Abstract
Aims/hypothesis
Individuals at a high risk of type 2 diabetes demonstrate moderate impairments in glucose metabolism years before the clinical manifestation of type 2 diabetes, a state called ‘prediabetes’. In order to elucidate the pathophysiological processes leading to type 2 diabetes, we aimed to identify protein biomarkers associated with prediabetes.
Methods
In a proteomics study, we used targeted selected reaction monitoring (SRM)-MS to quantify 23 candidate proteins in the plasma of 439 randomly selected men and women aged 47–76 years from the population-based German KORA F4 study. Cross-sectional associations of protein levels with prediabetes (impaired fasting glucose and/or impaired glucose tolerance), type 2 diabetes, glucose levels in both the fasting state and 2 h after an OGTT, fasting insulin and insulin resistance were investigated using regression models adjusted for technical covariables, age, sex, BMI, smoking, alcohol intake, physical inactivity, actual hypertension, triacylglycerol levels, total cholesterol/HDL-cholesterol ratio, and high-sensitivity C-reactive protein levels.
Results
Mannan-binding lectin serine peptidase 1 (MASP1; OR per SD 1.77 [95% CI 1.26, 2.47]), thrombospondin 1 (THBS1; OR per SD 1.55 [95% CI 1.16, 2.07]) and glycosylphosphatidylinositol-specific phospholipase D1 (GPLD1; OR per SD 1.40 [95% CI 1.01, 1.94]) were positively associated with prediabetes, and apolipoprotein A-IV (ApoA-IV; OR per SD 0.75 [95% CI 0.56, 1.00]) was inversely associated with prediabetes. MASP1 was positively associated with fasting and 2 h glucose levels. ApoA-IV was inversely and THBS1 was positively associated with 2 h glucose levels. MASP1 associations with prediabetes and fasting glucose resisted Bonferroni correction. Type 2 diabetes associations were partly influenced by glucose-lowering medication.
Conclusions/interpretation
We discovered novel and independent associations of prediabetes and related traits with MASP1, and some evidence for associations with THBS1, GPLD1 and ApoA-IV, suggesting a role for these proteins in the pathophysiology of type 2 diabetes.
Similar content being viewed by others
Introduction
Type 2 diabetes develops slowly over a long time period, and individuals at risk usually demonstrate moderate derangement in glucose metabolism years before the manifestation of type 2 diabetes [1]. Biomarkers associated with this ‘prediabetic’ state (impaired fasting glucose and/or impaired glucose tolerance) may help to elucidate the pathophysiological pathways leading to type 2 diabetes [2, 3].
Multiple omics technologies have been applied in biomarker research to date [4, 5]. Transcriptomics has the advantage that mRNA can be amplified after conversion to cDNA; in contrast, no amplification step is possible in proteomics or metabolomics, and sensitivity therefore remains a limiting factor. Recent advances in MS have moved the biomarker field forward from investigating transcriptional biomarkers towards protein and metabolite biomarkers. The latter have the advantage that they are downstream of gene expression and therefore reflect disease status and pathophysiological processes more directly compared with nucleic acid markers [6]. Several putative metabolomics biomarkers have recently been identified in diabetes research [7, 8]. Compared with metabolites, proteins have the advantage of being more stable and less variable. Classical approaches such as clinical chemistry assays, especially antibody-based methods for protein measurement, are impaired by time-consuming and costly development, and vary in quality and availability [9]. Targeted MS approaches such as selected reaction monitoring (SRM) have the potential to overcome these limitations. SRM assays can be generated for any detectable protein in a tailored fashion, thus allowing a high degree of multiplexing [10]. Established SRM assays can be measured on any benchtop triple quadrupole instrument and can therefore be transferred to a clinical setting and used across different clinical laboratories [11]. We used a nano-liquid chromatography (LC) MS approach in SRM mode (SRM-MS) to quantify the relative levels of 23 proteins in 439 plasma samples from the population-based Cooperative Health Research in the Region of Augsburg (KORA) F4 study (see electronic supplementary material [ESM] Table 1). The candidate proteins were selected based on previous results from a mouse model for type 2 diabetes [12], literature mining and unpublished shotgun discovery proteomic analyses (von Toerne and Hauck, data not shown).
Our study aimed to identify novel proteins associated with prediabetes in order to help elucidate pathophysiological processes that drive development towards overt type 2 diabetes. We also compared protein alterations observed in the early stage of prediabetes with those in manifest type 2 diabetes. As a secondary aim, we investigated whether changed protein abundances are associated with the following prediabetes-related continuous traits: glucose levels in the fasting state or 2 h after an oral glucose load (2 h glucose), fasting insulin, and insulin resistance.
Methods
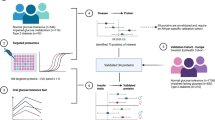
The complete study workflow is depicted in ESM Fig. 1.
Participants
KORA F4 study
We performed a cross-sectional analysis of participants in the population-based KORA F4 study. The KORA F4 study was carried out in 2006–2008 as a follow-up of the KORA S4 baseline study (1999–2001). In KORA S4, 4261 participants were recruited from a randomised two-stage cluster sample of 6640 individuals (with equal-sized sex and age strata) from the target population of all German residents in the region of Augsburg aged 25–74 years. The KORA F4 study included 3080 participants (response 79.6%). Investigations were carried out in accordance with the Declaration of Helsinki, and written informed consent was obtained from all participants. The study was approved by the Ethics committee of the Bavarian Chamber of Physicians [13].
Selection criteria and randomisation
Plasma samples from 250 male and 250 female KORA F4 participants were randomly selected for SRM-MS from 1705 individuals fulfilling the following criteria: (1) omics data were available, to allow extended analyses in future studies; (2) within the age range of 47–76 years in which most of the KORA F4 omics data were available; (3) BMI data were available; (4) at least nine frozen plasma straws were available; and (5) individuals were either non-diabetic with fasting glucose, 2 h glucose and fasting insulin data, or had type 2 diabetes.
The 500 samples were randomly distributed into six sample preparation batches. The complete SRM-MS workflow was blinded to disease status. The analysis order was randomised. The sixth batch did not pass quality control and was removed from the study (n = 60; see ESM methods section ‘Quality control of sample preparation and MS-measurement’). One additional individual was excluded because of missing data in the C-reactive protein (CRP) covariable, leaving data from 439 participants for analysis of the dichotomous outcomes: prediabetes and type 2 diabetes vs normal glucose metabolism. The analyses of continuous fasting glucose, fasting insulin and insulin resistance were restricted to 406 participants who were not taking glucose-lowering medication. Two hour glucose levels were investigated in only 398 participants because there were missing values for eight participants who did not receive OGTTs because they were known to have diabetes.
Non-SRM measurements
All participants underwent a standard physical and medical examination, which is described in detail elsewhere [14–16].
Participants classified as having known diabetes included all participants with self-reported diabetes that was validated by contacting the responsible physician or medical chart review, and those self-reporting a current intake of glucose-lowering medication. All participants without known diabetes underwent a standard 75 g OGTT. Blood samples were taken without stasis after an overnight fast of at least 8 h and 2 h after glucose solution intake. Serum glucose was analysed using a hexokinase method (GLU Flex, Dade Behring, Deerfield, IL, USA). Normal glucose metabolism (i.e. fasting glucose <6.1 mmol/l and 2 h glucose <7.8 mmol/l), prediabetes (fasting glucose ≥6.1 mmol/l but <7.0 mmol/l, and 2 h glucose <7.8 mmol/l [isolated impaired fasting glucose (IFG)] or fasting glucose <6.1 mmol/l and 2 h glucose ≥7.8 mmol/l but <11.1 mmol/l [isolated impaired glucose tolerance (IGT)], or both [IFG and IGT]), and newly diagnosed diabetes (fasting glucose ≥7.0 mmol/l or 2 h glucose ≥11.1 mmol/l) were defined according to the 1999 WHO criteria [17]. Newly diagnosed and known diabetic participants for whom the diabetes type could not be validated and for whom no contradictory information was given were assumed to have type 2 diabetes. Fasting insulin levels in frozen serum were assessed by ELISA (Invitrogen, Darmstadt, Germany). The HOMA-IR was calculated as fasting insulin (in pmol/l) × fasting glucose (in mmol/l) ÷ 135. Since insulin secretion is usually impaired at a later stage in diabetes development, the association with insulin secretion was not examined in our study.
Information about age, sex, smoking, alcohol intake and physical activity were assessed during a standardised interview, carried out by trained medical staff. Regular smoking was defined as smoking of at least one cigarette per day on average. Alcohol intake was categorised as no (0 g/day), moderate (men 0.1–39.9 g/day; women 0.1–19.9 g/day) and high (men ≥40 g/day; women ≥20 g/day). Individuals who did not participate in leisure time physical activity for at least one hour per week during both summer and winter were classified as physically inactive. Actual hypertension was defined as systolic blood pressure ≥140 mmHg and/or diastolic blood pressure ≥90 mmHg and/or use of antihypertensive medication given that the subjects were aware that they had hypertension. The laboratory methods used for measurement of the clinical chemistry variables are described in the ESM methods section ‘Clinical chemistry measurements’.
SRM-MS measurements
Plasma depletion and on-bead digestions using ProteoMiner beads
Depletion and tryptic digestion of plasma samples which had been stored at −80°C were performed as previously described in six processing batches [12]. For quality control of depletion, digestion and MS measurements, a final concentration of 50 fmol ribulose-1,5-bisphosphate carboxylase oxygenase protein (RuBisCO; Sigma-Aldrich, Taufkirchen, Germany) was spiked into each 10 μl plasma sample as an internal standard (see ESM methods section ‘Quality control of sample preparation and MS-measurement’). After digestion, samples were stored at −20°C.
SRM-MS measurement and quantification principle
Selected proteotypic peptides (so-called precursor ions) were fragmented into multiple collision-induced dissociation products of peptides (so-called fragment ions or transitions). Isotope-labelled synthetic peptides (heavy peptides; PEPotec Thermo Fisher Scientific, Ulm, Germany) were purchased for each selected peptide. These were used as internal controls to correct signal integration and for relative quantification.
Candidate protein selection
Candidate proteins were obtained from a previous pilot mouse study [12] and unpublished mouse data (von Toerne and Hauck, data not shown). These were complemented by results from shotgun measurements of human plasma samples from different sources (data not shown). The main selection criterion for a candidate protein was an observed >1.3-fold difference between diabetic and non-diabetic samples. These observed tendencies were considered for hypothesis generation independent of their statistical significance because of the lack of power resulting from small sample sizes. Finally, candidate proteins were added by literature mining.
Development of SRM assays
Skyline software (MacCoss Lab Software, Seattle, USA) was used to create SRM assays for 24 proteins (ESM Table 1), including the control protein [18]. Details are described in the ESM methods section.
Targeted MS measurement
The heavy peptide mix was added to the digested sample prior to MS measurement. LC-tandem MS (MSMS) analysis was performed on a Tempo nano MDLC system (Eksigent Technologies, Dublin, CA, USA) coupled online to a QTrap4000 (ABSCIEX, Framingham, MT, USA) mass spectrometer by a nano spray ion source, as previously described [12] with the following changes. Peptides were separated on a 90 min non-linear gradient of 2% acetonitrile and 3% DMSO in 0.1% formic acid (solution A) to 95% acetonitrile and 3% DMSO in 0.1% formic acid (solution B) at a flow rate of 250 nl/min. The gradient settings were: 5–65 min, 5–45% solution B; 65–70 min, 45–90% solution B; 70–72 min, 90% solution B; 72–80 min, 90–5% solution B; followed by re-equilibration for 10 min to starting conditions. The mass spectrometer was used in positive ion mode with a 2600 V ion spray voltage, curtain gas setting of 14, ion source gas setting of 30, nebuliser gas setting of 0 and an interface heater temperature of 170°C. Eluted peptides were measured using a scheduled method in a detection window of 360 s and a target scan time of 3 s. The resolution in Q1 und Q3 was set to unit (0.7 atomic mass units at full width at half maximum).
Preprocessing of SRM-MS data and quantification
The data preprocessing procedure (see ESM Fig. 2) was performed using R version 3.1.2 [19]. First, CVs were calculated for all transitions based on five replicate measurements of a pool containing all samples using AuDIT software [20]. Transitions with a CV of ≥30% were excluded. Transition measurements that were zero (in total 0.25% of all data values) were randomly assigned a number between 1 and the minimum observation of the respective transition. For each peptide with three transitions, the two transitions with the most similar information were selected for use in further calculations. The method used was adapted from Mani et al in 2012 [21] (see ESM methods section ‘Preprocessing of SRM data’). For the final 108 transitions (of 54 peptides from the 23 candidate proteins), light (endogenous) to heavy (synthetic) peptide ratios (LHRs) of the AUC values were calculated, log2-transformed, and averaged for all transitions of a single peptide. Finally, peptide level LHR information was averaged per protein to yield relative protein levels.
Statistical analysis
Statistical analysis was performed using R version 3.1.2. All SRM-MS protein level LHRs were log2-transformed and divided by their SDs to ensure comparability of effect estimates between different proteins in the regression models. Pearson’s correlation coefficients between standardised LHRs (SRM-MS protein levels) and available clinical chemistry results were investigated for proteins measured previously with alternative technology (high-sensitivity CRP [hsCRP], apolipoprotein A-IV (ApoA-IV), adiponectin [ADIPOQ] and retinol-binding protein 4 [RBP4]). Correlations between all SRM-MS measured proteins and all continuous covariables were also investigated.
Associations between standardised LHRs and both prediabetes and diabetes were analysed by logistic regression. In order to investigate the potential influence of glucose-lowering medication, all diabetic participants taking glucose-lowering medication were excluded in a sensitivity analysis. Continuous glucose and insulin outcomes were loge-transformed, z-standardised and analysed by linear regression. All regression models were adjusted for technical covariables (analysis batch, day of analysis as well as MS opening and cleaning routine), age and sex (model 1). Model 2 was additionally adjusted for BMI and model 3 for smoking status, alcohol intake, physical inactivity, actual hypertension, continuous triacylglycerol levels, total cholesterol/HDL-cholesterol ratio and hsCRP levels.
A p value of <0.05 was considered statistically significant; p values were calculated with and without Bonferroni correction for the number of measured candidate proteins.
Results
Descriptive statistics
Of the 219 men and 220 women with valid SRM-MS protein data, 281 were normoglycaemic, 95 were prediabetic and 63 had type 2 diabetes. Diabetic participants were on average older than the normoglycaemic participants, had a higher BMI and were more likely to suffer from actual hypertension. Furthermore, they had a higher total cholesterol/HDL-cholesterol ratio, as well as higher triacylglycerol and hsCRP levels. In general, the characteristics of prediabetic participants were intermediate (Table 1). Correlations between SRM-MS quantified proteins and continuous type 2 diabetes risk factors are shown in ESM Table 2.
Correlations between SRM-MS and clinical chemistry measurements
SRM-MS results correlated strongly with the previous ELISA measurements available for ADIPOQ (Pearson’s r = 0.89) and ApoA-IV (r = 0.77), as well as nephelometric measurements for hsCRP (r = 0.90) and RBP4 (r = 0.49; p < 0.001 for all; ESM Fig. 3).
MASP1, THBS1, GPLD1 and ApoA-IV are associated with prediabetes
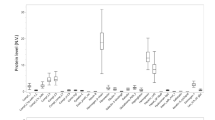
After adjustment for technical covariables, age and sex (model 1), seven SRM-MS quantified proteins were associated with prediabetes (vs normal glucose metabolism): positive association, apolipoprotein C-II (ApoC-II), CRP, lipopolysaccharide-binding protein (LBP), mannan-binding lectin serine peptidase 1 (MASP1), and thrombospondin 1 (THBS1 [also called TSP-1]); inverse association, ADIPOQ and ApoA-IV (ESM Fig. 4). After additional adjustment for BMI and other type 2 diabetes risk factors (model 3), MASP1, THBS1 and glycosylphosphatidylinositol-specific phospholipase D1 (GPLD1) were positively associated and ApoA-IV was inversely associated with prediabetes (Fig. 1). Of these four proteins, only ApoA-IV and GPLD1 were also positively associated with type 2 diabetes. After excluding the 33 diabetic participants taking glucose-lowering medication, the association between ApoA-IV levels and diabetes disappeared, the positive association between GPLD1 and diabetes remained, and MASP1 levels became positively associated with diabetes (Table 2).
ORs with 95% CIs for the association ‘prediabetes vs normal glucose metabolism’ per one SD increase in SRM-MS measured proteins (n = 376). Model 3 is adjusted for technical covariables, age, sex, BMI, smoking status, alcohol intake, physical inactivity, actual hypertension, triacylglycerol levels and the total cholesterol/HDL-cholesterol ratio. All proteins except CRP were additionally adjusted for hsCRP, as measured by latex-enhanced immunonephelometry. ApoC-III, apolipoprotein C-III; ApoE, apolipoprotein E; C7, complement component 7; C8A, complement component 8, α polypeptide; C8B, complement component 8, β polypeptide; CD14, CD14 molecule; CD5L, CD5 molecule-like; CLEC3B, C-type lectin domain family 3, member B; GSN, gelsolin; ITIH4, inter-α-trypsin inhibitor heavy chain family, member 4; ORM1, orosomucoid 1; PZP, pregnancy-zone protein; SHBG, sex hormone-binding globulin; *p < 0.05, **p < 0.01, ***p < 0.001 (uncorrected significance levels), † p < 0.05 (Bonferroni-corrected significance levels)
The analysis of diabetes-related continuous traits adjusted for type 2 diabetes risk factors in participants not taking glucose-lowering medication (model 3; Fig. 2) showed that ApoA-IV was inversely and THBS1 was positively associated with 2 h glucose levels. MASP1 was positively associated with fasting and 2 h glucose levels. After adjustment for only age, sex and BMI (model 2), MASP1 was also associated with HOMA-IR. The association between MASP1 and HOMA-IR was most strongly attenuated by adjustment for triacylglycerol levels (data not shown).
Estimated difference in continuous outcomes for 406 participants not taking glucose-lowering medication expressed as the SD change in the continuous outcome (standardised z-score β estimate with 95% CI) per one SD increase in the respective protein. (a) Model 2: adjusted for technical covariables, age, sex and BMI. (b) Model 3: adjusted for technical covariables, age, sex, BMI, smoking status, alcohol intake, physical inactivity, actual hypertension, triacylglycerol levels, total cholesterol/HDL-cholesterol ratio and hsCRP levels measured by latex-enhanced immunonephelometry. FG, fasting glucose; 2hG, 2 h glucose (n = 398); FI, fasting insulin. *p < 0.05, **p < 0.01, ***p < 0.001 (uncorrected significance levels), † p < 0.05, †† p < 0.01, ††† p < 0.01 (Bonferroni-corrected significance levels)
Correlations among all investigated SRM-MS quantified proteins are displayed in ESM Table 3. ApoA-IV, GPLD1 and MASP1 correlated positively (0.30 ≤ Pearson’s r < 0.50), while THBS1 was not strongly correlated with any of the other three proteins (0.04 ≤ r ≤ 0.21). In order to investigate whether the association results for ApoA-IV, GPLD1 and MASP1 were independent from one another, we extended all regression models by mutually adjusting these three proteins for each other. As displayed in Table 2, the inverse ApoA-IV–prediabetes association was strengthened, the GPLD1–prediabetes association was attenuated, and the MASP1–prediabetes association was not appreciably affected. Similarly, the inverse association between ApoA-IV and 2 h glucose levels was strengthened, and the positive MASP1 associations with fasting and 2 h glucose levels were virtually unchanged (data not shown).
When correcting p values for the number of measured candidate proteins, the positive MASP1 associations with prediabetes and fasting glucose remained statistically significant in all main models investigated (Table 2 and Fig. 2).
Discussion
We evaluated the association of prediabetes and type 2 diabetes with a novel set of 23 plasma protein candidates that had been assessed in a multiplexed, targeted MS approach in 439 random samples from the population-based KORA F4 study. Relative quantification based on SRM-MS measurements correlated highly with available clinical chemistry results for the three proteins, demonstrating the reliability of our multiplex approach.
In order to identify biomarkers associated with early derangements in glucose metabolism that might play a role in type 2 diabetes pathophysiology, we primarily focused on prediabetic individuals. Our approach revealed a strong and novel positive association between MASP1 and prediabetes. MASP1, the most abundant serine protease of the complement lectin pathway, is a major player in the complement cascade which is initiated when a complex comprising mannose-binding lectin (MBL), MBL-associated serine proteases (MASPs: MASP1, MASP2, MASP3) and MBL-associated proteins (MAp19 and MAp44) binds to its target carbohydrate-containing ligands, primarily derived from pathogens or damaged tissues [22, 23]. In our study, the adjusted odds of developing prediabetes was estimated to be 77% higher per SD elevation in MASP1 plasma levels; this association was consistently observed under conditions of both elevated continuous fasting and 2 h glucose levels, which together define the prediabetic state. Moreover, there was some evidence for a positive association with insulin resistance. This association was most strongly attenuated by adjustment for triacylglycerol levels, indicating that triacylglycerols might mediate part of the association between MASP1 levels and HOMA-IR. MASP1 plasma levels were also elevated in diabetic participants, but this association lost statistical significance after multivariable adjustment, possibly because of the smaller sample size. In a small pilot study, MASP1 plasma levels were associated with diabetes in individuals with coronary artery disease but not healthy controls; however, no associations with glucose or HbA1c levels were found [24].
For another member of the lectin pathway, the MBL2 protein, we found no evidence of a difference in abundance between normoglycaemic and prediabetic or diabetic participants, or an association with glucose or insulin variables. Elevated plasma MBL2 levels have been reported in type 1 diabetes adolescents [25, 26] and adults [27]. In addition, MBL2 levels correlated positively with HbA1c levels in a study of 391 individuals with type 1 diabetes and 100 healthy controls [28], and were inversely associated with HOMA-IR after adjustment for age, sex, BMI and waist-to-hip-ratio in a study of 113 non-diabetic participants [29]. Our null finding for MBL2 does not support a general upregulation of the lectin pathway in type 2 diabetes development. Instead, MASP1, which has a relaxed substrate specificity and was shown to cleave more than a dozen proteins in human blood, might play a role in type 2 diabetes pathogenesis via interacting with the coagulation cascade, the kinin generating system or proinflammatory signalling [23]. Alternatively, increased MASP1 levels might have been induced by the moderately elevated glucose levels present in the prediabetic state [30, 31].
In addition to MASP1, which was significantly associated with prediabetes after Bonferroni correction, we also observed some evidence for associations of GPLD1 and THBS1 (both positive) and ApoA-IV (inverse) with prediabetes.
GPLD1 also called GPI-PLD) is an amphiphilic protein, associated with HDLs in serum [32]. Increased plasma GPLD1 was associated with type 2 diabetes and (albeit less strongly) with prediabetes. The prediabetes association was attenuated after adjustment for MASP1 levels. Together with the strong correlation between GPLD1 and MASP1, this suggests an as yet undescribed relationship between the two proteins. Comparable studies in humans are lacking and the potential role of GPLD1 in diabetes and prediabetes is currently unknown. GPLD1 cleavage generates second messengers that can act as insulin-mimetic low molecular mass species [33, 34]. Suzuki et al demonstrated that treatment of mice with certain insulin-mimetic low molecular mass species led to decreased plasma glucose levels, increased glycogen content in the liver and skeletal muscles, and improved glucose tolerance in db/db mice and C57B6N mice on high-fat diets [34].
The amphiphilic protein ApoA-IV is produced in enterocytes of the small intestine and is secreted as one of the structural proteins of chylomicrons, VLDL or HDL; it is also found free in plasma [35, 36]. As summarised by Wang et al [37], ApoA-IV has potent antioxidant and anti-inflammatory properties, and has been suggested to reduce food intake by increasing satiety (at least in animal models). These properties all have the potential to prevent type 2 diabetes development [3, 38, 39]. In support of this notion, we observed associations between increased ApoA-IV levels and both decreased 2 h glucose levels and decreased prediabetes prevalence independent of all investigated covariables. Experimental studies in isolated pancreatic islets and Apoa4 knockout mice have already demonstrated that ApoA-IV has a glucose-lowering effect [40, 41]. Interestingly, consistent with two small type 2 diabetes studies, individuals with manifest type 2 diabetes had increased ApoA-IV levels in our study, [42, 43]. However, a sensitivity analysis which we conducted showed that ApoA-IV levels were not increased in diabetic individuals who did not take glucose-lowering medication. Owing to the limited sample size, future larger studies are needed to clarify the impact of glucose-lowering medication in more detail.
Finally, there was some evidence that plasma levels of the adipokine THBS1 are associated with increased 2 h glucose levels as well as higher prediabetes prevalence. THBS1 is reported to be upregulated in diverse tissues in humans and rodent models of diabetes and obesity [44]. Moreover, increased THBS1 mRNA levels were associated with increased insulin resistance, as measured by a frequently sampled IVGTT in 86 non-diabetic participants [45]. Experiments in Thbs1 knockout mice demonstrated THBS1 involvement in adipose inflammation and metabolic dysregulation in obesity and type 2 diabetes [46]. The elevated THBS1 levels we observed in prediabetic participants might thus be explained by the proinflammatory properties of THBS1.
Strengths and limitations
To our knowledge, this study is the largest MS-based proteomics study in the field of type 2 diabetes research. An obvious strength of our study is the highly reliable parallel measurement of 23 candidate proteins using targeted proteomics. One of the major advantages of tailored SRM-MS is that the measured peptide sequences are known, which cannot be guaranteed with other methods. Moreover, the use of isotope-labelled peptide standards for relative quantification represents the state of the art. Additional strengths are that we adjusted our analyses for several type 2 diabetes risk factors and present results on continuous diabetes-related traits in addition to categorical diabetes status. A limitation is that our study does not provide absolute protein concentrations; however, this should not affect the reported associations. Similarly, relative levels should not be strongly affected if protein amounts are reduced by freezing and thawing. HOMA-IR was used as a surrogate variable for the true insulin resistance status. Owing to use of only one OGTT, some participants in the normoglycaemic, prediabetic or newly diagnosed type 2 diabetic groups might be misclassified. Together with a sample size of only 439, this might lead to false negative results. We thus might have missed true protein–prediabetes associations. On the other hand, some of our findings might be false positives and future large, independent studies will be needed to replicate our results, especially for the three weaker protein signals from GPLD1, THBS1 and ApoA-IV. The present study is cross-sectional in nature; thus, our results might be influenced by reverse causation. To provide optimal data, future replication studies should be prospective.
Conclusions
Our SRM-MS approach revealed a novel, independent positive association of MASP1 with prediabetes and related continuous traits, suggesting a role for MASP1 in type 2 diabetes pathophysiology. There was also some evidence of higher GPLD1 and THBS1 levels and lower ApoA-IV levels in prediabetic participants. These associations, however, were not strong enough to resist Bonferroni correction. Future larger studies need to corroborate these findings.
Abbreviations
- ADIPOQ:
-
Adiponectin
- ApoA-IV:
-
Apolipoprotein A-IV
- ApoC-II:
-
Apolipoprotein C-II
- CRP:
-
C-reactive protein
- GPLD1:
-
Glycosylphosphatidylinositol-specific phospholipase D1
- hsCRP:
-
High-sensitivity C-reactive protein
- KORA:
-
Cooperative Health Research in the Region of Augsburg
- LBP:
-
Lipopolysaccharide-binding protein
- LC:
-
Liquid chromatography
- LHR:
-
Light to heavy ratio
- MASP1:
-
Mannan-binding lectin serine peptidase 1
- MBL2:
-
Mannose-binding lectin (protein C) 2
- MSMS:
-
Tandem MS
- RBP4:
-
Retinol-binding protein 4
- SRM:
-
Selected reaction monitoring
- THBS1:
-
Thrombospondin 1
References
Tabak AG, Herder C, Rathmann W, Brunner EJ, Kivimaki M (2012) Prediabetes: a high-risk state for diabetes development. Lancet 379:2279–2290
Huth C, Beuerle S, Zierer A et al (2015) Biomarkers of iron metabolism are independently associated with impaired glucose metabolism and type 2 diabetes: the KORA F4 study. Eur J Endocrinol 173:643–653
Kluppelholz B, Thorand B, Koenig W et al (2015) Association of subclinical inflammation with deterioration of glycaemia before the diagnosis of type 2 diabetes: the KORA S4/F4 study. Diabetologia 58:2269–2277
Herder C, Karakas M, Koenig W (2011) Biomarkers for the prediction of type 2 diabetes and cardiovascular disease. Clin Pharmacol Ther 90:52–66
Herder C, Kowall B, Tabak AG, Rathmann W (2014) The potential of novel biomarkers to improve risk prediction of type 2 diabetes. Diabetologia 57:16–29
Shao S, Guo T, Aebersold R (2015) MS-based proteomic quest for diabetes biomarkers. Biochim Biophys Acta 1854:519–527
Wang-Sattler R, Yu Z, Herder C et al (2012) Novel biomarkers for pre-diabetes identified by metabolomics. Mol Syst Biol 8:615
Yokoi N, Beppu M, Yoshida E et al (2015) Identification of putative biomarkers for prediabetes by metabolome analysis of rat models of type 2 diabetes. Metabolomics 11:1277–1286
Marx V (2013) Targeted proteomics. Nat Methods 10:19–22
Picotti P, Bodenmiller B, Aebersold R (2013) Proteomics meets the scientific method. Nat Methods 10:24–27
Gillette MA, Carr SA (2013) Quantitative analysis of peptides and proteins in biomedicine by targeted mass spectrometry. Nat Methods 10:28–34
von Toerne C, Kahle M, Schafer A et al (2013) Apoe, Mbl2, and Psp plasma protein levels correlate with diabetic phenotype in NZO Mice – an optimized rapid workflow for SRM-based quantification. J Proteome Res 12:1331–1343
Rathmann W, Kowall B, Tamayo T et al (2012) Hemoglobin A1c and glucose criteria identify different subjects as having type 2 diabetes in middle-aged and older populations: the KORA S4/F4 Study. Ann Med 44:170–177
Ruckert IM, Heier M, Rathmann W, Baumeister SE, Doring A, Meisinger C (2011) Association between markers of fatty liver disease and impaired glucose regulation in men and women from the general population: the KORA-F4-study. PLoS One 6:e22932
Stangl S, Kollerits B, Lamina C et al (2015) Association between apolipoprotein A-IV concentrations and chronic kidney disease in two large population-based cohorts: results from the KORA studies. J Intern Med 278:410–423
Meisinger C, Ruckert IM, Rathmann W et al (2011) Retinol-binding protein 4 is associated with prediabetes in adults from the general population: the Cooperative Health Research in the Region of Augsburg (KORA) F4 Study. Diabetes Care 34:1648–1650
WHO (1999) Definition, diagnosis and classification of diabetes mellitus and its complications. Report of a WHO consultation Part 1: Diagnosis and classification of diabetes mellitus. WHO Department of Noncommunicable Disease Surveillance, Geneva, pp 1–59
MacLean B, Tomazela DM, Shulman N et al (2010) Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26:966–968
R Development Core Team (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, available from www.r-project.org/
Abbatiello SE, Mani DR, Keshishian H, Carr SA (2010) Automated detection of inaccurate and imprecise transitions in peptide quantification by multiple reaction monitoring MS. Clin Chem 56:291–305
Mani DR, Abbatiello SE, Carr SA (2012) Statistical characterization of multiple-reaction monitoring mass spectrometry (MRM-MS) assays for quantitative proteomics. BMC Bioinformatics 13(Suppl 16):S9
Thiel S (2007) Complement activating soluble pattern recognition molecules with collagen-like regions, mannan-binding lectin, ficolins and associated proteins. Mol Immunol 44:3875–3888
Dobo J, Schroeder V, Jenny L, Cervenak L, Zavodszky P, Gal P (2014) Multiple roles of complement MASP-1 at the interface of innate immune response and coagulation. Mol Immunol 61:69–78
Frauenknecht V, Thiel S, Storm L et al (2013) Plasma levels of mannan-binding lectin (MBL)-associated serine proteases (MASPs) and MBL-associated protein in cardio- and cerebrovascular diseases. Clin Exp Immunol 173:112–120
Sildorf SM, Eising S, Hougaard DM et al (2014) Differences in MBL levels between juvenile patients newly diagnosed with type 1 diabetes and their healthy siblings. Mol Immunol 62:71–76
Bouwman LH, Eerligh P, Terpstra OT et al (2005) Elevated levels of mannose-binding lectin at clinical manifestation of type 1 diabetes in juveniles. Diabetes 54:3002–3006
Hansen TK, Thiel S, Knudsen ST et al (2003) Elevated levels of mannan-binding lectin in patients with type 1 diabetes. J Clin Endocrinol Metab 88:4857–4861
Hansen TK, Tarnow L, Thiel S et al (2004) Association between mannose-binding lectin and vascular complications in type 1 diabetes. Diabetes 53:1570–1576
Fernandez-Real JM, Straczkowski M, Vendrell J et al (2006) Protection from inflammatory disease in insulin resistance: the role of mannan-binding lectin. Diabetologia 49:2402–2411
Jenny L, Ajjan R, King R, Thiel S, Schroeder V (2015) Plasma levels of mannan-binding lectin-associated serine proteases MASP-1 and MASP-2 are elevated in type 1 diabetes and correlate with glycaemic control. Clin Exp Immunol 180:227–232
Fortpied J, Vertommen D, Van Schaftingen E (2010) Binding of mannose-binding lectin to fructosamines: a potential link between hyperglycaemia and complement activation in diabetes. Diabetes Metab Res Rev 26:254–260
Hoener MC, Brodbeck U (1992) Phosphatidylinositol-glycan-specific phospholipase D is an amphiphilic glycoprotein that in serum is associated with high-density lipoproteins. Eur J Biochem 206:747–757
Schofield JN, Stephens JW, Hurel SJ, Bell KM, deSouza JB, Rademacher TW (2002) Insulin reduces serum glycosylphosphatidylinositol phospholipase D levels in human type I diabetic patients and streptozotocin diabetic rats. Mol Genet Metab 75:154–161
Suzuki S, Suzuki C, Hinokio Y et al (2014) Insulin-mimicking bioactivities of acylated inositol glycans in several mouse models of diabetes with or without obesity. PLoS One 9:e100466
Green PH, Glickman RM, Riley JW, Quinet E (1980) Human apolipoprotein A-IV. Intestinal origin and distribution in plasma. J Clin Invest 65:911–919
Ezeh B, Haiman M, Alber HF et al (2003) Plasma distribution of apoA-IV in patients with coronary artery disease and healthy controls. J Lipid Res 44:1523–1529
Wang F, Kohan AB, Lo CM, Liu M, Howles P, Tso P (2015) Apolipoprotein A-IV: a protein intimately involved in metabolism. J Lipid Res 56:1403–1418
Maritim AC, Sanders RA, Watkins JB 3rd (2003) Diabetes, oxidative stress, and antioxidants: a review. J Biochem Mol Toxicol 17:24–38
Hellstrom PM (2013) Satiety signals and obesity. Curr Opin Gastroenterol 29:222–227
Wang F, Kohan AB, Kindel TL et al (2012) Apolipoprotein A-IV improves glucose homeostasis by enhancing insulin secretion. Proc Natl Acad Sci U S A 109:9641–9646
Pressler JW, Haller A, Sorrell J et al (2015) Vertical sleeve gastrectomy restores glucose homeostasis in apolipoprotein A-IV KO mice. Diabetes 64:498–507
Verges B (1995) Apolipoprotein A-IV in diabetes mellitus. Diabete Metab 21:99–105
Sun Z, Larson IA, Ordovas JM, Barnard JR, Schaefer EJ (2000) Effects of age, sex, and lifestyle factors on plasma apolipoprotein A-IV concentrations. Atherosclerosis 151:381–388
Kong P, Cavalera M, Frangogiannis NG (2014) The role of thrombospondin (TSP)-1 in obesity and diabetes. Adipocytes 3:81–84
Varma V, Yao-Borengasser A, Bodles AM et al (2008) Thrombospondin-1 is an adipokine associated with obesity, adipose inflammation, and insulin resistance. Diabetes 57:432–439
Kong P, Gonzalez-Quesada C, Li N, Cavalera M, Lee DW, Frangogiannis NG (2013) Thrombospondin-1 regulates adiposity and metabolic dysfunction in diet-induced obesity enhancing adipose inflammation and stimulating adipocyte proliferation. Am J Physiol Endocrinol Metab 305:E439–E450
Acknowledgements
We thank S. Becker, Research Unit Protein Science, Helmholtz Zentrum München, Germany, for her excellent technical assistance in assay development, sample preparation and machine maintenance; all members of the Institute of Epidemiology II, Helmholtz Zentrum München, Germany; and KORA field staff in Augsburg, Helmholtz Zentrum München, Germany, who planned and conducted the study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was supported by grants from the German Federal Ministry of Education and Research (BMBF) to the German Center for Diabetes Research (DZD e.V.), the Ministry of Science and Research of the State of North Rhine-Westphalia, and the German Federal Ministry of Health; and by intramural funding for Translational & Clinical Projects of the Helmholtz Zentrum München – GmbH, Germany, which is funded by the BMBF and the State of Bavaria. The KORA study was initiated and financed by the Helmholtz Zentrum München – GmbH, Germany. KORA research was also supported within the Munich Center of Health Sciences (MC-Health), Ludwig-Maximilians-Universität, Germany, as part of LMUinnovativ. The Diabetes Cohort Study was funded by the German Research Foundation (DFG; RA 459⁄2-1 to WR). The measurement of ApoA-IV by ELISA was supported by a grant from the ‘Standortagentur Tirol’ to FK.
Duality of interest statement
CvT, CHu, TdlHG, BT and SMH declare that they have a patent application ‘biomarkers of cardiometabolic diseases’ pending at the European Patent Office, comprising the biomarkers investigated here. All other authors declare that there is no duality of interest associated with their contribution to this manuscript.
Contribution statement
CHe, CM, WR, AP and BT contributed to the conception and design of the KORA F4 study; CHu, FK, CHe, WK, CM, WR, AP and BT were responsible for data acquisition; CvT, CHu, TdlHG, BT and SMH planned the investigation; CvT developed the SRM assays and performed and evaluated the SRM-MS measurements; TdlHG performed data preprocessing and statistical analysis; MW and MR contributed to data interpretation alongside the other authors. CvT, CHu and TdlHG drafted the manuscript and all authors, critically reviewed and edited the manuscript, and approved the version to be published. CvT, CHu and TdlHG are the guarantors of this work and, as such, had full access to all data in the study and take responsibility for data integrity and the accuracy of data analysis.
Additional information
Christine von Toerne and Cornelia Huth are joint first authors.
Barbara Thorand and Stefanie Hauck are joint senior authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM
(PDF 486 kb)
Rights and permissions
About this article
Cite this article
von Toerne, C., Huth, C., de las Heras Gala, T. et al. MASP1, THBS1, GPLD1 and ApoA-IV are novel biomarkers associated with prediabetes: the KORA F4 study. Diabetologia 59, 1882–1892 (2016). https://doi.org/10.1007/s00125-016-4024-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-016-4024-2