Abstract
Aims/hypothesis
We determined whether single nucleotide polymorphisms (SNPs) previously associated with diabetogenic traits improve the discriminative power of a type 2 diabetes genetic risk score.
Methods
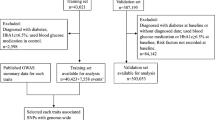
Participants (n = 2,751) were genotyped for 73 SNPs previously associated with type 2 diabetes, fasting glucose/insulin concentrations, obesity or lipid levels, from which five genetic risk scores (one for each of the four traits and one combining all SNPs) were computed. Type 2 diabetes patients and non-diabetic controls (n = 1,327/1,424) were identified using medical records in addition to an independent oral glucose tolerance test.
Results
Model 1, including only SNPs associated with type 2 diabetes, had a discriminative power of 0.591 (p < 1.00 × 10−20 vs null model) as estimated by the area under the receiver operator characteristic curve (ROC AUC). Model 2, including only fasting glucose/insulin SNPs, had a significantly higher discriminative power than the null model (ROC AUC 0.543; p = 9.38 × 10−6 vs null model), but lower discriminative power than model 1 (p = 5.92 × 10−5). Model 3, with only lipid-associated SNPs, had significantly higher discriminative power than the null model (ROC AUC 0.565; p = 1.44 × 10−9) and was not statistically different from model 1 (p = 0.083). The ROC AUC of model 4, which included only obesity SNPs, was 0.557 (p = 2.30 × 10−7 vs null model) and smaller than model 1 (p = 0.025). Finally, the model including all SNPs yielded a significant improvement in discriminative power compared with the null model (p < 1.0 × 10−20) and model 1 (p = 1.32 × 10−5); its ROC AUC was 0.626.
Conclusions/interpretation
Adding SNPs previously associated with fasting glucose, insulin, lipids or obesity to a genetic risk score for type 2 diabetes significantly increases the power to discriminate between people with and without clinically manifest type 2 diabetes compared with a model including only conventional type 2 diabetes loci.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Type 2 diabetes is a complex disease characterised by chronically elevated fasting or post-challenge systemic glucose concentrations [1]. Heritability studies suggest that genetic factors influence the risk of developing the disease. Indeed, multiple loci pre-disposing to type 2 diabetes have been discovered recently, many of which have emerged from genome-wide association studies [2].
Several studies have examined the clinical value of variants known to predispose to type 2 diabetes by analysing their ability to discriminate between people with or without pre-existing diabetes, or to predict development of the disease [3–5]. Although opinion is divided on the clinical value of these genetic risk scores (GRS), in their present form they do not meaningfully improve the predictive power over risk scores comprised solely of established non-genetic risk factors [3–5].
As with type 2 diabetes, major advances have also been made in identifying gene variants that influence some of the major risk factors for type 2 diabetes, e.g. chronic obesity, dyslipidaemia and hyperglycaemia. Indeed, we have previously studied the level of type 2 diabetes risk associated with several of these loci in the Northern Sweden Health and Disease Study (NSHDS) [6, 7]. However, to our knowledge, the discriminative or predictive power of multi-trait GRSs for type 2 diabetes have not yet been reported on.
The purpose of this study was to test whether gene variants that are not explicitly defined as loci predisposing to type 2 diabetes, but have been shown to influence antecedent traits (i.e. hyperglycaemia, hyperinsulinaemia, dyslipidaemia or obesity) can be used to improve the discriminative power of a GRS for type 2 diabetes compared with a score comprised solely of specific type 2 diabetes loci. We did not seek to establish the comparative power of this score with non-GRSs for type 2 diabetes, in part because cross-sectional studies are inadequate for this purpose.
Methods
Participants
Participants (effective n = 1,327 type 2 diabetic patients, 1,424 controls) were Swedish adults from the county of Västerbotten in northern Sweden, and selected from the NSHDS, a prospective cohort study of common diseases [8]. All living participants provided written informed consent and the Research Ethics Committee of Umeå University Hospital approved all aspects of the study.
Ascertainment of type 2 diabetes cases and controls
The case ascertainment methods have been described in detail previously [7]. In brief, cases were those participants with a documented clinical history of type 2 diabetes in addition to an independent OGTT result consistent with a diagnosis of type 2 diabetes, according to the WHO criteria [1]. Conversely, controls were those participants who did not have a documented clinical diagnosis of diabetes (of any type), were not taking glucose-lowering medications, and who had fasting and 2 h glucose values below the diagnostic thresholds for diabetes [1].
Clinical measures
The clinical methods have been described in detail previously [8]. Briefly, height, weight, glucose concentrations and lipid fractions were measured using standard methods (Table 1). The purpose of providing this information is to emphasise that type 2 diabetic patients and controls differed significantly in levels of the traits related to the single nucleotide polymorphisms (SNPs) focused on in this study. Blood was drawn after an overnight fast from an antecubital vein; a second sample was drawn 2 h after a 75 g oral glucose load.
Selection of SNPs and genetic analyses
The type 2 diabetes and lipid SNPs are those for which replication results were in the public domain as of May 2008 (Fig. 1a–d). Additional obesity and fasting glucose/insulin SNPs were identified through participation in the Genetic Investigation of ANthropometric Traits consortium [9, 10] and the Meta-Analyses of Glucose and Insulin-related traits Consortium [11], respectively. Thus, because of the timing of genotyping relative to progress in the field, and to a limited extent because of assay design limitations, several previously replicated SNPs could not be included.
Individual odds ratios (95% CIs) for type 2 diabetes for each of the a type 2 diabetes (n = 17), b fasting glucose/insulin (n = 13), c dyslipidaemia (n = 26) and d obesity (n = 17) SNPs included in these analyses (n = 73). Allele frequencies were calculated in the control group. All SNPs are located on the plus strand (HapMap CEU, Phase II + III, release 27, NCBI build 36). LOC287761 is discontinued, but was included here as it was documented as a replicated locus when this study began. Data were adjusted for age and sex. Odds ratios between the WFS1 rs10010131 and most obesity SNPs with type 2 diabetes have been previously reported for this sample (7, 8). The association between the TCF7L2 SNP and type 2 diabetes has been previously reported for a sub-sample of the case–control cohort examined here [35]
DNA was extracted from peripheral white blood cells [6, 7]. Genomic DNA samples were subsequently diluted to 4 ng/µl. Genotyping was performed using Taqman MGB chemistry (Applied Biosystems, Foster City, CA, USA) or Sequenom iPLEX (Sequenom, Hamburg, Germany). Genotyping success rates were >95%.
Statistical analysis
Analyses were conducted in SAS version 9.2 (SAS Institute, Cary, NC, USA). A likelihood ratio test with 1 df was used to test Hardy–Weinberg equilibrium (all SNPs fulfilled Hardy–Weinberg expectations; Bonferroni corrected p > 0.05). SNPs were individually tested (additive SNP models) for association with type 2 diabetes using unconditional logistic regression from which ORs and 95% CI were estimated (Fig. 1a–d). In the discriminative power comparison models, effect alleles for all SNPs are coded in a manner consistent with the Diabetes Genetics Replication and Meta-Analysis (DIAGRAM) database [12] or the findings from MAGIC [11]. Regression models were adjusted for age and sex. In an ethnically homogeneous population such as this, biological traits such as obesity and dyslipidaemia are unlikely to confound the effects of gene variants on diabetes risk, once age has been accounted for. Therefore, because we sought to exploit such effects, no adjustments for intermediate diabetes risk factors were made. The basic genetic model (model 1) included 17 variants previously associated with type 2 diabetes. Comparison models included previously associated fasting glucose/insulin (n = 13) (model 2), lipid (n = 26) (model 3) [13] or obesity (n = 17) (model 4) [9, 10, 14, 15] SNPs. Finally, a model containing all 73 SNPs (model 5) was compared with the null model and with model 1. The discriminative power of the five different SNP models was estimated by comparing the area under the receiver operator characteristic curves (ROC AUC) for each model. Because the majority of SNPs studied here are in low linkage disequilibrium, we were unable to accurately impute missing genotypes using methods based on linkage disequilibrium. Therefore, we calculated the mean genotype at each locus in cases and controls separately, and exchanged missing genotypes for the relevant mean value for that SNP. Alleles were rounded to the nearest whole unit. Prior to imputing genotypes, we tested whether genotyping failure rates differed between the type 2 diabetes group and controls, as this could have biased tests of association using imputed data. There was no evidence of such selection bias (association of missing genotypes with diabetes: OR 1.00, 95% CI 0.98–1.01). ROC AUCs were compared using the methods described by DeLong et al. [16]. In these analyses, the null model included no predictor variables. Prior to entering the SNPs into the ROC models, we ensured each risk allele was coded in a manner consistent with the DIAGRAM database [12] and used the relevant random effects ORs from this dataset to derive weightings for each risk allele. This was achieved by multiplying each risk allele by the log of its OR in the DIAGRAM dataset. Four SNPs were unavailable in DIAGRAM. In these cases, we used the average effect estimate for the SNPs within the relevant trait group (i.e. diabetes, glucose, lipid or obesity SNPs). The GRSs were computed by summing the weighted risk alleles across all loci for each trait (or for the full model for all traits). Overall, weighting SNP models did not materially alter the discriminative power compared with the unweighted models.
Results
Participant characteristics are shown in Table 1. Figure 1a–d shows ORs (95% CIs) for each of the 73 SNPs. In general, the risk estimates in this cohort for SNPs previously associated with type 2 diabetes were directionally consistent with previous reports. As shown in Fig. 1a–d, few SNPs were individually statistically associated with type 2 diabetes (at p < 0.05).
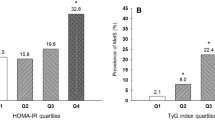
Figure 2 shows the relationships between each GRS (expressed in quartiles of the GRS) and type 2 diabetes risk. For each of the GRSs, statistically significant relationships with type 2 diabetes risk were observed (p < 0.05). The odds of type 2 diabetes per quartile of the score was: for the type 2 diabetes GRS OR 1.25 (95% CI 1.17–1.34); for the glucose GRS OR 1.08 (95% CI 1.01–1.15); for the lipid GRS OR 1.07 (95% CI 1.00–1.14); for the obesity GRS OR 1.14 (95% CI 1.07–1.22); and for the full GRS OR 1.33 (95% CI 1.24–1.43). With the exceptions of the glucose and lipid SNP GRSs (p = 0.09 and p = 0.06, respectively), individuals in the highest quartile of each GRS were at statistically greater risk of type 2 diabetes than those in the first quartile. For example, those in the highest quartile of the full GRS had a 2.40-fold higher odds of type 2 diabetes than those in the lowest quartile (p = 3.50 × 10−16); for the type 2 diabetes GRS, the respective odds of type 2 diabetes was 1.96 (p = 3.42 × 10−8).
Odds ratios (95% CI) for type 2 diabetes relative to the number of risk alleles across 73 SNP loci. a Glucose and insulin SNPs, b obesity SNPs, c lipid SNPs, d type 2 diabetes SNPs and e all SNPs. Data are adjusted for age and sex. Missing genotypes were imputed as described in the Methods. Type 2 diabetes patients, n = 1,327, controls, n = 1,424
Five separate ROC models were run to compare the discriminative power of the different SNP sets. Model 1, including only type 2 diabetes-associated SNPs, had a discriminative power of 0.591 (p < 1.00 × 10−20 vs null model) as estimated by the ROC AUC. Model 2, including only fasting glucose/insulin SNPs, had significantly higher discriminative power than the null model (ROC AUC 0.543; p = 9.38 × 10−6 vs null model), but lower discriminative power than model 1 (p = 5.92 × 10−5 vs model 1). Model 3, with only lipid-associated SNPs, had significantly higher discriminative power than the null model (ROC AUC 0.565; p = 1.44 × 10−9) and was not statistically different from model 1 (p = 0.083). The ROC AUC of model 4, which included only obesity SNPs, was 0.557 (p = 2.30 × 10−7 vs null model), which was smaller than model 1 (p = 0.025). Finally, the model including all SNPs yielded a significant improvement in discriminative power compared with the null model (p < 1.0 × 10−20) and model 1 (p = 1.32 × 10−5); its ROC AUC was 0.626. Figure 3 shows the ROC AUCs for all SNPs compared with only conventional type 2 diabetes SNPs.
Power to discriminate between type 2 diabetes cases and controls for GRSs comprised of type 2 diabetes variants (dashed line) or all variants (solid line) expressed as ROC AUCs. ROC AUCs for type 2 diabetes loci and for all loci are 0.591 and 0.626, respectively (p = 1.32 × 10−5 for difference). Data are unadjusted
Discussion
Our findings show that inclusion of genetic information from loci previously associated with quantitative risk factors for type 2 diabetes, but not primarily with diabetes, significantly increases the power to discriminate between people with and without clinically manifest type 2 diabetes. This emphasises the multi-factorial nature of type 2 diabetes and highlights the important potential role in disease development played by loci that do not reach a level of genome-wide significance in type 2 diabetes scans.
Our study was based on the premise that some loci capable of influencing diabetes risk and thus contributing to the discriminative power of type 2 diabetes GRSs have weak effects on type 2 diabetes individually, falling, as a result, below the stringent significance thresholds used in genome-wide scans, which means they have not previously been identified as diabetes-predisposing loci. We hypothesised that some of the loci reliably associated with traits that predispose to type 2 diabetes might, by virtue of this association, also raise the risk of type 2 diabetes. Hyperglycaemia is the cardinal feature of type 2 diabetes, providing sufficient justification for including glucose-raising alleles in a GRS for type 2 diabetes. However, it is worth noting that not all glucose-raising loci appear to influence type 2 diabetes risk [11], possibly because some loci may cause modest elevations in glucose concentrations that do not worsen over time, as observed in maturity-onset diabetes of the young [17]. Obesity is also a well established risk factor for diabetes, as illustrated in clinical trials where weight loss interventions have substantially reduced the incidence of the disease in high risk individuals [18, 19]. For dyslipidaemia, the mechanisms of association with type 2 diabetes primarily involve insulin resistance caused by the infiltration of insulin-sensitive tissues by triacylglycerol and other lipid metabolites [20, 21]. Two important organs in this regard are muscle and liver, the former being important because of its predominance as a site for glucose uptake and metabolism, and the latter because of its major role in glucose production. Studies in non-obese individuals with a strong family history of type 2 diabetes have provided experimental evidence that elevations in NEFA directly impair muscle glycogen synthesis and glucose uptake, and induce muscle, hepatic and adipose tissue insulin resistance in a genetically determined manner [3]. Prospective epidemiological studies indicate that dyslipidaemia early in life [22, 23] or during adulthood [24] raises the risk of developing type 2 diabetes later in life, but such associations may be driven by obesity [22] rather than a lipid-specific genetic defect. Nevertheless, animal and human studies suggest a shared genetic basis for diabetes and dyslipidaemia. For example, expression of the HDL-associated apolipoprotein M is completely abolished in the liver of mice lacking the HNF1A gene [25]; mutations in HNF1A also cause maturity-onset diabetes of the young class 3 [26]. Epidemiological studies have also identified genetic loci that influence dyslipidaemia and glucose homeostasis or type 2 diabetes [25, 27–30]. Although these joint relationships are unlikely to result from confounding, it remains unclear whether they reflect causal relationships between dyslipidaemia and diabetes, or pure genetic pleiotropy. Similarly, one cannot easily determine whether the cumulative association between lipid loci and diabetes in the present study is attributable to (1) dyslipidaemia mediating the effects of the genotypes on diabetes risk; (2) purely pleiotropic effects; or (3) a combination of these explanations. Notwithstanding these limitations of interpretation, the use of a priori biological information to help filter genome-wide scan results minimises the multiple testing burden inherent in hypothesis-free whole-genome genetic association studies and may raise the prior probability of association, hence helping to preserve statistical power.
To minimise over-fitting of our models, prior evidence of association from the DIAGRAM dataset [12] was used to code the effect alleles in the ROC analyses presented here. Fitting the alleles in this way did not result in markedly different ROC AUCs than when alleles were fitted directly to the current dataset, indicating that our data are unlikely to be markedly over- or under-fitted. We were unable to include all currently identified risk alleles for the traits of interest, partly because the rate at which new risk variants have been discovered out-paced our study and partly because resources were limited. Although initially presumed otherwise [31], it is unlikely that LOC387761 is a true diabetes locus and could thus have been excluded from our models without diminishing the discriminative power. It is also important to highlight that there are many other antecedent traits for type 2 diabetes beyond those studied here, e.g. HbA1c, fibrinogen and adiponectin; if variants associated with such traits were to be included in a GRS, the discriminative power would probably increase further.
The derivation of GRSs using the approach applied here requires complete genotype data in the population in which the score is computed. Because the genotype success rates were less than perfect in our study (as in virtually all studies) and genotyping failures were randomly distributed across the selection of SNPs in this cohort, it was necessary and appropriate to impute missing genotypes. The alternative would have been to use a sample set in which directly genotyped data were available for all SNPs. However, because missing genotype data were random across the study sample, around half of all participants were missing data on at least one of the 73 SNPs. Thus, use of only the complete directly genotyped subgroups would have resulted in a considerable loss of statistical power and could have led to biased conclusions about the magnitude of association for the GRSs.
A further consideration is whether our findings are likely to be attributable to confounding. With the exception of linkage disequilibrium between the non-functional observed and functional unobserved loci, statistical associations between germline genetic variants such as SNPs and phenotypes are generally robust to confounding in ethnically homogeneous cohorts such as that studied here. Therefore, the associations reported here are unlikely to be prone to confounding.
Our study is clearly a hypothesis-generating effort and robust type 2 diabetes effect sizes for most of the GRSs of interest in this report are absent from the published literature. As such, meaningful a priori power calculations could not be performed for this study and post-hoc power calculations would be inappropriate, as discussed at length elsewhere [32–34]. The fact that most of the associations reported for the GRS models are highly statistically significant indicates that our study was well powered to detect the observed effects (which is a circular argument and one important reason why post hoc power calculations are often discouraged).
Finally, owing to the cross-sectional study design, we were unable to calculate the reclassification index attributable to the different genetic models, which would be valuable when considering a possible clinical application. One should also consider that in cross-sectional studies, in which cases and controls are phenotypically highly distinct, estimates of discriminative power may exceed estimates of predictive power derived from prospective studies.
In conclusion, polymorphisms that affect diabetogenic traits, but which are not conventionally considered to be diabetes-predisposing loci, significantly improve the discriminative power of a conventional GRS for type 2 diabetes. This is the case even though, on an individual basis, most variants have weak effects that were not statistically associated with type 2 diabetes in our study. Nevertheless, the discriminative power of the GRS remains below a level many would consider clinically useful; thus, validated non-genetic prediction algorithms remain the most appropriate tools for predicting type 2 diabetes in the clinical setting.
Abbreviations
- DIAGRAM:
-
Diabetes Genetics Replication and Meta-Analysis
- GRS:
-
Genetic risk score
- NSHDS:
-
Northern Sweden Health and Disease Study
- ROC AUC:
-
Area under the receiver operator characteristic curve
- SNP:
-
Single nucleotide polymorphism
References
World Health Organization (1999) Definitions, diagnosis and classification of diabetes mellitus and its complications. Part 1: diagnosis & classification of diabetes mellitus. World Health Organization, Geneva
Prokopenko I, McCarthy MI, Lindgren CM (2008) Type 2 diabetes: new genes, new understanding. Trends Genet 24:613–621
Lango H, Palmer CN, Morris AD et al (2008) Assessing the combined impact of 18 common genetic variants of modest effect sizes on type 2 diabetes risk. Diabetes 57:3129–3135
Lyssenko V, Jonsson A, Almgren P et al (2008) Clinical risk factors, DNA variants, and the development of type 2 diabetes. N Engl J Med 359:2220–2232
Meigs JB, Shrader P, Sullivan LM et al (2008) Genotype score in addition to common risk factors for prediction of type 2 diabetes. N Engl J Med 359:2208–2219
Renstrom F, Payne F, Nordstrom A et al (2009) Replication and extension of genome-wide association study results for obesity in 4923 adults from northern Sweden. Hum Mol Genet 18:1489–1496
Franks PW, Rolandsson O, Debenham SL et al (2008) Replication of the association between variants in WFS1 and risk of type 2 diabetes in European populations. Diabetologia 51:458–463
Hallmans G, Agren A, Johansson G et al (2003) Cardiovascular disease and diabetes in the Northern Sweden Health and Disease Study Cohort—evaluation of risk factors and their interactions. Scand J Public Health Suppl 61:18–24
Willer CJ, Speliotes EK, Loos RJ et al (2009) Six new loci associated with body mass index highlight a neuronal influence on body weight regulation. Nat Genet 41:25–34
Lindgren CM, Heid IM, Randall JC et al (2009) Genome-wide association scan meta-analysis identifies three loci influencing adiposity and fat distribution. PLoS Genet 5:e1000508
Dupuis J, Langenberg C, Prokopenko I et al (2010) New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat Genet 42:105–116
Zeggini E, Scott LJ, Saxena R et al (2008) Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat Genet 40:638–645
Kathiresan S, Willer CJ, Peloso GM et al (2009) Common variants at 30 loci contribute to polygenic dyslipidemia. Nat Genet 41:56–65
Thorleifsson G, Walters GB, Gudbjartsson DF et al (2009) Genome-wide association yields new sequence variants at seven loci that associate with measures of obesity. Nat Genet 41:18–24
Benzinou M, Creemers JW, Choquet H et al (2008) Common nonsynonymous variants in PCSK1 confer risk of obesity. Nat Genet 40:943–945
DeLong ER, DeLong DM, Clarke-Pearson DL (1988) Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44:837–845
Fajans SS, Bell GI, Polonsky KS (2001) Molecular mechanisms and clinical pathophysiology of maturity-onset diabetes of the young. N Engl J Med 345:971–980
Knowler WC, Barrett-Connor E, Fowler SE et al (2002) Reduction in the incidence of type 2 diabetes with lifestyle intervention or metformin. N Engl J Med 346:393–403
Tuomilehto J, Lindstrom J, Eriksson JG et al (2001) Prevention of type 2 diabetes mellitus by changes in lifestyle among subjects with impaired glucose tolerance. N Engl J Med 344:1343–1350
Randle PJ, Garland PB, Hales CN, Newsholme EA (1963) The glucose fatty-acid cycle. Its role in insulin sensitivity and the metabolic disturbances of diabetes mellitus. Lancet 1:785–789
Petersen KF, Shulman GI (2006) Etiology of insulin resistance. Am J Med 119:S10–S16
Franks PW, Hanson RL, Knowler WC et al (2007) Childhood predictors of young-onset type 2 diabetes. Diabetes 56:2964–2972
Morrison JA, Glueck CJ, Horn PS, Wang P (2010) Childhood predictors of adult type 2 diabetes at 9- and 26-year follow-ups. Arch Pediatr Adolesc Med 164: 53–60
Wilson PW, Meigs JB, Sullivan L, Fox CS, Nathan DM, D'Agostino RB Sr (2007) Prediction of incident diabetes mellitus in middle-aged adults: the Framingham Offspring Study. Arch Intern Med 167:1068–1074
Richter S, Shih DQ, Pearson ER et al (2003) Regulation of apolipoprotein M gene expression by MODY3 gene hepatocyte nuclear factor-1alpha: haploinsufficiency is associated with reduced serum apolipoprotein M levels. Diabetes 52:2989–2995
Frayling TM, Evans JC, Bulman MP et al (2001) Beta-cell genes and diabetes: molecular and clinical characterization of mutations in transcription factors. Diabetes 50(Suppl 1):S94–S100
Hicks AA, Pramstaller PP, Johansson A et al (2009) Genetic determinants of circulating sphingolipid concentrations in European populations. PLoS Genet 5:e1000672
Beer NL, Tribble ND, McCulloch LJ et al (2009) The P446L variant in GCKR associated with fasting plasma glucose and triglyceride levels exerts its effect through increased glucokinase activity in liver. Hum Mol Genet 18:4081–4088
Orho-Melander M, Melander O, Guiducci C et al (2008) Common missense variant in the glucokinase regulatory protein gene is associated with increased plasma triglyceride and C-reactive protein but lower fasting glucose concentrations. Diabetes 57:3112–3121
Vaxillaire M, Cavalcanti-Proenca C, Dechaume A et al (2008) The common P446L polymorphism in GCKR inversely modulates fasting glucose and triglyceride levels and reduces type 2 diabetes risk in the DESIR prospective general French population. Diabetes 57:2253–2257
Sladek R, Rocheleau G, Rung J et al (2007) A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 445:881–885
Goodman SN, Berlin JA (1994) The use of predicted confidence intervals when planning experiments and the misuse of power when interpreting results. Ann Intern Med 121:200–206
Smith AH, Bates MN (1992) Confidence limit analyses should replace power calculations in the interpretation of epidemiologic studies. Epidemiology 3:449–452
Detsky AS, Sackett DL (1985) When was a “negative” clinical trial big enough? How many patients you needed depends on what you found. Arch Intern Med 145:709–712
Mayans S, Lackovic K, Lindgren P et al (2007) TCF7L2 polymorphisms are associated with type 2 diabetes in northern Sweden. Eur J Hum Genet 15:342–346
Acknowledgements
We thank the study participants, the staff of the Umeå Medical Biobank for the preparation of materials and staff of the Västerbottens Intervention Programme for data collection. We also thank M. Sjögren and M. Orho-Melander for facilitating aspects of the Sequenom genotyping, and both the Västerbotten Diabetes Registry (DIVE; chaired by O. R. Rolandsson) for access to phenotypic data and S. Steiginga for assistance with the figures. We thank S. Lindström for helpful feedback on genotype imputation methods. The study was funded by project grants from Novo Nordisk, the Swedish Heart–Lung Foundation, the Swedish Diabetes Association, Påhlssons Foundation, the Swedish Research Council, Umeå University Career Development Award and The Heart Foundation of Northern Sweden (all to P. W. Franks). Other project grants were from Tore Nilsons Foundation (to F. Renström) and the Wellcome Trust grant 077016/Z/05/Z (to I. Barroso). F. Renström was supported by a postdoctoral stipend from the Swedish Heart–Lung Foundation.
Duality of interest
I. Barroso owns stock in Incyte and GlaxoSmithkline. All other authors declare that there is no duality of interest associated with this manuscript.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Consortia
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Fontaine-Bisson, B., Renström, F., Rolandsson, O. et al. Evaluating the discriminative power of multi-trait genetic risk scores for type 2 diabetes in a northern Swedish population. Diabetologia 53, 2155–2162 (2010). https://doi.org/10.1007/s00125-010-1792-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-010-1792-y