Abstract
Aims/hypothesis
Little is known about the heritable basis of gene–environment interactions in humans. We therefore screened multiple cardiometabolic traits to assess the probability that they are influenced by genotype–environment interactions.
Methods
Fourteen established environmental risk exposures and 11 cardiometabolic traits were analysed in the VIKING study, a cohort of 16,430 Swedish adults from 1682 extended pedigrees with available detailed genealogical, phenotypic and demographic information, using a maximum likelihood variance decomposition method in Sequential Oligogenic Linkage Analysis Routines software.
Results
All cardiometabolic traits had statistically significant heritability estimates, with narrow-sense heritabilities (h 2) ranging from 24% to 47%. Genotype–environment interactions were detected for age and sex (for the majority of traits), physical activity (for triacylglycerols, 2 h glucose and diastolic BP), smoking (for weight), alcohol intake (for weight, BMI and 2 h glucose) and diet pattern (for weight, BMI, glycaemic traits and systolic BP). Genotype–age interactions for weight and systolic BP, genotype–sex interactions for BMI and triacylglycerols and genotype–alcohol intake interactions for weight remained significant after multiple test correction.
Conclusions/interpretation
Age, sex and alcohol intake are likely to be major modifiers of genetic effects for a range of cardiometabolic traits. This information may prove valuable for studies that seek to identify specific loci that modify the effects of lifestyle in cardiometabolic disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cardiometabolic diseases are the predominant cause of mortality, morbidity and healthcare spending globally [1, 2], and are believed to result in part from the combined additive and synergistic effects of genetic and environmental risk factors. Environmental exposures such as diet and physical activity have enormous potential for prevention and treatment of these diseases, but no single therapy works well in all individuals. Determining whether susceptibility to adverse environmental exposures is genetically determined (i.e. gene–environment interactions [3]) and elucidating the specific nature of these interactions may facilitate the stratification of patient populations into subgroups that can be treated with optimal therapies.
In contemporary population genetics research, the heritability of a given trait is usually assessed by quantitative genetics approaches to make inferences about the extent to which polygenic variation influences the trait. Assessing heritability is usually done prior to embarking on studies that seek to discover specific loci influencing the trait. While it is equally logical to use quantitative genetics to determine whether traits are influenced by genotype–environment interactions as a prelude to studies focused on specific environmental exposures and genetic loci, this is rarely done in practice [4–8]. The dearth of such studies may be because large, well-characterised cohorts including genealogies, which are necessary for genotype–environment quantitative genetic studies, are rare.
Here we sought to screen for genotype–environment interactions across a number of environmental exposures and cardiometabolic traits using quantitative genetic analyses in extended pedigrees. Accordingly, we characterised the genealogical structure of a large northern Swedish population, within which detailed measures of environmental exposures, cardiometabolic traits and other personal characteristics exist [9].
Methods
Study participants
The Västerbotten Imputation Databank of Near-Complete Genomes (VIKING) study is nested in a population-based cohort from the county of Västerbotten in northern Sweden. The study capitalises on the extensively mapped genealogies in this low admixture population, in combination with an ongoing health survey in the population that makes available extensive phenotypic data in the cohort [9]. The genealogical information stems from the POPLINK database at the Demographic Database/Centre for Demographic and Ageing Research (CEDAR) at Umeå University, Umeå, Sweden. Data are based on detailed Swedish population registers, covering the period 1700–1950, linked to population data from Statistics Sweden from 1950 to the present day. Lifestyle and clinical data were collected within the framework of the Västerbottens Health Survey (also called the Västerbottens Intervention Project) initiated in 1985 [10]. In the Västerbottens Health Survey, residents within the county are invited to attend an extensive health examination in the years of their 40th, 50th and 60th birthdays. For the current analysis, health examinations were performed between 1985 and 2013. All participants provided written informed consent as part of the Västerbottens Health Survey, and the study was approved by the regional ethics review board in Umeå, Sweden.
The current study includes 1682 extended pedigrees comprising 193,060 people of whom 16,430 have detailed phenotype data. The most extended genealogy descends from 4255 founders and contains 160,533 people of whom 10,498 are phenotyped. The phenotyped sample includes 8908 first-degree relative pairs, 5794 second-degree relative pairs and 29,706 third-degree relative pairs, in addition to other more distant relatives (electronic supplementary material [ESM] Table 1).
Cardiometabolic traits
The assessment of clinical measures in the Västerbottens Health Survey has been described in detail elsewhere [10, 11]. Briefly, weight (to the nearest 0.1 kg) and height (to the nearest 1 cm) were measured with a calibrated balance-beam scale and a wall-mounted stadiometer, respectively, and with participants wearing indoor clothing and without shoes. BMI was calculated as weight (kg)/height (m)2. In a subgroup, waist circumference was measured using a non-stretchable nylon tape at the midpoint between the 12th rib and the iliac crest. Systolic and diastolic BPs (SBP and DBP) were measured once using a mercury sphygmomanometer following a 5 min rest. Capillary blood was drawn after an overnight fast and again 2 h after administration of a standard 75 g oral glucose load [12]. Before the first blood draw, 83% of participants had fasted for a minimum of 8 h. Capillary plasma glucose concentrations, total cholesterol and triacylglycerols were measured with a Reflotron bench-top analyser (Roche Diagnostics Scandinavia, Umeå, Sweden). HDL-cholesterol (HDL-C) was measured in a subgroup of participants. LDL-cholesterol (LDL-C) was calculated by applying the Friedewald formula: LDL-C=total cholesterol−HDL-C−(triacylglycerol/2.2) [13]. The analysis methods for total cholesterol, triacylglycerol, SBP and DBP changed in 2009: from Reflotron to a clinical chemical analysis at the laboratory for total cholesterol and triacylglycerol, and from BP measurements taken once in the supine position to being taken twice in a sitting position (the average of these two values being used in analyses). Lipid and BP values taken after 2009 were therefore corrected to make them comparable to values taken before 2009. Lipid and BP traits were also corrected for the use of lipid-lowering and antihypertensive medication using published constants (total cholesterol +1.347 mmol/l, triacylglycerol +0.208 mmol/l, HDL-C −0.060 mmol/l, LDL-C +1.290 mmol/l, SBP +15 mmHg, DBP +10 mmHg) [14, 15].
Lifestyle assessment
Participants completed a self-administered questionnaire that queried physical activity levels and diet and asked additional questions about tobacco use and alcohol consumption. Diet was assessed using a validated semi-quantitative food-frequency questionnaire designed to capture habitual dietary intake over the last year [16, 17]. The initial food-frequency questionnaire (used from 1985) covered 84 independent or aggregated food items but was reduced in 1996 to 66 food items by combining several questions related to similar foods and deleting some. Participants with ≥10% of the food-frequency questionnaire missing or a seemingly implausible total energy intake (<2093 or >18,841 kJ/day; <500 or >4500 kcal/day) were excluded from the analyses.
In order to obtain a summary factor representing the overall dietary pattern, a principal component analysis including all macronutrients (i.e. carbohydrate, protein, total fat, saturated fat, monounsaturated fatty acids [MUFA], polyunsaturated fatty acids [PUFA], essential fatty acids [n-3 and n-6 fatty acids], and fibre intakes expressed as per cent of total energy intake [E%]) was conducted, as previously described [18]. A single factor that contrasted carbohydrate and fibre intake against fat intake and accounted for 53.8% of the variance of all macronutrients was retained (ESM Table 2).
A validated modified version of the International Physical Activity Questionnaire [19] was used to gather information on leisure time physical activity for the past 3 months categorised as: (1) never; (2) occasionally; (3) 1–2 times/week; (4) 2–3 times/week; or (5) more than 3 times/week. For the current analyses, categories were combined into physically inactive (never and occasionally) and physically active (≥1–2 times/week).
Statistical analyses
All cardiometabolic traits were first adjusted for age, age2, sex and their interactions (age–sex and age2–sex) by conducting a multiple regression analysis using R software (version 3.1.1) [20] and retaining the residuals. Models with glycaemic and lipid traits as the dependent variables were additionally adjusted for fasting status. Models were also adjusted for the environmental exposure that was later tested in the genotype–environment interaction analyses; when the environmental exposure was alcohol intake or a dietary variable the model was also adjusted for the food-frequency questionnaire version. Retained residuals were then normalised by inverse normal transformation and used in the subsequent quantitative genetic analyses as recommended elsewhere [21, 22].
Kinship matrix
Kinship coefficients of the 16,430 participants with phenotype data were obtained based on the genealogical information gathered for the whole sample (193,060 individuals) using the CFC program [23], as Sequential Oligogenic Linkage Analysis Routines (SOLAR [24]) software is not designed to analyse such a large sample size.
Heritability estimation
Quantitative genetic analyses were conducted using the maximum likelihood-based variance components decomposition method implemented in SOLAR.
In the standard model, the observed covariance of a complex trait (Ω, cardiometabolic trait), assuming that dominance and epistasis are negligible, is defined as:
Here, Ω is an N-by-N matrix of the observed covariance of the cardiometabolic trait for each pair of the N individuals in the dataset, 2Φ gives the expected coefficient of relationship (Φ, kinship coefficient), σ 2 G is the additive genetic variance (i.e. genetic variation attributed to additive effects of the multiple genes affecting the cardiometabolic trait), I is the identity matrix of the unique unshared environmental component and σ 2 E is the environmental variance. This model is used to estimate narrow-sense heritability (h 2), i.e. the proportion of the cardiometabolic trait variance attributable to additive genetic effects:
where σ 2P is the total cardiometabolic trait variance.
Genotype–environment interactions
Genotype–environment interactions describe a relationship between genetic variation and changes in the cardiometabolic trait that is conditional on an environmental exposure. The presence of genotype–environment interactions can be tested with an extension of the standard model [equation (1)] [24, 25], which can be adapted for both discrete and continuous environmental exposures [5].
-
(a)
For a discrete (dichotomous) environmental exposure: Adaptation can be made by modelling environment-specific additive genetic and environmental standard deviations and a genetic correlation across the two exposure groups (i.e.. the proportion of variance in a trait explained by the same genetic factors in the two different exposure groups):
$$ \Omega =2\upphi {\uprho}_G{\upsigma}_{G1}{\upsigma}_{G2}+I{\upsigma}_{E1}{\upsigma}_{E2} $$(3)Additive genetic variance σ 2 G in equation (1) is decomposed as a product of additive genetic standard deviations for the two different environmental exposure groups (\( {\upsigma}_{G_1} \) and \( {\upsigma}_{G_2} \)) and a genetic correlation across the two groups denoted by ρ G [7, 25], i.e. \( {\upsigma}_G^2={\uprho}_G{\upsigma}_{G_1}{\upsigma}_{G_2} \). In the same way, environmental variance is decomposed into the environmental standard deviations for the two different environmental groups (\( {\upsigma}_{E_1} \) and \( {\upsigma}_{E_2} \)), i.e. σ 2 E = σ E1σ E2. Because the statistical genetic model assumes that the genetic and environmental effect estimates are uncorrelated, the function does not include an environmental correlation term. In the presence of genotype–environment interactions, narrow-sense heritability in k-th (k = 1,2) discrete environmental exposure group can then be estimated as: \( {h}_{Ek}^2=\frac{\upsigma_{Gk}^2}{\upsigma_{Gk}^2+{\upsigma}_{Ek}^2} \) [7].
-
(b)
For a continuous environmental exposure: Both additive genetic variance σ 2 G and genetic correlation ρ G can be modelled as exponential functions of the levels of the continuous environmental exposure [5, 26]. Genetic variance is modelled as:
$$ {\upsigma}_G^2= \exp \left({\upalpha}_G+{\upgamma}_G\left({e}_i-\overline{e}\right)\right) $$(4)where α G and γ G are parameters to be estimated, and e i is the value of the environmental exposure e of the i-th individual standardised against the sample mean (ē). Genetic correlation is modelled as an exponential decay function of the absolute difference of the pair-wise environmental exposure differences for the i-th and j-th individuals as:
$$ {\uprho}_G= \exp \left(-\uplambda \left|{e}_i-{e}_j\right|\right) $$(5)where λ is the parameter to be estimated.
The null hypothesis of genotype–environment interaction is that the expression of the genotype is independent of the environment. It can be shown that in the absence of a genotype–environment interaction (null hypothesis): (1) the genetic variance (σ 2 G ) will be homogenous across the levels of environmental exposure; and (2) the same quantitative trait measured in participants living in different levels of environmental exposure (e.g. active vs inactive or different ages) will have a genetic correlation (ρ G ) of 1.0 [5, 25, 27]. Hence, the presence of genotype–environment interactions is determined by testing two null hypotheses, which for the sake of simplicity will be referred to as class 1 and class 2 interactions from here on.
-
(a)
Class 1 interaction: The extended model is restricted by assuming homogenous genetic variance (σ 2 G ) across the levels of the environmental exposure. For a discrete environmental exposure [equation (3)], this means that the genetic standard deviations in the two exposure groups are equal, i.e. \( {\upsigma}_{G_1}={\upsigma}_{G_2} \). For a continuous environmental exposure [equation (4)], genetic variance (σ 2 G ) is homogenous across the different environmental levels when it is independent of the level of the environmental exposure, i.e. γ G = 0.
Rejection of the model constraining the genetic variance of the groups to be equal (i.e. presence of a significant class 1 interaction) would imply that the magnitude of the genetic effect on the cardiometabolic trait is significantly different depending on the level of the environmental exposure.
-
(b)
Class 2 interaction: The extended model is restricted by constraining the genetic correlation to 1. For a discrete environmental exposure [equation (3)], this means that the same cardiometabolic trait measured in individuals living in the different levels of the environmental exposure will have a genetic correlation of 1.0, i.e. ρ G = 1. For a continuous environmental exposure [equation (5)], genetic correlation (ρ G ) is equal to 1.0 if: (1) individuals i and j have the same level of the environmental exposure; or (2) λ=0. Thus, the null hypothesis of a class 2 interaction (i.e. genetic correlation is equal to 1) is equivalent to λ=0.
Rejection of the model constraining the genetic correlation between the environmental exposure groups to equal 1 (i.e. presence of a significant class 2 interaction) implies that a different gene or different set of genes are contributing to the variance of the cardiometabolic trait depending on the level of the environmental exposure.
To test the null hypothesis, each restricted model is compared with the extended model using the likelihood ratio test (LRT). The LRT statistic to test the null hypothesis of variance homogeneity (\( {\upsigma}_{G_1}={\upsigma}_{G_2} \) or γ G = 0) is distributed as a χ2 random variable with one degree of freedom (χ 21 ); the LRT to test the null hypothesis of genetic correlation equal to 1 (ρ G = 1 or λ = 0) is distributed as a 50:50 mixture of a χ2 random variable with a point mass at zero and one degree of freedom (0.5χ 20 + 0.5χ 21 ) [5].
In the figures representing class 1 and class 2 interactions for continuous environmental exposures, additive genetic variances and genetic correlations were calculated based on equations (4) and (5) and the estimates obtained for α G , γ G and λ parameters.
Multiple testing correction
The Bonferroni method assumes that the individual tests are independent of each other. However, the tests conducted in this study were not independent, so we estimated the total number of effective cardiometabolic traits and environmental exposures by accounting for the collective correlation of each set of clinical and environmental variables [28, 29]. The method utilises the estimates of variance of the eigenvalues (λs) derived from the correlation matrix of the set of variables and uses the following formula:
where M eff is the number of effective factors and M is the total number of variables (either clinical or environmental) included in the correlation matrix.
For the 14 environmental exposures and 11 cardiometabolic traits, 12.407 and 10.507 effective factors were obtained, respectively. Considering that we tested for both class 1 and class 2 interactions, the total number of effective tests are 260.721 (12.407 × 10.507 × 2). The Bonferroni corrected level of statistical significance for a threshold of 0.05 is thus 0.00019 (0.05/260.721).
Results
The characteristics of the 16,430 study participants are presented in Table 1.
Heritability estimates
All 11 cardiometabolic traits showed statistically significant narrow-sense heritability estimates (h 2 range 0.24–0.47; p < 0.001) (Table 2). Waist circumference conveyed the highest heritability estimate, followed by the remaining anthropometric, lipid and BP traits. Glycaemic traits conveyed the lowest heritability estimates.
Genotype–environment interactions
To test whether cardiometabolic traits are modulated by genotype–environment interactions the full model was compared with its constrained alternatives (i.e. genetic variance homogeneity and genetic correlation equal to 1). Statistically significant class 1 and class 2 interactions are summarised in Fig. 1.
Heat plot showing p values for (a) class 1 and (b) class 2 interactions. Experiment-wise significance threshold is p ≤ 1 × 10−4 (darkest blue in the heat plot). All environmental exposures are continuous variables except for sex, physical activity and smoking, which are dichotomous variables. TC, total cholesterol; TG, triacylglycerol
Genotype–age interactions
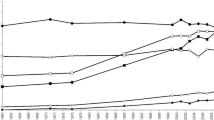
All the cardiometabolic traits except waist circumference, HDL-C and triacylglycerol showed significant genotype–age interactions. For fasting glucose, 2 h glucose, SBP and DBP significant class 1 interactions were observed (Fig. 2a). Class 2 interactions were observed for weight, BMI, total cholesterol, LDL-C, 2 h glucose, SBP and DBP (Fig. 2b), suggesting that different sets of genes influence the index traits in older compared with younger participants (ESM Table 3).
Genotype–age interactions: (a) class 1 and (b) class 2 interactions. Dark blue full square, weight; dark blue empty square, BMI; dark blue full triangle, fasting glucose; dark blue empty triangle, 2 h glucose; light blue full square, SBP; light blue empty square, DBP; light blue full triangle, total cholesterol; light blue full circle, LDL-C. Only significant traits are represented in the figure. Experiment-wise significant interactions (p ≤ 1 × 10−4) are marked with an asterisk (SBP for class 1 interactions, and weight for class 2 interactions). α G , γ G and λ parameters were calculated based on individuals 30–60 years of age, as this was the age range in the dataset. A broader age range curve (0–80 years) based on estimates above is displayed in the x-axis to improve the visualisation
Genotype–sex interactions
Genotype–sex interactions were observed for eight of the 11 cardiometabolic traits. For BMI, LDL-C, triacylglycerol, SBP and DBP class 1 interactions were observed (Fig. 3). The additive genetic effects for BMI, DBP, LDL-C and SBP were greater in women than in men, suggesting that the expression of these cardiometabolic traits is under greater genetic influence in women than in men (h 2 = 0.44, 0.38, 0.68 and 0.43 in women vs 0.35, 0.26, 0.23 and 0.29 in men for BMI, DBP, LDL-C and SBP, respectively). The additive genetic effects for triacylglycerol were greater in men than in women (h 2 = 0.49 in men vs 0.44 in women). Class 2 interactions were observed for body weight (ρ G = 0.86 ± 0.08; p = 0.049), total cholesterol (ρ G = 0.79 ± 0.08; p = 0.008), triacylglycerol (ρ G = 0.55 ± 0.07; p = 2 × 10−10), fasting glucose (ρ G = 0.73 ± 0.13; p = 0.03) and SBP (ρ G = 0.84 ± 0.08; p = 0.03) (ESM Table 4).
Class 1 genotype–sex interactions for (a) BMI, (b) DBP, (c) LDL-C, (d) SBP and (e) triacylglycerol. Only significant traits are represented in the figure; *p < 0.05, **p < 0.01, ***p < 0.001. Experiment-wise significant class 1 interactions (p ≤ 1 × 10−4) are BMI (p ≤ 4 × 10−6) and triacylglycerol (p ≤ 4 × 10−7)
Genotype–physical activity interactions
Class 1 interactions were observed for DBP and 2 h glucose, with the estimated heritabilities being higher in physically inactive (h 2 = 0.36 and 0.28, respectively) than in active individuals (h 2 = 0.20 and 0.16, respectively) (ESM Fig. 1). A class 2 interaction was observed for triacylglycerol (ρ G = 0.77 ± 0.11; p = 0.03) (ESM Table 5).
Genotype–smoking interactions
A class 2 genotype–smoking interaction was observed for body weight (ρ G = 0.79 ± 0.11; p = 0.04) (ESM Table 6).
Genotype–alcohol intake interactions
Body weight, BMI and 2 h glucose concentrations were influenced by genotype–alcohol intake interactions. For 2 h glucose, the interaction was a class 1 interaction (Fig. 4). Both class 1 and class 2 interactions were observed for body weight and BMI, suggesting that the interaction is a joint function of genetic effects that differ in magnitude and of different sets of genes influencing the body composition traits at different levels of alcohol intake (ESM Table 7).
Genotype–alcohol intake interactions: (a) class 1 and (b) class 2 interactions. Dark blue full square, weight; dark blue empty square, BMI; dark blue empty triangle, 2 h glucose. Only significant traits are represented in the figure. Experiment-wise significant interactions (p ≤ 1 × 10−4) are marked with an asterisk (weight for class 1 and class 2 interactions)
Genotype–diet interactions
In order to quantify genotype–diet interactions, we constructed a score representing the global dietary intake (i.e. diet pattern), as described in the Methods section. In a second step, we analysed the interactions with each macronutrient intake variable separately.
Genotype–diet pattern interactions
Body weight, BMI, glycaemic traits and SBP were influenced by genotype–diet pattern interactions. For SBP, the additive genetic variance decreased as the dietary fat/carbohydrate–fibre ratio increased (class 1 interaction) (ESM Fig. 2a). Class 2 genotype–diet pattern interactions were observed for body weight, BMI, and fasting and 2 h glucose concentrations (ESM Fig. 2b; ESM Table 8).
Genotype–carbohydrate intake interactions
LDL-C and SBP showed class 1 genotype–carbohydrate intake interactions (ESM Fig. 2c), whereas class 2 genotype–carbohydrate intake interactions were observed for BMI, waist circumference and fasting glucose (ESM Fig. 2d; ESM Table 9).
Genotype–protein intake interactions
For triacylglycerol and 2 h glucose, class 1 genotype–protein intake interactions were inferred (ESM Fig. 2e). For body weight and BMI, class 2 genotype–protein intake interactions were observed (ESM Fig. 2f; ESM Table 10).
Genotype–fibre intake interactions
SBP was the only cardiometabolic trait where a genotype–fibre intake interaction (class 1) was evident (ESM Fig. 2g; ESM Table 11).
Genotype–fat intake interactions
Body weight, BMI, fasting glucose and 2 h glucose showed significant genotype–total fat intake interactions (class 2) (ESM Fig. 3a; ESM Table 12).
Apart from total fat intake, four additional fat intake variables were analysed (saturated fat, essential fatty acids, PUFA and MUFA). Fasting glucose showed a class 1 genotype–saturated fat interaction (ESM Fig. 3b; ESM Table 13). Body weight, BMI and fasting glucose showed a significant class 2 genotype–saturated fat intake interaction (ESM Fig. 3c). Triacylglycerol, fasting glucose and DBP showed significant genotype–essential fatty acids and genotype–PUFA interactions. For triacylglycerol and DBP, the interactions were class 1 interactions, whereas for fasting glucose these interactions were class 2 (ESM Fig. 3d–g; ESM Tables 14 and 15). All anthropometric and glycaemic traits showed significant genotype–MUFA interactions, all of which were class 2 interactions (ESM Fig. 3h; ESM Table 16).
Multiple testing correction
Seven analyses withstood multiple testing correction: genotype–age interactions for body weight (class 2) and SBP (class 1); genotype–sex interactions for BMI (class 1) and triacylglycerol (class 1 and class 2) and genotype–alcohol intake interactions for body weight (class 1 and class 2).
There was no material change to the interpretation of these results when participants who were not fully fasted were excluded from the interaction analyses for lipid and glycaemic traits (ESM Table 17).
Discussion
To our knowledge, this is the first compendium of genotype–environment interactions for cardiometabolic traits to be reported. The purpose of doing so is to provide a foundation for subsequent locus-specific analyses of interaction effects and to aid the interpretation of published locus-specific interaction studies. After accounting for multiple testing, we observed robust evidence of genotype–age interactions for body weight and SBP, genotype–sex interactions for BMI and triacylglycerol, and genotype–alcohol intake interaction for body weight.
There are many published reports concerning interactions of environmental exposures with genetic factors in cardiometabolic traits (reviewed in [30–33]). Approaches include quantitative genetics studies, usually undertaken in twin or family-based cohorts [4, 6, 7, 34–38] and candidate gene studies, focused on individual genetic variants, haplotypes, or genetic risk scores constructed from variants with high biological priors for interactions or those conveying genome-wide significant marginal effects [39–47]. Several quantitative genetic studies have shown that physical activity attenuates the influence of genetic effects on cardiometabolic traits [4, 6, 34, 35, 37, 38]. However, only FTO–physical activity interactions in obesity [39–42] have been adequately replicated in candidate gene studies. In the present study, we observed evidence of genotype–physical activity interactions for DBP, 2 h glucose (class 1) and triacylglycerol (class 2), but not for obesity-related traits. This may be because analyses of the kind reported here account for the overall modifying effect of genetic variation (polygenic interactions), whereas gene–physical activity interactions in obesity may be oligogenic in nature.
According to our analyses, variation in the intake of macronutrients (whether modelled together or separately) may interact with genetic variation to affect body composition and glycaemic control. Although several candidate gene studies have focused on gene–diet interactions (e.g. [43–47]), there are few quantitative genetics studies on this topic, and these were restricted in scope and conducted in relatively small cohorts [35]. On the other hand, family-based studies have reported class 2 genotype–smoking interactions with serum leptin levels (an important endophenotype of adiposity) [7, 36], and those findings are consistent with the current analyses for body weight.
Although this is a hypothesis-generating study, and as such one might argue against multiple test adjustments owing to the risk of a type II error [48], we adopted a conservative approach to minimise the number of false-positives reported. Nevertheless, as described in the Results section, many of the statistical models yielded nominal evidence of interactions for the environmental exposures and cardiometabolic traits assessed. We present those findings, as the approach used here is orthogonal to standard approaches used to model genotype–environment interactions; thus, the combination of these approaches may help verify the presence or absence of interaction effects. Despite the relatively large sample size used here, it is of course likely that some of the hypothesis tests were underpowered. Statistical power may be diminished by the imprecise nature of the self-reported methods used to assess many of the environmental exposures and the need to dichotomise some of these variables for analysis. Survival bias is a further possible limitation, as people with the most deleterious genetic and/or environmental risk characteristics might have been excluded from the cohort because of early mortality. Systematic error (bias), on the other hand, may lead to false-positive or false-negative conclusions: for example, if an environmental exposure is over-reported at high or low levels of the cardiometabolic trait, or a strong correlate [49], an observed genotype–environment interaction may be false-positive. However, this limitation clearly does not impact our strongest findings (for age and sex), as these were objectively assessed. Additionally, as in other studies including genealogical information from registries (without genetic validation), the pedigrees are unlikely to be completely accurate due, for example, to false paternity. A further consideration is that some environmental exposures assessed here are to a limited extent influenced by genetic background [50, 51]; hence, it is possible that what might on the surface appear to be a genotype–environment interaction reflects, at least in part, epistasis.
In conclusion, our results suggest that cardiometabolic traits are heavily influenced by the interactions between the genotype and environmental exposures. Our data indicate that future studies focused on identifying specific genetic variants underlying genotype–environment interactions should focus on the exposures of age, sex and alcohol intake on body composition. Numerous other exposures and outcomes defined here are also plausible candidates for genotype–environment interaction.
Abbreviations
- E%:
-
Per cent of total energy intake
- DBP:
-
Diastolic blood pressure
- HDL-C:
-
HDL-cholesterol
- LDL-C:
-
LDL-cholesterol
- LRT:
-
Likelihood Ratio Test
- MUFA:
-
Monounsaturated fatty acids
- PUFA:
-
Polyunsaturated fatty acids
- SBP:
-
Systolic blood pressure
- SOLAR:
-
Sequential Oligogenic Linkage Analysis Routines
- VIKING:
-
Västerbotten Imputation Databank of Near-Complete Genomes
References
WHO (2015) Cardiovascular diseases (CVDs). Available from www.who.int/mediacentre/factsheets/fs317/en/index.html. Accessed 15 May 2016
Leal J, Luengo-Fernandez R, Gray A, Petersen S, Rayner M (2006) Economic burden of cardiovascular diseases in the enlarged European Union. Eur Heart J 27:1610–1619
Hunter DJ (2005) Gene–environment interactions in human diseases. Nat Rev Genet 6:287–298
Santos DM, Katzmarzyk PT, Diego VP et al (2014) Genotype by energy expenditure interaction and body composition traits: the Portuguese Healthy Family Study. Biomed Res Int 2014:845207
Santos DM, Katzmarzyk PT, Diego VP et al (2014) Genotype by sex and genotype by age interactions with sedentary behavior: the Portuguese Healthy Family Study. PLoS One 9, e110025
Santos DM, Katzmarzyk PT, Diego VP et al (2013) Genotype by energy expenditure interaction with metabolic syndrome traits: the Portuguese healthy family study. PLoS One 8, e80417
Martin LJ, Kissebah AH, Sonnenberg GE, Blangero J, Comuzzie AG (2003) Genotype–by–smoking interaction for leptin levels in the metabolic risk complications of obesity genes project. Int J Obes Relat Metab Disord 27:334–340
Towne B, Siervogel RM, Blangero J (1997) Effects of genotype–by–sex interaction on quantitative trait linkage analysis. Genet Epidemiol 14:1053–1058
Kurbasic A, Poveda A, Chen Y et al (2014) Gene–lifestyle interactions in complex diseases: design and description of the GLACIER and VIKING studies. Curr Nutr Rep 3:400–411
Hallmans G, Agren A, Johansson G et al (2003) Cardiovascular disease and diabetes in the Northern Sweden Health and Disease Study Cohort – evaluation of risk factors and their interactions. Scand J Publ Health Suppl 61:18–24
Ahmad S, Poveda A, Shungin D et al (2016) Established BMI-associated genetic variants and their prospective associations with BMI and other cardiometabolic traits: the GLACIER Study. Int J Obes 40:1346–1352
WHO (1999) Definition, diagnosis and classification of diabetes mellitus and its complications: Part 1: diagnosis and classification of diabetes mellitus. World Health Organization, Geneva
Friedewald WT, Levy RI, Fredrickson DS (1972) Estimation of the concentration of low-density lipoprotein cholesterol in plasma, without use of the preparative ultracentrifuge. Clin Chem 18:499–502
Wu J, Province MA, Coon H et al (2007) An investigation of the effects of lipid-lowering medications: genome-wide linkage analysis of lipids in the HyperGEN study. BMC Genet 8:60
Tobin MD, Sheehan NA, Scurrah KJ, Burton PR (2005) Adjusting for treatment effects in studies of quantitative traits: antihypertensive therapy and systolic blood pressure. Stat Med 24:2911–2935
Johansson I, Hallmans G, Wikman A, Biessy C, Riboli E, Kaaks R (2002) Validation and calibration of food-frequency questionnaire measurements in the Northern Sweden Health and Disease cohort. Public Health Nutr 5:487–496
Wennberg M, Vessby B, Johansson I (2009) Evaluation of relative intake of fatty acids according to the Northern Sweden FFQ with fatty acid levels in erythrocyte membranes as biomarkers. Public Health Nutr 12:1477–1484
Poveda A, Koivula RW, Ahmad S et al (2016) Innate biology versus lifestyle behaviour in the aetiology of obesity and type 2 diabetes: the GLACIER Study. Diabetologia 59:462–471
InterAct Consortium, Peters T, Brage S et al (2012) Validity of a short questionnaire to assess physical activity in 10 European countries. Eur J Epidemiol 27:15–25
R Development Core (2008) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Blangero J, Diego VP, Dyer TD et al (2013) A kernel of truth: statistical advances in polygenic variance component models for complex human pedigrees. Adv Genet 81:1–31
Beasley TM, Erickson S, Allison DB (2009) Rank-based inverse normal transformations are increasingly used, but are they merited? Behav Genet 39:580–595
Sargolzaei M, Iwaisaki H, Colleau JJ (2006) CFC (contribution, inbreeding (F), coancestry) release 1.0. A software package for pedigree analysis and monitoring genetic diversity. In: Proceedings of the 8th World Congress on Genetics Applied to Livestock Production, Belo Horizonte, Brazil
Almasy L, Blangero J (1998) Multipoint quantitative-trait linkage analysis in general pedigrees. Am J Hum Genet 62:1198–1211
Blangero J (1993) Statistical genetic approaches to human adaptability. Hum Biol 65:941–966
Blangero J (2009) Statistical genetic approaches to human adaptability, 1993. Hum Biol 81:523–546
Diego VP, Almasy L, Dyer TD, Soler JM, Blangero J (2003) Strategy and model building in the fourth dimension: a null model for genotype x age interaction as a Gaussian stationary stochastic process. BMC genetics 4 Suppl 1: S34
Nyholt DR (2004) A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am J Hum Genet 74:765–769
Patel CJ, Chen R, Kodama K, Ioannidis JP, Butte AJ (2013) Systematic identification of interaction effects between genome- and environment-wide associations in type 2 diabetes mellitus. Hum Genet 132:495–508
Franks PW, Pearson E, Florez JC (2013) Gene–environment and gene–treatment interactions in type 2 diabetes: progress, pitfalls, and prospects. Diabetes Care 36:1413–1421
Ahmad S, Varga TV, Franks PW (2013) Gene x environment interactions in obesity: the state of the evidence. Hum Hered 75:106–115
Franks PW, Poveda A (2011) Gene–lifestyle and gene–pharmacotherapy interactions in obesity and its cardiovascular consequences. Curr Vasc Pharmacol 9:401–456
Joseph PG, Pare G, Anand SS (2013) Exploring gene–environment relationships in cardiovascular disease. Can J Cardiol 29:37–45
Mustelin L, Silventoinen K, Pietilainen K, Rissanen A, Kaprio J (2009) Physical activity reduces the influence of genetic effects on BMI and waist circumference: a study in young adult twins. Int J Obes (2005) 33:29–36
Silventoinen K, Hasselbalch AL, Lallukka T et al (2009) Modification effects of physical activity and protein intake on heritability of body size and composition. Am J Clin Nutr 90:1096–1103
Martin LJ, Cole SA, Hixson JE et al (2002) Genotype by smoking interaction for leptin levels in the San Antonio Family Heart Study. Genet Epidemiol 22:105–115
Bouchard C, Tremblay A, Despres JP et al (1994) The response to exercise with constant energy intake in identical twins. Obes Res 2:400–410
Karnehed N, Tynelius P, Heitmann BL, Rasmussen F (2006) Physical activity, diet and gene–environment interactions in relation to body mass index and waist circumference: the Swedish young male twins study. Public Health Nutr 9:851–858
Kilpelainen TO, Qi L, Brage S et al (2011) Physical activity attenuates the influence of FTO variants on obesity risk: a meta-analysis of 218,166 adults and 19,268 children. PLoS Med 8, e1001116
Andreasen CH, Stender-Petersen KL, Mogensen MS et al (2008) Low physical activity accentuates the effect of the FTO rs9939609 polymorphism on body fat accumulation. Diabetes 57:95–101
Rampersaud E, Mitchell BD, Pollin TI et al (2008) Physical activity and the association of common FTO gene variants with body mass index and obesity. Arch Intern Med 168:1791–1797
Young AI, Wauthier F, Donnelly P (2016) Multiple novel gene–by–environment interactions modify the effect of FTO variants on body mass index. Nat Commun 7:12724
Sonestedt E, Roos C, Gullberg B, Ericson U, Wirfalt E, Orho-Melander M (2009) Fat and carbohydrate intake modify the association between genetic variation in the FTO genotype and obesity. Am J Clin Nutr 90:1418–1425
Rukh G, Sonestedt E, Melander O et al (2013) Genetic susceptibility to obesity and diet intakes: association and interaction analyses in the Malmö Diet and Cancer Study. Genes Nutr 8:535–547
Sanchez-Moreno C, Ordovas JM, Smith CE, Baraza JC, Lee YC, Garaulet M (2011) APOA5 gene variation interacts with dietary fat intake to modulate obesity and circulating triglycerides in a Mediterranean population. J Nutr 141:380–385
Lamri A, Abi Khalil C, Jaziri R et al (2005) Dietary fat intake and polymorphisms at the PPARG locus modulate BMI and type 2 diabetes risk in the DESIR prospective study. Int J Obes 36:218–224
Nettleton JA, Follis JL, Ngwa JS et al (2015) Gene × dietary pattern interactions in obesity: analysis of up to 68 317 adults of European ancestry. Hum Mol Genet 24:4728–4738
Bender R, Lange S (2001) Adjusting for multiple testing—when and how? J Clin Epidemiol 54:343–349
Murakami K, Livingstone MB (2015) Prevalence and characteristics of misreporting of energy intake in US adults: NHANES 2003-2012. Br J Nutr 114:1294–1303
Nikolaidis PT (2011) Familial aggregation and maximal heritability of exercise participation: a cross-sectional study in schoolchildren and their nuclear families. Sci Sports 26:157–165
Hasselbalch AL, Heitmann BL, Kyvik KO, Sorensen TI (2008) Studies of twins indicate that genetics influence dietary intake. J Nutr 138:2406–2412
Acknowledgements
The authors thank the participants, the health professionals, data managers and other staff involved in the Västerbotten Health Survey. We also thank V. Diego, C. Peterson and J. Blangero (Department of Genetics, Texas Biomedical Research Institute, San Antonio, TX, USA) for sharing program scripts for interaction analyses in the SOLAR program, and J. Armelius Lindberg and Å. Ågren (Department of Biobank Research, Umeå University, Umeå, Sweden), M. Sandström and P. Vikström (Centre for Demographic and Ageing Research, Umeå University, Umeå, Sweden) for their work on data extraction. Some of the computations were performed on resources provided by the Swedish National Infrastructure for Computing at HPC2N in Umeå, Sweden.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Data availability
Access to the data that support the findings of this study is restricted according to the permissions accorded by the institutional review board (www.epn.se/umeaa/om-naemnden/) and data access committee (www.biobank.umu.se/biobank/biobank---for-researchers/expert-panels/); requests to access these data are reviewed by the respective committees and should be directed to them, after first contacting the corresponding author of this paper (paul.franks@med.lu.se).
Funding
The study was funded by the Swedish Heart–Lung Foundation, Novo Nordisk, the Swedish Research Council, and the European Research Council (CoG-2015_681742_NASCENT). AP was funded in part by a postdoctoral grant from the Basque Government and a research grant from the Swedish Research Council.
Duality of interest
PWF was a member of advisory boards for Sanofi Aventis and Eli Lilly and has received research funding from Sanofi Aventis, Eli Lilly, Servier, Jansson, Boehringer Ingelheim and Novo Nordisk. The study sponsor was not involved in the design of the study; the collection, analysis, and interpretation of data; writing the report; or the decision to submit the report for publication. All other authors declare no competing financial interest associated with their contribution to this manuscript.
Contribution statement
PWF designed and obtained funding for the VIKING study. AP, AK and PWF conceived and designed the analyses. AP, AK and YC analysed the data. AB, EE, GH and IJ contributed to data acquisition. AP, FR, AK and PWF interpreted the data and wrote the manuscript. All the authors revised the manuscript and contributed to the final version. AP, AK and PWF are the guarantors of this work. All authors approved the version to be published.
Additional information
Azra Kurbasic and Paul W. Franks contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM
(PDF 4575 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Poveda, A., Chen, Y., Brändström, A. et al. The heritable basis of gene–environment interactions in cardiometabolic traits. Diabetologia 60, 442–452 (2017). https://doi.org/10.1007/s00125-016-4184-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-016-4184-0