Abstract
Aims/hypothesis
The aim of this work was to explore molecular amino acids (AAs) and related structures of HLA-DQA1-DQB1 that underlie its contribution to the progression from stages 1 or 2 to stage 3 type 1 diabetes.
Methods
Using high-resolution DQA1 and DQB1 genotypes from 1216 participants in the Diabetes Prevention Trial-Type 1 and the Diabetes Prevention Trial, we applied hierarchically organised haplotype association analysis (HOH) to decipher which AAs contributed to the associations of DQ with disease and their structural properties. HOH relied on the Cox regression to quantify the association of DQ with time-to-onset of type 1 diabetes.
Results
By numerating all possible DQ heterodimers of α- and β-chains, we showed that the heterodimerisation increases genetic diversity at the cellular level from 43 empirically observed haplotypes to 186 possible heterodimers. Heterodimerisation turned several neutral haplotypes (DQ2.2, DQ2.3 and DQ4.4) to risk haplotypes (DQ2.2/2.3-DQ4.4 and DQ4.4-DQ2.2). HOH uncovered eight AAs on the α-chain (−16α, −13α, −6α, α22, α23, α44, α72, α157) and six AAs on the β-chain (−18β, β9, β13, β26, β57, β135) that contributed to the association of DQ with progression of type 1 diabetes. The specific AAs concerned the signal peptide (minus sign, possible linkage to expression levels), pockets 1, 4 and 9 in the antigen-binding groove of the α1β1 domain, and the putative homodimerisation of the αβ heterodimers.
Conclusions/interpretation
These results unveil the contribution made by DQ to type 1 diabetes progression at individual residues and related protein structures, shedding light on its immunological mechanisms and providing new leads for developing treatment strategies.
Data availability
Clinical trial data and biospecimen samples are available through the National Institute of Diabetes and Digestive and Kidney Diseases Central Repository portal (https://repository.niddk.nih.gov/studies).
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.

Introduction
Type 1 diabetes is a life-long chronic autoimmune disease with an annual incidence among young people (under 20 years) of around 130,000 worldwide [1]. Its development typically runs from seroconversion (stage 1) to hyperglycaemia (stage 2), to onset of symptomatic type 1 diabetes (stage 3) [2]. The recent development of biochemically defined autoantibody markers allows us to screen for individuals at stage 1 or stage 2 disease (henceforward shortened to stage 1/2), facilitating secondary prevention [3] to interrupt the progression phase from stage 1/2 to stage 3 type 1 diabetes. This is the focus of the current investigation. Earlier, Butty and colleagues used archived clinical and genetic data from the Diabetes Prevention Trial-Type 1 (DPT-1) to study genetic determinants in the disease progression [4]. Their study uncovered the role of DQB1 alleles in disease progression; carriers of the allele DQB1*03:02 were faster progressors, while DQB1*03:01 carriers were slower progressors [4]. To gain molecular insights into the role of specific amino acids (AAs) and related protein structures, we sequenced DQA1 and DQB1 genes in DNA samples from participants in the DPT-1 and uncovered two AAs (−18β, β57) that associated with the disease progression [5]. Recently, we acquired DNA samples from participants in the Oral Insulin Prevention Trial (TrialNet07 [TN07]) for sequencing and combined both DPT-1 and TN07 to investigate DQ associations; the results indicated a new association of DQ2.5 with disease progression [6].
This new finding and the larger sample size provide an impetus to investigate further the molecular structure of HLA-DQ. The HLA system is critical to host immunity [7] through binding antigenic peptide fragments and presenting to T lymphocytes in the thymus (self-presentation) to avoid obvious autoimmune reactivity. Unfortunately, immune tolerance to certain autoantigenic peptides is incomplete and may, under certain circumstances and stimuli, lead to self-destruction of otherwise normal cells (i.e. autoimmune diseases). The vast majority of autoimmune diseases are associated with alleles from the HLA class II (HLAII) molecules that are each composed of an α-chain and a β-chain. The structure of these molecules is essentially preserved with minor but important differences: the so-called MHC fold (α1β1) consists of two anti-parallel α-helices on top of a β-sheet platform that shape an antigen-binding groove, open at both ends [8, 9]. The groove has five depressions, called pockets, positioned at relative positions 1, 4, 6, 7 and 9. The polymorphic residues of the various HLAII allelic molecules mainly concern several of the residues occupying these pockets. The second extracellular domain, α2β2, of HLAII molecules participates in accessory functions, such as binding to co-receptor molecule CD4 and presumed cognate T cell receptor (TCR)-mediated homodimerisation [10]. Identifying specific AAs within α- and β-chains, encoded by DQA1/DQB1, could shed new insights into genetic mechanisms that may be useful to understand the DQ binding process of autoantigen-derived peptides and hence to develop secondary prevention strategies. In this investigation, we integrated DPT-1 and TN07 cohorts into a single cohort known as iCohort, and studied the molecular structure of DQA1-DQB1 in the context of the disease progression from stage 1/2 to stage 3 type 1 diabetes.
Methods
Diabetes prevention trials (DPT-1 and TN07)
DPT-1 and TN07 were randomised clinical trials to assess efficacy of oral/parenteral insulin for preventing type 1 diabetes or slowing disease progression among participants at moderate/high risk and have been detailed elsewhere [4, 6, 11]. Briefly, the integrated iCohort includes 408 type 1 diabetes onset events among 1216 participants with an accumulated 3916 person-years of observation. Demographically, most participants were white and there were 200 more male participants than female participants (see electronic supplementary material [ESM] Table 1). Most participants were under 20 years of age, while the rest were spread across the age range 20–46 years (ESM Fig. 1). Associations of risk factors with diabetes progression were assessed (ESM Table 1) and associated risk factors were treated as potential confounding variables.
Clinical trial data and biospecimen samples are available through the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) Central Repository portal (https://repository.niddk.nih.gov/studies).
Genotyping HLA using next-generation targeted sequencing technology
HLA typing was carried out using the Scisco HLA v6 typing kit (Scisco Genetics, Seattle, WA, USA) following the kit protocol. Briefly, the method employs an amplicon-based two-stage PCR, followed by sample pooling and sequencing using a MiSeq v2 PE500 (Illumina, San Diego, CA, USA). The protocol yielded three-field coverage of HLAII genes (DRB1, DRB3, DRB4, DRB5, DQA1, DQB1, DPA1 and DPB1). The phase within each gene was determined in part by bridging amplicons and, when not available, by database lookup [12].
DQA1 and DQB1 are in strong linkage disequilibrium (LD), which is used to infer their haplotypes, referred to as DQA1-DQB1. Of the eight cases with ambiguously inferred haplotypes (six with posterior probability of 0.96 and two with less than 0.95 posterior probabilities), the haplotypes with the highest posterior probabilities were chosen for further analysis.
To maintain structural equivalence in DQ residues, we adopted the numbering system of Fremont et al for H2-A [13], as modified for DQ molecules [14]. In this manner, essentially all residues that occupied identical or near identical positions in the respective structures had the same residue numbers in the sequence, thus taking into account all possible insertions and deletions in certain alleles. AA abbreviations are presented mostly in the one-letter convention and occasionally in the three-letter convention.
Within the iCohort, the observed 43 DQA1-DQB1 haplotypes were referred to a cis- or empirical haplotypes, in contrast to potentially new trans-haplotypes that are generated by ‘heterodimeric recombination’ and referred to as heterodimers. Computationally, we enumerated all possible cis- and trans-pairing of two DQA1 alleles with two DQB1 alleles, resulting in four heterodimers. For doubly heterozygous DQA1 and DQB1, heterodimeric recombination could create two new trans-haplotypes, referred to as ‘heterodimeric recombinants’ (ESM Fig. 2). Some of heterodimeric recombinants were observed not to have any surface expressions, probably due to structural incompatibility of corresponding α-helices and β-sheet [15, 16], and many ‘forbidden’ haplotypes were found to have limited functional roles in hematopoietic cell transplantation [17].
We encourage all interested and qualified researchers to contact us for independent or collaborative research, with sequenced HLA genotype data, together with clinical data from NIDDK Central Repository.
Molecular rendering and depiction
We used the coordinates of the HLA-DQ8 hybrid peptide (HP) insulin C-peptide–islet amyloid polypeptide (IAPP) [18] in order to demonstrate the various AA residues shown by hierarchically organised haplotype association analysis (HOH) to be linked to progression towards type 1 diabetes or protection therefrom, among the DQ molecules found in the iCohort participants [6].
Statistical analysis
All analyses relied on statistical functions and packages in R (https://www.r-project.org/). For haplotyping DQA1 and DQB1, we used the ‘haplo.em’ function from the R package ‘haplo.stats’. For assessing genetic associations with the time-to-onset of type 1 diabetes, we used the ‘coxph’ function. There are three specific analytical approaches that are worth mentioning. First, analysis generally centres on associations of an individual allele (or haplotype) vs all others, to avoid the need for choosing a reference allele. Second, HOH consists of three analytical steps: (1) phylogenic analysis of AA sequences to hierarchically organise DQ alleles/haplotypes and to identify one or more relevant clusters (RCs) that are informative for identifying type 1 diabetes-associated AAs; (2) identifying polymorphic AAs within RCs and assessing AA-specific association analysis; and (3) selecting associated AAs to form motifs, evaluating the motif associations and interpreting associations in the context of DQ molecular structures. Finally, the current investigation, even though relying on RCTs, should be considered as an exploratory study, given multiple analyses concerning multiple alleles/haplotypes, multiple AAs in genes and multiple adjustments of covariates. In this analysis, we used the threshold of p≤0.05 to indicate a ‘nominal significance’ of corresponding statistic, to highlight risk, protective or neutral association without arbitrary correction of multiple comparisons.
Results
Replication of (−18β, β57) associations with type 1 diabetes progression
An earlier study from the DPT-1 uncovered two AAs (−18β and β57) in DQB1 that associated with type 1 diabetes progression [5] (see DPT-1 [discovery] in ESM Table 2). Available TN07 data permitted an independent replication study of these two AAs. The AA β57A had a positive association with the onset of type 1 diabetes (HR 1.56, p=2.52×10−4) (ESM Table 2), a positive replication. Interestingly, β57D negatively associated with the type 1 diabetes onset (HR 0.50, p=9.83×10−3), in support of the marginal protective association observed in the discovery cohort (HR 0.70, p=0.074). When pooling the DPT-1 and TN07 data, β57A and β57D demonstrated associations with greater significance (HR 1.41 and 0.62, p=1.35×10−5 and 2.15×10−3, respectively) (ESM Table 2). On the other hand, the associations of −18βA and −18βV were not replicated in TN07 (HR 1.28 and 1.08, p=0.05 and 0.53, respectively). In fact, the association for −18βA was in the opposite direction from the discovery association (HR 0.82, p=0.06). Despite lacking this replication, the genetic association with −18βV in the pooled DPT-1 and TN07 sample remained significant (HR 1.30, p=0.013) (see Discussion for a possible explanation).
Role of AA ‒18β in type 1 diabetes progression
The association with type 1 diabetes progression of AA ‒18βV in the discovery cohort (HR 1.47, p=2.81×10−4) was not replicated in TN07 (HR 1.08, p=0.53) (ESM Table 2), even though its association in the iCohort was positive (HR 1.30, p=0.0013) (Table 2, ESM Tables 2, 5). To decipher this peculiar phenomenon, we performed a stratified analysis in two groups of participants who had lower and higher risk (ESM Table 10) and found that the regulatory association of ‒18β was present only among high-risk individuals with both risk and resistance associations with ‒18βV and ‒18βA (HR 1.65 and 0.75, p=2.22×10−4 and 0.033, respectively). While this association needs to be independently validated, it is of importance to investigate the biological significance of ‒18β [19].
Haplotypic associations of HLA-DQ with type 1 diabetes progression
Leveraging LD between DQA1 and DQB1, we inferred their empirical haplotypes and assessed their association with the onset of type 1 diabetes and found that DQA1*03:01:01-DQB1*03:02:01 (DQ8.1) and DQA1*05:01:01-DQB1*02:01:01 (DQ2.5) had significant associations (HR 1.25 and 1.19, p=3.47×10−3 and 0.044, respectively), while DQA1*03:03:01-DQB1*03:01:01 had the opposite association (HR 0.55, p=1.19×10−3) (ESM Table 3).
Based on AA sequences specific to each DQ haplotype, HOH hierarchically organised 43 empirical haplotypes, including 17 common haplotypes (ten or more observed copies) and 26 rare haplotypes (with fewer than ten copies), and represented organised haplotypes in a fan-based tree. This tree is shown in ESM Fig. 3 in which DQ haplotypes were highlighted as resistance and risk haplotypes if p<0.05; haplotypes with insignificant associations (p>0.05) and all rare haplotypes are also shown. A single cluster, created by a phylogenic tree cutting threshold (indicated by the arrow in ESM Fig. 3), included two risk, one protective, six neutral and five rare haplotypes (ESM Table 4). Eliminating monomorphic AAs among nine common haplotypes (excluding five rare haplotypes) within this cluster led to seven polymorphic AAs in the α-chain (−6α, α22, α23, α31, α44 and α157) and 11 polymorphic AAs in the β-chain (−18β, −10β, β9, β13, β23, β26, β55, β57, β66, β71 and β135) (ESM Table 4; AAs in DQ8.1 were considered as reference). Three AAs (α76, β9 and β23) were polymorphic, with only rarely occurring AAs in these positions among rare haplotypes. Additionally, β26 was in absolute LD with β13, and its association was represented by β13. Similarly β55, β66 and β71 were in absolute LD with −10β, and their associations were also represented by −10β. For every polymorphic AA, HOH directly assessed their associations with the onset of type 1 diabetes among those who carried those potentially risk haplotypes (ESM Table 5). AAs −18βV, β13G, β57A and β135D on the β-chain were found to positively associate with type 1 diabetes onset (HR 1.30, 1.39, 1.41 and 1.29, p=1.29×10−3, 3.09×10−5, 1.35×10−5 and 1.70×10−3, respectively), and so were AAs −6αM, α22Y, α31Q, α44C and α157A on the α-chain (HR 1.28, 1.31, 1.19, 1.19 and 1.30, p=3.83×10−3, 1.05×10−3, 0.044, 0.044 and 2.83×10−4, respectively). On the other hand, AAs β13A and β57D negatively associated with type 1 diabetes onset (HR 0.62 and 0.62, p=4.30×10−3 and 2.15×10−3, respectively).
Heterodimeric associations of DQ with type 1 diabetes progression
As noted above, DQ heterodimers (ESM Fig. 2) consist of 186 unique haplotypes (ESM Table 6), which is more diverse than the empirically observed 43 DQ haplotypes (ESM Table 3). As shown in ESM Table 6, we created an ID number for each haplotype with a prefix of ‘e’ or ‘h’, corresponding to an empirical haplotype or a heterodimeric recombinant. Besides the formal nomenclature, we abbreviated these haplotypes in ESM Table 6 by eliminating ‘DQA1’ and ‘DQB1’ but adding ‘e’ or ‘h’ as a suffix to indicate empirical haplotype or heterodimeric recombinant, respectively. We also determined whether the corresponding heterodimeric recombinant was structurally forbidden (designated as ‘F’, or otherwise as ‘N’) [14,15,16]. In total, there were 74 ‘forbidden’ heterodimeric recombinants, 55 of which were considered rare, and 19 heterodimeric recombinants were observed 10–52 times in the iCohort. Note that a DQ haplotype from a heterodimerisation is the same as one of the empirical haplotypes and is still identified as an empirical haplotype. Without prejudicing against ‘forbidden’ DQ heterodimeric recombinants, we included these in further analyses.
Out of 186 haplotypes, there were 57 common haplotypes and 129 rare haplotypes that accounted for 321 observed heterodimers (ESM Table 7). Among common haplotypes, five of those listed in Table 1 (e1 [*03:01:01-*03:02:01], h3 [*05:01:01-*03:02:01], h4 [*03:01:01-*02:01:01], h46 [*02:01:01-*04:02:01] and h57 [*04:01:01-*02:02:01]) were positively associated with the type 1 diabetes (HR 1.11, 1.35, 1.32, 2.41 and 2.94, p=0.013, 0.0046, 0.012, 0.033 and 0.017, respectively). On the other hand, two DQ haplotypes (e6 [*03:03:01-*03:01:01] and h19 [*05:01:01-*03:01:01]) were negatively associated with the onset of type 1 diabetes (HR 0.66 and 0.42, p=0.0043 and 0.036, respectively).
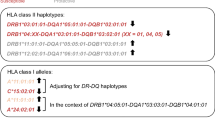
To gain insights into type 1 diabetes associations with both empirical haplotypes and heterodimeric recombinants, DQ haplotypes were extracted from ESM Table 7 and organised into three groups (Table 1). The first group comprised two empirical haplotypes (e1 [*03:01:01-*03:02:01e] and e2 [*05:01:01-*02:01:01e], commonly known as DQ8.1 and DQ2.5, respectively) and two heterodimeric recombinants (DQ8.1 and DQ2.5: h4 [*03:01:01-*02:01:01h] and h3 [*05:01:01-*03:02:01h], denoted as DQ8.1-DQ.2.5 and DQ.2.5-DQ8.1, respectively). Both empirical haplotypes (DQ8.1 and DQ2.5) retained positive associations with type 1 diabetes onset (HR 1.11 and 1.10, p=0.013 and 0.075, respectively), largely consistent with haplotypic associations (ESM Table 3). Not necessarily unexpectedly, the two heterodimeric recombinants, h4 (*03:01:01-*02:01:01h) and h3 (*05:01:01-*03:02:01h), exhibited stronger associations with type 1 diabetes onset (HR 1.32 and 1.35, p=0.012 and p=0.0046, respectively).
The second group shown in Table 1 (e9, e45, e18, h46 and h57) included three empirical haplotypes (*02:01:01-*02:02:01e, *02:01:01-*03:03:02e and *04:01:01-*04:02:01e) and two heterodimeric recombinants (*02:01:01-*04:02:01h and *04:01:01-*02:02:01h). A surprising pattern of haplotypic associations emerged, as while the three empirical haplotypes had no significant associations with the onset of type 1 diabetes (HR 0.88, 0.42 and 0.93, p=0.445, 0.23 and 0.79, respectively), the two heterodimeric recombinants had strong positive associations with the type 1 diabetes onset (HR 2.41 and 2.94, p=0.031 and 0.017, respectively).
In the third group shown in Table 1 (e6, e8 and h19), DQ7.3/DQ7.5 haplotypes and their heterodimeric recombinants with DQ2.5 appeared to retain their negative associations with type 1 diabetes onset (HR 0.66, 0.78 and 0.42, p=0.004, 0.10 and 0.036, respectively).
DQ haplotype clusters and related AAs
With diverse DQ heterodimers and emerging associations (Table 1 and ESM Table 7), it is expected that additional AAs contribute to their associations with type 1 diabetes. Here we applied HOH directly to hierarchically organise 186 DQ heterodimers (ESM Table 6) and display them in a fan-like hierarchical tree (Fig. 1) in which risk and protective haplotypes (p values <0.05), neutral haplotypes (p values ≥0.05) and rare haplotypes (fewer than ten copies) were highlighted. By choosing a threshold (brown arrow in Fig. 1), one has netted four clusters of DQ haplotypes labelled as clusters 1–4, and the cluster ID was placed as a prefix for each haplotype in the tree. From Fig. 1, it is clear that cluster 1 included all associated DQ heterodimers (both risk and protective haplotypes) and was naturally designated as an RC. This cluster included 29 common DQ haplotypes, in which 19 AAs (nine in the α-chain and ten in the β-chain) were polymorphic among these common haplotypes (ESM Table 8), excluding AAs pertaining to rare DQ haplotypes. Note that identified AAs of DQ heterodimers (ESM Table 8) are slightly different from those with DQ empirical haplotypes (ESM Table 4).
Fan-representation from phylogenic analysis of all heterodimeric HLA-DQ haplotypes based on similarity measurements between AA sequences of all individual haplotypes. A DQ haplotype is highlighted green or red if corresponding p values are <0.05 and HRs are ≥1, with red representing a risk haplotype and green representing a protective haplotype; a DQ haplotype is highlighted black if p values are ≥0.05 and the remaining DQ haplotypes are highlighted grey if they have fewer than ten copies. The brown arrow indicates the threshold for identifying a sub-phylogenic tree, resulting in clusters of heterodimeric DQ haplotypes. Note that all possible heterodimeric haplotypes are included, even though some may not be permissible
AAs involved in heterodimeric associations with stage 3 type 1 diabetes onset
To tease out which AAs were involved in the type 1 diabetes-associated DQ heterodimers, we performed AA associations among carriers of those haplotypes within the type 1 diabetes-associated cluster (Table 2). Other than AA α31 and ‒10β, β55, β66 and β71, the remaining eight AAs in the α-chain (‒16α, ‒13α, ‒6α, α22, α23, α44, α72 and α157) and six AAs in the β-chain (‒18β, β9, β13, β26, β57 and β135) were found to associate with type 1 diabetes onset (p<0.05) (Table 2). Note that two or more AAs of HLA-DQ are in absolute LD with each other within the RC, and only one is selected to represent them (ESM Table 9, ESM Fig. 4). To gain insights into these LD groups of AAs, we placed them on a protein structure of HLA-DQ8 in complex with an HP autoantigen (insulin C-peptide fragment plus IAPP fragment) (Fig. 2a, b). Quite a number of the residues in the α1β1 domain of the involved HLAII molecules are directly involved in antigenic peptide binding via any one of the five residue-anchoring pockets, or are TCR contact residues, or participate in the β49–56 homodimerisation stretch (Fig. 2a and ESM Table 9) [8, 10, 14, 20]. In the α2β2 domain of the HLA-DQ molecules under examination, we notice significant linkage to residues involved in binding to CD4 (β135 and β140), as well as to α157, a residue involved in the putative homodimerisation of HLA-DQ, and to β167 a part of the β167–69RGD loop whose function is still unknown [8, 10, 14, 20].
(a) Detailed view of the α1β1 domain of the HLA-DQ8-hybrid peptide insulin C-peptide−IAPP GQVELGGGNAVEVCK (anchors in bold, p11C mutated to form a disulphide bond with mutated DQα72C) complex (from https://www.rcsb.org/structure/6XCP, accessed 29 March 2024), showing all residues involved in type 1 diabetes progression or prevention thereof, among participants of the DPT-1 and TN07 trials, as revealed by HOH. Included are all residues in absolute LD to the involved residues among all the HLA-DQs. Residue β135D is added for orientation purposes. Depiction convention: protein backbone is shown in flat ribbons (α-helix, red; β-sheet, turquoise; β-turn, green; random coil, grey); antigenic peptide is shown in space-filling mode, and DQ residues are shown in stick mode (carbon, grey; oxygen, red; nitrogen, purple; hydrogen, white; sulphur, yellow). It is quite significant, in our view, that most of the depicted DQ8α1β1 residues line up pockets 4, 6, 7 and 9, or potentially can make contact with cognate TCR residues. Residues β26L, β30Y, β38A, β52P, β55P and β71T are covered mostly by the antigenic peptide or other DQ residues, in this orientation, hence cannot be seen. (b) Detailed view of α2β2 domain of HLA-DQ8–insulin C-peptide−IAPP hybrid peptide GQVELGGGNAVEVCK (anchors in bold) complex (from same source as above), in an orientation where the presumed interaction with CD4 would be such that the latter molecule has its long symmetry axis in a near vertical position. Residues α83T, α137F and β94R are in space-filling mode and shown only for orientation purposes; otherwise, the molecular depiction conventions are as in (a). The loop β105–113 is missing in the crystal structure, so the two immediately preceding residues, β104S and β114L, are shown in space-filling mode. Of the residues shown by HOH to be important for progression to type 1 diabetes or protection therefrom, plus those in absolute LD with the former, we note that α157 is involved in the putative cognate TCR-induced homodimerisation of MHC II molecules that promotes the binding of homodimer CD4 to such a complex, leading to T cell activation [10, 21]. In the respective β2 domain, residues β135 and β140 are part of the β134–148 stretch involved in CD4 binding, while residue β167R is part of the RGD loop found in all the known structures of HLA-DQ molecules possessing this tripeptide sequence. It is likely that residues involved in CD4 binding in the α2β2 domain, or others very close to them, interact with the inhibitory CD4 T cell membrane molecule LAG-3, which is inhibitory to the action of MHC II molecules [24]. Note that residues α172E and β167R have only partially determined crystallographic structure. Both figures drawn via the DSViewer Pro v.6 of Accelrys
Heterodimeric motif associations with type 1 diabetes onset
Based on eight α-chain and six β-chain AAs, HOH extracted motifs and assessed their associations with the onset of stage 3 type 1 diabetes (Table 3). There were 31 common motifs out of 53 motifs, and they were organised hierarchically and represented in a fan-like tree, in which tree leaves are denoted by one or more DQ heterodimer IDs sharing the same motif (ESM Fig. 5). By the hierarchical organisation, HOH suggested three clusters of heterodimers that included risk or protective haplotypes. Cluster c1 included two risk motifs (MAMYSQIAVYGLAD [e1, h101] and MAMYSQIAAYGLAD [h4], HR 1.11 and 1.32, p=0.014 and 0.012, respectively). Cluster c2 included a single risk motif (MAMYTCSAVYGLAD HR 1.35, p=0.0052), corresponding to two haplotypes [h3, h117]. Cluster c3 included one risk motif (MAMYTCSAAYAYAD [h69], HR 2.44, 0.049) and two protective motifs (MAMYSQIDAYAYDD [e6] and MAMYTCSAAYAYDD [h19], HR 0.66 and 0.42, p=0.004 and 0.036, respectively). It is of great interest in this respect, that protective h19 (DQA1*05:01/B1*03:01) and susceptible h69 (DQA1*05:01/B1*03:02) molecules differ by only one residue in their entire sequence (signal peptide plus mature chain) (i.e. β57Asp vs Ala, respectively. This is a case where the role of β57Asp becomes paramount in protection from progression to type 1 diabetes. Indeed, all motifs associated with resistance against progression to type 1 diabetes are β57Asp+ while all those associated with susceptibility to progression in this study are β57Asp−.
Discussion
Using AA sequences of DQ heterodimers in the iCohort, we have carried out an exploratory analysis on genetic associations of residues on the α-chain and β-chain, and identified AAs that provide new insights into their genetic mechanisms. In addition, it is of interest to note that four AAs (−16α, −13α, −6α and −18β) concern residues in the respective signal peptides that could be involved in the efficiency of translation of the respective polypeptide chain. This is a subject that has not been adequately studied for HLAII [19]. The list of identified residues, plus residues in absolute LD to them in the alleles at hand, essentially contains residues involved in all known functions of HLAII molecules: antigen binding in the α1β1 domain groove; TCR contact; putative TCR-induced (αβ)2 homodimerisation; CD4 co-receptor binding (and perhaps lymphocyte-activation gene 3 binding); and cholesterol-raft clustering (see ESM Table 9) [8,9,10, 14, 20,21,22,23,24,25].
HOH with and without inclusion of structurally forbidden heterodimers
As noted, the list of 186 DQ heterodimers in iCohort included 74 ‘forbidden’ heterodimeric recombinants, although 19 heterodimers were common, with the rest being rare heterodimers (ESM Table 6). Rare heterodimers make a limited contribution to type 1 diabetes. For those common forbidden DQ heterodimers, none of them were significantly associated with type 1 diabetes (p>0.05, ESM Table 7). Collectively, all ‘forbidden’ heterodimers made no contribution to type 1 diabetes progression, an observation consistent with that made in haematopoietic stem cell transplantation [17]. Further, we repeated HOH with D heterodimers, excluding 74 ‘forbidden’ heterodimers, and obtained a list of AAs the same as those presented in Table 2.
Structural properties of identified AAs
It comes as no surprise, once again, that the HLA-DQ molecules constituting the main gene product leading to type 1 diabetes, are essential in people at risk: alleles predisposing towards type 1 diabetes to begin with, are predisposing in people at risk (with the exception of DQA1*05:01-DQB1*02:01), while alleles conferring protection do so in people at risk (see Conclusions below for explanation of lower HR values for progression/protection). It is remarkable that most AAs shown to be important by HOH for progression to type 1 diabetes or protection therefrom (ESM Table 9), as well as those in absolute LD in the alleles at hand, are involved in one or more of the known functions of HLAII molecules. In particular, the most highly statistically significant correlations involve four residues: at α22 (Tyr for susceptibility, Phe for resistance alleles, but without significance); at β13 (Gly for susceptibility, Ala for protective alleles); at β26 (Leu for susceptibility and Tyr for protective alleles); and β57 (Ala for susceptibility and Asp for protective alleles) (Table 2). These four residues participate in the formation of pockets 1 (α22, formation of a water-mediated hydrogen bond, in the case of Tyr, with the antigenic peptide backbone near pocket 1), 4 (β13 and β26) and 9 (β57), with sharply distinct anchor preferences. Specifically, β13Gly/β26Leu in the susceptible DQ8/cis and trans molecules (DQA1*03:01-DQB1*03:02 and DQA1*05:01-DQB1*03:02, respectively, and perhaps other DQA1*03/05 molecules with one or two substitutions) allow for large aliphatic and aromatic anchors in pocket 4. By contrast, the susceptible molecules DQ2/cis and trans (DQA1*05:01-DQB1*02:01 and DQA1*03:01-DQB1*02:01, respectively, and likewise other DQA1*03/05 alleles with one or two substitutions), and also β13Gly/β26Leu, prefer primarily acidic residues at p4 because of the β70Arg/71Lys/74Ala combination in this pocket, and secondarily large aliphatic and aromatic anchors [26,27,28,29]. On the other hand, β13Ala/β26Tyr in the resistant DQ7/cis and trans (DQA1*03:01-DQB1*03:01 and DQA1*05:01-DQB1*03:01, and likewise for other DQA1*03/05 molecules with one or two substitutions) allow only small anchors, such as Gly, Ala, Ser and Cys at pocket 4, while peptides with larger p4 anchors cannot bind [26]. In the case of pocket 9, the β57Asp/Ala dimorphism seen, respectively, in the vast majority of type 1 diabetes resistant vs susceptible HLA-DQB1 molecules is responsible for the anchor choice of small aliphatic vs acidic residues. Furthermore, the DQ2 cis/trans molecules show an even higher preference of aromatic residues at p9 (even Trp) because their respective p9 pocket is the largest for all known MHC II molecules, due to the presence of β30Ser/β37Ile instead of β30Tyr/β37Tyr in DQ8 cis/trans molecules [27, 28, 30]. In addition to these residues, shown to be associated with significant predispositions for progression to disease or protection therefrom, other such residues in the α2β2 domain as well as the intramembranous domain are associated with different HLA-DQ functions (ESM Table 9), that might also make a difference in type 1 diabetes progression.
Limitations
This is a post hoc analysis of an integrated iCohort from two randomised clinical trials, with newly generated HLA genotypes from all participants. While the randomisation obviates biases in allocations of participants, the post hoc nature implies that association results from HLA-DQ need to be independently replicated.
Conclusions
The current investigation of participants in the DPT-1 and TN07 who probably have ongoing autoimmunity suggests that autoimmune processes involving HLA-DQ molecules are necessary for islet beta cell destruction, together with other subsequent processes (such as CD8 T cell attacks), leading at-risk individuals to progress to clinical type 1 diabetes. Insights on DQ molecular structures could stimulate further immunological investigation on beta cell destruction and hence new immunotherapies to halt or revert the type 1 diabetes progression [31].
Abbreviations
- AA:
-
Amino acid
- DPT-1:
-
Diabetes Prevention Trial-Type 1
- HLAII:
-
HLA class II genes (DR, DQ and DP)
- HOH:
-
Hierarchically organised haplotype association analysis
- HP:
-
Hybrid peptide
- IAPP:
-
Islet amyloid polypeptide
- LD:
-
Linkage disequilibrium
- NIDDK:
-
National Institute of Diabetes and Digestive and Kidney Diseases
- RC:
-
Relevant cluster
- TCR:
-
T cell receptor
- TN07:
-
TrialNet07 ‘Randomised Diabetes Prevention Trial with Oral Insulin
References
Patterson CC, Karuranga S, Salpea P et al (2019) Worldwide estimates of incidence, prevalence and mortality of type 1 diabetes in children and adolescents: Results from the International Diabetes Federation Diabetes Atlas, 9th edition. Diabetes Res Clin Pract 157:107842. https://doi.org/10.1016/j.diabres.2019.107842
Katsarou A, Gudbjornsdottir S, Rawshani A et al (2017) Type 1 diabetes mellitus. Nat Rev Dis Primers 3:17016. https://doi.org/10.1038/nrdp.2017.16
Skyler JS (2013) Primary and secondary prevention of Type 1 diabetes. Diabet Med 30(2):161–169. https://doi.org/10.1111/dme.12100
Butty V, Campbell C, Mathis D, Benoist C, Group DPTS (2008) Impact of diabetes susceptibility loci on progression from pre-diabetes to diabetes in at-risk individuals of the diabetes prevention trial-type 1 (DPT-1). Diabetes 57(9):2348–2359. https://doi.org/10.2337/db07-1736
Zhao LP, Skyler J, Papadopoulos GK et al (2022) Association of HLA-DQ heterodimer residues -18beta and beta57 with progression from islet autoimmunity to diabetes in the diabetes prevention trial-type 1. Diabetes Care 45(7):1610–1620. https://doi.org/10.2337/dc21-1628
Zhao LP, Papadopoulos GK, Skyler JS et al (2024) HLA class II (DR, DQ, DP) genes were separately associated with the progression from seroconversion to onset of type 1 diabetes among participants in two diabetes prevention trials (DPT-1 and TN07). Diabetes Care 47(5):826–834. https://doi.org/10.2337/dc23-1947
Murphy KM, Weaver C (2017) Janeway’s Immunobiology, Chapter 6: Antigen presentation to T lymphocytes. 9th edn. Garland Science, New York
Stern LJ, Brown JH, Jardetzky TS et al (1994) Crystal structure of the human class II MHC protein HLA-DR1 complexed with an influenza virus peptide. Nature 368(6468):215–221. https://doi.org/10.1038/368215a0
Lee KH, Wucherpfennig KW, Wiley DC (2001) Structure of a human insulin peptide-HLA-DQ8 complex and susceptibility to type 1 diabetes. Nat Immunol 2(6):501–507. https://doi.org/10.1038/88694
Brown JH, Jardetzky TS, Gorga JC et al (1993) Three-dimensional structure of the human class II histocompatibility antigen HLA-DR1. Nature 364(6432):33–39. https://doi.org/10.1038/364033a0
Diabetes Prevention Trial-Type 1 Diabetes Study Group (2002) Effects of insulin in relatives of patients with type 1 diabetes mellitus. N Engl J Med 346(22):1685–1691. https://doi.org/10.1056/NEJMoa012350
Nelson WC, Pyo CW, Vogan D et al (2015) An integrated genotyping approach for HLA and other complex genetic systems. Hum Immunol 76(12):928–938. https://doi.org/10.1016/j.humimm.2015.05.001
Fremont DH, Monnaie D, Nelson CA, Hendrickson WA, Unanue ER (1998) Crystal structure of I-Ak in complex with a dominant epitope of lysozyme. Immunity 8(3):305–317. https://doi.org/10.1016/s1074-7613(00)80536-1
Bondinas GP, Moustakas AK, Papadopoulos GK (2007) The spectrum of HLA-DQ and HLA-DR alleles, 2006: a listing correlating sequence and structure with function. Immunogenetics 59(7):539–553. https://doi.org/10.1007/s00251-007-0224-8
Kwok WW, Kovats S, Thurtle P, Nepom GT (1993) HLA-DQ allelic polymorphisms constrain patterns of class II heterodimer formation. J Immunol 150(6):2263–2272. https://doi.org/10.4049/jimmunol.150.6.2263
Tollefsen S, Hotta K, Chen X et al (2012) Structural and functional studies of trans-encoded HLA-DQ2.3 (DQA1*03:01/DQB1*02:01) protein molecule. J Biol Chem 287(17):13611–13619. https://doi.org/10.1074/jbc.M111.320374
Petersdorf EW, Bengtsson M, Horowitz M et al (2022) HLA-DQ heterodimers in hematopoietic cell transplantation. Blood 139(20):3009–3017. https://doi.org/10.1182/blood.2022015860
Tran MT, Faridi P, Lim JJ et al (2021) T cell receptor recognition of hybrid insulin peptides bound to HLA-DQ8. Nat Commun 12(1):5110. https://doi.org/10.1038/s41467-021-25404-x
Hegde RS (2022) The function, structure, and origins of the ER membrane protein complex. Annu Rev Biochem 91:651–678. https://doi.org/10.1146/annurev-biochem-032620-104553
Paliakasis K, Routsias J, Petratos K, Ouzounis C, Kokkinidis M, Papadopoulos GK (1996) Novel structural features of the human histocompatibility molecules HLA-DQ as revealed by modeling based on the published structure of the related molecule HLA-DR. J Struct Biol 117(2):145–163. https://doi.org/10.1006/jsbi.1996.0079
Cochran JR, Cameron TO, Stern LJ (2000) The relationship of MHC-peptide binding and T cell activation probed using chemically defined MHC class II oligomers. Immunity 12(3):241–250. https://doi.org/10.1016/s1074-7613(00)80177-6
König R (2002) Interactions between MHC molecules and co-receptors of the TCR. Curr Opin Immunol 14(1):75–83. https://doi.org/10.1016/s0952-7915(01)00300-4
Zhou W, König R (2003) T cell receptor-independent CD4 signalling: CD4-MHC class II interactions regulate intracellular calcium and cyclic AMP. Cell Signal 15(8):751–762. https://doi.org/10.1016/s0898-6568(03)00037-8
Silberstein JL, Du J, Chan KW et al (2024) Structural insights reveal interplay between LAG-3 homodimerization, ligand binding, and function. Proc Natl Acad Sci U S A 121(12):e2310866121. https://doi.org/10.1073/pnas.2310866121
Dixon AM, Roy S (2019) Role of membrane environment and membrane-spanning protein regions in assembly and function of the Class II Major Histocompatibility complex. Hum Immunol 80(1):5–14. https://doi.org/10.1016/j.humimm.2018.07.004
Kwok WW, Domeier ML, Raymond FC, Byers P, Nepom GT (1996) Allele-specific motifs characterize HLA-DQ interactions with a diabetes-associated peptide derived from glutamic acid decarboxylase. J Immunol 156(6):2171–2177. https://doi.org/10.4049/jimmunol.156.6.2171
van de Wal Y, Kooy YM, Drijfhout JW, Amons R, Papadopoulos GK, Koning F (1997) Unique peptide binding characteristics of the disease-associated DQ(alpha 1*0501, beta 1*0201) vs the non-disease-associated DQ(alpha 1*0201, beta 1*0202) molecule. Immunogenetics 46(6):484–492. https://doi.org/10.1007/s002510050309
Vartdal F, Johansen BH, Friede T et al (1996) The peptide binding motif of the disease associated HLA-DQ (alpha 1* 0501, beta 1* 0201) molecule. Eur J Immunol 26(11):2764–2772. https://doi.org/10.1002/eji.1830261132
van Lummel M, van Veelen PA, Zaldumbide A et al (2012) Type 1 diabetes-associated HLA-DQ8 transdimer accommodates a unique peptide repertoire. J Biol Chem 287(12):9514–9524. https://doi.org/10.1074/jbc.M111.313940
Stepniak D, Wiesner M, de Ru AH et al (2008) Large-scale characterization of natural ligands explains the unique gluten-binding properties of HLA-DQ2. J Immunol 180(5):3268–3278. https://doi.org/10.4049/jimmunol.180.5.3268
Fugger L, Jensen LT, Rossjohn J (2020) Challenges, progress, and prospects of developing therapies to treat autoimmune diseases. Cell 181(1):63–80. https://doi.org/10.1016/j.cell.2020.03.007
Funding
Open access funding provided by Lund University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Acknowledgements
The authors would like to express appreciation to the Steering Committee of TN07 in TrialNet for approving our request to access DNA samples and to thank NIDDK Central Repository for providing clinical data from both the DPT-1 and TN07. Equally, we would like to thank reviewers whose comments have substantially improved the presentation of this work.
Data availability
Clinical trial data and biospecimen samples are available through the NIDDK Central Repository portal (https://repository.niddk.nih.gov/studies); clinical data from participants in DPT-1 and TN07 are available from NIDDK Central Repository (https://repository.niddk.nih.gov/home/). Interested researchers are encouraged to contact corresponding authors for collaborative research on HLA-DQ genotype data.
Funding
The study was supported by a grant (R01 DK132406) from National Institute of Health/National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK). The funder played no role in study design, analysis or interpretation.
Authors’ relationships and activities
The authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work.
Contribution statement
LPZ, ÅL and GKP researched and analysed the data and wrote the manuscript. JSS and AP conducted the DPT-1 trial and were involved in analysis conceptualisation and result interpretation and both reviewed and edited the manuscript. HMP reviewed and analysed data, and also edited the manuscript. WWK reviewed the analysis results, expanded on implications of findings in the context of DRB3 binding peptides and contributed to the manuscript writing. TPL reviewed results and contributed to the writing of the manuscript. DEG, RW, C-WP and WCN sequenced HLA genes using next-generation sequencing, analysed sequence data and reviewed the manuscript. GKP, AKM and GPB performed structural analysis, constructed graphical representations of selected DQ molecules and contributed to the Discussion section of the manuscript. All authors approved the version of the manuscript submitted for publication. LPZ and ÅL are guarantors of this work, had full access to all the data, and take full responsibility for the integrity of data and accuracy of data analysis.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
George K. Papadopoulos has retired from the Technological Educational Institute (TEI) of Epirus, Arta, Greece effective 1 September 2018. The affiliation is provided for identification purposes only. As of 1 October 2018, the TEI of Epirus has been absorbed by the University of Ioannina. The respective department is now called the Department of Agriculture.
George P. Bondinas is an adjunct member of the Department of Food Science and Technology, Faculty of Environmental Sciences, Ionian University.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, L.P., Papadopoulos, G.K., Skyler, J.S. et al. Progression to type 1 diabetes in the DPT-1 and TN07 clinical trials is critically associated with specific residues in HLA-DQA1-B1 heterodimers. Diabetologia (2024). https://doi.org/10.1007/s00125-024-06274-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00125-024-06274-6