Abstract
Aims/hypothesis
Treatment change following a genetic diagnosis of MODY is frequently indicated, but little is known about the factors predicting future treatment success. We therefore conducted the first prospective study to determine the impact of a genetic diagnosis on individuals with GCK-, HNF1A- or HNF4A-MODY in the UK, and to identify clinical characteristics predicting treatment success (i.e. HbA1c ≤58 mmol/mol [≤7.5%]) with the recommended treatment at 2 years.
Methods
This was an observational, prospective, non-selective study of individuals referred to the Exeter Molecular Genetic Laboratory for genetic testing from December 2010 to December 2012. Individuals from the UK with GCK- or HNF1A/HNF4A-MODY who were not on recommended treatment at the time of genetic diagnosis, and who were diagnosed below the age of 30 years and were currently aged less than 50 years, were eligible to participate.
Results
A total of 44 of 58 individuals (75.9%) changed treatment following their genetic diagnosis. Eight individuals diagnosed with GCK-MODY stopped all diabetes medication without experiencing any change in HbA1c (49.5 mmol/mol [6.6%] both before the genetic diagnosis and at a median of 1.25 years’ follow-up without treatment, p = 0.88). A total of 36 of 49 individuals (73.5%) diagnosed with HNF1A/HNF4A-MODY changed treatment; however, of the 21 of these individuals who were being managed with diet or sulfonylurea alone at 2 years, only 13 (36.1% of the population that changed treatment) had an HbA1c ≤58 mmol/mol (≤7.5%). These individuals had a shorter diabetes duration (median 4.6 vs 18.1 years), lower HbA1c (58 vs 73 mmol/mol [7.5% vs 8.8%]) and lower BMI (median 24.2 vs 26.0 kg/m2) at the time of genetic diagnosis, compared with individuals (n = 23/36) with an HbA1c >58 mmol/mol (>7.5%) (or <58 mmol/mol [<7.5%] on additional treatment) at the 2 year follow-up. Overall, 64% (7/11) individuals with a diabetes duration of ≤11 years and an HbA1c of ≤69 mmol/mol (≤8.5%) at time of the genetic test achieved good glycaemic control (HbA1c ≤58 mmol/mol [≤7.5%]) with diet or sulfonylurea alone at 2 years, compared with no participants with a diabetes duration of >11 years and an HbA1c of >69 mmol/mol (>8.5%) at the time of genetic diagnosis.
Conclusions/interpretation
In participants with GCK-MODY, treatment cessation was universally successful, with no change in HbA1c at follow-up. In those with HNF1A/HNF4A-MODY, a shorter diabetes duration, lower HbA1c and lower BMI at genetic diagnosis predicted successful treatment with sulfonylurea/diet alone, supporting the need for early genetic diagnosis and treatment change. Our study suggests that, in individuals with HNF1A/HNF4A-MODY with a longer duration of diabetes (>11 years) at time of genetic test, rather than ceasing current treatment, a sulfonylurea should be added to existing therapy, particularly in those who are overweight or obese and have a high HbA1c.
Similar content being viewed by others
Avoid common mistakes on your manuscript.

Introduction
In the UK, MODY accounts for 3.6% of diabetes cases in individuals diagnosed younger than 30 years [1]. Diagnosis of MODY has significant implications for diabetes management. GCK-MODY causes asymptomatic, mild fasting hyperglycaemia (usually 5.4–8.3 mmol/l) [5]. The glucose level is regulated at a higher level in GCK-MODY, making glucose-lowering treatment ineffective [6], and therefore treatment is not recommended [4]. Individuals with HNF1A- or HNF4A-MODY are optimally treated with low-dose sulfonylureas [7,8,9,10] because of an increased pancreatic insulin secretory response to sulfonylureas and increased insulin sensitivity to the insulin secreted [7].
There is a significant delay from the diagnosis of diabetes to the correct molecular genetic diagnosis of MODY [2, 11,12,13,14]. The majority of individuals are initially misdiagnosed with type 1 or type 2 diabetes and inappropriately treated [12, 15,16,17,18,19,20].
Current data on the success of transfer to sulfonylurea treatment in individuals with HNF1A/HNF4A-MODY following genetic diagnosis are limited and retrospective [11, 21]. In one case, a study focused on individuals in a single centre with expertise in monogenic diabetes [10]. There have been no prospective studies that have assessed the success of treatment change, glycaemic control and maintenance on recommended treatment following genetic diagnosis in individuals with MODY in non-specialist centres.
The aims of our study were to determine the impact of a genetic diagnosis on diabetes treatment in UK individuals with GCK-, HNF1A- or HNF4A-MODY, and to identify clinical characteristics that predict successful management (i.e. HbA1c ≤58 mmol/mol [≤7.5%]) with no treatment in those with GCK-MODY or sulfonylureas in those with HNF1A and HNF4A-MODY at 2 years after genetic diagnosis.
Methods
Study design
This was an observational, prospective, non-selective study of all individuals with HNF1A/HNF4A- or GCK-MODY identified from routine UK referrals to the Exeter Molecular Genetic Laboratory for genetic testing from December 2010 to December 2012. Ethics approval was granted by the NRES Committee South West–Central Bristol (REC no. 10/H0106/03). This study was part of the UNITED (Using pharmacogeNetics to Improve Treatment in Early-onset Diabetes) study which aimed to determine prevalence of monogenic diabetes in those diagnosed with diabetes below the age of 30 years [1]. All study participants gave informed consent (with parental consent and children’s assent gained for those younger than 16 years, n = 9).
Individual characteristics
Individuals were eligible to participate if: (1) genetic testing confirmed HNF1A-, HNF4A- or GCK-MODY; (2) they were not on recommended treatment at time of genetic diagnosis; and (3) they had been diagnosed with diabetes when younger than 30 years and were younger than 50 years at time of genetic testing. Treatment was considered ‘non-recommended’ if those with HNF1A/HNF4A-MODY were treated with medication other than sulfonylureas and those with GCK-MODY were taking any diabetes therapy.
Overall, 305 individuals referred from across the UK were confirmed to have GCK-MODY (n = 112), HNF1A-MODY (n = 143) or HNF4A-MODY (n = 50) within the duration of this study. A total of 244 individuals did not meet eligibility criteria and were not followed up: 101 were excluded on age criteria (37 with GCK-MODY, 43 with HNF1A-MODY and 21 with HNF4A-MODY) and 143 were excluded based on treatment criteria (63 with GCK-MODY, 61 with HNF1A-MODY and 19 with HNF4A-MODY).
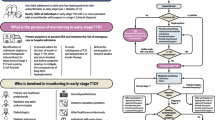
Therefore, 61 individuals fulfilled the eligibility criteria. Of these, 58 were contactable and agreed to participate (39 with HNF1A-MODY, 10 with HNF4A-MODY and nine with GCK-MODY; Fig. 1). This included 11 related individuals from five families: two parent–child pairs with HNF1A-MODY; one family in which the mother, her son and her sister had HNF1A-MODY; one sibling pair with HNF1A-MODY; and one sibling pair with GCK-MODY (see electronic supplementary material [ESM] Table 1). All the individuals in the study were white, except one who was of mixed white and East Asian ethnicity. There were 41 women. The median age at the diagnosis of diabetes was 17 (interquartile range [IQR] 13–21] years; at the time of genetic testing median BMI was 24.8 (IQR 21.9–28.2) kg/m2, duration of diabetes 10 (IQR 2–20) years and baseline HbA1c 59.5 (IQR 50–73) mmol/mol (7.6% [IQR 6.7–8.8%]). At the time of genetic diagnosis, 50 individuals (86.2%) were being treated with insulin (43 with HNF1A/HNF4A-MODY and seven with GCK-MODY) and eight (13.8%) were taking metformin alone (six with HNF1A/HNF4A-MODY and two with GCK-MODY). Of those on insulin, 46 were on insulin alone and four took metformin in addition. The BMI for children under the age of 19 years was adjusted to the adult equivalent using the Child Growth Foundation Reference Standards [22].
Flow chart indicating recruitment, treatment at genetic diagnosis and treatment at 2 years after the genetic diagnosis. GLA, glucose-lowering agent; MF, metformin; SU, sulfonylurea. aThis individual with GCK-MODY was initially treated with insulin and metformin but did not stop all treatment following the genetic test
Follow-up and treatment
Individuals were telephoned at baseline (i.e. the time of the genetic test result) and at 3, 6, 12 and 24 months. Self-reported diabetes treatment was recorded. HbA1c was measured at baseline (prior to treatment change) and at 3, 6 and 12 months from ‘finger-prick’ blood samples that were collected at home and posted to the Blood Sciences laboratory at the Royal Devon and Exeter NHS Foundation Trust. HbA1c results at 24 months were accessed from the individual’s local laboratory. Genetic reports included a statement indicating the recommended treatment for GCK-, HNF1A- and HNF4A-MODY, but all decisions regarding diabetes management after genetic diagnosis were made by local clinicians.
Statistical analysis
Non-parametric tests (Mann–Whitney test for continuous variables, χ2 or Fisher’s exact test for categorical variables) were used to compare the characteristics of treatment groups. The Wilcoxon matched-pairs signed-ranks test was used to compare HbA1c results before and after the genetic diagnosis. Continuous data are expressed as medians (IQR). A p value <0.05 was considered significant. For two individuals, a single HbA1c value was imputed assuming a linear trend between two available HbA1c points. Analysis was conducted using Stata/SE 14 (StataCorp, College Station, TX, USA).
Results
A total of 44 of 58 individuals (75.9%) changed treatment following the genetic diagnosis (Fig. 1, ESM Table 1). Eleven of the 44 participants (25%) were younger than 18 years (four with GCK-MODY, six with HNF1A-MODY and one with HNF4A-MODY) at the time of genetic diagnosis. Fourteen individuals (24.1%) did not change treatment and were not followed-up. Reasons for continuing with the previous treatment were pregnancy (n = 3), individual choice (n = 5) and clinician choice (n = 5); this included individuals with retinopathy and nephropathy or concomitant confirmed type 1 diabetes (n = 1, GAD-antibody positive, urine C-peptide creatinine ratio 0.12 nmol/mol) (Fig. 1).
Eight of nine individuals with GCK-MODY (including seven previously treated with insulin) stopped all diabetes treatment following their genetic diagnosis, irrespective of diabetes duration (median 1.8 [IQR 0.6–7.2] years) and BMI (median 19.8 [IQR 17.9–22.7] kg/m2). HbA1c remained the same at a median of 1.25 (IQR 1–2) years’ follow-up without any treatment (49.5 [IQR 47–52] mmol/mol [6.6%, IQR 6.4–6.9%] at the genetic diagnosis vs 49.5 [IQR 47–50.5] mmol/mol [6.6%, IQR 6.5–6.8%] at follow-up, p = 0.88) (ESM Fig. 1). One individual with GCK-MODY stopped insulin but remained on metformin through the clinician’s choice; however, all recorded HbA1c values were 52–57 mmol/mol (6.9–7.4%), which are consistent with levels seen in GCK-MODY.
A total of 36 of 49 (73.5%) individuals diagnosed with HNF1A/HNF4A-MODY changed treatment following the genetic diagnosis (Fig. 1). Of these, 21 of 36 (58%) were treated with diet (n = 3) or sulfonylurea (n = 18) alone at 2 years. Thirteen of these 21 individuals (62%) had HbA1c ≤58 mmol/mol (≤7.5%) at 2 years (Table 1).
We next compared the clinical characteristics of the 13 individuals (36.1%, 13/36) with HNF1A/HNF4A-MODY being managed with sulfonylurea/diet alone who achieved an HbA1c ≤58 mmol/mol (≤7.5%) with those of the 23 individuals with an HbA1c >58 mmol/mol (>7.5%) (n = 22) or ≤58 mmol/mol (≤7.5%) on additional treatment (n = 1) at 2 year follow-up (Table 1). The individuals with an HbA1c ≤58 mmol/mol (≤7.5%) on sulfonylurea/diet alone at 2 years had a shorter diabetes duration (median 4.6 vs 18.1 years), lower BMI (median 24.2 vs 26.0 kg/m2) and lower HbA1c (58 vs 73 mmol/mol [7.5 vs 8.8%]) at treatment transfer compared with those with an HbA1c >58 mmol/mol (>7.5%) or the single individual with an HbA1c <58 mmol/mol (<7.5%) on additional treatment (Table 1). There was no difference in genetic aetiology between these groups (ESM Table 1). Those managed with sulfonylurea/diet alone at 2 years improved their HbA1c from a median of 58 mmol/mol (7.5%) pre-genetic diagnosis to 46 mmol/mol (6.4%) at 2 years (p = 0.001). This contrasted with the other group, in which HbA1c increased (median 73 vs 77 mmol/mol [8.8 vs 9.2%]), p = 0.03) (Table 1). Individuals in the latter group who were taking sulfonylureas were on maximum recommended dose (gliclazide 160 mg twice daily).
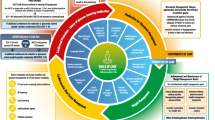
We also assessed the combined effect of diabetes duration and HbA1c at genetic diagnosis on the ability to achieve good glycaemic control with diet/sulfonylurea alone in individuals with HNF1A/HNF4A-MODY. We divided the cohort by median diabetes duration (≤11 vs >11 years) and median HbA1c (≤69 vs >69 mmol/mol [≤8.5% vs >8.5%]) at genetic diagnosis (Fig. 2). A total of 10/18 individuals (56%) with a shorter diabetes duration achieved optimal control, compared with 3/18 (17%) with longer diabetes duration (p = 0.03). Similarly, 10/18 (56%) with lower HbA1c at genetic diagnosis achieved optimal control compared with 3/18 (17%) with higher HbA1c at genetic diagnosis (p = 0.03). A total of 7 of 11 individuals (64%) with shorter diabetes duration and lower HbA1c at genetic diagnosis achieved optimal control, while none of the individuals (0/11) with longer duration and higher HbA1c at genetic diagnosis achieved an HbA1c ≤58 mmol/mol (≤7.5%) with diet/sulfonylurea alone (p = 0.02). Similar results were seen for diabetes duration and BMI at genetic diagnosis (ESM Fig. 2).
Effect of duration of diabetes and HbA1c at genetic diagnosis on the ability to achieve good glycaemic control with diet/sulfonylurea alone at 2 years following genetic diagnosis in individuals with HNF1A/HNF4A-MODY. Participants were divided into groups according to HbA1c (≤69 or >69 mmol/mol [≤8.5% or >8.5%], n = 18 in each group) and duration of diabetes (≤11 or >11 years, n = 18 in each group) using the median values of the HNF1A/HNF4A-MODY cohort (n = 36). The participant numbers in each of the groups were: HbA1c ≤69 mmol/mol (≤8.5%) and duration ≤11 years, n = 11; HbA1c ≤69 mmol/mol (≤8.5%) and duration >11 years, n = 7; HbA1c >69 mmol/mol (>8.5%) and duration ≤11 years, n = 7; and HbA1c >69 mmol/mol (>8.5%) and duration >11 years, n = 11. The number of individuals who had an HbA1c ≤58 mmol/mol (≤7.5%) with diet or sulfonylurea alone at 2 years in these four groups was n = 7, n = 3, n = 3 and n = 0, respectively
Discussion
This national, prospective, non-selective study demonstrates that most individuals with MODY commence the recommended treatment after a genetic diagnosis has been confirmed. However, only 58% of individuals with HNF1A/HNF4A-MODY were on diet or sulfonylurea alone at 2 years and, overall, just 36% of individuals with HNF1A/HNF4A-MODY who changed treatment achieved the good glycaemic control (≤58 mmol/mol [≤7.5%]) needed to avoid diabetes complications. Our study suggests that successful treatment with diet/sulfonylurea alone was most likely in those with HNF1A/HNF4A-MODY who had a shorter duration of diabetes, healthy BMI and lower HbA1c at the time of genetic diagnosis. All participants with GCK-MODY were able to stop insulin or oral hypoglycaemic agents without deterioration in glycaemic control, as previously shown [6]. Identifying those with GCK-MODY is important, as all diabetes treatments can be discontinued and follow-up is not required [2,3,4, 6].
Improvement in glycaemic control among individuals with HNF1A/HNF4A-MODY is needed to prevent diabetes complications. Individuals with HNF1A/HNF4A-MODY are at increased, or at least the same, risk of developing diabetes-related complications compared with those with other diabetes subtypes [23, 24]. Our study showed that despite transfer to the recommended treatment, only 36% of individuals achieved an HbA1c ≤58 mmol/mol (≤7.5%) on sulfonylurea/diet alone. The lack of optimal glycaemic control in our study may have resulted from clinical inertia or limited experience among local clinicians in managing HNF1A/HNF4A-MODY and previous advice advocating a trial of sulfonylureas even in those with longstanding diabetes [11]. The lack of standardised treatment guidelines for individuals needing additional second-line therapy may also contribute to suboptimal glycaemic control. Our results are similar, albeit lower, than those of previous studies, which found that around 50–62% of participants attained an HbA1c ≤58 mmol/mol (≤7.5%) with sulfonylurea therapy alone [10, 11]. The difference in the results may be a result of differences in the duration of diabetes at genetic diagnosis.
Progressive loss of pancreatic beta cell function is a feature of HNF1A/HNF4A-MODY, resulting in increasing glycaemia and increasing treatment requirements over time [25]. Successful treatment change and achieving good glycaemic control is more likely to be achieved if the genetic diagnosis is made early. Prompt transfer to sulfonylureas, enabling optimal glycaemic control soon after diabetes diagnosis, may reduce the risk of future complications in those with HNF1A/HNF4A-MODY, as seen with type 1 and type 2 diabetes [26, 27]. If individuals are transferred to optimal treatment early, then it may be easier to achieve good control and to maintain it. This is reflected by our data showing that individuals with lower HbA1c levels at genetic diagnosis are more likely to achieve good glycaemic control at 2 years. In contrast to this, individuals with higher HbA1c levels at genetic diagnosis rarely achieved good glycaemic control with sulfonylureas alone. As a consequence of these data, we now recommend that a sulfonylurea should be added to existing treatment, rather than replacing it, in individuals with HNF1A/HNF4A-MODY with a longer diabetes duration (>11 years), especially in those with higher HbA1c levels at genetic diagnosis and a BMI >25 kg/m2.
In this study, we found that higher HbA1c levels and BMI at genetic diagnosis were associated with reduced success on sulfonylurea treatment in participants with HNF1A/HNF4A-MODY. Similar results have been seen in a previous retrospective study [10]. In our data, a higher BMI at genetic diagnosis markedly reduced the success of sulfonylurea therapy in those with a longer duration of diabetes. This is likely to reflect the impact of increased insulin resistance in those with more severe beta cell dysfunction. These data raise the question of whether weight loss may aid glycaemic control in individuals with HNF1A/HNF4A-MODY.
Our study has limitations. Treatment decisions were made via local clinicians and were not standardised. We did not collect data regarding changes in BMI over time and any effect this had on treatment requirements, which has previously been shown to negatively affect glycaemic control [10]. It was not appropriate to use multiple regression analysis of factors predicting successful long-term treatment with sulfonylureas alone to identify the relative contribution of each factor because of the small size of our study. We did not measure endogenous insulin secretion at time of genetic diagnosis in our participants and were therefore unable to assess its role in treatment response. Finally, our study did not have a large enough sample size to detect whether specific genetic mutations had an effect on the response to treatment over and above the strongly associated clinical features we identified. Despite these limitations, our study provides the first national prospective data regarding treatment change following genetic diagnosis in non-specialised centres across the UK.
In summary, our national prospective study identified that the majority of individuals changed treatment following a genetic diagnosis of MODY. Those with GCK-MODY were able to stop all diabetes treatment with no deterioration in HbA1c, highlighting the significance of identifying individuals with GCK-MODY as diabetes medication is unnecessary and follow-up is not required. In participants with HNF1A/HNF4A-MODY, only 58% were maintained on sulfonylurea/diet alone at 2 years and just 36% of participants with HNF1A/HNF4A-MODY who changed treatment achieved an HbA1c ≤58 mmol/mol (≤7.5%) 2 years following genetic diagnosis. A shorter duration of diabetes, lower HbA1c level and lower BMI at genetic diagnosis predicted successful treatment with sulfonylurea/diet alone in participants with HNF1A/HNF4A-MODY, supporting the need for early genetic diagnosis and treatment change.
Data availability
The datasets generated and analysed from the study are available from the corresponding author on reasonable request.
Abbreviations
- IQR:
-
Interquartile range
References
Shields BM, Shepherd M, Hudson M et al (2017) Population-based assessment of a biomarker-based screening pathway to aid diagnosis of monogenic diabetes in young-onset patients. Diabetes Care 40:1017–1025
Carmody D, Lindauer KL, Naylor RN (2015) Adolescent non-adherence reveals a genetic cause for diabetes. Diabet Med 32:e20–e23
Carmody D, Naylor RN, Bell CD et al (2016) GCK-MODY in the US National Monogenic Diabetes Registry: frequently misdiagnosed and unnecessarily treated. Acta Diabetol 53:703–708
Chakera AJ, Steele AM, Gloyn AL et al (2015) Recognition and management of individuals with hyperglycemia because of a heterozygous glucokinase mutation. Diabetes Care 38:1383–1392
Steele AM, Shields BM, Wensley KJ, Colclough K, Ellard S, Hattersley AT (2014) Prevalence of vascular complications among patients with glucokinase mutations and prolonged, mild hyperglycemia. JAMA 311:279–286
Stride A, Shields B, Gill-Carey O et al (2014) Cross-sectional and longitudinal studies suggest pharmacological treatment used in patients with glucokinase mutations does not alter glycaemia. Diabetologia 57:54–56
Pearson ER, Starkey BJ, Powell RJ, Gribble FM, Clark PM, Hattersley AT (2003) Genetic cause of hyperglycaemia and response to treatment in diabetes. Lancet 362:1275–1281
Murphy R, Ellard S, Hattersley AT (2008) Clinical implications of a molecular genetic classification of monogenic beta-cell diabetes. Nat Clin Pract Endocrinol Metab 4:200–213
Rubio-Cabezas O, Hattersley AT, Njolstad PR et al (2014) ISPAD Clinical Practice Consensus Guidelines 2014. The diagnosis and management of monogenic diabetes in children and adolescents. Pediatr Diabetes 15(Suppl 20):47–64
Bacon S, Kyithar MP, Rizvi SR et al (2016) Successful maintenance on sulphonylurea therapy and low diabetes complication rates in a HNF1A-MODY cohort. Diabet Med 33:976–984
Shepherd M, Shields B, Ellard S, Rubio-Cabezas O, Hattersley AT (2009) A genetic diagnosis of HNF1A diabetes alters treatment and improves glycaemic control in the majority of insulin-treated patients. Diabet Med 26:437–441
Pihoker C, Gilliam LK, Ellard S et al (2013) Prevalence, characteristics and clinical diagnosis of maturity onset diabetes of the young due to mutations in HNF1A, HNF4A, and glucokinase: results from the SEARCH for Diabetes in Youth. J Clin Endocrinol Metab 98:4055–4062
Thanabalasingham G, Owen KR (2011) Diagnosis and management of maturity onset diabetes of the young (MODY). BMJ 343:d6044
Irgens HU, Molnes J, Johansson BB et al (2013) Prevalence of monogenic diabetes in the population-based Norwegian Childhood Diabetes Registry. Diabetologia 56:1512–1519
Shields BM, Hicks S, Shepherd MH, Colclough K, Hattersley AT, Ellard S (2010) Maturity-onset diabetes of the young (MODY): how many cases are we missing? Diabetologia 53:2504–2508
Gandica RG, Chung WK, Deng L, Goland R, Gallagher MP (2015) Identifying monogenic diabetes in a pediatric cohort with presumed type 1 diabetes. Pediatr Diabetes 16:227–233
Lambert AP, Ellard S, Allen LI et al (2003) Identifying hepatic nuclear factor 1α mutations in children and young adults with a clinical diagnosis of type 1 diabetes. Diabetes Care 26:333–337
Thirumalai A, Holing E, Brown Z, Gilliam LK (2013) A case of hepatocyte nuclear factor-1β (TCF2) maturity onset diabetes of the young misdiagnosed as type 1 diabetes and treated unnecessarily with insulin. J Diabetes 5:462–464
Thanabalasingham G, Pal A, Selwood MP et al (2012) Systematic assessment of etiology in adults with a clinical diagnosis of young-onset type 2 diabetes is a successful strategy for identifying maturity-onset diabetes of the young. Diabetes Care 35:1206–1212
Johansson BB, Irgens HU, Molnes J et al (2017) Targeted next-generation sequencing reveals MODY in up to 6.5% of antibody-negative diabetes cases listed in the Norwegian Childhood Diabetes Registry. Diabetologia 60:625–635
Shepherd M, Pearson ER, Houghton J, Salt G, Ellard S, Hattersley AT (2003) No deterioration in glycemic control in HNF-1α maturity-onset diabetes of the young following transfer from long-term insulin to sulphonylureas. Diabetes Care 26:3191–3192
Freeman JV, Cole TJ, Chinn S, Jones PR, White EM, Preece MA (1995) Cross sectional stature and weight reference curves for the UK, 1990. Arch Dis Child 73:17–24
Steele AM, Shields BM, Shepherd M, Ellard S, Hattersley AT, Pearson ER (2010) Increased all-cause and cardiovascular mortality in monogenic diabetes as a result of mutations in the HNF1A gene. Diabet Med 27:157–161
Isomaa B, Henricsson M, Lehto M et al (1998) Chronic diabetic complications in patients with MODY3 diabetes. Diabetologia 41:467–473
Pearson ER, Velho G, Clark P et al (2001) Beta-cell genes and diabetes: quantitative and qualitative differences in the pathophysiology of hepatic nuclear factor-1alpha and glucokinase mutations. Diabetes 50(Suppl 1):S101–S107
Holman RR, Paul SK, Bethel MA, Matthews DR, Neil HA (2008) 10-year follow-up of intensive glucose control in type 2 diabetes. N Engl J Med 359:1577–1589
Nathan DM, Cleary PA, Backlund JY et al (2005) Intensive diabetes treatment and cardiovascular disease in patients with type 1 diabetes. N Engl J Med 353:2643–2653
Acknowledgements
We are grateful to the individuals who took part in this study and the clinicians involved in their care. We thank K. Colclough (Department of Molecular Genetics, Royal Devon and Exeter NHS Foundation Trust, Exeter, UK) for his help retrieving data regarding genotypes. Some of the data were presented as an abstract at the Diabetes UK Professional Conference in 2018.
Funding
This work presents independent research commissioned by the Health Innovation Challenge Fund (grant number HICF-1009-041), a parallel funding partnership between the Wellcome Trust and the Department of Health; and was supported by the National Institute for Health Research (NIHR) Exeter Clinical Research Facility. MS, BS and MH are supported by the NIHR Exeter Clinical Research Facility. KAP has a postdoctoral fellowship funded by the Wellcome Trust (110082/Z/15/Z). CH is supported by the NIHR Collaboration for Leadership in Applied Health Research and Care South West Peninsula. SE and ATH are Wellcome Trust Senior Investigators (WT098395/Z/12/Z), and ATH is an NIHR Senior Investigator. ERP is a Wellcome Trust New Investigator (102820/Z/13/Z).
The views expressed are those of the author(s) and not necessarily those of the Wellcome Trust, Department of Health, NHS or NIHR.
Author information
Authors and Affiliations
Consortia
Contributions
All authors contributed to the concept, design, acquisition of data or analysis and interpretation of data and drafting/revising the article and final approval of the article. MS is responsible for the integrity of the work as a whole.
Corresponding author
Ethics declarations
The authors declare that there is no duality of interest associated with this manuscript.
Electronic supplementary material
ESM
(PDF 170 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Shepherd, M.H., Shields, B.M., Hudson, M. et al. A UK nationwide prospective study of treatment change in MODY: genetic subtype and clinical characteristics predict optimal glycaemic control after discontinuing insulin and metformin. Diabetologia 61, 2520–2527 (2018). https://doi.org/10.1007/s00125-018-4728-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-018-4728-6