imaged. It detects inter-dependencies between spatially close pixels, and by estimating how much of them remain unchanged between the networks’ denoising predictions and the ground truth, is able to measure the perceptual improvement in the quality.
4.5 Pseudocode for our method
Based on the information provided by the earlier subsections, Algorithm 1 shows the pipeline for the training, validation and testing processes of our network Stacked-DenseUSeg.

Algorithm 1 also describes how we train, validate and test DenseUSeg and the other networks that we use as baselines in the following section. The only difference is the omission of the input \({\varvec{D}}\) if the network will be used for segmentation or input \({\varvec{A}}\) if will be used for denoising. Also, depending on the operation (denoising or segmentation) the corresponding loss function is used (mean square error or weighted cross-entropy, respectively). Algorithm 2 describes the inference process for Stacked-DenseUSeg.

Training for 100 epochs takes approximately 18 hours in PyTorch [55] on 4 NVIDIA Tesla V100s. Naturally, a termination condition can be employed during training/fine-tuning with lower amounts of training samples, since the best performing epoch comes early (see Fig. 7c–e), and so reduce the time needed for training even further.
5 Experiments
5.1 DenseUSeg and Stacked-DenseUSeg comparison to the state-of-the-art using real-world data
In this subsection, we present and compare the results obtained from our Stacked-DenseUSeg and DenseUSeg [37] architectures to the state-of-the-art methods of DenseSeg [5], 3D-UNet [3] and VoxResNet [4]. In the following experiments, we increase the growth rate of DenseSeg [5] to \(g=64\), as with the use of the original growth rate (16 in [5]), we observed very low performance in preliminary tests and additionally this setting (\(g=64\)) allows us to produce results comparable to DenseUSeg (which also has \(g=64\)).
The denoising networks are trained networks using the Mean Square Error (MSE) as the loss criterion, except for Stacked-DenseUSeg which is trained based on the combined loss described in Eq. 3 in Sect. 4. The input to the networks is the low-quality reconstruction [10], and the high-quality reconstruction [9] acts as ground truth. For the denoising task, the final Softmax layer in the networks is removed or replaced with a \(1 \times 1 \times 1\) convolution, if there is no one present before it.
This experiment is performed in order to determine which architecture is best for the task of denoising. As shown in Table 1 our DenseUSeg architecture performs similar to, or better than, other state-of-the-art methods across both PSNR and SSIM [54] metrics. As can be seen in Fig. 6 from mark (F), the DenseUSeg is able to restore the air pocket with the correct intensity and size. Furthermore, from mark (G) it can be seen that it is better able to restore the difficult case in which the base material consists of water and not aluminium than the other networks. Table 1 also displays the performance of Stacked-DenseUSeg’s denoised output, and while it is not as accurate as a single DenseUSeg trained for the task of denoising, it still outperforms DenseSeg [5] on PSNR and 3D-UNet [3] on both PSNR and SSIM. However, given that Stacked-DenseUSeg’s denoised output is its secondary and not primary output, its performance is noteworthy. Furthermore, it can be seen in Fig. 5a,b that both of our networks converge stably, and based on the standard deviation in Table 1 our networks’ predictions vary similarly to the other state-of-the-art methods.
Nevertheless, it is important to point out that in Fig. 6i–m, compared to Fig. 6h, the exact structure of the small internal components cannot be recovered. This is due to large undersampling of projections in the input (the undersampled tomogram) and, as can be observed in Fig. 6a, spatial information is permanently lost.
a Validation PSNR results for the denoising experiments, b Validation SSIM results for the denoising experiments, c Validation mean IoU for the segmentation experiments where the input is the high-quality reconstruction of the real-world tomogram, d Validation mean IoU for the segmentation experiments where the input is the low-quality reconstruction of the real-world tomogram. All values are averages between the 3 iterations of the threefold validation/testing and they are also averaged based on 10 epoch moving average. The error bar heights refer to the standard deviation that the approaches achieved based om a (3\({\times }\))10 epoch moving standard deviation
For the task of segmentation, DenseUSeg and the state-of-the-art methods are trained using as loss criterion the weighted cross-entropy described in Eq. 3 in Sect. 4. Two segmentation experiments are conducted. In the first, the high-quality reconstruction [9] (see Table 2a and Fig. 5c) is the input, and in the second the low-quality reconstruction [10] is the input.
In the first, the goal is to learn the accuracy of the different networks under optimal conditions. This is attained by the high-quality reconstruction due to the absence of noise and with potentially fewer of the artefacts which appear when using a smaller number of projections. In this experiment both DenseUSeg and DenseSeg are set with a Dropout rate of \(p=0\), as in preliminary tests it was seen that Dropout was not helpful when noise is absent. In addition, Stacked-DenseUSeg is not listed in Table 2a as this network is designed to upscale the low-quality reconstruction [10] by using the high-quality reconstruction [9] and the annotations of Fig. 1a,e, respectively, as ground truths; in this first segmentation experiment, we are already given optimal data as input.
Based on Table 2a and Fig. 5c, it can be deduced that under optimal data input conditions, DenseUSeg is able to outperform or match performance of the other methods in the IoU of individual classes and in the mean IoU. Also the same can be observed for the F1 score, the only exception being for class 0, where it is only slightly behind the F1 score for the class 0 of DenseSeg. Looking ahead to Table 2b and Fig. 5d, where the data are not optimal, it can be seen that the accuracy gap between the DenseUSeg and the second best approach, 3D-UNet [3] is greater, which provides clear evidence regarding the elevated accuracy that the DenseUSeg offers compared to the state-of-the-art. Additionally, as it can be seen in Fig. 5c DenseUSeg is similarly stable during training as the rest of the state-of-the-art methods. The confusion matrices of the methods presented by Table 2a can be seen in the Supplementary Fig. S1.
Moving to the second experiment, here we are using as input the low-quality reconstructions [37]. As can be seen from Table 2b and Fig. 5d, the DenseUSeg and Stacked-DenseUSeg achieve higher accuracy compared to the other state-of-the-art methods. Stacked-DenseUSeg performs notably better than the other approaches, presumably thanks to the denoised output. As can be seen in Table 2b, our DenseUSeg, and particularly our Stacked-DenseUSeg approach, segments more accurately the more challenging \(2^{nd}\) and \(3^{rd}\) labels, which due to the small size of objects that they represent lose much of their spatial details during the projection undersampling. Moreover, from Fig. 5d and the standard deviations presented in Table 2b, both approaches are similar in terms of stability to other state-of-the-art methods. In terms of qualitative results, examining the marks (A–E) in Fig. 6 shows that Stacked-DenseUSeg can offer more accurate segmentation predictions, presumably due to utilising its denoising module. Specifically, it is the only approach that accurately separates objects of the same class that are in close proximity [marks (B), (D) and (E)], and does not overestimate the size of the internal components in the tomogram [mark (A)]. Finally, the confusion matrices of the methods presented by Table 2b can be seen in the Supplementary Fig. S2.
As it was shown earlier, DenseUSeg performs similar to, or better than, other state-of-the-art methods regarding the denoise of the undersampled tomograms. This combined with DenseUSeg’s exceptional performance regarding the segmentation of undersampled tomograms, justifies our selection of two stacked-DenseUSegs for the formation of our proposed Stacked-DenseUSeg. Additionally, since stacking multiple networks is easier when these networks have the same structure (no complications connecting them), our selection of DenseUSeg for the denoising part is also justified despite the slightly better performance of the VoxResNet [4].
5.2 Experiments regarding knowledge transfer from synthetic tomograms
In the previous subsection, we demonstrated that our networks have similar, if not better, results to the state-of-the-art in regard to denoising low-quality reconstructions of undersampled tomograms, and outperform them in regards of semantic segmentation. However, typical to deep learning approaches, they require a sizeable amount of annotated real-world data for successful training before they can be applied to real-world applications. This is a challenge. Annotated micro-CT data are hard to come by and expensive to produce as they require many hours of manual labour. Nevertheless, through the development of software tools like the TomoPhantom by Kazantsev et al.’s [13], it is possible to automatically create both realistic synthetic tomograms and their corresponding annotations. Specifically, their annotations [45] (phantoms) are first designed and created by a user, and then their simulated tomographic counterparts [41,42,43,44], similar to capturing the described ground truth by a physical CT scanner, are produced by the software. This is a physical-based rendering, which, as we shall see (Fig. 3a–d), can even include imaging artefacts appropriate to CT scanners. This makes them ideal for use in conjunction with a deep network as they are physically viable, and are generated with their annotations. In this subsection, we will present how these tomograms can be used within a knowledge transfer scheme, use of our proposed Stacked-DenseUSeg network (the best performing in the previous section), to further increase its overall segmentation accuracy. Additionally, by simulating data we can run effective training using only a small amount of real-world annotated data.
Segmentation and Denoising Results on the real-world tomogram. a Input to the denoising and segmentation networks, the CGLS reconstruction from 91 projections [10], b Annotations: Label 0 in black refers to the air outside the base material, Label 1 in red refers to the base material, Label 2 in yellow are the magnesium deposits and Label 3 in white are the air pockets, c–g Segmentation output of the respective networks, h the FBP reconstruction from 3601 projections that is ground truth for the denoising networks, i–m are the denoised outputs of the respective networks when they work to denoise (a) using (h) as ground truth. For Stacked-DenseUSeg c and i are produced simultaneously and there is no need for retraining. Marks (A–G) are described in text
Inspired by Badrinarayanan et al. [22] where parameters of networks trained in similar tasks are used for parameter initialisation of new networks, and recent advances regarding of the use of synthetic data to increase the network’s performance [24, 25], we propose a knowledge transfer scheme where initial instances of Stacked-DenseUSeg are pretrained using synthetic tomograms, and these learned parameters are utilised in the parameter initialisation of new instances of Stacked-DenseUSeg, trained to perform the true task of denoising and segmenting real-world data. Using this process, the advantages are twofold. First, it enables the transfer of knowledge to the new real-world instances of Stacked-DenseUSeg, which allows for better generalisation in the way the network infers its predictions, leading to higher accuracy. Second, the transfer of knowledge allows for low-level information regarding the nature of X-ray tomography, which is easily simulated by synthetic analogues, to pass into the new networks, permitting the use of less annotated data to sufficiently train them for use in real-world applications.
As can be seen in Fig. 3, there are two different kinds of synthetic tomograms used during the knowledge transfer experiments, resulting in two different networks that are trained on them. The first synthetic tomogram lacks rotating artefacts [42, 44] (see Fig. 3e–h) and was generated to increase performance; however, it was suspected that the addition of rotating artefacts (see Fig. 3a–d) similar to the ones present in real-world tomograms, may yield a noteworthy improvement in the segmentation accuracy, leading to the creation of a second synthetic tomogram [41, 43]. As can be seen in Table 3, by initialising the network’s parameters using a Stacked-DenseUSeg already trained with a synthetic tomogram with rotating artefacts (which we refer to as “pretrained nets”), our network (Stacked-DenseUSeg as well), that predicts the real-world data is able to achieve a higher accuracy score in segmentation, versus no use of parameter initialisation.
Additionally, the network that is trained/fine-tuned with real-world data in order to offer predictions for the real-world use-cases is able to do so only using a small number of real-world annotated samples, without any accuracy sacrifices as the data annotated samples lessen. Specifically, it is only after training/fine-tuning with 1% of the real-world training samples, namely 37 \(64 \times 64 \times 64\) samples (approximately a \(213 \times 213 \times 213\) of annotated volume), that the accuracy is comparable to when nearly all the available real-world data is used, even for the more challenging 2 and 3 class labels. Even in the case where no fine-tuning with real-world data is employed, it is still able to achieve a noteworthy segmentation accuracy, which is testament to the quality of the synthetic data. Finally, by observing the standard deviation in Table 3 and the error bars in Fig. 7, it can be deduced that the network achieves more stable predictions, compared to the case where no pretrained net is used to initialise the parameters. The confusion matrices of the methods presented by Table 3 can be seen in the Supplementary Figs. S3,S4.
Regarding the parameter initialisation using the pretrained net derived from the synthetic tomogram without rotating artefacts [42, 44], it is can be observed both in Table 3 and in Fig. 7 that its use is not as advantageous as the pretrained net from synthetic data with rotating artefacts [41, 43]. This is especially true by examining the last rows of Table 3. When using the pretrained nets directly without fine-tuning, the performance of the pretrained net from the synthetic without rotating artefacts is especially poor. This is to be expected as the physical rendering is less realistic without these artefacts. However, in cases where the available amount of real-world training data is fairly small, the parameter initialisation using any pretrained net is essential. In any case, the improved performance via the use of any pretrained net (with or without rotation artefacts) compared to random parameter initialisation (not pretrained), signifies that is worth investing, creating the synthetic tomograms and training the pretrained nets despite of the additional time that both processes require.
Specifically, the dictation of a file that describes how TomoPhantom [13] should generate a synthetic tomogram, may take a only few minutes from the user to be completed. Alternatively, for more complex synthetic tomograms the above file can be partially generated via code. In contrast, the required partial annotation of a real-world tomogram, using special software like SuRVoS [50], may take from a few hours to a day for an experienced user to annotate, depending also on the complexity of the tomogram. In comparison, the complete annotation of a tomogram \(2160 \times 2560 \times 2560\) in size similar to ones used here, may require multiple days to weeks by a single annotator, or even multiple annotators. This means that the human involvement when using the knowledge transfer scheme may be for just one day compared to weeks in the case of a complete tomogram annotation. Combined with the couple of minutes, mentioned earlier, to dictate the file that would generate the synthetic tomogram, the human involvement would probably not exceed that of one day, compared to weeks required for a complete tomogram annotation. The additional computational time required for the synthetic tomogram rendering and the training for the generation of pretrained nets, may take close to 24 hours depending of course on the available hardware. During this time, no human involvement is required and again compared to weeks of manual work required for a complete tomogram annotation there is an obvious benefit. Furthermore, since the synthetic datasets can be general enough to simulate a variety of similar real-world tomograms, the aforementioned additional time required render new synthetic tomograms and create pretrained nets, might happen less frequently. Overall, below are all the steps for obtaining a Stacked-DenseUSeg with less real-world annotated data via knowledge transfer.
-
1.
Generation from the user of a file with the placement, orientation and absorption rate of virtual objects within a virtual stage that will generate a phantom.
-
2.
Generation of the earlier phantom by TomoPhantom [13]. The phantom is also a segmentation map.
-
3.
Generation with TomoPhantom of synthetic tomograms based on the earlier phantom. From this phantom, the following are generated:
-
(a)
A high-quality tomogram reconstruction using many projections (emulating a highly sampled tomogram) and also the FBP reconstruction algorithm [8].
-
(b)
A low-quality tomogram reconstruction using a limited number of projections (emulating an undersampled tomogram) and also the CGLS reconstruction algorithm [7].
-
(a)
-
4.
Training with Stacked-DenseUSeg using:
-
(a)
As input the previous low-quality synthetic reconstruction.
-
(b)
As denoising ground truth the previous high-quality synthetic reconstruction.
-
(c)
As segmentation ground truth the synthetic phantom/segmentation map.
-
(a)
-
5.
From the above, a pretrained net has been acquired.
-
6.
Partial manual segmentation of a real-world representative highly sampled tomogram using SuRVoS [50] or any other annotation tool.
-
7.
Parameter initialisation of a Stacked-DenseUSeg the earlier pretrained net. This Stacked-DenseUSeg is being training/fine-tuned with real-world data using:
-
(a)
As input, a low-quality reconstruction (produced with CGLS) of the earlier real-world highly sampled tomogram that is undersampled artificially.
-
(b)
As denoising ground truth, a high-quality reconstruction (produced with FBP) of the earlier highly sampled tomogram.
-
(c)
As segmentation ground truth, the tomogram annotations obtained the highly sampled tomogram via SuRVoS or any other tool.
-
(a)
-
8.
The final Stacked-DenseUSeg trained/fine-tuned with real-world data and initialised with a pretrained net generated using on synthetic data, will have high segmentation accuracy despite using only a small portion of real-world annotated data.
Validation mean IoU for the segmentation experiments where the input is the low-quality reconstruction of the real-world tomogram, and by initialising the parameters using pretrained nets produced from synthetic data with rotating artefacts [41, 43], without rotating artefacts [42, 44] or without the use of a pretrained net. a The network is then fine-tuned using 100% of available real-world training samples, b 50% of available real-world training samples, c 10% of available real-world training samples, d 5% of available real-world training samples, e 1% of available real-world training samples and lastly (f) mean IoU of the nets during their training with synthetic data, but tested on real-world data. As it can be seen using the pretrained net produced from training with a synthetic tomogram with rotating artefacts offers overall higher accuracy, even in when using very small amounts of real-world data for training/fine-tuning, while it is also very stable based on the error bars. All values are averages between the 3 iterations of the threefold validation/testing and they are also averaged based on 10 epoch moving average. The error bar heights refer to the standard deviation that the approaches achieved based om a (3\({\times }\))10 epoch moving standard deviation
Segmentation and Denoising results of Stacked-DenseUSeg with progressively lower amounts of the real-world training samples and by initialising the parameters using pretrained nets produced from synthetic data with rotating artefacts [41, 43], without rotating artefacts [42, 44] or without the use of a pretrained net. a The CGLS reconstruction input from 91 projections [10] that is the networks input, b the FBP reconstruction [9] from 3601 projections that is ground truth for the denoising part of Stacked-DenseUSeg, c Annotations: Label 0 in black refers to the air outside the base material, Label 1 in red refers to the base material, Label 2 in yellow are the magnesium deposits and Label 3 in white are the air pockets, d–f Using 100% of training samples for the 3 parameter initialisation options, g–i Using 50% of training samples, j–l Using 10% of training samples, m–o Using 5% of training samples, p–r Using 1% of training samples, s–t Inferring directly using the pretrained nets without any training/fine-tuning. Marks H–M are described in text
Moving on by viewing, Fig. 8 and specifically marks (H, I, K, M) we can also qualitatively infer the advantage of using a pretrained net from a synthetic tomogram with rotating artefacts [41, 43]. The obtained accuracy using only 5% and 10% of the available real-world training data is comparable to the use of 100% of the available real-world training data, but without any parameter initialisation with a pretrained net. Also, from marks (J, L) we can see that the network is more accurate for such difficult cases as these, where a great loss of spatial information is present due to the projection undersampling.
Finally, looking into Stacked-DenseUSeg’s secondary output, the denoised output, from Table 4 we can see that with the use of a pretrained net, the restoration quality is also increased. Nevertheless, the improvement is only slight and it is dependant on the amount of real-world training data used and the nature of the pretrained net utilised for parameter initialisation. It seems that for low amounts of real-world training data, both PSNR and SSIM [54] reach higher scores using the pretrained net from the synthetic tomogram with rotating artefacts [41, 43], which for the 5% and 10% of real-world used for fine-tuning is by a slight amount. Nevertheless, for the use of 1% or no fine-tuning the improvement is noteworthy, due to the synthetic data with rotating artefacts resembling more closely the real-world data. When higher amounts of real-world training data are available for fine-tuning, the pretrained net from synthetic tomogram without rotating artefacts [42, 44] achieves slightly higher performance, which is unexpected. One hypothesis is that this is because while training with the synthetic tomogram without rotating artefact, the high-quality reconstruction better represents the ground truth (no additional artefacts) than the high-quality reconstruction of the synthetic tomogram with rotating artefacts. It could therefore be a good idea for the future to combine the two synthetic tomograms during the training of a pretrained net. For that, the low-quality synthetic reconstruction which is the input would have rotating artefacts and the high-quality synthetic reconstruction (that is the denoising ground truth) would not have rotating artefacts. But to note, we can deduce from Table 4 the potential improvement is expected to be only slight. Also, in Table 4 we can observe that the standard deviation of both metrics is comparable between the use of a pretrained map or not, which indicates the denoised output remains consistent.
6 Discussion
Based on the earlier results of Sect. 5, our approaches DenseUSeg and Stacked-DenseUSeg (code available from [37]) outperform the state-of-the-art accuracy of DenseSeg [5], 3D-UNet [3] and VoxResNet [4] regarding the volumetric semantic segmentation of undersampled tomograms and the highly sampled tomograms (except from Stacked-DenseUSeg that is only applied on undersampled tomograms). Especially, our Stacked-DenseUSeg network, by producing a second denoised output, is able to utilise it to achieve higher segmentation accuracy compared to a single DenseUSeg dedicated for segmentation. Furthermore, DenseUSeg, in the task of denoising, is able to match VoxResNet in the SSIM metric, and be only slightly behind it in PSNR metric. This means that our DenseUSeg is well-suited for both the tasks of denoising and segmentation of undersampled CT tomograms, and therefore for the 4D datasets collected at Synchrotron facilities such as Diamond Light Source. Moreover, this is also the underlying reason that Stacked-DenseUSeg is composed of two stacked-DenseUSegs. Finally, Stacked-DenseUSeg denoising output is better than the other two methods, DenseUSeg and 3D-UNet, in both the PSNR and SSIM metric being the third best model in the denoising accuracy after DenseUSeg and VoxResNet in both metrics. This is a significant achievement considering that Stacked-DenseUSeg is focused on the dual tasks of denoising and segmentation.
Furthermore, using a knowledge transfer scheme described in Sect. 5.2, the use of synthetic tomograms to train initial versions of Stacked-DenseUSeg (pretrained nets) to initialise that parameters of Stacked-DenseUSeg versions operating on real-world data offers multiple advantages. Firstly, both the segmentation and denoising accuracy are increased. Using the pretrained net training on synthetic data with rotating artefacts is the most beneficial for the improvement of the segmentation accuracy. It seems the addition of further realistic elements in the form of rotating artefacts facilitates the creation of better quality pretrained nets. This means that networks that are trained/fine-tuned on real-data and use these pretrained nets for parameter initialisation, are better prepared for the task of segmentation. Additionally, with the use of knowledge transfer, a second advantage becomes apparent, that the same segmentation accuracy can be acquired with significantly less real-world highly sampled annotated data. This is highly significant as manual annotation of real-world CT reconstructions is difficult and time-consuming to obtain.
The last argument brings forward the third advantage of the use of knowledge transfer which is that time needed to employ the knowledge transfer scheme is considerably less that the time needed to manually annotated real-world CTs. In addition, the time needed for application of knowledge transfer is mainly computational time and the need for human involvement is minimal. Finally, knowledge transfer also improves the denoising accuracy; however, the improvement is not as noteworthy as that reported for segmentation. As reported at the end of Sect. 5.2, it may be a good future idea to use both synthetic tomograms for the training of a pretrained net. For each of the synthetic tomograms, there are two reconstructions, one low-quality reconstruction used as input and one high-quality reconstruction used as the denoising ground truth. It would therefore be a good idea to use the low-quality reconstruction of the synthetic tomogram with rotating artefacts as input and the high-quality reconstruction of the synthetic tomogram without rotating artefacts as the denoising ground truth. The use as input of the low-quality reconstruction of the synthetic tomogram with rotating artefacts would create pretrained nets that expect rotating artefacts in the input as they also appear in real-world data. The use as the denoising ground truth of the high-quality reconstruction of the synthetic tomogram without rotating artefacts would create pretrained nets that produce denoising outputs without any rotating artefacts. After the parameter initialisation of the resulting pretrained nets, the network operating on real-world data may potentially be better prepared and able to denoise more accurately, which may also improve its segmentation accuracy.
7 Conclusions
Concluding, in this paper, we propose the DenseUSeg (code available from [37]) architecture which is able to denoise and segment undersampled tomogram reconstructions in an end-to-end fashion. Inspired by Newell et al. [12], our Stacked-DenseUSeg architecture is constructed by sequentially stacking two of our proposed DenseUSeg networks and applying an intermediate inspection. Contrary, though, to Newell et al.’s approach our intermediate inspection is not regarding an early version of the segmentation outputs for a secondary denoising output. With this approach, our network is able to increase the low signal-to-noise ratio of the input undersampled tomogram and improve its segmentation accuracy. Furthermore, our proposed DenseUSeg network expands Bui et al.’s [5] architecture with the addition of a decoder module similar to the one proposed by Ronneberger et al. [14]. For the experiments of Sect. 5, our network performs well on both the tasks of denoising and segmentation, with the Stacked-DenseUSeg outperforming the other approaches in segmentation by utilising its denoising output.
Moreover, through the use of procedurally simulated synthetic tomograms, we propose a knowledge transfer scheme that can offer an increase on the semantic segmentation accuracy of Stacked-DenseUSeg, and produce a high level of accuracy. Specifically, the procedurally generated synthetic tomograms are used to train instances of Stacked-DenseUSeg, the pretrained nets. The parameters of the pretrained nets are then used to initialise the parameters of Stacked-DenseUSeg that will be trained/fine-tune in order to operate on real-world data. The resulting Stacked-Dense-USeg’s will have more generalisable knowledge for how to accurately segment the real-world tomograms, resulting in more consistent and accurate predictions. This knowledge also reduces the need for large amounts of annotated real-world data. This is also translated to a time reduction for the application of Stacked-DenseUSeg in new varieties of real-world data, and especially the reduction of time regarding human involvement.
References
Hounsfield, G.N.: Computed medical imaging. Science 210(4465), 22 (1980). https://doi.org/10.1126/science.6997993
Roth, H.R., Farag, A., Lu, L., Turkbey, E.B., Summers, R.M.: Deep convolutional networks for pancreas segmentation in CT imaging. In: Ourselin, S., Styner, M.A. (eds.) Medical Imaging 2015: Image Processing, vol. 9413 (SPIE, 2015), vol. 9413, pp. 378–385. https://doi.org/10.1117/12.2081420
Cicek, O., Abdulkadir, A., Lienkamp, S.S., Brox, T., Ronneberger, O.: 3D U-Net: learning dense volumetric. In: Ourselin, S., Joskowicz, L., Sabuncu, M.R., Unal, G., Wells, W. (eds.) Medical Image Computing and Computer-Assisted Intervention—MICCAI 2016, pp. 424–432. Springer (2016). https://doi.org/10.1007/978-3-319-46723-8
Chen, H., Dou, Q., Yu, L., Qin, J., Heng, P.A.: VoxResNet: deep voxelwise residual networks for brain segmentation from 3D MR images. NeuroImage 170, 446 (2018). https://doi.org/10.1016/j.neuroimage.2017.04.041
Bui, T.D., Shin, J., Moon, T.: 3D Densely convolutional networks for volumetric segmentation. arXiv:1709.03199 [cs] (2017)
Zhang, R., Zhao, L., Lou, W., Abrigo, J.M., Mok, V.C., Chu, W.C., Wang, D., Shi, L.: Automatic segmentation of acute ischemic stroke from DWI using 3D fully convolutional DenseNets. IEEE Trans. Med. Imaging 37(9), 2149 (2018). https://doi.org/10.1109/TMI.2018.2821244
Zhu, W., Wang, Y., Yao, Y., Chang, J., Graber, H.L., Barbour, R.L.: Iterative total least-squares image reconstruction algorithm for optical tomography by the conjugate gradient method. J. Opt. Soc. Am. A 14(4), 799 (1997). https://doi.org/10.1364/JOSAA.14.000799
Dudgeon, D.E., Mersereau, R.M.: Multidimensional digital signal processing (1985). https://doi.org/10.1109/MCOM.1985.1092416
Bellos, D., Basham, M., French, A.P., Pridmore, T.: Stacked dense denoise-segmentation FBP reconstruction from 3601 projection (2019). https://doi.org/10.5281/ZENODO.2645965. https://zenodo.org/record/2645965#.XLiEKnVKiA0
Bellos, D., Basham, M., French, A.P., Pridmore, T.: Stacked dense denoise-segmentation CGLS reconstruction from 91 projection (2019). https://doi.org/10.5281/ZENODO.2645838. https://zenodo.org/record/2645838#.XLiEKHVKiA0
Bellos, D., Basham, M., French, A.P., Pridmore, T.: Stacked dense denoise-segmentation volumetric annotations (2019). https://doi.org/10.5281/ZENODO.2646117
Newell, A., Yang, K., Deng, J.: Stacked hourglass networks for human pose estimation. In: Lecture Notes in Computer Science (Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) (2016). https://doi.org/10.1007/978-3-319-46484-8_29
Kazantsev, D., Pickalov, V., Nagella, S., Pasca, E., Withers, P.J.: TomoPhantom, a software package to generate 2D–4D analytical phantoms for CT image reconstruction algorithm benchmarks. SoftwareX 7, 150 (2018). https://doi.org/10.1016/j.softx.2018.05.003
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015, pp. 234–241. Springer, Cham (2015)
Ciresan, D., Giusti, A., Gambardella, L., Schmidhuber, J.: Deep neural networks segment neuronal membranes in electron microscopy images. In: Pereira, F., Burges, C.J.C., Bottou, L., Weinberger, K.Q. (eds.) Advances in Neural Information Processing Systems, vol. 25. Curran Associates, Inc., 2012, pp. 2843–2851 (2012). https://doi.org/10.1016/S1673-8527(09)60051-5. http://papers.nips.cc/paper/4741-deep-neural-networks-segment-neuronal-membranes-in-electron-microscopy-images.pdf
Pereira, S., Pinto, A., Alves, V., Silva, C.A.: Brain tumor segmentation using convolutional neural networks in MRI images. IEEE Trans. Med. Imaging (2016). https://doi.org/10.1109/TMI.2016.2538465
Moeskops, P., Viergever, M.A., Mendrik, A.M., De Vries, L.S., Benders, M.J., Isgum, I.: Automatic segmentation of MR brain images with a convolutional neural network. IEEE Trans. Med. Imaging (2016). https://doi.org/10.1109/TMI.2016.2548501
Havaei, M., Davy, A., Warde-Farley, D., Biard, A., Courville, A., Bengio, Y., Pal, C., Jodoin, P.M., Larochelle, H.: Brain tumor segmentation with deep neural networks. Med. Image Anal. 35, 18 (2017). https://doi.org/10.1016/j.media.2016.05.004
Kamnitsas, K., Ledig, C., Newcombe, V.F., Simpson, J.P., Kane, A.D., Menon, D.K., Rueckert, D., Glocker, B.: Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 36, 61 (2017). https://doi.org/10.1016/j.media.2016.10.004
Mohseni Salehi, S.S., Erdogmus, D., Gholipour, A.: Auto-context convolutional neural network (auto-net) for brain extraction in magnetic resonance imaging. IEEE Trans. Med. Imaging (2017). https://doi.org/10.1109/TMI.2017.2721362
Shelhamer, E., Long, J., Darrell, T.: Fully convolutional networks for semantic segmentation. IEEE Trans. Pattern Anal. Mach. Intell. (2015). https://doi.org/10.1109/TPAMI.2016.2572683
Badrinarayanan, V., Kendall, A., Cipolla, R.: SegNet: a deep convolutional encoder–decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. (2017). https://doi.org/10.1109/TPAMI.2016.2644615
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition. In: 3rd International Conference on Learning Representations, ICLR 2015—Conference Track Proceedings (2015). arXiv:1409.1556
Tsai, C.J., Tsai, Y.W., Hsu, S.L., Wu, Y.C.: Synthetic training of deep CNN for 3D hand gesture identification. In: Proceedings—2017 International Conference on Control, Artificial Intelligence, Robotics and Optimization, ICCAIRO 2017 (2017). https://doi.org/10.1109/ICCAIRO.2017.40
Lindgren, K., Kalavakonda, N., Caballero, D.E., Huang, K., Hannaford, B.: Learned hand gesture classification through synthetically generated training samples. In: IEEE International Conference on Intelligent Robots and Systems (2018). https://doi.org/10.1109/IROS.2018.8593433
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (IEEE, 2016), pp. 770–778 (2016). https://doi.org/10.1109/CVPR.2016.90. http://ieeexplore.ieee.org/document/7780459/
Milletari, F., Navab, N., Ahmadi, S.A.: V-Net: fully convolutional neural networks for volumetric medical image segmentation. In: 2016 4th International Conference on 3D Vision, 3DV 2016, pp. 565–571 (2016).https://doi.org/10.1109/3DV.2016.79
Lee, C.Y., Xie, S., Gallagher, P., Zhang, Z., Tu, Z.: Deeply-supervised nets. In: Lebanon, G., Vishwanathan, S.V.N. (eds.) Proceedings of the 18th International Conference on Artificial Intelligence and Statistics, Proceedings of Machine Learning Research, vol. 38 (PMLR, San Diego, California, USA, 2015), Proceedings of Machine Learning Research, vol. 38, pp. 562–570 (2015). http://proceedings.mlr.press/v38/lee15a.html
Huang, G., Liu, Z., Van Der Maaten, L., Weinberger, K.Q.: Densely connected convolutional networks. In: 30th IEEE Conference on Computer Vision and Pattern Recognition, CVPR 2017, pp. 2261–2269 (2017). https://doi.org/10.1109/CVPR.2017.243
Li, X., Chen, H., Qi, X., Dou, Q., Fu, C.W., Heng, P.A.: H-DenseUNet: hybrid densely connected UNet for liver and tumor segmentation from CT volumes. IEEE Trans. Med. Imaging 37(12), 2663 (2018). https://doi.org/10.1109/TMI.2018.2845918
Liu, Y., Zhang, Y.: Low-dose CT restoration via stacked sparse denoising autoencoders. Neurocomputing 284, 80 (2018). https://doi.org/10.1016/j.neucom.2018.01.015
Chen, H., Zhang, Y., Zhang, W., Liao, P., Li, K., Zhou, J., Wang, G.: Low-dose CT denoising with convolutional neural network. In: Proceedings—International Symposium on Biomedical Imaging, pp. 143–146 (2017). https://doi.org/10.1109/ISBI.2017.7950488
Chen, H., Zhang, Y., Kalra, M.K., Lin, F., Chen, Y., Liao, P., Zhou, J., Wang, G.: Low-dose CT with a residual encoder–decoder convolutional neural network (red-CNN). IEEE Trans. Med. Imaging 36(12), 2524 (2017). https://doi.org/10.1109/TMI.2017.2715284
Kang, E., Min, J., Ye, J.C.: A deep convolutional neural network using directional wavelets for low-dose X-ray CT reconstruction. Med. Phys. 44(10), e360 (2017). https://doi.org/10.1002/mp.12344
Kang, E., Chang, W., Yoo, J., Ye, J.C.: Deep convolutional framelet denosing for low-dose CT via wavelet residual network. IEEE Trans. Med. Imaging 37(6), 1359 (2018). https://doi.org/10.1109/TMI.2018.2823756
You, C., Yang, Q., Shan, H., Gjesteby, L., Li, G., Ju, S., Zhang, Z., Zhao, Z., Zhang, Y., Cong, W., Wang, G.: Structurally-sensitive multi-scale deep neural network for low-dose CT denoising. IEEE Access 6, 41839 (2018). https://doi.org/10.1109/ACCESS.2018.2858196
Bellos, D., Bellos, D.: DimitriosBellos/Stacked-DenseDenSeg: knowledge transfer and HMMs (2021). https://doi.org/10.5281/ZENODO.4416013. https://doi.org/10.5281/zenodo.4416013#.X_Nath28S1k.mendeley
Ioffe, S., Szegedy, C.: Batch normalization: accelerating deep network training by reducing internal covariate shift. In: Bach, F., Blei, D. (eds.) Proceedings of the 32nd International Conference on Machine Learning (PMLR, 2015), pp. 448–456 (2015). https://doi.org/10.1007/s13398-014-0173-7.2. http://proceedings.mlr.press/v37/ioffe15.html
Glorot, X., Bordes, A., Bengio, Y.: Deep sparse rectifier neural networks. In: AISTATS’11: Proceedings of the 14th International Conference on Artificial Intelligence and Statistics, vol. 15, pp. 315–323 (2011). http://proceedings.mlr.press/v15/glorot11a.html
Sutskever, I., Hinton, G., Krizhevsky, A., Salakhutdinov, R.R.: Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. (2014). https://doi.org/10.1214/12-AOS1000
Bellos, D., French, A.P., Basham, M., Pridmore, T.: Stacked dense denoise-segmentation FBP synthetic reconstruction from 3601 projection with ring artefacts (2020). https://doi.org/10.5281/ZENODO.3986502. https://doi.org/10.5281/zenodo.3986502#.Xzq2kjpcFEo.mendeley
Bellos, D., French, A.P., Basham, M., Pridmore, T.: Stacked dense denoise-segmentation FBP synthetic reconstruction from 3601 projection without ring artefacts (2020). https://doi.org/10.5281/ZENODO.3986504. https://doi.org/10.5281/zenodo.3986504#.Xzq2xIJZsp0.mendeley
Bellos, D., French, A.P., Basham, M., Pridmore, T.: Stacked dense denoise-segmentation CGLS synthetic reconstruction from 91 projection with ring artefacts (2020). https://doi.org/10.5281/ZENODO.3986506. https://doi.org/10.5281/zenodo.3986506#.Xzq2YTp4xNY.mendeley
Bellos, D., French, A.P., Basham, M., Pridmore, T.: Stacked dense denoise-segmentation CGLS synthetic reconstruction from 91 projection without ring artefacts (2020). https://doi.org/10.5281/ZENODO.3986508. https://doi.org/10.5281/zenodo.3986508#.Xzq2SfAiX4E.mendeley
Bellos, D., French, A.P., Basham, M., Pridmore, T.: Stacked dense denoise-segmentation synthetic annotations (2020). https://doi.org/10.5281/ZENODO.3986494
Bodey, A.J., Rau, C.: Launch of the I13–2 data beamline at the diamond light source synchrotron. J. Phys. Conf. Ser. 849(1), 012038 (2017). https://doi.org/10.1088/1742-6596/849/1/012038
Bellos, D., Basham, M., Pridmore, T., French, A.P.: A convolutional neural network for fast upsampling of undersampled tomograms in X-ray CT time-series using a representative highly sampled tomogram. J. Synch. Radiat. (2019). https://doi.org/10.1107/S1600577519003448
Eckermann, F., Suter, T., Uggowitzer, P.J., Afseth, A., Davenport, A.J., Connolly, B.J., Larsen, M.H., Carlo, F.D., Schmutz, P.: In situ monitoring of corrosion processes within the bulk of AlMgSi alloys using X-ray microtomography. Corros. Sci. 50(12), 3455 (2008). https://doi.org/10.1016/j.corsci.2008.09.015
Wadeson, N., Basham, M.: Savu: a Python-based, MPI framework for simultaneous processing of multiple, N-dimensional, large tomography datasets (2016). arXiv:1610.08015 [cs]
Luengo, I., Darrow, M.C., Spink, M.C., Sun, Y., Dai, W., He, C.Y., Chiu, W., Pridmore, T., Ashton, A.W., Duke, E.M., Basham, M., French, A.P.: SuRVoS: super-region volume segmentation workbench. J. Struct. Biol. 198(1), 43 (2017). https://doi.org/10.1016/j.jsb.2017.02.007
Hecksel, C.W., Darrow, M.C., Dai, W., Galaz-Montoya, J.G., Chin, J.A., Mitchell, P.G., Chen, S., Jakana, J., Schmid, M.F., Chiu, W.: Quantifying variability of manual annotation in cryo-electron tomograms. Microsc. Microanal. 22(3), 487 (2016). https://doi.org/10.1017/S1431927616000799
Kingma, D.P., Ba, J.: Adam: a method for stochastic optimization. arXiv:1412.6980 [cs] (2014)
Krogh, A., Hertz, J.A., Connect, A.K.: A simple weight decay can improve generalization. Adv. Neural Inf. Process. Syst. 4, 950 (1992)
Wang, Z., Bovik, A.C., Sheikh, H.R., Simoncelli, E.P.: Image quality assessment: from error visibility to structural similarity. IEEE Trans. Image Process. 13(4), 600 (2004). https://doi.org/10.1109/TIP.2003.819861
Paszke, A., Chintala, S., Chanan, G., Lin, Z., Gross, S., Yang, E. Antiga, L., Devito, Z., Lerer, A., Desmaison, A.: Automatic differentiation in PyTorch. In: 31st Conference on Neural Information Processing Systems (2017). https://doi.org/10.1017/CBO9781107707221.009
Acknowledgements
We thank Diamond Light Source for access to beamline I13-2 (MT9396) that contributed to the results presented here, and Alison Davenport and her team at the University of Birmingham for the sample preparation and experimental method employed. We also want to thank Diamond Light Source and the School of Computer Science at Nottingham for jointly funding Dimitrios Bellos under Ph.D. STU0185.
Funding
This research is jointly funded by Diamond Light Source and the University of Nottingham under Dimitrios Bellos’s Ph.D. studentship with code STU0185.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Code availability
Code for our proposed networks [37] is available online in GitHub and Zenodo.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bellos, D., Basham, M., Pridmore, T. et al. A stacked dense denoising–segmentation network for undersampled tomograms and knowledge transfer using synthetic tomograms. Machine Vision and Applications 32, 75 (2021). https://doi.org/10.1007/s00138-021-01196-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00138-021-01196-4