Abstract
Genome-wide association studies have revealed that single-nucleotide polymorphisms in the first intron of the gene encoding fat mass and obesity-associated protein (FTO) are robustly associated with BMI and obesity. Subsequently, this association with body weight, which is replicable across multiple populations and different age groups, has been unequivocally linked to increased food intake. Although evidence from a number of animal models with perturbed FTO expression indicates a role for FTO in energy homeostasis, to date, no conclusive link has been made between the risk alleles and FTO expression or its physiological role. FTO is a nucleic acid demethylase, and a deficiency in FTO leads to a complex phenotype highlighted by postnatal growth retardation, pointing to some fundamental developmental role. Recent emerging data now points to a role for FTO in the sensing of nutrients and the regulation of translation and growth. In this review, we explore the in vivo and in vitro evidence detailing the complex biology of FTO and discuss how these might link to the regulation of body weight.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Until quite recently, obesity was largely confined to industrialised countries of the developed world, but its prevalence is now increasing rapidly in the urban centres of emerging economies. In fact, it would not be an exaggeration to classify obesity as a major public health issue now reaching worldwide epidemic proportions [1]. The underlying cause of obesity is clearly the consumption of more energy than is expended, and the rapid changes in lifestyle and food availability over the past 30 years have undeniably driven this rise in obesity. The more difficult and far more interesting question, however, is why some people eat more than others. Individuals respond differently to ‘obesigenic’ environmental changes and an increasing weight of evidence points to a potent genetic element underlying this variation in response. Indeed, studying the correlations of BMI in monozygotic, dizygotic, biological and adopted siblings reveals the heritability of fat mass to be between 30% and 70% [2–5]. Consequently, genetic approaches offer a powerful tool for characterising the molecular and physiological mechanisms of food intake and body weight control, and allow us to understand how these may become ineffective in the obese state.
Over the past 20 years, insights from human and mouse genetics have illuminated multiple pathways within the brain that play a key role in the control of food intake (reviewed by [6]). We now know, for example, that the hypothalamic leptin–melanocortin signalling pathway is crucial for the appropriate control of food intake, with genetic disruption of most components of the pathway resulting in severe obesity in both mouse and man. However, these and other monogenic syndromes of severe obesity remain very rare. The major burden of disease is carried by those with ‘common obesity’, which until recently, largely as a result of a lack of sufficient statistical power, resisted yielding meaningful biological insights.
In 2007, the advent of technology enabling the eponymous genome-wide association studies (GWAS) led to the identification of the gene encoding fat mass and obesity-associated protein (FTO) [7–9], the first of the so-called ‘post-genomics’ obesity genes. Since then more than a hundred other candidate genes that are associated with obesity-related traits have been identified [10]. FTO remains statistically the most robust gene and with the largest effect size. However, to date, FTO’s physiological role and its link to body weight regulation remain unclear. Emerging data from our laboratory now points to a role for FTO in the sensing of nutrients and the regulation of translation and growth [11]. In this review, we explore the in vivo and in vitro evidence detailing the complex biology of FTO and discuss how this might link to the regulation of body weight.
Identification of FTO
In 2007, Frayling and colleagues performed a GWAS to identify type 2 diabetes susceptibility genes and reported that multiple single-nucleotide polymorphisms (SNPs) in the first intron of FTO were associated with disease [8]. However, the association between these FTO SNPs and type 2 diabetes disappeared after adjustment for BMI. Given that body weight is a risk factor for type 2 diabetes, it was suggested that these SNPs are actually associated with BMI [8]. Two subsequent reports, a French study [7] and a Sardinian study [9], closely followed, confirming a robust association between FTO SNPs and body weight. Many other studies have now examined and confirmed the influence of FTO variants on BMI across multiple populations and age groups (reviewed by [12]).
Recent studies have reported that the association of the FTO risk allele with the odds of obesity is attenuated by 27% in adults who are more physically active [13], highlighting the importance of exercise in influencing the ‘genetic burden’ of the FTO risk alleles. This is consistent with observations that between-person variability in obesity can also partly be explained by the genotype at the FTO locus [14]. In other words, although carriers of the FTO risk allele have an increased mean BMI, there is also increased variability of effect size around the mean, which is likely to be influenced by the impact of exercise and other environmental factors.
When the influence of FTO on body weight is studied in more detail, particularly with respect to the component aspects of the energy balance equation, it is unequivocal that the major effect of FTO ‘risk alleles’ is an increase in energy intake [15–20], with a reduction in satiety [21, 22] and possibly with an increased daily fat intake [19, 23]. In contrast, the evidence for involvement of mechanisms involving energy expenditure in the effect of FTO on human adiposity, either as altered basal energy utilisation [16, 18, 24–26] or physical activity [24, 27–29] is, to date, less compelling.
FTO or not FTO?
However, although GWAS has been incredibly powerful, providing valuable insights into the genetic architecture of complex diseases and traits, it is important to consider the limitations of what is, in effect, a gene-agnostic approach. SNPs reaching the appropriate statistical threshold for association with disease can appear anywhere in the genome, and very rarely occur within any coding sequence. The current assumption that the closest coding region, which is sometimes hundreds of kilobases away, is the likely candidate is certainly a reasonable first guess, but until empirically proven, is not necessarily true [30, 31].
In this instance, the FTO risk alleles are actually located in the first intron of the FTO gene. However, they are also very close to the transcriptional start site of RPGRIP1L (the human orthologue of mouse Ftm), which is adjacent to and coded for on the opposite DNA strand to FTO [32]. There were two main reasons why the focus of study began with FTO and not RPGRIP1L: (1) unlike FTO, expression of RPGRIP1L, which is known to localise in the primary cilia and centrosomes of ciliated cells, is not nutritionally regulated [33]; Second, defects in human RPGRIP1L exist and cause Joubert syndrome type 7 (JBTS), which presents clinically with a peculiar cerebellar and brainstem malformation and renal failure [34]. The patients do not present with any obvious bodyweight-related phenotypes, with the caveat that any potential ‘lean’ phenotype is difficult to ascertain in a healthy individual, let alone someone who is severely ill. Deletion of the mouse orthologue Ftm recapitulates the cerebral, renal and hepatic defects seen in JBTS patients [34].
There is, however, evidence for possible co-regulatory mechanisms between FTO and RPGRIP1L. Stratigopoulos et al report an overlapping regulatory region within intron 1 of FTO that contains at least two putative transcription factor binding sites (CUTL1), one of which overlaps with another obesity-associated SNP (rs8050136) [32]. It certainly is a possibility that the association between FTO SNPs and body weight regulation is mediated through changing the expression of both FTO and RPGRIP1L.
From an ‘unknown unknown’ to a nucleic acid demethylase
Because of its ‘hypothesis-free’ nature, the power of GWAS lies in uncovering potentially new biology that would not have been possible using a candidate gene approach. The problem with new biology, of course, is that by definition, little or nothing at all might be known about the gene. This was certainly true for FTO, which, when first identified, to memorably quote Frayling and colleagues from 2007, was ‘a gene of unknown function in an unknown pathway’ [8].
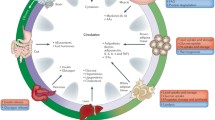
Using bioinformatic tools we and our colleagues reported that FTO belongs to the superfamily of Fe(II) 2-oxoglutarate (2-OG)-dependent dioxygenases, with highest sequence similarity to the Escherichia coli DNA repair enzyme AlkB and its mammalian AlkB homologues 2 and 3 (ABH2 and 3) [33]. We and others showed that, in an in vitro assay, FTO is able to catalyse the Fe(II)- and 2-OG-dependent demethylation of 3-methylthymine and 3-methyluracil (3meU) in single-stranded DNA and RNA to thymine and uracil, respectively (Fig. 1) [33, 35]. Subsequently, Jia et al reported that N 6-methyladenosine (6meA) in both DNA and RNA is yet another substrate of FTO, but one with 50-fold greater affinity for the enzyme compared with 3meU [36]. While 3meU is found largely in ribosomal RNA (rRNA) [37], 6meA is the most common modified nucleoside found in mRNA [38]. However, because rRNA accounts for the vast majority of total RNA, in absolute terms, there is actually a hundredfold more 3meU than 6meA in any given cell. It is still not entirely clear whether one or both of these modified bases is the endogenous substrate/s for FTO.
Biochemical function of FTO. FTO is able to catalyse the Fe(II)- and 2-OG-dependent demethylation of 3-methyluracil and N 6-methyladenine, with concomitant production of succinate, formaldehyde and carbon dioxide. 3-Methyluracil is found largely in ribosomal RNA, while N 6-methyladenine is found in mRNA, and the proposed downstream consequences of appropriate demethylation are indicated
The crystal structure of FTO is available, providing a basis for the study of structure–function relationships of FTO [39]. The FTO protein comprises two domains, an N-terminal domain containing the catalytic core, and a C-terminal domain with an as yet unknown function/role. The catalytic domain of FTO contains five obligate amino acid residues found in all members of this enzyme superfamily, which are required for 2-OG and Fe(II) binding. FTO also possesses an additional region, referred to as Loop 1, which is highly conserved among FTO proteins from different species but is absent from other members of the ALKBH family of proteins. Through steric hindrance of the substrate binding pocket, the Loop 1 region is thought to give FTO specificity for single-stranded nucleic acids during substrate binding [39].
FTO deficiency
There are several examples in metabolic disease where common variants close to a particular gene are associated with alterations in risk of common phenotypes such as fat mass or risk of obesity, whereas rare loss- or gain-of-function mutations in the same gene are associated with a more severe version of the same metabolic phenotype (e.g. MCR4 [40–43], POMC [44, 45], BDNF [46–48] and PCSK1 [49, 50]). As it turns out, however, the phenotype seen in FTO deficiency proved far more complicated.
FTO was actually first identified as part of a 1.6 Mb deletion in the Fused toes mouse mutant [51]. This deletion included not only Fto, but also Ftm, Ftl and the Iroquois B cluster, consisting of Irx3, Irx5, and Irx6. Fused toes mutants display severe developmental defects, such as left–right asymmetry [52], polydactyly and defects in hypothalamus development [52, 53]. While the homozygotes are embryonically lethal, heterozygous mice have fused digits and hyperplasia of the thymus without any metabolic alterations. In contrast, mice with a specific targeted deletion of Fto did not display these severe developmental abnormalities, but exhibited a complex phenotype of postnatal growth retardation, decreased fat and lean body mass and elevated food intake when corrected for lean body mass [54]. There was also a significant level of postnatal lethality, with only 50% of homozygous pups reaching weaning age [54, 55].
Humans with a homozygous loss-of-function R316Q mutation in FTO displayed an equally complex phenotype [56]. Like the Fto-null mice, these patients exhibited severe postnatal growth retardation. In addition, however, they also displayed microcephaly, psychomotor delay, facial dysmorphism, certain brain malformations and cardiac abnormalities. Tragically, none of the patients survived beyond the age of 3 years. The arginine at position 316 is one of the critical amino acid residues that interacts with 2-OG, which when mutated renders FTO catalytically inactive and unable to demethylate [56]. The phenotypes seen in the patients with the R316Q mutation and in the Fto-deficient mice provide direct evidence that FTO, in particular its demethylase activity, plays some fundamental role in the control of growth and development.
Fto overexpression
Church and colleagues have generated a ‘knock-in’ mouse model that carries one or two additional copies of Fto [57]. Mice overexpressing Fto showed a dramatic increase in food intake resulting in a marked increase in body weight and fat mass when they were fed either chow or a high-fat diet. There was no significant change in either energy expenditure or physical activity. Although the increase in weight with the overexpression of Fto seems consistent with FTO deficiency resulting in a ‘lean’ phenotype, the increase in food intake seen in these mice is not. Curiously, at the age of 8 weeks, mice overexpressing Fto had reduced leptin levels after an overnight fast compared with wild-type controls [57]. It should be noted that these are obese mice with increased fat mass, that in the wild-type state should normally respond with increased leptin levels. The authors of the study do comment that the hyperphagic phenotype is possibly due to an FTO-dependent reduction of leptin [57], although no mechanistic details are provided.
Tissue-specific perturbation of FTO expression
FTO is expressed ubiquitously in human and animal tissues, with the highest expression seen in the brain. Within the brain, FTO expression is relatively high in a number of hypothalamic nuclei, namely, the arcuate (ARC), paraventricular, dorsomedial, and ventromedial nuclei, where control of energy homeostasis is centred [33]. Within the ARC, expression of Fto is bi-directionally regulated as a function of nutritional status, decreasing following a 48 h fast and increasing after 10 weeks of exposure to a high-fat diet [33, 58]. If FTO had a direct action on the control of energy intake, one would predict that reducing its expression would recapitulate the fasting state, thereby resulting in an increase in food intake. This does indeed seem to be the case. Tung et al knocked down and overexpressed FTO specifically in the ARC in rats, using adeno-associated virus vectors coupled with stereotactic injections. When FTO is overexpressed, food intake is reduced. Conversely, reducing FTO expression increases food intake [58].
The region-specific manipulation of FTO provides further support for the notion that, by acting specifically in the ARC of the hypothalamus, FTO itself can influence energy homeostasis by having a direct effect on food intake. These findings are consistent with the robust association between the FTO risk alleles and increase in food intake. Mice with central nervous system (CNS)-specific deletion of Fto have now been generated [59]. Surprisingly, these brain-specific Fto-deficient mice recapitulate the phenotype of the whole body knockouts, although this has yet to be exhaustively examined [59]. This suggests that an important proportion of the function of FTO, particularly its link to the regulation of energy homeostasis, is mediated in the brain.
On a related note, in addition to BMI, the risk alleles in FTO have also been associated with a number of other, what could be considered neurological phenotypes. These include decreased brain volume both in adolescents [60] and the elderly [61], increased incidence of attention deficit-hyperactivity disorder [62], depression [63] and alcohol dependence [64], as well as an increased risk of Alzheimer’s disease [65]. It is difficult to really know at this point whether these associations are dependent or independent of FTO’s effect on BMI.
FTO is not a sensor for 2-OG
Given the evidence above that FTO can influence food intake, we began to explore whether FTO could be acting as a nutrient (in its broadest sense) sensor. Because 2-OG, a key intermediate in the citric acid cycle, is a co-substrate of FTO, we considered whether FTO functions as a sensor for intracellular concentrations of this metabolite and, thus, cellular metabolism. To this end, our laboratory developed a high-throughput fluorescence-based FTO activity assay [66]. This assay is based on the principle that RNase A selectively cleaves the phosphodiester bond 3- of unpaired pyrimidine residues and that a methyl group on N3 of the uracil base (3-meU) would inhibit the ability of RNase A to cleave. We engineered a stem–loop structure with a DNA-based stem and a loop derived from RNA, which contains a 3-meU as the sole pyrimidine in the middle of the loop. Utilising this stem–loop structure as a substrate in the FTO activity assay, we demonstrated that FTO can demethylate 3-meU in this microlitre-scale assay. As 2-OG acts as a co-substrate, we could then apply this assay to examine the enzyme kinetics of FTO with regards to its usage of 2-OG, arriving at a 2-OG K m value for FTO of 2.88 μmol/l. Since typical intracellular concentrations of 2-OG are measured to be more than tenfold higher, at around 50–100 μmol/l, it is clear that, although required for FTO activity, it is unlikely that the physiological role of FTO is to sense 2-OG [66].
FTO levels are regulated by availability of essential amino acids
In another study, we used an in vitro approach to determine which nutrients could regulate FTO concentrations at a cellular level [67]. We found that FTO mRNA and protein levels are dramatically downregulated by total amino acid deprivation in mouse hypothalamic N46 cells, mouse embryonic fibroblasts (MEFs) and in human embryonic kidney 293 (HEK 293) cells. The fall in expression of Fto mRNA is faster than its rate of natural degradation, pointing to some regulation at the transcriptional level, which is reversible upon amino acid replacement. Strikingly, this downregulation was seen only with deprivation of essential amino acids. These data suggest that there might be some interplay between FTO levels and essential amino acid availability [67].
A role for FTO in cell growth and mRNA translation
Could the regulation of FTO expression by amino acids then be linked to the growth retardation phenotype seen in FTO deficiency? In a recent study, we showed that MEFs derived from Fto −/− mice exhibit slower rates of growth and have reduced mRNA translation compared with wild-type MEFs [11]. Aminoacyl-tRNA synthetases (AARSs), which tether free AAs to their cognate tRNAs in a process known as ‘tRNA charging’, are one of the key modulators of mRNA translation. During tRNA charging, at least nine different AARSs work together as a part of large multimer complex known as the multi-synthetase complex (MSC) [68]. The maintenance of the MSC is essential for mRNA translation, and disruption of the MSC leads to a reduction in protein levels of member AARSs and a concomitant drop in AARS activity, leading to reduced mRNA translation rates [69]. Consistent with the reduced rates of translation, Fto −/− MEFs have reduced protein levels of MSC components. The defects in mRNA translation and reduced levels of MSC components in Fto −/− MEFs are rescued by re-expressing exogenous FTO in these cells [11]. These results indicate a role for FTO in the regulation of mRNA translation through maintenance of cellular MSC protein levels.
FTO links amino acid availability to mTORC1 signalling
As a key regulator of both cell growth and mRNA translation that is sensitive to amino acid levels, we measured the status of mammalian target of rapamycin complex 1 (mTORC1) signalling in Fto −/− MEFs. Cells lacking FTO display decreased activation of the mTORC1 pathway, decreased rates of mRNA translation and increased autophagy [11]. It makes mechanistic sense that these changes are likely to contribute to the stunted growth phenotype seen in Fto −/− mice [54] and in humans homozygous for loss-of-function FTO mutations [56]. The dramatic regulation of FTO by amino acids [67] appears to be necessary for the physiological response to fluctuating amino acid levels, as expression of exogenous FTO in cells renders them insensitive to amino acid deprivation by preventing the expected reduction in mTORC1 signalling [11]. Critically, using the catalytically inactive R316Q mutant of FTO [56], we showed that the link between amino acid availability and mTORC1 signalling is dependent upon the demethylase activity of FTO [11].
The role of AARSs in mTORC1 activation has recently taken centre stage, with recent reports implicating leucyl-tRNA synthetase (LRS) as a key amino acid sensor upstream of Ras-related GTPase (RAG) and mTORC1 [70, 71]. When we measured LRS levels in Fto −/− MEFs, we found that, as with the other AARSs, expression of LRS is reduced in response to a lack of FTO [11]. Thus, taken in context with these recently published data, we suggest a model whereby FTO is upstream of one of the AARSs (LRS) in the link between amino acid levels and mTORC1 signalling, leading to appropriate regulation of mRNA translation rates and cell growth (Fig. 2a). In the absence of FTO, mTORC1 signalling is dysregulated, putting cells into amino acid deprivation-like conditions even in the presence of amino acids (Fig. 2b).
Proposed model for FTO as an amino acid sensor. We suggest a model whereby FTO is upstream of AARS (LRS) in the link between amino acid levels and mTORC1 signalling, leading to appropriate levels of translation and growth (a). In the absence of FTO, mTORC1 signalling is dysregulated, putting cells into amino acid deprivation-like conditions even in the presence of amino acids (b). Solid outlines around FTO, AARS, MSC and LRS indicate maintenance of levels, while dashed outlines indicate reduced levels of expression. AAs, amino acids; AARS, amino-acyl tRNA synthetase; LRS, leucyl-tRNA synthetase; MSC, multi-synthetase complex; RAG, Ras-related GTPase; mTORC1, mammalian target of rapamycin complex 1
Adult deletion of FTO leads to loss of lean mass
Additional evidence supporting a role for FTO in the regulation of translation and growth has recently been provided [55]. When FTO is deleted in adult mice, using a tamoxifen-driven Cre recombinase conditional deletion approach, the mice begin to lose weight almost immediately. Intriguingly, there is no reduction of fat mass, with all of the change in weight down to a dramatic loss in lean mass [55]. The sudden loss of FTO expression deprives a cell of the ability to respond appropriately to amino acids, in effect depriving a cell of amino acids, which, as discussed above, would reduce mTORC1 signalling and increase autophagy (Fig. 2b). Autophagy is the basic catabolic mechanism that involves the cellular degradation of unnecessary or dysfunctional cellular components through the lysosomal machinery. This breakdown of cellular components, including proteins, can also ensure cell survival during starvation by maintaining cellular energy levels. As skeletal muscle is the largest depot of protein in the body, it appears to be the most sensitive to the sudden removal of FTO and subsequent increase in autophagy, leading to a dramatic loss of muscle mass. In contrast, when FTO is postnatally deleted specifically from the mediobasal hypothalamus, there is a more subtle weight loss, resulting from a change in food intake [55].
The 6meA methylome
A critical question not answered by these latest findings is: how does the catalytic function of FTO as a demethylase lead to its regulation of mTORC1 signalling? One possibility could involve FTO’s ability to demethylate 6meA. Two recently published studies have used antibodies against 6meA to immunoprecipitate human and mouse transcripts that carry the 6meA modification, and subjected the ‘pulled-down’ material to RNA sequencing, allowing both groups to map the presence of 6meA in a transcriptome-wide manner [72, 73]. As it turns out, 6meAs are common and are highly conserved between mice and man. Crucially, 6meAs are dynamically regulated, both in terms of development and tissue specificity. These studies have far-reaching implications, as the presence of 6meA at appropriate mRNA sites appears to play a fundamental regulatory role in mRNA translation, in addition to exerting varying effects on mRNA splicing and transport [72, 73].
To date, only one enzyme—methyltransferase like 3 (METTL3)—has been identified as catalysing the conversion of adenosine to 6meA [74]. Critically, and relevant to our discussion, two enzymes are currently known to catalyse the removal of this methyl group: ALKBH5 [75] and FTO [36, 73]. In fact, Meyer and colleagues have shown that transient overexpression of FTO in HEK 293 cells decreases the total amount of 6meAs found in the transcriptome [73]. An area of intense study is now aimed at understanding the repertoire of mRNAs that are demethylated by FTO and whether these lead to changes in rates of mRNA translation.
What is the role of FTO in obesity?
So, for the 64 million dollar question: how do these findings relate to the association between FTO SNPs and obesity? The short answer is, we still do not know. The biggest hurdle remains the fact that we do not know if or how the FTO risk alleles influence FTO. Considering the intronic location of all the FTO obesity-related SNPs (spanning across ∼40 kb), they are unlikely to cause functional mutations. Instead, the SNPs are more likely to be playing a transcriptional regulatory role, either to up- or downregulate FTO expression.
What is clear, as has been demonstrated in multiple studies, is that the association of FTO with BMI is largely driven by effects on appetite and food intake [15–20]. However, unlike the comparatively unambiguous examples from the leptin–melanocortin pathway, the study of humans and mice with disruptions in FTO has not been a panacea for our understanding of FTO function. That said, enough evidence has emerged from the complex phenotype seen in in vivo models where its expression is perturbed to support a role for FTO in the regulation of energy homeostasis. For example, Fto +/− mice are resistant to high-fat-diet-induced obesity [54]; ‘knocking-in’ one or two extra copies of Fto dose-dependently increases body weight [57]; and manipulating the expression of FTO within the hypothalamus, where it is nutritionally regulated, can influence food intake [58].
How does this in vivo evidence relate to the cellular role of FTO in the sensing of amino acids? FTO is ubiquitously expressed, and all evidence to date indicates that its role in linking amino acid levels to mTORC1 signalling would be conserved in all cell types. We argue that any apparent ‘tissue-specific’ action of FTO would thereby reflect the key functions of the tissue being studied, as opposed to any different role for FTO itself. The sudden loss of muscle mass in response to the adult-specific deletion of FTO in the mouse (see above) is one case in point. Likewise, we hypothesise that the role of FTO in the regulation of food intake is also linked to its function in the sensing of amino acids. FTO is most highly expressed in the hypothalamus [33], where the sensing of amino acid levels has been demonstrated to affect orexigenic and anorexigenic pathways controlling food intake [76]. Keeping in mind the modest effect size of around 1.5 kg per FTO risk allele in humans (with homozygote carriers being on average 3 kg heavier), we suggest that the risk alleles result in subtle variations in the expression of FTO in key appetite control centres within the brain, which are likely to influence the way in which we sense and therefore respond to amino acid levels. Human carriers for the FTO risk SNPs actually display quite a nuanced feeding phenotype compared with non-carriers. In test meals, they not only consume more calories in absolute terms, but also show an alteration in nutrient preference [77], supporting our contention that perhaps one’s FTO status can influence the direct sensing of dietary macronutrient content by the brain. The proposed cellular role for FTO in amino acid sensing provides some clues towards understanding the basis for this physiological phenomenon.
Future perspectives: is FTO a drug target?
Given the fact that 50% of the world’s population are carriers of FTO ‘risk’ alleles, and therefore have their body weight subtly influenced by FTO, a reasonable question to ask is: can FTO ever be considered a viable pharmaceutical target? Given its ubiquitous expression, the severity of the phenotype seen in human and murine FTO deficiency, and the fact that adult deletion of FTO results in a dramatic loss of muscle mass, we believe that this is unlikely. However, understanding the biology of FTO and its downstream actions could potentially reveal novel therapeutic targets in our battle against the increasing epidemic of obesity. We believe that our efforts to date in turning the statistical association of FTO into a deeper understanding of its biology could form a template of how to approach the many other emerging GWAS obesity genes of unknown function.
Abbreviations
- AARS:
-
Aminoacyl-tRNA synthetase
- FTO:
-
Fat mass and obesity-associated protein
- GWAS:
-
Genome-wide association studies
- HEK:
-
Human embryonic kidney
- JBTS:
-
Joubert syndrome type 7
- LRS:
-
Leucyl-tRNA synthetase
- 6meA:
-
N 6-Methyladenosine
- MEF:
-
Mouse embryonic fibroblast
- 3meU:
-
3-Methyluracil
- MSC:
-
Multi-synthetase complex
- mTORC1:
-
Mammalian target of rapamycin complex 1
- 2-OG:
-
2-Oxoglutarate
- SNP:
-
Single-nucleotide polymorphism
References
Zimmet P, Alberti KG, Shaw J (2001) Global and societal implications of the diabetes epidemic. Nature 414:782–787
MacDonald A, Stunkard A (1990) Body-mass indexes of British separated twins. N Engl J Med 322:1530
Maes HH, Neale MC, Eaves LJ (1997) Genetic and environmental factors in relative body weight and human adiposity. Behav Genet 27:325–351
Stunkard AJ, Foch TT, Hrubec Z (1986) A twin study of human obesity. JAMA 256:51–54
Stunkard AJ, Sorensen TI, Hanis C et al (1986) An adoption study of human obesity. N Engl J Med 314:193–198
Yeo GS, Heisler LK (2012) Unraveling the brain regulation of appetite: lessons from genetics. Nat Neurosci 15:1343–1349
Dina C, Meyre D, Gallina S et al (2007) Variation in FTO contributes to childhood obesity and severe adult obesity. Nat Genet 39:724–726
Frayling TM, Timpson NJ, Weedon MN et al (2007) A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316:889–894
Scuteri A, Sanna S, Chen WM et al (2007) Genome-wide association scan shows genetic variants in the FTO gene are associated with obesity-related traits. PLoS Genet 3:e115
Loos RJ (2012) Genetic determinants of common obesity and their value in prediction. Best Pract Res Clin Endocrinol Metab 26:211–226
Gulati P, Cheung MK, Antrobus R et al (2013) Role for the obesity-related FTO gene in the cellular sensing of amino acids. Proc Natl Acad Sci U S A 110:2557–2562
Fawcett KA, Barroso I (2010) The genetics of obesity: FTO leads the way. Trends Genet 26:266–274
Kilpelainen TO, Qi L, Brage S et al (2011) Physical activity attenuates the influence of FTO variants on obesity risk: a meta-analysis of 218,166 adults and 19,268 children. PLoS Med 8:e1001116
Yang J, Loos RJ, Powell JE et al (2012) FTO genotype is associated with phenotypic variability of body mass index. Nature 490:267–272
Cecil JE, Tavendale R, Watt P, Hetherington MM, Palmer CN (2008) An obesity-associated FTO gene variant and increased energy intake in children. N Engl J Med 359:2558–2566
Haupt A, Thamer C, Staiger H et al (2009) Variation in the FTO gene influences food intake but not energy expenditure. Exp Clin Endocrinol Diabetes 117:194–197
Jonsson A, Franks PW (2009) Obesity, FTO gene variant, and energy intake in children. N Engl J Med 360:1571–1572, author reply 1572
Speakman JR, Rance KA, Johnstone AM (2008) Polymorphisms of the FTO gene are associated with variation in energy intake, but not energy expenditure. Obesity (Silver Spring) 16:1961–1965
Timpson NJ, Emmett PM, Frayling TM et al (2008) The fat mass- and obesity-associated locus and dietary intake in children. Am J Clin Nutr 88:971–978
Wardle J, Llewellyn C, Sanderson S, Plomin R (2009) The FTO gene and measured food intake in children. Int J Obes (Lond) 33:42–45
den Hoed M, Westerterp-Plantenga MS, Bouwman FG, Mariman EC, Westerterp KR (2009) Postprandial responses in hunger and satiety are associated with the rs9939609 single nucleotide polymorphism in FTO. Am J Clin Nutr 90:1426–1432
Wardle J, Carnell S, Haworth CM, Farooqi IS, O’Rahilly S, Plomin R (2008) Obesity associated genetic variation in FTO is associated with diminished satiety. J Clin Endocrinol Metab 93:3640–3643
Tanofsky-Kraff M, Han JC, Anandalingam K et al (2009) The FTO gene rs9939609 obesity-risk allele and loss of control over eating. Am J Clin Nutr 90:1483–1488
Berentzen T, Kring SI, Holst C et al (2008) Lack of association of fatness-related FTO gene variants with energy expenditure or physical activity. J Clin Endocrinol Metab 93:2904–2908
Do R, Bailey SD, Desbiens K et al (2008) Genetic variants of FTO influence adiposity, insulin sensitivity, leptin levels, and resting metabolic rate in the Quebec Family Study. Diabetes 57:1147–1150
Goossens GH, Petersen L, Blaak EE et al (2009) Several obesity- and nutrient-related gene polymorphisms but not FTO and UCP variants modulate postabsorptive resting energy expenditure and fat-induced thermogenesis in obese individuals: the NUGENOB study. Int J Obes (Lond) 33:669–679
Hakanen M, Raitakari OT, Lehtimaki T et al (2009) FTO genotype is associated with body mass index after the age of seven years but not with energy intake or leisure-time physical activity. J Clin Endocrinol Metab 94:1281–1287
Jonsson A, Renstrom F, Lyssenko V et al (2009) Assessing the effect of interaction between an FTO variant (rs9939609) and physical activity on obesity in 15,925 Swedish and 2,511 Finnish adults. Diabetologia 52:1334–1338
Liu G, Zhu H, Lagou V et al (2010) FTO variant rs9939609 is associated with body mass index and waist circumference, but not with energy intake or physical activity in European- and African-American youth. BMC Med Gen 11:57
Cantor RM, Lange K, Sinsheimer JS (2010) Prioritizing GWAS results: a review of statistical methods and recommendations for their application. Am J Hum Genet 86:6–22
Yeo GS (2011) Where next for GWAS? Brief Funct Genom 10:51
Stratigopoulos G, Padilla SL, LeDuc CA et al (2008) Regulation of Fto/Ftm gene expression in mice and humans. Am J Physiol Regul Integr Comp Physiol 294:R1185–R1196
Gerken T, Girard CA, Tung YC et al (2007) The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 318:1469–1472
Delous M, Baala L, Salomon R et al (2007) The ciliary gene RPGRIP1L is mutated in cerebello-oculo-renal syndrome (Joubert syndrome type B) and Meckel syndrome. Nat Genet 39:875–881
Jia G, Yang CG, Yang S et al (2008) Oxidative demethylation of 3-methylthymine and 3-methyluracil in single-stranded DNA and RNA by mouse and human FTO. FEBS Lett 582:3313–3319
Jia G, Fu Y, Zhao X et al (2011) N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol 7:885–887
Kowalak JA, Pomerantz SC, Crain PF, McCloskey JA (1993) A novel method for the determination of post-transcriptional modification in RNA by mass spectrometry. Nucleic Acids Res 21:4577–4585
Desrosiers R, Friderici K, Rottman F (1974) Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc Natl Acad Sci U S A 71:3971–3975
Han Z, Niu T, Chang J et al (2010) Crystal structure of the FTO protein reveals basis for its substrate specificity. Nature 464:1205–1209
Chambers JC, Elliott P, Zabaneh D et al (2008) Common genetic variation near MC4R is associated with waist circumference and insulin resistance. Nat Genet 40:716–718
Loos RJ, Lindgren CM, Li S et al (2008) Common variants near MC4R are associated with fat mass, weight and risk of obesity. Nat Genet 40:768–775
Vaisse C, Clement K, Guy-Grand B, Froguel P (1998) A frameshift mutation in human MC4R is associated with a dominant form of obesity. Nat Genet 20:113–114
Yeo GS, Farooqi IS, Aminian S, Halsall DJ, Stanhope RG, O’Rahilly S (1998) A frameshift mutation in MC4R associated with dominantly inherited human obesity. Nat Genet 20:111–112
Challis BG, Pritchard LE, Creemers JW et al (2002) A missense mutation disrupting a dibasic prohormone processing site in pro-opiomelanocortin (POMC) increases susceptibility to early-onset obesity through a novel molecular mechanism. Hum Mol Genet 11:1997–2004
Krude H, Biebermann H, Luck W, Horn R, Brabant G, Gruters A (1998) Severe early-onset obesity, adrenal insufficiency and red hair pigmentation caused by POMC mutations in humans. Nat Genet 19:155–157
Gray J, Yeo GS, Cox JJ et al (2006) Hyperphagia, severe obesity, impaired cognitive function, and hyperactivity associated with functional loss of one copy of the brain-derived neurotrophic factor (BDNF) gene. Diabetes 55:3366–3371
Han JC, Liu QR, Jones M et al (2008) Brain-derived neurotrophic factor and obesity in the WAGR syndrome. N Engl J Med 359:918–927
Thorleifsson G, Walters GB, Gudbjartsson DF et al (2009) Genome-wide association yields new sequence variants at seven loci that associate with measures of obesity. Nat Genet 41:18–24
Benzinou M, Creemers JW, Choquet H et al (2008) Common nonsynonymous variants in PCSK1 confer risk of obesity. Nat Genet 40:943–945
Jackson RS, Creemers JW, Ohagi S et al (1997) Obesity and impaired prohormone processing associated with mutations in the human prohormone convertase 1 gene. Nat Genet 16:303–306
Peters T, Ausmeier K, Ruther U (1999) Cloning of Fatso (Fto), a novel gene deleted by the Fused toes (Ft) mouse mutation. Mamm Genome Off J Int Mamm Genome Soc 10:983–986
Anselme I, Laclef C, Lanaud M, Ruther U, Schneider-Maunoury S (2007) Defects in brain patterning and head morphogenesis in the mouse mutant Fused toes. Dev Biol 304:208–220
van der Hoeven F, Schimmang T, Volkmann A, Mattei MG, Kyewski B, Ruther U (1994) Programmed cell death is affected in the novel mouse mutant Fused toes (Ft). Development 120:2601–2607
Fischer J, Koch L, Emmerling C et al (2009) Inactivation of the Fto gene protects from obesity. Nature 458:894–898
McMurray F, Church CD, Larder R et al (2013) Adult onset global loss of the fto gene alters body composition and metabolism in the mouse. PLoS Genet 9:e1003166
Boissel S, Reish O, Proulx K et al (2009) Loss-of-function mutation in the dioxygenase-encoding FTO gene causes severe growth retardation and multiple malformations. Am J Hum Genet 85:106–111
Church C, Moir L, McMurray F et al (2010) Overexpression of Fto leads to increased food intake and results in obesity. Nat Genet 42:1086–1092
Tung YC, Ayuso E, Shan X et al (2010) Hypothalamic-specific manipulation of Fto, the ortholog of the human obesity gene FTO, affects food intake in rats. PLoS One 5:e8771
Gao X, Shin YH, Li M, Wang F, Tong Q, Zhang P (2010) The fat mass and obesity associated gene FTO functions in the brain to regulate postnatal growth in mice. PLoS One 5:e14005
Melka MG, Gillis J, Bernard M et al (2013) FTO, obesity and the adolescent brain. Hum Mol Genet 22:1050–1058
Ho AJ, Stein JL, Hua X et al (2010) A commonly carried allele of the obesity-related FTO gene is associated with reduced brain volume in the healthy elderly. Proc Natl Acad Sci U S A 107:8404–8409
Choudhry Z, Sengupta SM, Grizenko N et al (2013) Association between obesity-related gene FTO and ADHD. Obesity (Silver Spring). doi:10.1002/oby.20444
Samaan Z, Anand S, Zhang X et al (2012) The protective effect of the obesity-associated rs9939609 A variant in fat mass- and obesity-associated gene on depression. Mol Psychiatry. doi:10.1038/mp.2012.160
Wang L, Liu X, Luo X, Zeng M, Zuo L, Wang KS (2013) Genetic variants in the fat mass- and obesity-associated (FTO) gene are associated with alcohol dependence. J Mol Neurosci. doi:10.1007/s12031-013-0044-2
Reitz C, Tosto G, Mayeux R, Luchsinger JA, Group N-LNFS, Alzheimer’s Disease Neuroimaging I (2012) Genetic variants in the Fat and Obesity Associated (FTO) gene and risk of Alzheimer’s disease. PLoS One 7:e50354
Ma M, Harding HP, O’Rahilly S, Ron D, Yeo GS (2012) Kinetic analysis of FTO (fat mass and obesity-associated) reveals that it is unlikely to function as a sensor for 2-oxoglutarate. Biochem J 444:183–187
Cheung MK, Gulati P, O’Rahilly S, Yeo GS (2012) FTO expression is regulated by availability of essential amino acids. Int J Obes (Lond) 37:744–747
Quevillon S, Robinson JC, Berthonneau E, Siatecka M, Mirande M (1999) Macromolecular assemblage of aminoacyl-tRNA synthetases: identification of protein-protein interactions and characterization of a core protein. J Mol Biol 285:183–195
Park SG, Ewalt KL, Kim S (2005) Functional expansion of aminoacyl-tRNA synthetases and their interacting factors: new perspectives on housekeepers. Trends Biochem Sci 30:569–574
Bonfils G, Jaquenoud M, Bontron S, Ostrowicz C, Ungermann C, De Virgilio C (2012) Leucyl-tRNA synthetase controls TORC1 via the EGO complex. Mol Cell 46:105–110
Han JM, Jeong SJ, Park MC et al (2012) Leucyl-tRNA synthetase is an intracellular leucine sensor for the mTORC1-signaling pathway. Cell 149:410–424
Dominissini D, Moshitch-Moshkovitz S, Schwartz S et al (2012) Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485:201–206
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR (2012) Comprehensive analysis of mRNA methylation reveals enrichment in 3' UTRs and near stop codons. Cell 149:1635–1646
Bokar JA, Shambaugh ME, Polayes D, Matera AG, Rottman FM (1997) Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA 3:1233–1247
Zheng G, Dahl JA, Niu Y et al (2013) ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell 49:18–29
Cota D, Proulx K, Smith KA et al (2006) Hypothalamic mTOR signaling regulates food intake. Science 312:927–930
Tanaka T, Ngwa JS, van Rooij FJ et al (2013) Genome-wide meta-analysis of observational studies shows common genetic variants associated with macronutrient intake. Am J Clin Nutr 97:1395–1402
Funding
The authors are financially supported by the UK Medical Research Council, the European Commission Seventh Framework Programme (EU FP7- HEALTH- 2009- 241592 EurOCHIP, EU FP7- FOOD- 266408 Full4Health) and the Helmholtz Alliance for Imaging and Curing Environmental Metabolic Diseases (ICEMED).
Duality of interest
The authors declare that there is no duality of interest associated with this manuscript.
Contribution statement
PG and GSHY were responsible for the conception and design of the manuscript, drafting the article, revising it critically for intellectual content and approving the final version to be published.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Gulati, P., Yeo, G.S.H. The biology of FTO: from nucleic acid demethylase to amino acid sensor. Diabetologia 56, 2113–2121 (2013). https://doi.org/10.1007/s00125-013-2999-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-013-2999-5