Abstract
Aims/hypothesis
Dipeptidyl peptidase 4 (DPP-4) inhibitors are agents designed to increase the half-life of incretins. Although they are administered orally, little is known about their effects on the gut microbiota and functions, despite the fact that some bacteria present in the gut microbiota exhibit DPP-4-like activity. Our objective was to study the impact of the DPP-4 inhibitor vildagliptin on gut functions and the intestinal ecosystem in a murine model of obesity induced by a Western diet (WD).
Methods
Twenty seven male C57BL/6J mice were randomised to receive a control diet, a WD (45% kJ from fat and 17% kJ from sucrose) or a WD + vildagliptin (0.6 mg/ml in drinking water) for 8 weeks.
Results
Vildagliptin significantly reduced DPP-4 activity in the caecal content and faeces. Vildagliptin impacted on the composition of the gut microbiota and its metabolic activity. It mainly decreased Oscillibacter spp. (a direct effect independent of DPP-4 activity was shown on cultured O. valericigenes), increased Lactobacillus spp. and propionate, and reduced the ligands of Toll-like receptors 2 and 4. Vildagliptin protected against the reductions in crypt depth and ileal expression of antimicrobial peptides induced by the WD. In the liver, the expression of immune cell populations (Cd3g and Cd11c [also known as Itgax]) and cytokines was decreased in the WD + vildagliptin-fed mice compared with the WD-fed group. Ex vivo exposure of precision-cut liver slices to vildagliptin showed that this response was not related to a direct effect of the drug on the liver tissue.
Conclusions/interpretation
Our study is the first to consider the DPP-4-like activity of the gut microbiota as a target of DPP-4 inhibition. We propose that vildagliptin exerts beneficial effects at the intestinal level in association with modulation of gut microbiota, with consequences for hepatic immunity. If relevant in humans, this could open new therapeutic uses of DPP-4 inhibition to tackle gut dysfunctions in different pathophysiological contexts.
Data availability
The sequences used for analysis can be found in the MG-RAST database under the project name MYNEWGUT3.
Similar content being viewed by others
Avoid common mistakes on your manuscript.

Introduction
Enteroendocrine cells release incretins in response to nutrient ingestion. The two main incretins—glucagon-like peptide (GLP) 1 and glucose-dependent insulinotropic peptide (GIP)—regulate the postprandial secretion of insulin [1]. One limitation of GLP-1 and GIP activities is their short half-life, because they are rapidly cleaved and inactivated by dipeptidyl peptidase 4 (DPP-4). DPP-4 exists as a membrane-anchored cell surface peptidase and as a soluble form in the circulation [2]. The expression and circulation levels of DPP-4 in people with type 2 diabetes are higher than in those without diabetes [3]. Additionally, DPP-4 is considered an adipokine positively correlated to body weight, adipose tissue inflammation and insulin resistance in obese individuals [4, 5].
Sitagliptin, anagliptin and vildagliptin are a new type of DPP-4 inhibitors which are administered orally and are well tolerated [6]. Although first approved in 2006, their effects on essential intestinal functions remain poorly studied. In 2011, Waget et al described how inhibition of DPP-4 activity in the intestine was involved in regulating glycaemia [7]. Recently, Mulvihill et al demonstrated that glucose tolerance depended on inhibition of DPP-4 activity in haematopoietic and endothelial cells, but there was no involvement of the DPP-4 expressed by enterocytes [8]. DPP-4 activity modulates the functionality of more than 40 potential substrates, including cytokines, chemokines and growth factors, some of which are particularly relevant for gut homeostasis [9]. For example, the preservation of GLP-2 with DPP-4 inhibitors (valine pyrrolidide) has been proposed to stimulate intestinal epithelial growth [10, 11].
Besides cleaving gut peptides, DPP-4 plays a role in the immune response. Soluble DPP-4 increases the expression of Toll-like receptors (TLRs) and activates NFκB signalling, resulting in the secretion of proinflammatory cytokines [12, 13]. These effects can be reduced by DPP-4 inhibitors [14,15,16]. For instance, in macrophages vildagliptin counteracts the production of cytokines and expression of TLR-2 and TLR-4 induced by soluble DPP-4 and lipopolysaccharide [13]. As TLRs are part of the innate immune response and play a crucial role in the maintenance of gut homeostasis, investigating the effect of vildagliptin in the context of intestinal health may produce particularly relevant results [17].
Vildagliptin is principally absorbed in the small, and to a lesser extent in the large, intestine [18]. During its passage through the intestinal tract, eukaryotic cells as well as the gut microbiota are exposed to vildagliptin. It has been widely described that intestinal microbes respond to environmental factors such as drugs and food, generating protective or detrimental effects [19, 20]. Interestingly, some genera of bacteria in the gut microbiota, such as Prevotella and Lactobacillus, have been reported to exert DPP-4-like activity [21,22,23]. Furthermore, some strains of Bifidobacterium and Lactobacillus can produce DPP-4 inhibitors [24, 25]. To our knowledge, the DPP-4-like activity of the gut microbiota has never been studied. One study (without assessment of DPP-4 activity) reported that vildagliptin caused changes in the gut microbiota and that the changes were more significant in rats fed a high-fat diet than in those fed a control diet [26]. Furthermore, a recent report showed that high-fat, high-sugar diets altered the activation of enteric neurons by GLP-1 regulating insulin secretion, an effect that seemed to be also dependent on the gut microbiota [27]. We therefore assessed the effect of vildagliptin on the gut microbiota and intestinal functions of a murine model of obesity induced by a Western diet (WD), a model that induces gut dysfunction and related hepatic inflammation.
Methods
Animals and treatments
Two animal experiments were performed. In experiment 1, 27 male 9-week-old C57BL/6J mice (Janvier Labs, Saint-Berthevin, France) were purchased. Three mice were housed in one individually ventilated cage. In experiment 2, four male 12-week-old C57BL/6J mice (Janvier Labs) were housed in one individually ventilated cage. Mice were kept in a pathogen-free environment with a 12 h daylight cycle and free access to food and water. The acclimatisation period lasted 1 week on a standard diet (AIN-93M; ssniff, Soest, Germany). The experiments were approved by and performed in accordance with the guidelines of the local ethics committee of Université catholique de Louvain. Housing conditions were as specified by the Belgian Law of 29 May 2013 regarding the protection of laboratory animals (Agreement no LA 1230314).
In experiment 1, mice were randomised based on body composition assessed by NMR (LF50 minispec; Bruker, Rheinstetten, Germany) to minimise differences (initial mean body weight ± SEM was 24.46 ± 0.23 g). No blinding procedure was followed. The groups (n = 9) were: (1) group-fed a control diet (D12450K; Research Diets, New Brunswick, NJ, USA) containing 10% kJ fat; (2) group-fed a WD (D12451; Research Diets) containing 45% kJ fat and 17% kJ sucrose; and (3) group-fed a WD plus vildagliptin (Cayman, Ann Arbor, MI, USA [supplied by Sanbio, Uden, the Netherlands]) in the drinking water (0.6 mg/ml according to previous studies [13], corresponding to approximately 50 mg kg−1 day−1). To discriminate the effect of vildagliptin from the effect of the WD on DPP-4 activity, we pretreated with vildagliptin for 2 weeks. In the third week, the WD was introduced. A scheme of the experimental design is shown in electronic supplementary material (ESM) Fig. 1.
After 8 weeks and 6 h of fasting, glycaemia was measured using a glucometer (Roche Diagnostics, Basel, Switzerland) on blood from the tail. Mice were anaesthetised with isoflurane gas (Abbot, Lake Bluff, IL, USA). Portal blood was collected. Mice were necropsied after cervical dislocation. Liver, adipose tissue and caecal content and tissue were weighed. Blood, liver, caecal content and tissue, ileum and colon were collected, frozen in liquid nitrogen and stored at −80°C until analysed.
Details of experiment 2 are given below in the section Precision-cut liver slices.
Biochemical analysis
Portal active GLP-1 (7-36 amide and 7-37), insulin and total GIP were determined in plasma collected in an EDTA tube with a DPP-4 inhibitor (DDP4-010; Millipore, Darmstadt, Germany) using a multiplex immunoassay kit (Bio-Plex; Bio-Rad, Nazareth, Belgium).
Analysis of gut microbiota composition
Genomic DNA was extracted from the caecal content using a QIAamp DNA Stool Mini Kit (Qiagen, Hildren, Germany), including a bead-beating step. The composition of the gut microbiota was analysed by Illumina sequencing of the 16S rRNA gene (Illumina, San Diego, CA, USA). Illumina sequencing was performed as previously described [28, 29]. The V5–V6 region of the 16S rRNA gene was amplified by PCR using modified primers. The amplicons were purified, quantified and sequenced using the Illumina MiSeq system to produce 2 × 300 bp sequencing products, at the University of Minnesota Genomics Center. Subsequent bioinformatics and biostatistics analyses were performed as previously described [28]. The full protocol and statistical analyses are described in ESM Methods.
In addition, in the caecal content, some components of the gut microbiota were analysed by quantitative real-time PCR (qPCR) as previously described [28]. Primers are presented in ESM Table 1.
Gene expression analyses
Total RNA was isolated from tissues and qPCR was performed using the StepOne System (Applied Biosystems, Waltham, MA, USA) to analyse expression of the following genes: Cd3g, Cd11c (also known as Itgax), Cd68, Cd163, Claudin2 (also known as Cldn2), DefA, F4/80 (also known as Adgre1), Il1b, Il6, Il10, Ki67 (also known as Mki67), Lyz1, Mcp1 (also known as Ccl2), Muc2, Ocln, Pla2g2a, Reg3g, Tcf4, Tnfα (also known as Tnf) and Zo1 (also known as Tjp1). Data were analysed using the 2-ΔΔCT method and expression was normalised to Rpl19 (see ESM Methods and ESM Table 2 for further details).
Short-chain fatty acids
Caecal content was diluted 1:6 in LAL reagent water (Lonza, Walkersville, MD, USA) and homogenised with a tissue lyser (Qiagen) for 4 min without beads to avoid bacteria disruption. Samples were centrifuged (8000 g, 2 min). Cell-free supernatants were used for the quantification of acetate, butyrate and propionate as previously described [30] and for the analyses of TLR agonists as described below.
TLR-2 and TLR-4 agonists
TLR-2 and TLR-4 agonists were measured using HEK-Blue reporter cell lines according to the manufacturer’s instructions (InvivoGen, San Diego, CA, USA). Cells were exposed to the supernatants of caecal content and the protocol previously described was followed [31].
Histological analysis of the intestine
Crypt depth and villus length were measured after H&E staining. Sections were digitised (Leica SCN400; Leica, Wetzlar, Germany) and images were captured using Leica Image Viewer software, version 4.0.4. At least ten measurements per mouse were made.
Precision-cut liver slices
After 1 week of acclimatisation, mice in experiment 2 were anesthetised with ketamine (100 mg/kg body weight) (Nimatek; Eurovet, Bladel, Netherlands) and xylazine (100 mg/kg body weight) (Rompun; Bayer, Leverkusen, Germany). Precision-cut liver slices (PCLS) were prepared as previously described [32, 33]. PCLS were incubated without (control medium) or with vildagliptin (0.6 mg/ml) in oxygenated and supplemented Waymouth’s medium for 4 h at 37°C under agitation. PCLS were frozen in dry ice until analysed.
Bacterial growth conditions
Growth curves of Oscillibacter valericigenes DSM18026 and L. reuteri 100-23 in the presence of vildagliptin (0.6 mg/ml) or absence of vildagliptin (control broth) were determined (ESM Methods).
To measure DPP-4 activity 5 ml of overnight cultures were harvested and processed as described below.
DPP-4 activity
DPP-4 activity was measured by the cleavage of para-nitroanilide (PNA) from Gly-Pro-PNA (Sigma, St. Louis, MO, USA). Samples (20–50 mg) were suspended in TRIS-base buffer (50 mmol/l, 1% N-octylglucoside, pH 8.3), homogenised for 2 min with a tissue lyser and centrifuged (3000 g, 20 min). The supernatant (20 μl) was incubated with Gly-Pro-PNA. The enzymatic activity was measured in a kinetic analysis of 30 min at 37°C (380 nm) (SpectraMax M2; Molecular Devices, San Jose, CA, USA) and quantified with a standard curve of free PNA. In tissues and bacterial samples, the values were normalised by the amount of protein (Bradford method).
Statistical analyses
The number of mice per group was based on previous experiments to measure the primary outcomes (the WD increased the body weight) with the minimum number of animals [34]. Analyses were performed by one-way ANOVA followed by post hoc Tukey’s tests (GraphPad Prism, version 5; GraphPad, La Jolla, CA, USA). For the growth curves, two-way ANOVA followed by Bonferroni’s post hoc test was performed. A χ2 test was used for categorical data (active GLP-1 and TLR-4 agonist) (Statgraphics Plus, version 5.1; Statgraphics, The Plains, VA, USA). Data of microbiota composition were analysed using Welch’s t test and the false discovery rate was applied for p value correction (q value) according to the Benjamini–Hochberg procedure. Ecological descriptors and data from the PCLS experiment were analysed using Welch’s t test. Multiple correlation analyses were performed in R version 3.1 (www.R-project.org), with adjustment of p values according to the Benjamini–Hochberg procedure Results were considered statistically significant at p < 0.05. Any exclusion decision was supported by Grubbs’ test. Plots were generated using GraphPad showing the mean ± SEM. Each dot represents one biological replicate.
Results
Vildagliptin decreased systemic and intestinal DPP-4 activity
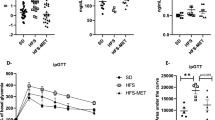
Vildagliptin reduced DPP-4 activity in the portal vein and increased the concentration of active GLP-1 (Fig. 1a, b). In addition, vildagliptin significantly reduced DPP-4 activity in the ileum, the caecum and the colon (Fig. 1c–e).
(a) DPP-4 activity in the portal vein, (b) concentration of active GLP-1 in the portal vein and (c) DPP-4 activity in the ileum, (d) caecum and (e) colon. Mice were fed a control diet, a WD or a WD + vildagliptin (WD+V). Significant differences between conditions are shown as *p < 0.05, **p < 0.01 and ***p < 0.001 according to one-way ANOVA followed by Tukey’s post hoc test
Organ weights and food intake are presented in ESM Table 3. WD-fed mice showed a significant increase in body weight and adiposity. Body weight evolution, glycaemia, insulin and GIP are shown in ESM Fig. 2. Neither WD nor vildagliptin modified the markers related to glucose homeostasis.
Vildagliptin abolished DPP-4 activity in faeces and caecal content
DPP-4 activity in the faeces did not differ between groups before the introduction of the treatments. The administration of vildagliptin for 2 weeks completely abolished DPP-4 activity in the faeces (Fig. 2a). The effect persisted after the introduction of the WD, as shown by the measurement of DPP-4 activity in the caecal content at the end of the experiment (Fig. 2b).
DPP-4 activity (a) in the faeces before and after administration of vildagliptin for 2 weeks (14 days) and (b) in the caecal content (8 weeks). Mice were fed a control diet, a WD or a WD + vildagliptin (WD+V). Significant differences between conditions are shown as **p < 0.01 and ***p < 0.001 according to one-way ANOVA followed by Tukey’s post hoc test
Vildagliptin modified gut microbiota composition and activity in vivo and inhibited growth of O. valericigenes in vitro, independently of DPP-4 activity
Vildagliptin increased the weight of the caecal content and caecal tissue compared with its control group (the WD-fed mice) (Fig. 3a, b). To assess whether these observations were linked to bacterial fermentation, we quantified the production of short-chain fatty acids (SCFA) in the caecal content. Vildagliptin significantly increased the amount of propionate compared with the WD group (Fig. 3e). There were no differences in total amount of SCFA, acetate or butyrate (Fig. 3c, d, f).
Weight of (a) caecal tissue and (b) caecal content. (c–f) Amount of SCFA in the caecal content (CC): (c) total SCFA, (d) acetate, (e) propionate and (f) butyrate. Mice were fed a control diet, a WD or a WD + vildagliptin (WD+V). Significant differences between conditions are shown as *p < 0.05 and **p < 0.01 according to one-way ANOVA followed by Tukey’s post hoc test
We sequenced the 16S rRNA gene of the two WD-fed groups with or without addition of vildagliptin. Principal coordinate analysis (PCoA) of beta diversity based on Morisita–Horn distance showed that vildagliptin accounted for 38% of the variation in the dataset (Adonis method, 1000 permutations; p < 0.001). The majority of the variation explained by the treatment was further confirmed by PCoA using binary Jaccard and Bray–Curtis indexes (14% and 26%, respectively; p < 0.001) (Fig. 4a–c). No differences were observed in any of the six indexes of alpha diversity analysed (ESM Fig. 3). A total of 293 operational taxonomic units (OTUs) were identified, of which 67 differed significantly between the two groups at the p < 0.05 threshold (ESM Table 4). When the false discovery rate was applied for p value correction, only two OTUs corresponding to the genus Oscillibacter spp. (OTU 4, p < 0.001, q = 0.016) (Fig. 4d) and unclassified Ruminococcaceae (OTU 241, p < 0.001, q = 0.016) (Fig. 4e) remained significantly reduced by vildagliptin.
Principal coordinate (PCo) plots of beta diversity based on (a) Morisita–Horn, (b) binary Jaccard and (c) Bray–Curtis indexes. (d, e) Relative (Rel.) abundance of OTUs modified by vildagliptin assessed by Illumina sequencing in DNA extracted from caecal content. Mice were fed a WD (black symbols) or a WD + vildagliptin (WD+V; grey symbols). Data were analysed using Welch’s t test and p values were corrected using the Benjamini–Hochberg method. Significant differences are shown as *q < 0.05
Furthermore, some specific bacteria were quantified using qPCR to confirm the reductions in Oscillibacter spp. and to quantify the bacteria initially hypothesised to be affected by DPP-4 inhibition. This included bacteria with DPP-4-like activity (Prevotella and Lactobacillus) [21,22,23] and those that produce DPP-4 inhibitors (Bifidobacterium and Lactobacillus) [24, 25]. Vildagliptin caused a significant reduction in Oscillibacter/Oscillospira and an increase in Lactobacillus spp. (Fig. 5). The latter observation is consistent with the sequencing data of L. johnsonii that tends to increase with vildagliptin (OTU 177, p = 0.016, q = 0.155) (ESM Table 4).
Gut microbiota composition analysed by qPCR in DNA extracted from caecal content. (a) Total bacteria, (b) Oscillibacter/Oscillospira, (c) Lactobacillus spp., (d) Bifidobacterium spp. and (e) Bacteroides/Prevotella. Mice were fed a WD or a WD + vildagliptin (WD+V). Significant differences are shown as *p < 0.05 according to Welch’s t test
To study whether vildagliptin explained these changes, we exposed O. valericigenes to vildagliptin in vitro and observed that the drug inhibited its growth (Fig. 6a). In parallel, we performed the same test with L. reuteri, which is a carrier of DPP-4-like activity in the gut microbiota. L. reuteri grew similarly in both conditions (Fig. 6b). No DPP-4 activity was detected in the cell-free extracts of O. valericigenes. Therefore, the effect of vildagliptin on bacterial growth is not truly dependent on the inhibition of bacterial DPP-4 activity (ESM Fig. 4).
Vildagliptin reduced TLR ligands in caecal content and restored expression of antimicrobial peptides and crypt depth in ileum
The caecal extracts of mice treated with vildagliptin presented lower TLR-2 and TLR-4 ligands compared with those of WD-fed mice (Fig. 7a, b).
Ligands of (a) TLR-2 and (b) TLR-4 in caecal content (CC). Mice were fed a control diet, a WD or a WD + vildagliptin (WD+V). Data for TLR-2 were analysed using one-way ANOVA followed by Tukey’s post hoc test. Categorical data (positive/total) for TLR-4 were analysed using the χ2 test. Significant differences between conditions are shown as *p < 0.05 and **p < 0.01. FSL-1, fibroblast-stimulating lipopeptide; LPS, lipopolysaccharide
In the ileum, the expression of antimicrobial peptides (AMPs) such as C-type lectins (Reg3g), phospholipases (Pla2g2a) and α-defensin (DefA) were downregulated in WD-fed mice (Fig. 8a). Vildagliptin counteracted the WD-induced downregulation of Reg3g, Pla2g2a and DefA. We studied whether vildagliptin could also have impacted the morphology and some proliferation markers associated with Paneth cells. Vildagliptin significantly restored crypt depth (Fig. 8b) without changing villus length (Fig. 8c). The histological images are shown in ESM Fig. 5. No differences were observed in Ki67 or Tcf4 expression (Fig. 8d, e).
(a) Gene expression of AMPs Reg3g, Pla2g2a, DefA and Lyz1. (b) Crypt depth, (c) villus length, (d) proliferation marker Ki67 and (e) Paneth cell differentiation marker Tcf4 in the ileum. Mice were fed a control diet, a WD or a WD + vildagliptin (WD+V). Significant differences between conditions are shown as *p < 0.05 and **p < 0.01 according to one-way ANOVA followed by Tukey’s post hoc test
No major changes due to vildagliptin were observed in the expression of markers related to gut barrier function or cytokines; vildagliptin only increased Claudin2 and Mcp1 (ESM Fig. 6). In the colon, we analysed some variables previously shown to be affected by vildagliptin in the ileum. Only a significant increase in crypt depth was observed, indicating that the major changes induced by vildagliptin occur in the ileum (ESM Fig. 7).
Finally, we performed a Spearman correlation analysis to evaluate the associations between gut microbes and host variables (ESM Fig. 8). We observed positive correlations between TLR-4 agonists and three OTUs (OTU 4 [Oscillibacter spp.], OTU 241 [unclassified Ruminococcaceae] and OTU 40 [Catabacter spp.]) and negative correlations between TLR-4 agonists and two OTUs (OTU 117 [unclassified Lachnospiraceae] and OTU 70 [Parabacteroides goldsteinii]). Also a positive association was found between Reg3g and OTU 16 (unclassified Porphyromonadaceae).
Vildagliptin indirectly reduced gene expression of proinflammatory cytokines in liver
Vildagliptin reduced DPP-4 activity in the portal vein (Fig. 1a), suggesting that the liver is exposed to an efficient dose. Consistent with this, we show that vildagliptin decreased DPP-4 activity in the liver (ESM Fig. 9). Vildagliptin was also associated with a significant reduction in the expression of Cd3g, Cd11c and the proinflammatory markers Mcp1, Tnfα, Il1b and Il6 (Fig. 9a, b). To assess whether the changes responded to a direct effect, we performed an ex vivo experiment in which PCLS were exposed to vildagliptin. In contrast to our observations in vivo, no changes in the expression of any cytokines were observed (Fig. 9c).
In an in vivo experiment, mice were fed a control diet, a WD or a WD + vildagliptin (WD+V). In an ex vivo experiment, PCLS were exposed to a control medium or medium with vildagliptin. Gene expression of (a) biomarkers of immune cell populations, and (b) cytokines in the liver in the in vivo experiment and (c) cytokines in PCLS in the ex vivo experiment. In (a, b), significant differences between conditions are shown as *p < 0.05 and **p < 0.01 according to one-way ANOVA followed by Tukey’s post hoc test. In (c), data were analysed with Welch’s t test
Discussion
Inhibition of DPP-4 activity to tackle metabolic disorders through the preservation of incretins has been widely described. However, the additional effects of DPP-4 inhibitors on the intestinal ecosystem remain poorly studied. In the present study, we evaluated whether the DPP-4 inhibitor vildagliptin could impact intestinal homeostasis—including the composition and activity of gut microbiota—using in vivo, ex vivo and in vitro approaches. Our work is the first to consider the DPP-4-like activity of the gut microbiota as a potential target of DPP-4 inhibitors and evaluate, in an animal model of diet-induced obesity, the effect of vildagliptin on gut barrier function, innate immune response and liver. Overall, our results show that vildagliptin improved gastrointestinal function judged by restoration of the expression of AMPs and the depth of the crypts in the ileum. In the liver, vildagliptin was associated with a reduction of cytokine expression. However, the ex vivo experiment with PCLS showed that reductions were not due to a direct effect of the drug, reinforcing the idea that reduced hepatic inflammatory tone might be related to changes at the intestinal level. Furthermore, vildagliptin was associated with changes in beta diversity indexes, reductions in Oscillibacter spp. and increases in Lactobacillus spp. and caecal propionate levels. Among the 67 OTUs changed by vildagliptin, we can pinpoint some unclassified Lachnospiraceae and Ruminococcaceae, as well as P. goldsteinii, as potential contributors to the increase in propionate levels [35], even if no statistically significant correlation backed up this fact. DPP-4 is a multifunctional protein and therefore it is plausible that its inhibition by vildagliptin has induced the broad changes that we observed through several mechanisms. In this regard, we demonstrated three effects of vildagliptin on the intestine, namely: (1) restoration of the active form of gut peptides such as GLP-1; (2) modulation of the innate immune response; and (3) modulation of the gut microbiota. There could be a link between these effects, as illustrated for instance in a study performed in obese prebiotic-fed mice which reported that the gut microbiota influenced the release of substrates of DPP-4 such as GLP-1 and GLP-2 [36]. Furthermore, a high-fat diet can impair enteroendocrine cell function and the release of GLP-1, as confirmed here [37]. We propose that the increase in active GLP-1 in mice treated with vildagliptin can also be partly explained by the increases in propionate in the caecal content [38]. The commutative relationship between microbiota and host is also supported by the fact that AMPs shape the composition of the gut microbiota, and, conversely, the gut microbiota influences host immunity [39].
Since we know that some bacteria exert DPP-4-like activity, one of our outcome measures was to assess whether vildagliptin targets DPP-4 activity of the gut microbiota [21,22,23]. We have shown for the first time that vildagliptin completely abolished DPP-4 activity in the caecal content and faeces. Analyses of the composition of the gut microbiota showed that Oscillibacter spp. was the most striking target of vildagliptin, but its reduction was not explained by DPP-4 inhibition. Only one study has considered the impact of the DPP-4 inhibitors saxagliptin, vildagliptin and sitagliptin on bacterial DPP-4-like activity [40]. Inhibition of the DPP-4-like activity of Streptococcus mutans found in the oral microbiota impaired its biofilm formation [40]. None of the DPP-4 inhibitors tested affected the growth of S. mutans in the concentration range considered (4–2048 μg/ml) [40]. In our study, however, vildagliptin (600 μg/ml) inhibited the growth of O. valericigenes, even though this bacterium does not exhibit DPP-4 activity. Taken together these results demonstrate the impact of vildagliptin on different members of the microbiota (intestinal and oral) via inhibition of DPP-4-like activity and through a mechanism still unknown. A recent study reported changes in the composition of the gut microbiota by vildagliptin in Sprague–Dawley rat models of diabetes [26]. In agreement with our results, vildagliptin caused a reduction in Oscillibacter that was interpreted as a beneficial effect [26]. Furthermore, other animal studies have associated increases in Oscillibacter with obese and diabetic phenotypes [41,42,43], and with increases in intestinal permeability, a condition involved in the development of metabolic disorders [41]. Consequently, vildagliptin induces beneficial changes in the composition of the intestinal microbiota that might become a therapeutic strategy beyond the preservation of incretins.
The gut is populated with multiple types of immune and epithelial cells that work together to maintain tolerance to the gut microbiota and food [17]. The interaction between microbiota and host cells is mediated by TLRs, which are sensors involved in the innate immune response. In addition, a chemical barrier consisting of the secretions of AMPs and other cell products (cytokines and immunoglobulins) is part of the first line of defence [44]. Here, on the one hand, we have shown that vildagliptin reduces levels of TLR-2 and TLR-4 agonists in the caecal content. Previously, it has been reported that vildagliptin suppresses TLR-2 and TLR-4 content in macrophages stimulated with soluble DPP-4 and lipopolysaccharide [13]. This observation is in agreement with the reduction in TLR-4 agonist activity (such as lipopolysaccharide) in the caecal content of mice treated with vildagliptin and demonstrates that vildagliptin impacts the crosstalk between the gut microbiota and the host. On the other hand, vildagliptin also influenced the innate immune response via restoration of AMP expression in the ileum. In agreement with previous studies carried out by our group, a fat-enriched diet caused a reduction in AMPs [34, 45]. Vildagliptin completely counteracted this reduction. We hypothesise that this effect might be linked to GLP-2. As GLP-2 is inactivated by DPP-4, vildagliptin could have preserved its functionality [10, 11]. The role of GLP-2 in inducing antimicrobial products has been shown in GLP-2 receptor knockout mice which had reduced AMPs and impaired mucosal bactericidal activity [46]. Also, the restoration of crypt depth in mice that received vildagliptin could be explained, at least in part, by GLP-2 stimulation of crypt cell proliferation and inhibition of apoptosis [47]. Unfortunately, we could not confirm experimentally this hypothesis due to the lack of commercially available methods to measure active GLP-2.
Finally, we investigated whether the changes observed at the intestinal level could also have impacted the inflammatory tone in the liver. Mice treated with vildagliptin showed a reduction in the expression of lymphocytes (Cd3g), macrophage activation (Cd11c) and proinflammatory markers. The ex vivo experiment in which PCLS were exposed to vildagliptin showed no changes in any of the inflammatory markers analysed. Even if there is a bias between the in vivo and ex vivo approaches, our observations indicate that the changes induced by vildagliptin at the intestinal level might have also impacted hepatic homeostasis. Specifically, the reinforcement of gut immunity induced by vildagliptin could result in a lower exposure of the liver to microbial stimuli. It would be useful to evaluate how vildagliptin influences gut microbial activity and the gut–liver axis, since microbial metabolites that are prone to reach the liver through the portal vein, such as bile acids (via the farnesoid X receptor) or SCFA (like butyrate), modulate hepatic inflammation and thereby the occurrence of non-alcoholic fatty liver disease [48, 49].
For our study, we selected vildagliptin for its potential effect on inflammation (reduction of TLR expression) [14]. The pharmacokinetics of vildagliptin is different from that of the other DPP-4 inhibitors (being more lipophilic). For instance, the percentage of vildagliptin excreted in the faeces is 4.5%, whereas for sitagliptin and saxagliptin it is reported to be 13% and 22%, respectively [50]. Therefore, the effects of other gliptins on the microbiota—and related metabolic effects—could be relevant, since they are found in higher quantities in the lower intestine. Human data may confirm this hypothesis.
In conclusion, we show that in WD-fed mice presenting gut disorders, vildagliptin affected the composition of the gut microbiota and improved intestinal homeostasis. Due to the multifunctional roles of DPP-4, further studies are needed to decipher the precise molecular mechanisms. If the beneficial effect of vildagliptin on the intestinal tract is confirmed in human studies, it might represent a novel mechanism of vildagliptin to improve human health beyond its standard use.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. The sequences used for analysis can be found in the MG-RAST database under the project name MYNEWGUT3.
Abbreviations
- AMP:
-
Antimicrobial peptide
- DPP-4:
-
Dipeptidyl peptidase 4
- GIP:
-
Glucose-dependent insulinotropic peptide
- GLP:
-
Glucagon-like peptide
- OTU:
-
Operational taxonomic unit
- PCLS:
-
Precision-cut liver slices
- PCoA:
-
Principal coordinate analysis
- PNA:
-
Para-nitroanilide
- qPCR:
-
Quantitative real-time PCR
- SCFA:
-
Short-chain fatty acids
- TLR:
-
Toll-like receptor
- WD:
-
Western diet
References
Seino Y, Fukushima M, Yabe D (2010) GIP and GLP-1, the two incretin hormones: similarities and differences. J Diabetes Investig 1:8–23
Mulvihill EE, Drucker DJ (2014) Pharmacology, physiology, and mechanisms of action of dipeptidyl peptidase-4 inhibitors. Endocr Rev 35:992–1019
Lee SA, Kim YR, Yang EJ et al (2013) CD26/DPP4 levels in peripheral blood and T cells in patients with type 2 diabetes mellitus. J Clin Endocrinol Metab 98:2553–2561
Stengel A, Goebel-Stengel M, Teuffel P et al (2014) Obese patients have higher circulating protein levels of dipeptidyl peptidase IV. Peptides 61:75–82
Sell H, Bluher M, Kloting N et al (2013) Adipose dipeptidyl peptidase-4 and obesity: correlation with insulin resistance and depot-specific release from adipose tissue in vivo and in vitro. Diabetes Care 36:4083–4090
Kushwaha RN, Haq W, Katti SB (2014) Sixteen-years of clinically relevant dipeptidyl peptidase-IV (DPP-IV) inhibitors for treatment of type-2 diabetes: a perspective. Curr Med Chem 21:4013–4045
Waget A, Cabou C, Masseboeuf M et al (2011) Physiological and pharmacological mechanisms through which the DPP-4 inhibitor sitagliptin regulates glycemia in mice. Endocrinology 152:3018–3029
Mulvihill EE, Varin EM, Gladanac B et al (2017) Cellular sites and mechanisms linking reduction of dipeptidyl peptidase-4 activity to control of incretin hormone action and glucose homeostasis. Cell Metab 25:152–165
Mortier A, Gouwy M, Van Damme J, Proost P, Struyf S (2016) CD26/dipeptidylpeptidase IV-chemokine interactions: double-edged regulation of inflammation and tumor biology. J Leukoc Biol 99:955–969
Yazbeck R, Howarth GS, Abbott CA (2009) Dipeptidyl peptidase inhibitors, an emerging drug class for inflammatory disease? Trends Pharmacol Sci 30:600–607
Hartmann B, Thulesen J, Kissow H et al (2000) Dipeptidyl peptidase IV inhibition enhances the intestinotrophic effect of glucagon-like peptide-2 in rats and mice. Endocrinology 141:4013–4020
Wronkowitz N, Gorgens SW, Romacho T et al (2014) Soluble DPP4 induces inflammation and proliferation of human smooth muscle cells via protease-activated receptor 2. Biochim Biophys Acta 1842:1613–1621
Lee DS, Lee ES, Alam MM et al (2016) Soluble DPP-4 up-regulates toll-like receptors and augments inflammatory reactions, which are ameliorated by vildagliptin or mannose-6-phosphate. Metabolism 65:89–101
Makdissi A, Ghanim H, Vora M et al (2012) Sitagliptin exerts an antinflammatory action. J Clin Endocrinol Metab 97:3333–3341
Ta NN, Li Y, Schuyler CA, Lopes-Virella MF, Huang Y (2010) DPP-4 (CD26) inhibitor alogliptin inhibits TLR4-mediated ERK activation and ERK-dependent MMP-1 expression by U937 histiocytes. Atherosclerosis 213:429–435
Ervinna N, Mita T, Yasunari E et al (2013) Anagliptin, a DPP-4 inhibitor, suppresses proliferation of vascular smooth muscles and monocyte inflammatory reaction and attenuates atherosclerosis in male apo E-deficient mice. Endocrinology 154:1260–1270
Santaolalla R, Abreu MT (2012) Innate immunity in the small intestine. Curr Opin Gastroenterol 28:124–129
He YL (2012) Clinical pharmacokinetics and pharmacodynamics of vildagliptin. Clin Pharmacokinet 51:147–162
Delzenne NM, Neyrinck AM, Backhed F, Cani PD (2011) Targeting gut microbiota in obesity: effects of prebiotics and probiotics. Nat Rev Endocrinol 7:639–646
Shin NR, Lee JC, Lee HY et al (2014) An increase in the Akkermansia spp. population induced by metformin treatment improves glucose homeostasis in diet-induced obese mice. Gut 63:727–735
Walker ND, McEwan NR, Wallace RJ (2003) Cloning and functional expression of dipeptidyl peptidase IV from the ruminal bacterium Prevotella albensis M384(T). Microbiology 149:2227–2234
Stressler T, Eisele T, Schlayer M, Lutz-Wahl S, Fischer L (2013) Characterization of the recombinant exopeptidases PepX and PepN from Lactobacillus helveticus ATCC 12046 important for food protein hydrolysis. PLoS One 8:e70055
Sanz Y, Toldra F (2001) Purification and characterization of an X-prolyl-dipeptidyl peptidase from Lactobacillus sakei. Appl Environ Microbiol 67:1815–1820
Zeng Z, Luo JY, Zuo FL et al (2016) Bifidobacteria possess inhibitory activity against dipeptidyl peptidase-IV. Lett Appl Microbiol 62:250–255
Zhu Zeng JL, Zuo F, Chen S Screening for potential novel probiotic Lactobacillus strains based on high dipeptidyl peptidase IV and α-glucosidase inhibitory activity. J Funct Foods 20:486–495
Zhang Q, Xiao X, Li M et al (2017) Vildagliptin increases butyrate-producing bacteria in the gut of diabetic rats. PLoS One 12:e0184735
Grasset E, Puel A, Charpentier J et al (2017) A specific gut microbiota dysbiosis of type 2 diabetic mice induces GLP-1 resistance through an enteric NO-dependent and gut–brain axis mechanism. Cell Metab 25:e1075
Bindels LB, Neyrinck AM, Claus SP et al (2016) Synbiotic approach restores intestinal homeostasis and prolongs survival in leukaemic mice with cachexia. ISME J 10:1456–1470
Bindels LB, Segura Munoz RR, Gomes-Neto JC et al (2017) Resistant starch can improve insulin sensitivity independently of the gut microbiota. Microbiome 5:12
Salazar N, Dewulf EM, Neyrinck AM et al (2015) Inulin-type fructans modulate intestinal Bifidobacterium species populations and decrease fecal short-chain fatty acids in obese women. Clin Nutr 34:501–507
Neyrinck AM, Taminiau B, Walgrave H et al (2017) Spirulina protects against hepatic inflammation in aging: an effect related to the modulation of the gut microbiota? Nutrients 9
Neyrinck AM, Alexiou H, Delzenne NM (2004) Kupffer cell activity is involved in the hepatoprotective effect of dietary oligofructose in rats with endotoxic shock. J Nutr 134:1124–1129
Neyrinck AM, Gomez C, Delzenne NM (2004) Precision-cut liver slices in culture as a tool to assess the physiological involvement of Kupffer cells in hepatic metabolism. Comp Hepatol 3(Suppl 1):S45
Suriano F, Bindels LB, Verspreet J et al (2017) Fat binding capacity and modulation of the gut microbiota both determine the effect of wheat bran fractions on adiposity. Sci Rep 7:5621
Reichardt N, Duncan SH, Young P et al (2014) Phylogenetic distribution of three pathways for propionate production within the human gut microbiota. ISME J 8:1323–1335
Cani PD, Possemiers S, Van de Wiele T et al (2009) Changes in gut microbiota control inflammation in obese mice through a mechanism involving GLP-2-driven improvement of gut permeability. Gut 58:1091–1103
Richards P, Pais R, Habib AM et al (2016) High fat diet impairs the function of glucagon-like peptide-1 producing L-cells. Peptides 77:21–27
Psichas A, Sleeth ML, Murphy KG et al (2015) The short chain fatty acid propionate stimulates GLP-1 and PYY secretion via free fatty acid receptor 2 in rodents. Int J Obes 39:424–429
Vaishnava S, Behrendt CL, Ismail AS, Eckmann L, Hooper LV (2008) Paneth cells directly sense gut commensals and maintain homeostasis at the intestinal host–microbial interface. Proc Natl Acad Sci U S A 105:20858–20863
De A, Pompilio A, Francis J et al (2018) Antidiabetic drugs ‘gliptins’ affect biofilm formation by Streptococcus mutans. Microbiol Res 209:79–85
Lam YY, Ha CW, Campbell CR et al (2012) Increased gut permeability and microbiota change associate with mesenteric fat inflammation and metabolic dysfunction in diet-induced obese mice. PLoS One 7:e34233
Moya-Perez A, Neef A, Sanz Y (2015) Bifidobacterium pseudocatenulatum CECT 7765 reduces obesity-associated inflammation by restoring the lymphocyte–macrophage balance and gut microbiota structure in high-fat diet-fed mice. PLoS One 10:e0126976
Geurts L, Lazarevic V, Derrien M et al (2011) Altered gut microbiota and endocannabinoid system tone in obese and diabetic leptin-resistant mice: impact on apelin regulation in adipose tissue. Front Microbiol 2:149
Scaldaferri F, Pizzoferrato M, Gerardi V, Lopetuso L, Gasbarrini A (2012) The gut barrier: new acquisitions and therapeutic approaches. J Clin Gastroenterol 46(Suppl):S12–S17
Everard A, Lazarevic V, Gaia N et al (2014) Microbiome of prebiotic-treated mice reveals novel targets involved in host response during obesity. ISME J 8:2116–2130
Lee SJ, Lee J, Li KK et al (2012) Disruption of the murine Glp2r impairs Paneth cell function and increases susceptibility to small bowel enteritis. Endocrinology 153:1141–1151
Drucker DJ, Erlich P, Asa SL, Brubaker PL (1996) Induction of intestinal epithelial proliferation by glucagon-like peptide 2. Proc Natl Acad Sci U S A 93:7911–7916
Briand F, Brousseau E, Quinsat M, Burcelin R, Sulpice T (2018) Obeticholic acid raises LDL-cholesterol and reduces HDL-cholesterol in the diet-induced NASH (DIN) hamster model. Eur J Pharmacol 818:449–456
Sheng L, Jena PK, Hu Y et al (2017) Hepatic inflammation caused by dysregulated bile acid synthesis is reversible by butyrate supplementation. J Pathol 243:431–441
Scheen AJ (2010) Pharmacokinetics of dipeptidylpeptidase-4 inhibitors. Diabetes Obes Metab 12:648–658
Acknowledgements
We thank V. Allaeys, I. Blave, R. Selleslagh and B. Es Saad (Louvain Drug Research Institute, Brussels, Belgium) for their skilful technical assistance. We thank B. Pot (Vrije Universiteit Brussel, Brussels, Belgium) for supplying the strain of L. reuteri 100-23.
Funding
This work was supported by EU grant 613979 (MyNewGut). MO is a beneficiary of a MOVE-IN Louvain Incoming Post-Doctoral Fellowship co-funded by the Marie Curie Actions of the European Commission. NS benefits from a Juan de la Cierva post-doctoral contract granted by the Spanish Ministry of Economy, Industry and Competitiveness (MINECO). PDC is a senior research associate of the Fonds de la Recherche Scientifique (FRS-FNRS) and is supported by FRFS-WELBIO through grant WELBIO-CGR-2017, the Fonds Baillet-Latour (Grant for Medical Research 2015) and European Research Council (ERC) Starting Grant 2013 (grant 336452-ENIGMO). LBB is the recipient of FSR subsidies (Fonds Spécial de la Recherche, Université catholique de Louvain).
Author information
Authors and Affiliations
Contributions
MO, AMN and NMD conceived and designed the experiments. MO and AMN performed the animal experiments. MB performed the ex vivo experiment. MO, AMN, SAP, MB, NS, PDC and LBB were responsible for the acquisition of data, or analysis and interpretation of data. LBB performed the correlation analyses. MO and NMD wrote the paper. AMN, SAP, MB, NS, PDC and LBB revised the article and provided important intellectual content. NMD planned and supervised all experiments and manuscript preparation. All authors read the manuscript and approved the version to be published. NMD is the guarantor of this work.
Corresponding author
Ethics declarations
The authors declare that there is no duality of interest associated with this manuscript.
Electronic supplementary material
ESM
(PDF 775 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Olivares, M., Neyrinck, A.M., Pötgens, S.A. et al. The DPP-4 inhibitor vildagliptin impacts the gut microbiota and prevents disruption of intestinal homeostasis induced by a Western diet in mice. Diabetologia 61, 1838–1848 (2018). https://doi.org/10.1007/s00125-018-4647-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-018-4647-6