Abstract
Model fitting is a fundamental component in computer vision for salient data selection, feature extraction and data parameterization. Conventional approaches such as the RANSAC family show limitations when dealing with data containing multiple models, high percentage of outliers or sample selection bias, commonly encountered in computer vision applications. In this paper, we present a novel model evaluation function based on Gaussian-weighted Jensen–Shannon divergence, and integrate into a particle swarm optimization (PSO) framework using ring topology. We avoid two problems from which most regression algorithms suffer, namely the requirements to specify inlier noise scale and the number of models. The novel evaluation method is generic and does not require any estimation of inlier noise. The continuous and meta-heuristic exploration facilitates estimation of each individual model while delivering the number of models automatically. Tests on datasets comprised of inlier noise and a large percentage of outliers (more than 90 % of the data) demonstrate that the proposed framework can efficiently estimate multiple models without prior information. Superior performance in terms of processing time and robustness to inlier noise is also demonstrated with respect to state of the art methods.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction and related work
Model fitting techniques are widely used in computer vision applications, in which the data to be modeled usually contains a significant number of outliers. RANSAC [1] is commonly applied to estimate a model with noisy data and numerous extensions and modifications have been derived from it (see [2, 3] for a good overview of different extensions of RANSAC). These RANSAC-based algorithms evaluate every hypothesis using a cost function \( f(.).\) Then the best hypothesis is selected as \(\mathbf{argmin }\,f(.).\)
The truncation cost functions used by RANSAC and its extensions facilitate toleration of more than 50 % outlier rate. However, this attractive capability also brings the requirement of a application-dependent parameter, i.e. the inlier noise scale (also referred to as bandwidth or truncation threshold). Since this prior information usually is not available for many practical computer vision applications, the development of nonparametric cost functions for robust estimators has received widespread attention in recent literature [4–9]. They analyze the distribution of residuals of individual data points w.r.t. the hypotheses, either with a histogram or with kernel methods. A problem for histogram analysis-based approaches is that the small pseudo-models and severe outliers can easily generate many false peaks and valleys which obscure the genuine modes. The kernel function-based approaches estimate the residual histogram density using a smoothness parameter, and they therefore can tackle small pseudo-structures, but instead they have the well-known density problem, i.e. over- or under-estimation of the proportion of inliers. Both of these two categories of hypothesis evaluation function still require parameters (kurtosis or skewness threshold for histogram approaches and bandwidth of the kernel approaches), although these parameters are less sensitive than the inliner noise scale used in RANSAC.
RANSAC-based approaches apply a simple and straightforward optimization for searching the best hypothesis. This optimization can be represented as \(\mathbf{argmin }\,f(\theta )=f(\hat{\theta })\) with \(f(\hat{\theta })\le f(\theta _{i}), \forall \theta _{i}\in \mathcal H \), where \( \mathcal H = \{\theta _{j}\}_{j=1,\ldots ,M} \) are the model hypotheses. This straightforward optimization mechanism facilitates preemptive capability [10] while confining the data to a single structure only. Several extensions of RANSAC are capable of tackling the problem of multi-model estimation and remaining preemptive [11, 12], but they introduce more sophisticated user-dependent parameters and therefore are less robust in real applications. Recently, the holistic multi-model estimators combine the hypotheses evaluation and optimization search strategy into a complete framework to reveal the number and the parameters of the models simultaneously. All these approaches discover the inherent model number using residual distribution analysis [4, 5, 13–15]. However, these holistic multi-model estimators lose the preemptive capability and are too slow to be employed in real-time (at least soft real-time) tasks, such as scene understanding for mobile robot or homography detection for autonomous vehicle navigation.
In this paper, we present a novel framework for multi-model estimation, which facilitates both nonparametric and preemptive characteristics, and can deal with multi-structure estimation. We make the assumption that data are constrained to a specific known input space, such as a 3D volume of space several meters across for estimating planar structures in indoor scenes. With this assumption, we distribute data points into the input space uniformly to form an ideal uniform dataset. By comparing the residual distribution of data from the ideal uniform dataset and from the real dataset using Gaussian-weighted Jensen–Shannon divergence (GJSD), an effective and nonparametric cost function for hypothesis evaluation is generated. An optimization framework based on evaluating every hypotheses with GJSD is then introduced, and uses the ring-topological particle swarm optimization [16] to find multiple structures simultaneously. The algorithm presented in this paper obtains faster convergence while being robust to the presence of large amounts of noise/outliers. Furthermore, it can handle data imbalance more effectively than conventional approaches. In summary, the algorithm presented provides a simple, generalizable framework for multi-model estimation that can be used in many practical tasks which need to estimate models in a limited time.
The most related work to our approach is the paper [17], which also applies PSO for plane fitting. However, it does not clearly consider the model fitting problem as two separate subtasks. It uses the same cost function as RANSAC which needs the prior estimation of inlier scale. The use of GJSD-based evaluation as alternative in this paper makes the prior inlier scale unnecessary. Also a “deflation” function is used in [17] to make the found optimum disappear in the subsequent PSO iterations; this requires more iterations and it is also difficult to control the invisible areas without any overlaps. Instead we adopt the simple but powerful lbest PSO method to select hypotheses and reveal the number of models simultaneously.
2 Gaussian-weighted Jensen–Shannon divergence
This section describes the details of the proposed evaluation function using Gaussian-weighted Jensen–Shannon divergence [18]. Jensen–Shannon divergence, which is based on Kullback–Leibler divergence (KLD) [19], is popularly used in probability theory and statistics for measuring the similarity between two distributions. The superiority of JSD over KLD on handling zero values in distributions and removal of nuisance in the use of KLD arising from its asymmetry has been demonstrated in [20].
We use JSD to measure the divergence between the residual distributions of data from the ideal uniform distribution dataset and from real dataset, which clearly differ for hypotheses near to and far from an actual optimum (see last column of Fig. 1). When the hypothesis is geometrically close to an optimum, the residual distribution of this hypothesis for the real dataset has a peak at zero (first row in Fig. 1). Conversely, the residual distribution of a hypothesis that is not close to the optimum, is similar to the residual distribution that is generated using ideal uniform dataset (second row in Fig. 1). Using a Gaussian-weighted variant, we could emphasize the divergence at the specific place (e.g. the peak position) and also depreciate the influence of the parallel models in the residual distribution (last figure of row 1 in Fig. 1).
The illustration of residual distribution of hypothesis that is generated using ideal uniform dataset and real dataset. The top row shows results for a correct hypothesis and the bottom row for an arbitrary hypothesis. From left to right, each column shows (left) input data and hypothesis (marked in red), (middle) residual distribution using ideal uniform dataset, (right) obtained histogram of distances between all data from real set to this hypothesis. The width of bins in histogram is 0.01 for this figure as well as for Fig. 10 (color figure online)
Given input data \(\mathcal L = \{x_{i}\}_{i=1,\ldots ,N}\) and model hypotheses \( \mathcal H = \{\theta _{j}\}_{j=1,\ldots ,M} \) which are generated by randomly sampling from \( \mathcal L \), we calculate the distance \( d_{j} \) of all the points to each hypothesis \( \theta _{j} \) in \( \mathcal L , d_{j}=\{d_{1}^{j},\ldots ,d_{N}^{j}\} \), where \( d_{n}^{j} \) is the distance of point \( x_{n} \) to hypothesis \( \theta _{j} \). Then the residual histogram of each hypothesis denoted as \( r_{j} \) can be computed using \( d_{j} \). The right-column images in Fig. 1 illustrate the obtained residual histogram with respect to the correct and arbitrary hypotheses (marked in red in left-column images of Fig. 1). Note that the evaluation function we present here is based on a signed distance metric. In other words, instead of using an absolute distance metric like the Euclidean, as in the case of RANSAC-based approaches, we use a signed distance metric that is weighted by the location of the candidate point whose distance is to be calculated, with respect to the reference hypothesis. Thus, the distances of candidate points from the reference line are negative for points left of the line and positive for points right of the line. Conventional residual distribution-based fitness measures [8, 13, 15] do not suppress non-maxima effectively, due to the use of unsigned metrics.
Observing the computed residual distribution histogram \( r_{j} \), it is obvious that there is a peak near \( d_{j}=0 \) for correct hypotheses. Instead of using user-dependent parameters such as kurtosis [21], bandwidth for kernel density estimation [7] or the number of points per structure (the height of the peak) [15], we detect this peak by comparing the residual histogram with an ideal distribution. Since the region where the data lie are available prior information, we can calculate the ideal distribution of the hypothesis to the uniformly distributed data. This histogram, denoted as \( t_{j} \), is used as a criterion to calculate the divergence to the real histogram of distances, for the scenario at hand. For instance, the middle-column images in Fig. 1 display the histogram of corresponding hypotheses (marked in red) to the assumed uniformly distributed dataset in a bounding box. Assuming a uniform distribution of candidate points, the histogram of distances obtained should be roughly flat for an isometric distribution of points in the original space. However, in practical applications, candidate points are distributed in a non-isometric space bounded by hyper-planes based on possible values the underlying parameters representing the space can take. Hence, it is necessary to estimate ideal histograms of distances based on these boundary considerations. For the example shown in the figure, for an isometric distribution, we would have a rectangular distribution for the histogram. But since the space of data points are bounded by the axes \(x = 1\) and \(y = 1\), we obtain distributions with slopes at either end. For the case of row 1, the hypothesis line is closer in inclination to one of the axes, the flat portion of the histogram is large while the slopes are small, corresponding to small regions beyond the boundary required to complete the isometric distribution. On the other hand, since in row 2, the inclination of the hypothesis line is farther away from both axes, the regions outside the boundary, required to complete the isometric distribution are much larger, resulting in smaller flat region of the histogram and larger slopes. Note that the ideal distribution \( t_{j} \) for each hypothesis \( \theta _{j} \) is only determined by the given input space, therefore, we can calculate this function and store it before we process the robust estimation.
We normalize \( t_{j} \) and \( r_{j} \) and represent them as \( t_{j} = \{ t_{j}^{1},\ldots ,t_{j}^{K} \} \) and \( r_{j} = \{ r_{j}^{1},\ldots ,r_{j}^{K} \} \), where \( K \) is the number of the bins in each histogram. The Jensen–Shannon divergence (JSD) of \( t_{j} \) and \( r_{j} \) is defined by
where \( m_{j}=\frac{1}{2} (r_{j}+t_{j}) \).
The JSD values of the hypotheses in Fig. 1 are 0.0511 for the correct one and 0.0175 for the arbitrary one. Thus, it is necessary to select hypotheses that produce larger differences between the ideal and test distributions, due to the occurrence of peaks, while discarding hypotheses that produce distributions that are similar to the ideal distribution. Thus maximization of the divergence or JSD is a necessary criteria for hypothesis validation. It can also be seen from the given example that the JSD resulting from correct and arbitrary hypotheses are significantly different. However, several parallel hypotheses (parallel with the correct one) can also produce a high JSD value because the divergence of the whole distribution is evenly taken into account. Hence, we weight the residual distribution using a standard Gaussian function \(g=\{g^{1},\ldots ,g^{K}\}\;(\mu =0, \sigma ^{2}=1)\) to generate the Gaussian-weighted JSD. The Gaussian function reinforces the influence of JSD for data points close to the hypothesis, and weakens the effect caused by the parallel model. In our tests, \( \sigma ^{2}=1 \) is robust enough to distinguish parallel line models with distance between each other larger than 0.2 in range of whole dataset [0 1]. If two parallel lines are closer, the inlier scale noise makes them even hard to be distinguished by a human and we will not consider these extreme cases in this paper. The GJSD is formulated as,
Using GJSD to evaluate the hypotheses in Fig. 1, we get 0.1612 for the correct one and 0.0075 for the arbitrary one. In fact, GJSD is a powerful evaluation tool for estimation of both a single model and multiple models, as demonstrated with the experiments in Sect. 5.1.
Figure 2 demonstrates when variance of Gaussian function varies, the average fitness value of all the particles can always converge to a stable value which means all the particles are clustered into several niches. Figure 2 depicts the independence of Gaussian parameter with respect to the algorithm performance.
The comparison of algorithm convergences with various Gaussian parameters. The result here is collected from Fig. 9a with inlier noise 0.01
3 PSO using ring topology
This section describes the details of the proposed solution to the model estimation problem using a particle swarm optimization based framework. Given the model can be formulated using \( D \) parameters, i.e. each optimal model can be estimated by searching in a \( D \) dimensional space. A cost function \(c(\theta )\) where \(c(\theta ) = 1/f(\theta )\) is used as the function to be minimized. This is similar to the cost function used with RANSAC. The goal here is to choose the hypothesis that maximizes the difference between the residue distribution \(r_j\) and the ideal distribution \(t_j\), which is given by the GJSD. Therefore, translating the maximization into a minimization function and choosing \( f(\theta ) = \mathrm{GJSD}(\theta ) \) produce the required hypothesis selection. Thus \(c(\theta )\) is used to evaluate the current position of model \( \theta \) in this space. The problem of model estimation is to find \( \mathbf{arg }\,\mathbf{min }\,\lbrace c(\theta ), \theta \in \mathbb R ^{D} \rbrace \).
Particle swarm optimization (PSO) is a meta-heuristic optimization method which was first intended for simulating social behaviour of birds [22]. The ring topology for PSO algorithm (known as lbest PSO) was actually not new and described in one of the first papers on PSO [22]. It was successfully applied in [16] with an aim to locate multiple optima. The stable and effective results displayed in [16] also inspire us to apply the ring topological PSO for the multiple structure selection problem. lbest PSO maintains a swarm of candidate solutions (called particles), and makes these particles fly around in the search-space according to their own personal best-found position and the best-known position in a neighbourhood niche. Each particle has a velocity that directs its movement and is adjusted iteratively. The lbest PSO is demonstrated in Fig. 3, the found optima are capable of attracting neighbouring particles to form a niche, and once two niches are overlapped, the “better” niche (better fitness value) will entice the connecting particles from another niche. Since the movement of a particle is affected not only by its neighbors but also by its inertia, personal best position and random parameters, the particles at the border of two niches moves slowly and the niches can stay stably and arrive at convergence [16].
where \( v_{j}=(v_{j1},v_{j1},\ldots ,v_{jd},\ldots ,v_{jD}) \) denotes the velocity of particle \( j \) in \( d \) dimensions \( (1\le d \le D) \), \( p_{j}=(p_{j1},p_{j1},\ldots ,p_{jD}) \) is the \( j{\mathrm{th}} \) particle’s own best position found so far, and \( l_{jd} \) is the best position in the neighbourhood of the \( j\)th particle. For all the particles in \( D \) dimensions, \( v^{t+1}, \theta ^{t+1} \) are the updated velocity and new position derived from this new velocity, respectively.
The demonstration of PSO using ring topology is shown. The colored nodes \( \{\theta _1,\theta _2,\ldots ,\theta _m,\!\!\} \) denote \( m \) particles, and the grey nodes \( \{M1,M2,\ldots ,Mm,\!\!\} \) represent the memory of each particle, i.e. particles’ personal best value p. Each particle connects with its own memory node as well as immediate left and right neighbours in the ring. Particles with the same color form a niche (color figure online)
As in [23], \( \chi \) is the “constriction factor” which is introduced to limit the maximum velocity and the dynamic search area in each dimension. \( w,c_{1},c_{2} \) are inertia weight and two acceleration constants, where \( c_{1} \) is the factor that influences the “cognitive” behaviour, i.e., how much the particle will follow its own best solution, and \( c_{2} \) is the factor for “social” behaviour, i.e. how much the particle will follow the swarm’s best solution. \( c_{1} \) and \( c_{2} \) are set to 2.1 and 2.0, respectively, as it was proved to be the best set [23]. \( \chi \) is a function of \( c_{1} \) and \( c_{2} \) as reflected in (Eq. 4). \( r_{1} \) and \( r_{2} \) are uniformly distributed random numbers in the interval \( [0,1] \).
4 Complexity
In this section, we will discuss the computational complexity associated to the proposed algorithm as well as the realization of preemptive model estimation. As the complexities of both our algorithm and RANSAC are dominated by two major factors, which are generation of hypothesis by drawing the minimal sets and evaluating the fitness of every candidate model, here we will only compare the complexities of these two algorithms. The superior performance of our algorithm over more state-of-the-art approaches will be illustrated by the experimental results in Sect. 5.1.
Given the cost of hypothesis generation \(C_1\) and evaluation \(C_2\), the overall complexities of the standard RANSAC and our approach in the worst case are:
where \( T, T^{\prime }\) and \( M \) are iteration number of RANSAC, number of the particles and iteration number of PSO. Note that the number of the particles \(T^{\prime }\) instead of the iterations of PSO \(M\) plays the similar role as iterations of RANSAC \(T\) in the algorithm since both these two parameters impact the runtime of hypothesis generation step. As displayed in Fig. 6, lbest PSO can quickly (after 15–30 iterations) cluster hypotheses into different niches and maintain a reasonable number of members in each niche for a long time (see results in Fig. 6 after 50 iterations). These stable niches facilitate using classical clustering technologies to obtain the parameters and the number of the models. We also notice that even after the small numbers of iterations (\(\sim \)15), the particles have been already clustered into several niches, the best solution of the model estimation can be provided by the niches’ means roughly. This is desirable that the system can provide the current best solution given the time constrains, hence the preemption can be achieved in the case of real time systems which require a solution in a specific time. Algorithm 1 lists the scheme of preemptive lbest PSO.

In all the following tests with synthetic data, the parameter \(T^{\prime }\) and \(M\) for PSO are set to 100 and 30. The required iteration of RANSAC (parameter \(T\)) is calculated using formula \(T=\log (1-\rho )/\log (1-\varepsilon ^2)\), where \(\varepsilon \) is the inlier rate and \(\rho \) is the desired possibility of successful estimation (\(\rho \) is set to 99 % in all the test, i.e. typical value of \(T\) is \(\sim \)750 if inlier rate is 5 %, therefore we can expect that in our tests RANSAC is approximately four times faster than the proposed algorithm).
For a more detailed demonstration that lbest PSO can effectively induce stable and robust niching for multiple global optima localization, we refer readers to paper [16, 24].
5 Experiments
We start to test our GJSD estimator with synthetic data for both single model and multi-model estimation, and then demonstrate the application of our methods in a real robotic task, indoor plane estimation. All the results of synthetic data processing are obtained on an Intel Quad Core 2.4 GHz CPU, 4 GB RAM and the real robotic task is operated on an Intel Mobile Core i7 2720QM CPU, 8 GB RAM laptop connected with a Microsoft Kinect.
5.1 Easy case: one model estimation
We first compare the speed of GJSD, RANSAC [1], MDPE [6], QMDPE [6] and pbM [7] to demonstrate the time complexity. Therefore, in this experiment only a single model will be estimated from a synthetic dataset. Note that we do not consider the RHA [8], J-linkage [15] and KF [13] methods in this experiment because all of them are originally designed for estimating multiple models. Note that each line in this experiment contains 100 inliers corrupted with Gaussian noise of standard deviation \( \sigma =0.02 \), which is high relative to the range of the whole dataset (i.e. [0 1] [0 1]). The outlier rate considers both gross outliers as well as pseudo outliers which actually are inliers for the other models. The times in the test do not take the random sampling procedure into account because all methods apply the same random sampling with replication and have same number (1,000) of sampling. It costs 0.174(s) in average (measured using Matlab clock function). All the methods are coded in Matlab except for pbM which is coded in C++. Every method is tested 20 times and average processing time is recorded. From Table 1, we notice that GJSD is slower than RANSAC but significantly faster than kernel-based methods since we do not run any kernel smoothing or residual sorting algorithms which have high computational cost.
The comparison of breakdown points among various methods is also interesting. From Fig. 4 we can observe that, given the correct scale of inlier error, RANSAC can provide correct estimation results and begins to break down at approximate 92 % outliers. Meanwhile, GJSD almost has the same breakdown points without the requirement of inlier scale. LMedS can only tolerate approximate 50 % outliers because it does not truncate the loss function. We also notice that the performance of RANSAC drops quickly when the outlier rate is more than 92 %, in contrast, even at relatively high outlier rate (\(>\)95 % ), GJSD is still, loosely speaking, able to correctly estimate 80 % of the models out of 20 times.
Breakdown point test with three methods. The input data is similar as Fig. 5a, but with the different outlier rates. All three methods are repeated 20 times to calculate the estimation error in average
Although RANSAC and GJSD start to break down at the same place, the incorrect estimations after the breakdown points of these two methods are significantly different as displayed in Fig. 5c. We see that RANSAC obtains completely wrong estimation of the model and meanwhile GJSD gives an incorrect estimation closely and parallel to the true model. This “slightly” wrong estimation result somehow is still useful in many practical applications and might be refined using some explicit restrictions (e.g. a plane estimation result can be refined with recognized objects which are supported by the plane).
Demonstration of false estimation under high outlier rate (99 %). a Input data with 100 inliers and 9,900 outliers, the inliers are perturbed with Gaussian noise (\( \sigma \!=\!0.01 \)) b inlier boundaries are drawn in the dataset. c Typical incorrect estimation results with RANSAC (marked in green) and GJSD (marked in red), note that RANSAC has a relatively high probability to obtain this kind of wrong estimation, while almost all the incorrect estimations from GJSD are similar to the displayed one (color figure online)
5.2 Challenging case: multi-model with high outlier rate
In this section, all the input data are contaminated with more than 90 % outliers and relatively high inlier noise (\( \sigma =0.01 \)). Figure 6 illustrates the convergence of lbest PSO in different line fitting problems.
Demonstration of single and multi-model fitting using ring topological PSO. Results in each row demonstrate fitting for piecewise linear shapes and circles, namely, an inclined line, inverted V (roof), W, a pentacle and multiple circles. Line and circle models have two (slope, intercept) and three (X, Y coordinates of center, radius) parameters to be estimated, respectively. The convergence of PSO for each input points dataset (left column with 100 inliers per line/circle and 900 gross outliers, each line/circle is also corrupted with Gaussian noise \( \sigma =0.01 \)) is illustrated in each column—initial 200 hypotheses and corresponding particles, hypotheses after 5, 15, 30 and 50 iterations (using 0.56, 1.49, 2.99 and 4.96 s for line fitting and 0.64, 1.57, 3.11, 5.03 for circle fitting in average), with the last figure showing the final estimation result using mean shift clustering (different clusters are marked in different colors). The mean of each cluster is the estimated model (color figure online)
For each line estimation result, we define \( \omega \!=\!\{\omega _1, \ldots , \omega _Q\}, \tilde{\omega }=\{\tilde{\omega }_1, \ldots , \tilde{\omega }_{Q^{\prime }}\} \), respectively, to be the true and estimated lines’ parameters, where \( \Vert \omega _q \Vert =1\) and \( \Vert \tilde{\omega }_{q^{\prime }} \Vert =1\). Thus the error metrics between the estimated and true model is calculated as \(\varepsilon =\Vert \omega _q - \tilde{\omega }_{q^{\prime }}\Vert /\sqrt{2}\). Also note that this error is measured only when the numbers of the model are correctly estimated, in other words, the incorrect estimation of the number of structures is not taken into account since we believe that this type of the error makes the system to produce the meaningless results thus the error measurement in this case is not necessary anymore. To measure the error of multiple line estimation results, the average error of \(\bar{\varepsilon }\) is computed as,
Results of multi-model fitting for the presented scheme in comparison to the state of th art Kernel Fitting (KF) [13] and J-Linkage [15] are shown in Figs. 7, 8.
Figure 7 shows the results of line fitting using KF, J-Linkage and the proposed approach. The results have been obtained for estimating an inclined line and a ‘W’ shape case (4 models); the outlier rates are 91.7 and 93.7 % for the single and multi-model cases. The inlier noise rate is 0.02. For KF the parameter step size \( h \) is fixed at 100.
Figure 8 compares the convergence rate of the presented PSO-based framework with the Kernel fitting method [13]. Analogous to Fig. 7, results have been presented for regression of a single model (inclined line) as well as for a multiple model scenario (shape of W in Fig. 9). It can be seen that the time taken for convergence is much higher for the KF in relation to the proposed framework. It was also been seen that while the time for convergence grows exponentially with the outlier rate in the case of the KF, it grows much slower for the PSO framework.
5.3 Discussion
Our experimental results demonstrate the superior performance in terms of better accuracy and stability under various challenging configurations (high outlier rats and inlier noise scale), whilst no prior information such as number of the models, inlier noise scale and outlier rate has been provided before hand. The main reason of this superiority of our method is that the models we attempt to estimate are not determined by the sampling from the minimal set of inliers but by the particles traversing in the solution parametric space. The advantage of this setup can be considered in two different points of the views, (1) even with no initial particles are generated from the minimal set of inliers (very often in high outlier rate case), the particles which are close to the true models in the parametric space will also hold the relatively better fitness value, thereby attracting the neighbouring particles to approach themselves and search this area together. This is the main reason that the proposed method provides the superior performance when the outlier rate is high. (2) When true models have been discovered by particles either during the initial step or after several traversing, these models with better fitness value could attract the neighbouring particles to approach themselves and search the nearby area together, thus the more accurate models might be discovered by the traversing of these particles in this niche. Usually, these more accurate models can only be determined by the inliers that are less disturbed due to the inlier noise. If only sampling of inliers can generate the correct models (this is probably most case in the aforementioned algorithms such as RANSAC family and J-linkage), obviously the chances of finding these models are much less than the proposed method which attempts to search the nearby area of the approximately correct models. This can explain why our method has the better accuracy in the case of the inlier noise scale increasing.
5.4 Influence of non-uniform outliers
As we mentioned in Sect. 2, we assume a uniform distribution of all the data points to produce the so-called ideal uniform dataset and compare the two residual distributions calculated based on this dataset and the real experimental dataset. Although in practice the dense point cloud and its bounding contour can be obtained with diverse de-noising and delineation techniques, investigation of the performance of our method under the condition of non-uniform outliers also presents an interesting avenue as addressed here.
To this end, non-uniform outliers have been generated by constraining the space of the outlier data points and residual distributions estimated using such a distribution. The effect of this constriction of the space is to shift the center of the ideal distribution along with a minimal change in the shape of the distribution. As it is shown in Fig. 10, even with the non-uniformly distributed outliers, the correct hypotheses can still generate reasonably higher GJSD value than that generated by the wrong hypotheses, which indicates that the differences between residual distributions of the correct hypothesis with the ideal uniform dataset and with the real dataset, is greater than the differences generated by the wrong hypothesis. As expected, it can be seen that these differences are not as obvious as in the case of uniformly distributed outliers. This effect is especially noticeable when the space of the wrong hypothesis is mutually disjoint to the restricted space of the outlier distribution [sub-figure (c) in Fig. 10]. However, since all hypotheses are generated using random sampling of the real data points (including outliers), the likelihood of generation of such hypotheses is not high. In summary, it can be seen that even with non-uniform distribution of outliers the GJSD-based hypothesis validation metric successfully detects the correct hypothesis.
The residual distributions of hypothesis that generated using ideal uniform dataset and real dataset with non-uniform outliers. The top row shows test data set with marked hypothesis. The middle row shows residual distributions of hypothesis that generated using ideal uniform dataset. The bottom row shows residual distributions of hypothesis that generated using real dataset with non-uniform outliers
5.5 Real data test
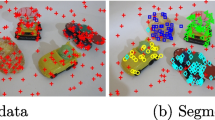
We mount a Microsoft Kinect camera on a Pioneer P3-DX mobile robot (1.2 m height) to acquire the RGB-D data. One challenging scene, which is typical for robotic search tasks, is demonstrated in Fig. 11. The robust regression task for our robot vision system is the estimation of multiple planes in the current scene in limited time.
Figure 11 shows multi-model plane fitting using the lbest PSO framework and GJSD estimator on real data. The point cloud captured from RGB-D sensor is firstly down-sampled to 2,000 for decreasing the computational complexity. Then the multiple plane models are estimated using the proposed method, wherein the ideal residual distribution is generated using the normally distributed points in the bounding box of point cloud. All the points are back-projected to one of the estimated plane which is the closest to the specific point. The estimated planes without significant number of projected inliers are then removed from the final result. The detected planes are represented in terms of different colors for the constituent data points.
6 Conclusion
Two major contributions, GJSD evaluation function and the application of ring topological PSO, have been proposed for estimating the multiple structures from the noisy data. A novel fitness evaluation metric, GJSD, has been presented in this paper which can clearly discriminate between correct and arbitrary hypotheses that are randomly generated in a specific region. The proposed framework, driven by lbest PSO and the GJSD estimation for measuring fitness of particles, facilitates the stable and robust localization of multiple optima. We have demonstrated that the performance of the presented technique significantly exceeds those based on the RANSAC family as well as the state-of-the-art Kernel-based techniques, while being able to detect multiple models robustly. Experimental results with synthetic data and real data validate the proposed approach. While the proposed method could be extended for applications requiring fitting to non-geometric error functions (such as in the estimation of fundamental matrices), the focus of this paper is on robust fitting for multi-modal data. Applications to non-geometric and higher order non-linear functions are exciting and promising avenues for future research as evidenced from [25]) that applies PSO to the estimation of fundamental matrix.
Furthermore, application of the GJSD evaluation function is limited to data constrained to a specific topological space. Although this in agreement with most real applications (or can be achieved with denoising techniques), it forms a good scope for future investigation. In addition, while we use the lbest PSO in a preemtive framework to estimate the stopping criterion in this iterative procedure, analysis of the convergence of PSO as well as the stopping criterion present further avenues of research.
References
Fischler, M.A., Bolles, R.C.: Random sample consensus: a paradigm for model fitting with applications to image analysis and automated cartography. Commun. ACM 24, 381–395 (1981)
Sunglok, C., Taemin, K., Wonpil, Y.: Performance evaluation of RANSAC family. In: Proceedings of the British Machine Vision Conference (BMVC’09) (2009)
Raguram, R., Frahm, J.M., Pollefeys, M.: A comparative analysis of ransac techniques leading to adaptive real-time random sample consensus. In: Proceedings of the European Conference on Computer Vision, vol. 5303, pp. 500–513 (2008)
Wong, H.S., Chin, T.J., Yu, J., Suter, D.: Dynamic and hierarchical multi-structure geometric model fitting. In: IEEE International Conference on Computer Vision (2011)
Wang, H., Chin, T.J., Suter, D.: Simultaneously fitting and segmenting multiple-structure data with outliers. In: IEEE Transactions on PAMI (2012)
Wang, H., Suter, D.: MDPE: a very robust estimator for model fitting and range image segmentation. Int. J. Comput. Vis. 59, 139–166 (2004)
Subbarao, R., Meer, P.: Beyond RANSAC: user independent robust regression. 25 Years of RANSAC, Workshop in conjunction with CVPR, New York (2006)
Zhang, W., Kosecka, J.: Nonparametric estimation of multiple structures with outliers. In: Dynamical Vision Workshop (2006)
Wang, H., Mirota, D., Ishii, M., Hager, G.: Robust motion estimation and structure recovery from endoscopic image sequences with an adaptive scale kernel consensus estimator. In: Proceedings of the 2001 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 2008, pp. 1–7 (2008)
Nister, D.: Preemptive ransac for live structure and motion estimation. Mach. Vis. Appl. 16, 321–329 (2005)
Zuliani, M., Kenney, C.S., Manjunath, B.S.: The multiransac algorithm and its application to detect planar homographies. In: IEEE International Conference on Image Processing (2005)
Stewart, C.V.: Bias in robust estimation caused by discontinuities and multiple structures. IEEE Trans. PAMI 19, 818–833 (1997)
Jun Chin, T., Wang, H., Suter, D.: Robust fitting of multiple structures: the statistical learning approach. In: IEEE International Conference on Computer Vision (2009)
Delong, A., Osokin, A., Isack, H.N., Boykov, Y.: Fast approximate energy minimization with label costs. In: Proceedings of the 2010 IEEE Computer Society Conference on Computer Vision and, Pattern Recognition (2010)
Toldo, R., Fusiello, A.: Robust multiple structures estimation with j-linkage. In: European Conference of Computer Vision, pp. 537–547 (2008)
Li, X.: Niching without niching parameters: particle swarm optimization using a ring topology. IEEE Trans. Evol. Comput. 14, 150–169 (2010)
Zhou, K., Zillich, M., Vincze, M., Vrečko, A., Skočaj, D.: Multi-model fitting using particle swarm optimization for 3d perception in robot vision. In: IEEE International Conference on Robotics and Biomimetics (ROBIO) (2010)
Fuglede, B., Topsoe, F.: Jensen–Shannon divergence and Hilbert space embedding. In: IEEE International Symposium on Information Theory (2004)
Kullback, S., Leibler, R.A.: On information and sufficiency. Ann. Math. Stat. 22, 79–86 (1951)
Lee, L.: On the effectiveness of the skew divergence for statistical language analysis. In: Artificial Intelligence and, Statistics, pp. 65–72 (2001)
Zhang, W., Kosecka, J.: Ensemble method for robust motion estimation. In: 25 Years of RANSAC, Workshop in conjunction with CVPR (2006)
Kennedy, J., Eberhart, R.: Particle swarm optimization. In: Proceedings of IEEE International Conference on, Neural Networks, pp. 1942–1948 (1995)
Eberhart, R., Shi, Y.: Comparing inertia weights and constriction factors in particle swarm optimization. In: Proceedings of the 2000 IEEE Congress on, Evolutionary Computation, vol. 1, pp. 84–88 (2000)
Jones, F., Soule, T.: Dynamic particle swarm optimization via ring topologies. In: Proceedings of the 11th Annual conference on Genetic and evolutionary computation. GECCO ’09, New York, NY, USA, ACM, pp. 1745–1746 (2009)
Chan, K.H., Tang, C.Y., Wu, Y.L., Hor, M.K.: Robust orthogonal particle swarm optimization for estimating the fundamental matrix. In: Visual Communications and Image Processing (VCIP), IEEE (Nov.), pp. 1–4 (2011)
Acknowledgments
The research leading to these results has received funding from the European Community’s Seventh Framework Programme [FP7/2007-2013] under grant agreement no.215181, CogX, the Austrian Science Fund (FWF): project TRP 139-N23, InSitu, and the Austrian Science Fund (FWF) project under grant agreement no. I513-N23.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Zhou, K., Varadarajan, K.M., Zillich, M. et al. Gaussian-weighted Jensen–Shannon divergence as a robust fitness function for multi-model fitting. Machine Vision and Applications 24, 1107–1119 (2013). https://doi.org/10.1007/s00138-013-0513-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00138-013-0513-1