Abstract
Aims/hypothesis
Pro-inflammatory cytokines involved in the pathogenesis of type 1 diabetes deplete endoplasmic reticulum (ER) Ca2+ stores, leading to ER-stress and beta cell apoptosis. However, the cytokine-induced ER-stress response in beta cells is atypical and characterised by induction of the pro-apoptotic PKR-like ER kinase (PERK)–C/EBP homologous protein (CHOP) branch of the unfolded protein response, but defective X-box binding protein 1 (XBP1) splicing and activating transcription factor 6 activation. The purpose of this study was to overexpress spliced/active Xbp1 (XBP1s) to increase beta cell resistance to cytokine-induced ER-stress and apoptosis.
Methods
Xbp1s was overexpressed using adenoviruses and knocked down using small interference RNA in rat islet cells. In selected experiments, Xbp1 was also knocked down in FACS-purified rat beta cells and rat fibroblasts. Expression and production of XBP1s and key downstream genes and proteins was measured and beta cell function and viability were evaluated.
Results
Adenoviral-mediated overproduction of Xbp1s resulted in increased XBP1 activity and induction of several XBP1s target genes. Surprisingly, XBP1s overexpression impaired glucose-stimulated insulin secretion and increased beta cell apoptosis, whereas it protected fibroblasts against cell death induced by ER-stress. mRNA expression of Pdx1 and Mafa was inhibited in cells overproducing XBP1s, leading to decreased insulin expression. XBP1s knockdown partially restored cytokine/ER-stress-driven insulin and Pdx1 inhibition but had no effect on cytokine-induced ER-stress and apoptosis.
Conclusions/interpretation
XBP1 has a distinct inhibitory role in beta cell as compared with other cell types. Prolonged XBP1s production hampers beta cell function via inhibition of insulin, Pdx1 and Mafa expression, eventually leading to beta cell apoptosis.
Similar content being viewed by others
Introduction
Type 1 diabetes is an autoimmune disease triggered by progressive destruction of beta cells during insulitis. Pro-inflammatory cytokines released by immune cells invading the islets contribute to beta cell dysfunction and apoptosis during insulitis [1]. We have previously shown that cytokines deplete endoplasmic reticulum (ER) Ca2+ stores, leading to activation of the unfolded protein response (UPR) [2]. The UPR seeks to restore normal ER function by attenuating translation, increasing folding capacity through upregulation of ER chaperones and promoting degradation of misfolded proteins via the ER-associated degradation (ERAD) pathway. If the UPR is unable to restore cell function, it eventually triggers the apoptosis cascade [3]. The UPR is mediated by the activation of three ER transmembrane sensor proteins: the activating transcription factor (ATF)6α, the PKR-like ER kinase (PERK) and the endoribonuclease inositol-requiring enzyme (IRE)1α [3].
IRE1α splices Xbp1 mRNA, leading to translation of the spliced/active form of the X-box binding protein-1 (XBP1s) transcription factor [4]. XBP1s induces production of chaperones and ERAD components, which respectively promote folding and degradation of misfolded proteins, thereby contributing to cellular recovery following ER-stress [5, 6].
Although it is well established that cytokines induce ER-stress in beta cells [3], the role of ER-stress in cytokine-induced beta cell apoptosis remains controversial. Thus while some studies suggest that C/EBP homologous protein (CHOP) [7] and binding immunoglobulin protein (BIP) [8] play pro- and anti-apoptotic roles in this process respectively, others failed to observe any association between ER-stress markers and cytokine-induced beta cell apoptosis [9–11]. We observed that the cytokine-induced ER-stress response is atypical and characterised by strong induction of the pro-apoptotic PERK–ATF4–CHOP branch of the UPR, but with defective XBP1 splicing and ATF6 activation [2, 12]. We also showed that IFN-γ alone decreases expression of Xbp1s and several ER chaperones [12]. We hypothesised that the defective ATF6 and XBP1s activation, and subsequent deficient production of ER proteins may deprive beta cells of a key mechanism for survival, increasing their susceptibility to cytokine/nitric oxide-mediated ER-stress and apoptosis.
The aim of this study was to increase XBP1s levels in order to improve protein-folding in the ER, and therefore protect beta cells against ER-stress-induced apoptosis. We generated adenoviruses encoding the spliced/active form of XBP1 (XBP1s) in rat dispersed islets and FACS-purified beta cells, and studied the molecular consequences of this overproduction, as well as its effects on beta cell function and viability. Surprisingly, XBP1s overproduction inhibited beta cell function and insulin expression, and triggered beta cell apoptosis, whereas it protected rat fibroblasts from cyclopiazonic acid (CPA)-induced apoptosis. XBP1s knockdown experiments suggest that this could be due to downregulation of the beta cell-specific transcription factors pancreatic and duodenal homeobox 1 (PDX1) and musculoaponeurotic fibrosarcoma oncogene homologue A (MAFA). Altogether our data suggest that XBP1s is not crucial for primary beta cell survival during ER-stress induced by cytokines or CPA, and that it plays a distinct inhibitory role in beta cells compared with other cell types.
Methods
Cell culture
Male Wistar rats (Charles River Laboratories, Brussels, Belgium) were housed and manipulated according to the guidelines of the Belgian Regulations for Animal Care; all performed experiments were approved by the local Ethical Committee. Rat islets were isolated by collagenase digestion followed by hand-picking under a stereomicroscope. Islets were then dispersed [13] and primary beta cells purified by autofluorescence-activated cell sorting (FACS Aria; BD Bioscience, San Jose, CA, USA) [14]. The preparations used in the present study contained 88.1 ± 4.5% beta cells (n = 5). Dispersed islets or FACS-purified beta cells were pre-cultured in complete beta cell medium supplemented with 5% heat-inactivated FBS from 20 to 48 h for recovery [15]. Experiments were then conducted in medium without serum [2, 13]. Rat fibroblast 208F cells (ECACC, Salisbury, UK) were cultured in DMEM supplemented with 10% FCS [16].
Cytokines and ER-stressors
The following cytokine concentrations were used: recombinant human IL-1β (a kind gift from C. W. Reinolds, National Cancer Institute, Bethesda, MD, USA) at 50 U/ml; and recombinant rat IFN-γ (R&D systems, Abingdon, UK) at 500 U/ml (0.036 µg/ml) [17, 18]. The sarco/endoplasmic reticulum calcium ATPase-2 inhibitor, cyclopiazonic acid (CPA) (Sigma-Aldrich, Bornem, Belgium), which induces rapid depletion of ER Ca2+ thus leading to ER-stress was dissolved in DMSO and used at a concentration of 25 μmol/l in beta cells and 50 μmol/l in 208F [2, 19]
Generation of recombinant adenoviruses and cell infection
To produce XBP1 in insulin-secreting cells, we generated recombinant adenoviruses comprising fragments of the mouse Xbp1 (mXbp1) mRNA (gi|18139942|gb|AF443192.1) [20]. The following primers (Electronic supplementary material [ESM] Fig. 1) containing the SalI (forward) and HindIII (reverse) cloning sites (indicated by underlining) were used to amplify the complete spliced mXBP1 (1,363 bp): forward 5'-ATCGTCGACCCGGGACTACAGGACCAATA-3′; reverse 5'-GCGCAAGCTT ATGTGATGGTCAGGGAAAGG-3′. Using an alternate reverse primer 5'-CGCAAGCTTGGCAAGAAGGTGGTCTCAGA-3′, we were able to amplify either 1,129 bp corresponding to a truncated fragment of the spliced mXbp1 or the unspliced mXbp1 (ESM Fig. 1). These PCR fragments were then inserted in the pcDNA3 vector. Truncated spliced mXbp1 and unspliced mXbp1 were distinguished by sequencing. The PCR fragment encoding the unspliced Xbp1 mRNA is 5′-truncated and its splicing by endogenous IRE1α activity results in translation of a truncated and less active mXBP1 (ESM Fig. 1). Adenoviruses were generated using the pShuttle/pAdEasy-1 vector system [17, 21]. Adenoviruses expressing the enhanced green fluorescent protein (Ad-eGFP) or adenoviruses expressing the luciferase protein (Ad-LUC) were used as controls [22]. Rat dispersed islets/FACS-purified beta cells were infected as previously described [22].
Transient transfection and luciferase assays
The reporter plasmid containing the luciferase gene under the control of five UPR elements was kindly provided by R. Prywes (Columbia University Biological Sciences, New York, USA). At 24 h after infection, dispersed islets cells were co-transfected with the UPR elements vector and the internal control pRL-CMV plasmid. Luciferase activities in the cell lysates were determined 48 h later in a luminometer (TD-20/20; Turner Designs, Sunnyvale, CA, USA) using the dual-luciferase reporter assay system according to the manufacturer’s instructions (Promega, Madison, WI, USA) [23]. Luciferase activities were normalised by renilla activities of the co-transfected plasmid (pRL-CMV).
Insulin secretion measurements
At 48 h after infection, dispersed islet cells were washed in KRBH–HEPES buffer and static insulin secretion was assessed for 2 h in KRBH supplemented with either 1.7 mmol/l glucose or 16.7 mmol/l glucose + 10 µmol/l forskolin. Insulin release was measured by radioimmunoassay [24].
RNA interference
Small interference RNA (siRNA) transfection in rat dispersed beta cells was conducted according to a protocol recently developed in our laboratory [15] using DharmaFECT 1 (Termo Scientific, Chicago, IL, USA) with a final concentration of 30 nmol/l siRNA. The efficiency of transfection was >90% [15]. A control siRNA (Allstars Negative Control; Qiagen, Venlo, the Netherlands) and two different BLOCK-iT Stealth Select siRNA (Invitrogen, Paisley, UK) designed against rat Xbp1 were used (sequences, see ESM Table 1). Cells were then cultured for a 48 h recovery period before being collected or exposed to CPA/cytokines as indicated.
Assessment of cell viability and caspase 3 activity
The percentage of viable, apoptotic and necrotic cells was determined using the DNA-binding dyes propidium iodide (5 µg/ml; Sigma-Aldrich) and Hoechst 33342 (5 µg/ml; Sigma-Aldrich) [2, 25]. The cells were examined by inverted fluorescence microscopy (Axiovert 200; Carl Zeiss, Zaventem, Belgium). A kit for live cells (NunView 488 Caspase-3 Kit; Biotium, Hayward, CA, USA) was used to determine caspase-3 activation [26]. A minimum of 500 cells were counted in each experimental condition and viability or caspase activity was evaluated by two independent observers, one of them unaware of sample identity.
mRNA extraction and quantitative RT-PCR
Poly(A) + mRNA was isolated from dispersed islet cells using a kit (Dynabead mRNA DIRECT; Invitrogen) and reverse-transcribed as previously described [2]. Quantitative PCR was performed using IQ SYBR Green Supermix (BioRad, Nazareth Eke, Belgium) in an IQ5 instrument (BioRad). Expression values were corrected to the housekeeping gene glyceraldehyde-3-phosphate dehydrogenase. Cytokine or CPA treatment does not modify Gapdh expression in insulin-producing cells under the present experimental conditions [18, 27]. The primers used in this study are described in the ESM Table 1.
Western blot analysis
Cells were washed with cold PBS and lysed with Laemmli buffer. Lysates were resolved by 12% SDS-PAGE and transferred to polyvinylidene fluoride membranes. Immunoblot analyses were performed using an antibody directed against XBP1 (sc-346; Santa Cruz Biotechnology, Santa Cruz, CA, USA) and normalised using anti-β-actin (Cell Signaling, Boston, MA, USA). Horseradish peroxidase-conjugated goat anti-rabbit IgG was used as a secondary antibody (Santa Cruz Biotechnology). Immunoreactive bands were revealed using a chemiluminescent substrate (SuperSignal West Femto; Termo Scientific, Chicago, IL, USA), detected using a LAS-3000 CCD camera (Fujifilm Europe-GE Healthcare Europe, Diegem, Belgium).
Immunofluorescence
Dispersed islet cells were grown on glass culture slides (BD Biosciences Europe, Erembodegem, Belgium), fixed for 15 min in fresh 4% paraformaldehyde, rinsed in PBS and permeabilised for 5 min in PBS-Triton X-100 0.1%. Slides were then blocked using PBS-goat serum 5% and incubated overnight at 4°C in the presence of polyclonal rabbit anti-BCL2-associated X protein (BAX) (1/1000; Santa Cruz Biotechnology) and monoclonal mouse anti-cytochrome c or anti-ATP Synthase β (1/1000; BD Biosciences). Cells were washed and further exposed for 1 h to appropriate Alexa fluor 488 or 555-conjugated antibodies (1/1000; N. V. Invitrogen, Merelbeke, Belgium). After washing, cells were stained with Hoechst 33342, mounted, analysed and photographed using fluorescence microscopy (Axio Imager; Carl Zeiss).
Statistical analysis
Data are presented as means ± SEM. Comparisons were performed by two-tailed paired Student’s t test or by ANOVA followed by t tests with Bonferroni correction for multiple comparison. A p value of ≤ 0.05 was considered statistically significant.
Results
Adenovirus-mediated Xbp1s overexpression induces XBP1 activity
To assess the role of XBP1s on beta cell function and survival, we generated several adenovirus vectors leading to expression of the following Xbp1 mRNAs (ESM Fig. 1): (1) a full-length active (spliced form) of mouse Xbp1 mRNA (Ad-XBP1s: 371 amino acids); (2) a 3′ truncated fragment of the spliced Xbp1 mRNA (Ad-tXBP1s: 326 amino acids); and (3) the unspliced Xbp1 mRNA (Ad-XBP1u: 267 amino acids). As additional controls, Ad-eGFP or Ad-LUC were also used.
Rat dispersed islets and FACS-purified beta cells infected with Ad-eGFP indicated an efficiency of infection >95% (data not shown). Transduction with Ad-XBP1s resulted in a strong dose-dependent expression of mouse Xbp1s mRNA (Fig. 1a). As expected, Ad-XBP1s infection resulted in the appearance of an approximately 55 kDa band, corresponding to the full-length active/spliced form of XBP1 (Fig. 1b). Ad-t-XBP1s infection led to strong production of an N-terminal truncated form of XBP1 about 50 kDa, while Ad-XBP1u infection resulted in weaker production of t-XBP1s, probably due to processing of a fraction of the unspliced Xbp1 mRNA by endogenous IRE1α activity. The unspliced form of the XBP1 protein was not detectable, which is in accordance with data indicating that the unspliced mRNA is unstable [28]. Western blot analyses of XBP1s after Ad-XBP1s infection (multiplicity of infection [MOI] 1, 2, 5 or 15) in dispersed islet cells demonstrated dose-dependent increase in the levels of the protein (Fig. 1c).
XBP1s overproduction in rat islet cells using an adenoviral strategy. Dispersed islet cells were infected with Ad-eGFP, Ad-LUC and adenoviruses encoding full-length spliced XBP1 (Ad-XBP1s), truncated spliced XBP1 (Ad-tXBP1s) or unspliced XBP1 (Ad-XBP1u) at increasing MOIs, as indicated. Samples were collected for analyses 72 h after infection. a Quantitative RT-PCR for mouse spliced Xbp1 and the housekeeping gene Gapdh indicates a dose-dependent increase in the expression of Xbp1s mRNA. Data are means ± SEM of four independent experiments. ***p < 0.001 vs control. b Western blot analyses using an antibody directed against XBP1 reveal the presence of the endogenous (wtXBP1s) spliced XBP1 (lanes 1–2, 9), the exogenous full-length XBP1 (arrowhead wtXBP1s, lanes 3–4) or the N-terminal truncated form of XBP1 (arrowhead tXBP1s, lanes 5–8). Blot is representative of three independent experiments. c Dose-dependent XBP1s protein overproduction in response to Ad-XBP1s infection. Blot is representative of four independent experiments. d 24 h after infection, cells were transfected with a plasmid expressing the reporter gene luciferase under the control of five UPR elements. Luciferase activity was evaluated 48 h later as described in “Methods”. Results are means ± SEM of seven independent experiments. ***p < 0.001 vs non-infected control. e Data represent mRNA expression of Pdi-p5 (black bars), Sec61α (striped bars), Edem (white bars) and P58 IPK (squared bars), normalised per Gapdh expression and expressed as fold change compared with non-infected cells. Results are means ± SEM of five independent experiments. *p < 0.05 and **p < 0.01 vs non-infected control
To check whether the viral XBP1 protein drove XBP1 activity in dispersed islet cells, we studied the expression of the reporter gene luciferase under the control of five UPR elements, known to be the target binding site for XBP1s [28]. Infection using Ad-XBP1s dose-dependently increased XBP1 activity, while infection with Ad-eGFP had no effect (Fig. 1d). Interestingly, infection with Ad-tXBP1s induced a two- to threefold activation of the reporter, indicating that the truncated form of XBP1 still conserved a residual transactivating capacity. Ad-XBP1u infection was unable to increase XBP1 activity. Importantly, XBP1s overproduction significantly increased mRNA expression of XBP1 target genes [5] Edem (also known as Edem1), Sec61α, Pdi-P5 and P58 IPK (Fig. 1e).
XBP1s overproduction induces beta cell apoptosis
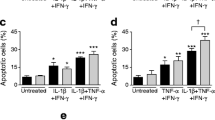
We next studied the effects of XBP1s overproduction on the viability of dispersed islet cells. Unexpectedly, XBP1s overproduction increased apoptosis in dispersed islet cells. This effect was already apparent 3 days after infection at MOI 15 (Fig. 2a), reaching significant levels at MOI 2, 5 and 15 after 5 days, with a dose-dependent effect of the virus (Fig. 2b). The control adenoviruses Ad-eGFP, Ad-LUC and Ad-XBP1u had no effect on viability, while Ad-tXBP1s-infected islet cells showed a 60% increase in cell death at MOI 15 (Fig. 2b). Linear regression analysis of all data revealed a clear correlation (R 2 = 0.65; p = 0.039) between the UPR activity driven by XBP1s and the prevalence of islet cells apoptosis (data not shown). Similar results were obtained in FACS-purified beta cells (Fig. 2c). XBP1s overproduction had no significant effect on necrosis in dispersed islets or FACS-purified beta cells (data not shown). XBP1s overproduction dose-dependently increased caspase 3 activity in dispersed islets, reaching significant levels at MOI 5 and 15 (Fig. 2d). We also assessed the effects of XBP1s overproduction on cytokine- and CPA- or tunicamycin-induced apoptosis, and observed that Ad-XBP1s-infected cells were neither sensitised nor protected against 48 h exposure to IL-1β + IFN-γ (Fig. 2e) or 24 h exposure to CPA or tunicamycin (Fig. 2f).
XBP1s overproduction induces beta cell apoptosis. Dispersed rat islets (a, b, d–f), FACS-purified beta cells (c) or rat fibroblasts (g) were infected or not (NI) with Ad-eGFP, Ad-LUC and adenoviruses encoding full-length spliced XBP1 (Ad-XBP1s) and truncated spliced XBP1 (Ad-tXBP1s) or unspliced XBP1 (Ad-XBP1u) at different MOIs, as indicated. Prevalence of apoptosis was evaluated by Hoechst 33342 and propidium iodide (HO-PI) 3 (a) or 5 (b, c) days after infection. Data are expressed as the percentage of apoptotic cells over the total number of cells counted ± SEM. d Caspase 3 activity was evaluated by fluorescence at 5 days of infection using the NuncView reporter system. Data are expressed as the percentage of caspase-positive cells over the total number of cells counted ± SEM. e 48 h after infection, cells were left untreated (black bars) or treated for 48 h with IL-1β + IFN- γ (white bars). f 48 h after infection, cells were left untreated (black bars) or treated for 24 h with CPA (white bars) or tunicamycin (striped bars). g Rat fibroblasts were either left untreated (black bars) or treated with CPA (white bars) for 48 h. Apoptosis was evaluated by HO-PI. Data (e–g) are expressed as the percentage of apoptotic cells over the total number of cells counted ± SEM. a–g Results are means of four to seven independent experiments. b–d *p < 0.05, **p < 0.01 and ***p < 0.001 vs respective non-infected control; (e–g) *p < 0.05 and ***p < 0.001 vs respective untreated condition; † p < 0.05 vs non-infected condition; ‡ p < 0.05 vs non-infected CPA-treated condition
In contrast to beta cells and in accordance with previous studies [29, 30], XBP1s overproduction did not induce apoptosis in the 208F rat fibroblast cell line. In fact, it protected these cells against apoptosis induced by 48 h of CPA treatment (Fig. 2g).
To exclude the possibility that XBP1s overproduction triggered apoptosis through non-specific ER-stress, we studied mRNA expression of Bip (also known as Hspa5), Chop (also known as Ddit3) and endogenous rat Xbp1 in infected cells. Ad-XBP1 infection (MOI 15) did not modify Chop or Bip expression, contrasting with the effect of 24 h exposure to CPA (ESM Fig. 2). Interestingly, expression of the endogenous rat Xbp1s mRNA was significantly decreased following Ad-XBP1s infection, suggesting a negative feedback action by the exogenous mXBP1s. Together, these data indicate that XBP1s does not induce apoptosis through increased ER-stress.
Upon induction of apoptosis through the intrinsic mitochondrial pathway, the pro-apoptotic protein BAX translocates to the mitochondria and oligomerises with BAK, forming a channel that promotes mitochondrial outer membrane permeabilisation and leads to the release of cytochrome c from the mitochondria to the cytosol [31]. We studied localisation of BAX and cytochrome c in Ad-XBP1s infected cells by immunofluorescence. Upon apoptosis (determined by nucleus morphology using DNA labelling by Hoechst), islet cells displayed a discrete punctate BAX staining that colocalised with the mitochondrial marker ATP-synthase β, compared with the widely spread staining observed in live cells (Fig. 3a). Interestingly, the mitochondrial morphology changed drastically between live and apoptotic cells, the latter showing disintegration of the tubular mitochondrial network and formation of punctiform, fragmented mitochondria. Cytochrome c staining was diffuse and cytosolic in apoptotic cells, whereas it was discrete and typically mitochondrial in live cells (similar to ATP synthase β staining). XBP1s overproduction clearly increased the number of cells displaying mitochondrial BAX and cytosolic cytochrome c staining (Fig. 3b). In contrast, XBP1s overproduction (ESM Fig. 3a) significantly decreased cytochrome c release by more than 60% in fibroblasts treated with CPA for 48 h (ESM Fig. 3b, c).
XBP1s overproduction increases BAX translocation and cytochrome c release in dispersed islets. Dispersed rat islet cells were infected or not (NI) with Ad-LUC or adenoviruses encoding full-length spliced XBP1 (Ad-XBP1s) at MOI 5. Five days after infection, cells were fixed and analysed by immunofluorescence. a Representative field showing a live (L) and an apoptotic cell (A) stained using anti-ATPsynthase β, anti-BAX and the DNA labelling chemical Hoechst 33342, as indicated. Magnification ×100; white scale bar, 10 µm. b Cells were stained using anti-cytochrome c and BAX, and counterstained with Hoechst 33342. Arrows indicate cells that have lost mitochondrial cytochrome c staining, cells with BAX mitochondrial staining, as opposed to normal cytosolic staining, and (Hoechst staining) typical apoptotic nuclei or cells which have lost DNA staining. Merged image combines cytochrome c, BAX and Hoechst staining. White scale bar, 40 µm. Images are representative fields of four independent experiments
XBP1s overproduction impairs insulin secretion and expression
We evaluated the effect of XBP1s overproduction on insulin secretion 3 days after infection, a time-point when XBP1s did not significantly increase apoptosis (Fig. 2a). Compared with non-infected or Ad-LUC-infected rat dispersed islets, Ad-XBP1s-infected cells displayed significantly reduced glucose-stimulated insulin secretion at MOIs 5 and 15 (Fig. 4a). XBP1s dose-dependently decreased insulin 2 (Fig. 4b) and insulin 1 mRNA expression (data not shown). Computational analysis of the insulin promoter did not reveal the presence of UPREs in its sequence (data not shown), suggesting that the effect of XBP1s is indirect. Insulin expression depends on three key transcription factors, namely PDX1, neurogenic differentiation 1 (NEUROD1) and MAFA [32, 33]. XBP1s overproduction downregulated Mafa and Pdx1 expression, but not Neurod1, although there was a tendency toward decreased expression at MOI 15 (Fig. 4b). The expression of Glut2 (also known as Slc2a2), a known PDX1/MAFA target gene [34], was also inhibited (Fig. 4b).
XBP1 overproduction impairs insulin secretion and decreases expression of insulin, Pdx1, Mafa and Glut2 mRNAs. Dispersed rat islets were infected with Ad-LUC or adenoviruses encoding full-length spliced XBP1 (Ad-XBP1s) at MOI 5 or 15, as indicated. a 48 h after infection, insulin secretion was measured under basal (1.7 mmol/l glucose; white bars) and stimulated (16.7 mmol/l glucose + 10 μmol/l forskolin; black bars) conditions. The results are means ± SEM of four independent experiments, each performed in duplicate. **p < 0.01 vs control stimulated secretion. b mRNA was extracted 72 h after infection and real-time PCR experiments performed to evaluate expression of insulin 2, Pdx1, Mafa, Neurod1 and Glut2 in Ad-LUC (MOI 15; black bars) or Ad-XBP1s-infected cells at MOI 5 (striped bars) or MOI 15 (white bars). Data were normalised per Gapdh expression and expressed as fold changes compared with non-infected cells. Results are means ± SEM of five independent experiments. **p < 0.01 and ***p < 0.001 vs non-infected control
XBP1 knockdown partially prevents cytokine/CPA-induced insulin mRNA decrease
To confirm the potential role of XBP1s in controlling expression of MafA and Pdx1, we knocked-down Xbp1 using a siRNA strategy. Two different siRNAs knocked-down Xbp1s mRNA and protein by >80% following 14 h of CPA treatment in INS-1E cells (data not shown). Subsequent experiments in dispersed islets were performed using the siRNA2. XBP1 siRNA transfection decreased basal Xbp1s mRNA expression by 40% and abolished CPA- or IL-1β + IFN-γ-induced Xbp1s overexpression (Fig. 5a). This resulted in complete blockade of CPA-driven XBP1s protein overproduction (Fig. 5b). We studied the impact of blocking XBP1s on the expression of insulin, Pdx1, MafA and Glut2. As previously described [17, 27], the expression of these genes was severely decreased upon treatment with CPA or IL-1β + IFN-γ. Knocking-down XBP1s partially restored insulin 2 (Fig. 5c) and Pdx1 expression (Fig. 5d), but failed to revert cytokine- and CPA-induced inhibition of Mafa (Fig. 5e) and Glut2 (Fig. 5f) expression. XBP1 knockdown did not affect CPA-induced Chop (Fig. 5g) and Bip expression (data not shown). It neither modified cytokine- nor CPA-induced islet cell apoptosis (Fig. 5h).
XBP1 knockdown partially restores cytokine- and CPA-mediated downregulation of insulin and Pdx1 mRNA. a, c–e Dispersed islet cells were left untransfected (black bars) or transfected using a control siRNA (striped bars) or a rat Xbp1 siRNA (white bars). 48 h after transfection, cells were treated with DMSO (control), CPA (25 μmol/l) or IL-1β (50 U/ml) + IFN-γ (500 U/ml) for 24 h. b Western blot analysis of XBP1s and β-actin levels. Blot is representative of four independent experiments. mRNA expression of rat (r)Xbp1s (a), insulin 2 (c), Pdx1 (d), Mafa (e), Glut2 (f) and Chop (g). Data normalised per Gapdh expression and expressed as fold changes compared with non-transfected cells. Results are means ± SEM of five independent experiments. a–g *p < 0.05, **p < 0.01 and ***p < 0.001 vs respective non-treated and non-transfected control; † p < 0.05 between cells transfected with XBP1 siRNA and the similarly treated non-transfected cells. h Dispersed islet cells were transfected using the control siRNA (black bars) or XBP1 siRNA2 (white bars). At 48 h after transfection, cells were treated with DMSO (control), CPA (25 μmol/l) or IL-1β (50 U/ml) + IFN-γ (500 U/ml) for 24 h. Apoptosis was evaluated by Hoechst 33342 and propidium iodide (HO-PI). Data are expressed as the percentage of apoptotic cells over the total number of cells counted. Results are means ± SEM of five independent experiments. *p < 0.05 and **p < 0.01 vs respective non-treated control
Discussion
In an attempt to better characterise the role of XBP1 in pancreatic beta cells, we used an adenoviral approach to overexpress the active/spliced form of XBP1 (Xbp1s) in rat islet cells, leading to increased XBP1 activity and expression of several XBP1s target genes. Surprisingly, prolonged XBP1s overproduction impaired insulin expression and glucose-stimulated insulin secretion, and led to BAX translocation, cytochrome c release, activation of caspase 3 and beta cell apoptosis. Neither Xbp1s overexpression nor knockdown modified cytokine- or CPA-induced ER-stress and apoptosis. In contrast, XBP1s overproduction had no effect on basal rat fibroblasts survival and protected these cells against CPA-induced apoptosis. XBP1s overproduction strongly decreased insulin, Pdx1 and Mafa expression in beta cells and XBP1 knockdown partially restored cytokine- or CPA-driven insulin and Pdx1 inhibition. Taken together, our results suggest that XBP1s does not play a key role in beta cell survival during ER-stress induced by CPA or cytokines. However, sustained production of XBP1s inhibits beta cell function and eventually triggers the mitochondrial pathway of apoptosis, probably secondary to inhibition of Pdx1 and Mafa expression.
Based on studies performed in other cell types, the IRE1α–XBP1 pathway is usually considered to be a protective pathway of the UPR [35–37]. We observed that XBP1s overproduction protects rat fibroblasts against CPA-induced apoptosis. In contrast, prolonged XBP1s overproduction induces beta cell apoptosis. This is not due to generation of non-specific ER-stress, since expression of the ER-stress markers Bip and Chop were not modified. In addition, viruses encoding truncated forms of XBP1 induced little (Ad-tXBP1s) or no apoptosis (Ad-XBP1u), and there was a close correlation between XBP1s activity and apoptosis, indicating that the increased apoptosis is specifically triggered by XBP1s. It is likely that the discrepancy between fibroblasts and beta cells is due to the role, discovered here, of XBP1s in inhibiting the expression of Pdx1 and Mafa, two key beta cell-specific transcription factors that are crucial for maintaining function [34] and, at least for Pdx1, survival [38, 39]. Xbp1 knockdown partially reversed Pdx1 downregulation, but failed to prevent cytokine- or CPA-induced apoptosis, suggesting that Pdx1 downregulation is not a major determinant of beta cell apoptosis in response to cytokines or CPA. Our data indicate that XBP1s overproduction triggers the intrinsic pathway of apoptosis, but further studies need to be done to elucidate the exact mechanism underlying the pro-apoptotic effect.
XBP1 knockdown had no effect on beta cell viability under control conditions or when the cells were treated with cytokines or CPA, suggesting that XBP1 does not affect the primary beta cell capacity to face some forms of ER-stress. This hypothesis is supported by the fact that Xbp1 knockdown had no effect on the expression of the ER-stress markers Bip and Chop, the latter a transcription factor shown to be increased in beta cells under pro-apoptotic ER-stress [15, 19]. This is in line with previous studies showing that XBP1s is not required for normal development of islets [40] while it is essential to development of liver [35] and plasma cells [41]. A recent report also suggests that XBP1s is dispensable in neurons [42], which share many features with beta cells [43, 44].
We and others have previously shown that the IRE1α–XBP1s pathway of the UPR controls insulin expression, mainly through IRE1α-mediated degradation of insulin mRNA [27, 45, 46]. In the present study we demonstrate another level of regulation of insulin expression by the UPR, via XBP1s-dependent downregulation of Pdx1 and Mafa. XBP1s did not affect stability of insulin 1 or 2 mRNA (data not shown), suggesting a transcriptional event. XBP1s strongly decreased Mafa and Pdx1 expression, thereby suppressing the cooperative action between these transcription factors and Neurod1 in stimulating insulin transcription [32]. Moreover, Xbp1 knockdown partially prevented cytokines and CPA-mediated decrease in insulin and Pdx1 mRNA expression [18, 27], supporting the hypothesis that XBP1s inhibits Pdx1 expression. XBP1s dose-dependently inhibited stimulated but not basal insulin secretion, suggesting that XBP1s interferes with the glucose signalling pathways leading to insulin release. XBP1s also downregulates expression of the glucose transporter Glut2. This finding is in accordance with a role of XBP1s in inhibiting Pdx1 and Mafa expression [34], and may also account for the effect of XBP1s on beta cell function, as GLUT2 plays a critical role in glucose sensing in beta cells [47].
While XBP1 knockdown partially restored insulin and Pdx1 expression in presence of CPA or cytokines, Mafa and Glut2 expression remained suppressed. Further studies are required to clarify the mechanisms involved in this apparent dissociation between the roles of PDX1 and MAFA in the regulation of key beta cell genes.
The classical view on the role of the IRE1α–XBP1s pathway in the UPR is that this pathway augments the folding capacity of the ER through increased expression of ER chaperones and stimulates degradation of misfolded proteins by the ERAD pathway [5, 6]. However, overexpression of Xbp1s in primary beta cells induced only a moderate (≤ twofold) increase in the expression of its target genes. A similar pattern was observed in fibroblasts [5], suggesting that XBP1s is necessary, but not sufficient to fully stimulate expression of its target genes. Accumulating evidence suggests that a cross-talk exists between the ATF6α and IRE1α–XBP1s pathways to achieve maximal induction of their downstream target genes [5, 6, 48]. A recent study revealed deleterious effects of ATF6α overexpression on beta cell function and expression of insulin, PDX1 and MAFA in INS-1 cells [49]. Together with our data in primary cells, this suggests that the ATF6α and XBP1 pathways may have redundant roles in the regulation of insulin expression and secretion.
Under stimulatory conditions, production of insulin may represent up to 50% of total protein production, placing an enormous burden on the ER [50]. As a consequence, beta cells are in a constant state of mild ER-stress, characterised by high basal expression of UPR transducers such as IRE1α, allowing them to adapt efficiently to the ever changing insulin demand [3]. In other cell types, activation of the IRE1α-XBP1s pathway increases production of ER-proteins aiming to fold and/or degrade misfolded proteins. Together with previous studies [27, 46], our data suggest that the IRE1α–XBP1s pathway has a distinct role in beta cells, relative to the fine-tuning of insulin mRNA expression through transcriptional and post-transcriptional mechanisms, allowing for rapid adjustment of insulin levels under ER-stress conditions [46]. However, due to the deleterious effects of prolonged increased XBP1s level, this mechanism has to be closely regulated in beta cells, resulting in a limited XPB1 splicing as shown by previous studies [2, 12] and the present data. These observations, together with other studies showing detrimental effects of prolonged eukaryotic translation initiation factor 2 alpha (eIF2α) or ATF6α activation in beta cells [24, 49], suggest that tight control of ER homeostasis is crucial to maintain beta cell function and survival.
Abbreviations
- Ad-eGFP:
-
Adenoviruses expressing the enhanced green fluorescent protein
- Ad-LUC:
-
Adenoviruses expressing the luciferase protein
- ATF:
-
Activating transcription factor
- BAX:
-
BCL2-associated X protein
- BIP:
-
Binding immunoglobulin protein
- CHOP:
-
C/EBP homologous protein
- CPA:
-
Cyclopiazonic acid
- ER:
-
Endoplasmic reticulum
- ERAD:
-
ER-associated degradation
- IRE:
-
Inositol-requiring enzyme
- MAFA:
-
Musculoaponeurotic fibrosarcoma oncogene homologue A
- MOI:
-
Multiplicity of infection
- mXBP1:
-
Mouse X-box binding protein 1
- NEUROD1:
-
Neurogenic differentiation 1
- PDX1:
-
Pancreatic and duodenal homeobox 1
- PERK:
-
PKR-like ER kinase
- siRNA:
-
Small interference RNA
- UPR:
-
Unfolded protein response
- XBP1:
-
X-box binding protein 1
- XBP1s:
-
Spliced/active X-box binding protein 1
References
Eizirik DL, Colli ML, Ortis F (2009) The role of inflammation in insulitis and beta-cell loss in type 1 diabetes. Nature Reviews Endocrinology 5:219–226
Cardozo AK, Ortis F, Storling J et al (2005) Cytokines downregulate the sarcoendoplasmic reticulum pump Ca2+ ATPase 2b and deplete endoplasmic reticulum Ca2+, leading to induction of endoplasmic reticulum stress in pancreatic beta-cells. Diabetes 54:452–461
Eizirik DL, Cardozo AK, Cnop M (2008) The role for endoplasmic reticulum stress in diabetes mellitus. Endocr Rev 29:42–61
Yoshida H (2007) Unconventional splicing of XBP-1 mRNA in the unfolded protein response. Antioxid Redox Signal 9:2323–2333
Lee AH, Iwakoshi NN, Glimcher LH (2003) XBP-1 regulates a subset of endoplasmic reticulum resident chaperone genes in the unfolded protein response. Mol Cell Biol 23:7448–7459
Yamamoto K, Sato T, Matsui T et al (2007) Transcriptional induction of mammalian ER quality control proteins is mediated by single or combined action of ATF6alpha and XBP1. Dev Cell 13:365–376
Oyadomari S, Takeda K, Takiguchi M et al (2001) Nitric oxide-induced apoptosis in pancreatic beta cells is mediated by the endoplasmic reticulum stress pathway. Proc Natl Acad Sci USA 98:10845–10850
Wang M, Zhao XR, Wang P et al (2007) Glucose regulated proteins 78 protects insulinoma cells (NIT-1) from death induced by streptozotocin, cytokines or cytotoxic T lymphocytes. Int J Biochem Cell Biol 39:2076–2082
Chambers KT, Unverferth JA, Weber SM, Wek RC, Urano F, Corbett JA (2008) The role of nitric oxide and the unfolded protein response in cytokine-induced beta-cell death. Diabetes 57:124–132
Huang CJ, Lin CY, Haataja L et al (2007) High expression rates of human islet amyloid polypeptide induce endoplasmic reticulum stress mediated beta-cell apoptosis, a characteristic of humans with type 2 but not type 1 diabetes. Diabetes 56:2016–2027
Akerfeldt MC, Howes J, Chan JY et al (2008) Cytokine-induced beta-cell death is independent of endoplasmic reticulum stress signaling. Diabetes 57:3034–3044
Pirot P, Eizirik DL, Cardozo AK (2006) Interferon-gamma potentiates endoplasmic reticulum stress-induced death by reducing pancreatic beta cell defence mechanisms. Diabetologia 49:1229–1236
Rasschaert J, Ladriere L, Urbain M et al (2005) Toll-like receptor 3 and STAT-1 contribute to double-stranded RNA + interferon-gamma-induced apoptosis in primary pancreatic beta-cells. J Biol Chem 280:33984–33991
Pipeleers DG, in’t Veld PA, van de Winkel M, Maes E, Schuit FC, Gepts W (1985) A new in vitro model for the study of pancreatic A and B cells. Endocrinology 117:806–816
Cunha DA, Hekerman P, Ladriere L et al (2008) Initiation and execution of lipotoxic ER stress in pancreatic beta-cells. J Cell Sci 121:2308–2318
Ortis F, Cardozo AK, Crispim D, Storling J, Mandrup-Poulsen T, Eizirik DL (2006) Cytokine-induced proapoptotic gene expression in insulin-producing cells is related to rapid, sustained, and nonoscillatory nuclear factor-kappaB activation. Mol Endocrinol 20:1867–1879
Cardozo AK, Heimberg H, Heremans Y et al (2001) A comprehensive analysis of cytokine-induced and nuclear factor-kappa B-dependent genes in primary rat pancreatic beta-cells. J Biol Chem 276:48879–48886
Cardozo AK, Kruhoffer M, Leeman R, Orntoft T, Eizirik DL (2001) Identification of novel cytokine-induced genes in pancreatic beta-cells by high-density oligonucleotide arrays. Diabetes 50:909–920
Pirot P, Ortis F, Cnop M et al (2007) Transcriptional regulation of the endoplasmic reticulum stress gene chop in pancreatic insulin-producing cells. Diabetes 56:1069–1077
Calfon M, Zeng H, Urano F et al (2002) IRE1 couples endoplasmic reticulum load to secretory capacity by processing the XBP-1 mRNA. Nature 415:92–96
He TC, Zhou S, da Costa LT, Yu J, Kinzler KW, Vogelstein B (1998) A simplified system for generating recombinant adenoviruses. Proc Natl Acad Sci USA 95:2509–2514
Heimberg H, Heremans Y, Jobin C et al (2001) Inhibition of cytokine-induced NF-kappaB activation by adenovirus-mediated expression of a NF-kappaB super-repressor prevents beta-cell apoptosis. Diabetes 50:2219–2224
Darville MI, Eizirik DL (1998) Regulation by cytokines of the inducible nitric oxide synthase promoter in insulin-producing cells. Diabetologia 41:1101–1108
Cnop M, Ladriere L, Hekerman P et al (2007) Selective inhibition of eukaryotic translation initiation factor 2 alpha dephosphorylation potentiates fatty acid-induced endoplasmic reticulum stress and causes pancreatic beta-cell dysfunction and apoptosis. J Biol Chem 282:3989–3997
Hoorens A, van de Casteele M, Kloppel G, Pipeleers D (1996) Glucose promotes survival of rat pancreatic beta cells by activating synthesis of proteins which suppress a constitutive apoptotic program. J Clin Invest 98:1568–1574
Ortis F, Pirot P, Naamane N et al (2008) Induction of nuclear factor-kappaB and its downstream genes by TNF-alpha and IL-1beta has a pro-apoptotic role in pancreatic beta cells. Diabetologia 51:1213–1225
Pirot P, Naamane N, Libert F et al (2007) Global profiling of genes modified by endoplasmic reticulum stress in pancreatic beta cells reveals the early degradation of insulin mRNAs. Diabetologia 50:1006–1014
Yoshida H, Matsui T, Yamamoto A, Okada T, Mori K (2001) XBP1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcription factor. Cell 107:881–891
Liu Y, Adachi M, Zhao S et al (2009) Preventing oxidative stress: a new role for XBP1. Cell Death Differ 16:847–857
Romero-Ramirez L, Cao H, Nelson D et al (2004) XBP1 is essential for survival under hypoxic conditions and is required for tumor growth. Cancer Res 64:5943–5947
Kroemer G, Galluzzi L, Brenner C (2007) Mitochondrial membrane permeabilization in cell death. Physiol Rev 87:99–163
Andrali SS, Sampley ML, Vanderford NL, Ozcan S (2008) Glucose regulation of insulin gene expression in pancreatic beta-cells. Biochem J 415:1–10
Melloul D, Marshak S, Cerasi E (2002) Regulation of insulin gene transcription. Diabetologia 45:309–326
Kaneto H, Miyatsuka T, Kawamori D et al (2008) PDX-1 and MafA play a crucial role in pancreatic beta-cell differentiation and maintenance of mature beta-cell function. Endocr J 55:235–252
Reimold AM, Etkin A, Clauss I et al (2000) An essential role in liver development for transcription factor XBP-1. Genes Dev 14:152–157
Sado M, Yamasaki Y, Iwanaga T et al (2009) Protective effect against Parkinson’s disease-related insults through the activation of XBP1. Brain Res 1257:16–24
Yoshida H, Nadanaka S, Sato R, Mori K (2006) XBP1 is critical to protect cells from endoplasmic reticulum stress: evidence from Site-2 protease-deficient Chinese hamster ovary cells. Cell Struct Funct 31:117–125
Gauthier BR, Brun T, Sarret EJ et al (2004) Oligonucleotide microarray analysis reveals PDX1 as an essential regulator of mitochondrial metabolism in rat islets. J Biol Chem 279:31121–31130
Johnson JD, Bernal-Mizrachi E, Alejandro EU et al (2006) Insulin protects islets from apoptosis via Pdx1 and specific changes in the human islet proteome. Proc Natl Acad Sci USA 103:19575–19580
Lee AH, Chu GC, Iwakoshi NN, Glimcher LH (2005) XBP-1 is required for biogenesis of cellular secretory machinery of exocrine glands. Embo J 24:4368–4380
Iwakoshi NN, Lee AH, Vallabhajosyula P, Otipoby KL, Rajewsky K, Glimcher LH (2003) Plasma cell differentiation and the unfolded protein response intersect at the transcription factor XBP-1. Nat Immunol 4:321–329
Hetz C, Lee AH, Gonzalez-Romero D et al (2008) Unfolded protein response transcription factor XBP-1 does not influence prion replication or pathogenesis. Proc Natl Acad Sci USA 105:757–762
Atouf F, Czernichow P, Scharfmann R (1997) Expression of neuronal traits in pancreatic beta cells. Implication of neuron-restrictive silencing factor/repressor element silencing transcription factor, a neuron-restrictive silencer. J Biol Chem 272:1929–1934
Martin D, Allagnat F, Chaffard G et al (2008) Functional significance of repressor element 1 silencing transcription factor (REST) target genes in pancreatic beta cells. Diabetologia 51:1429–1439
Han D, Lerner AG, Vande Walle L et al (2009) IRE1alpha kinase activation modes control alternate endoribonuclease outputs to determine divergent cell fates. Cell 138:562–575
Lipson KL, Ghosh R, Urano F (2008) The role of IRE1alpha in the degradation of insulin mRNA in pancreatic beta-cells. PLoS ONE 3:e1648
Thorens B, Wu YJ, Leahy JL, Weir GC (1992) The loss of GLUT2 expression by glucose-unresponsive beta cells of db/db mice is reversible and is induced by the diabetic environment. J Clin Invest 90:77–85
Yamamoto K, Yoshida H, Kokame K, Kaufman RJ, Mori K (2004) Differential contributions of ATF6 and XBP1 to the activation of endoplasmic reticulum stress-responsive cis-acting elements ERSE, UPRE and ERSE-II. J Biochem 136:343–350
Seo HY, Kim YD, Lee KM et al (2008) Endoplasmic reticulum stress-induced activation of activating transcription factor 6 decreases insulin gene expression via up-regulation of orphan nuclear receptor small heterodimer partner. Endocrinology 149:3832–3841
Schuit FC, In’t Veld PA, Pipeleers DG (1988) Glucose stimulates proinsulin biosynthesis by a dose-dependent recruitment of pancreatic beta cells. Proc Natl Acad Sci USA 85:3865–3869
Acknowledgements
This work was supported by grants from the Juvenile Diabetes Research Foundation International (1-2008-536), Fonds National de la Recherche Scientifique (FNRS-FRSM) Belgium, the Communauté Française de Belgique—Actions de Recherche Concertées (ARC), the European Union (Integrated Projects EuroDia in the Framework Programme 6 [FP6] and Novel Immunotherapies for Type 1 Diabetes [NAIMIT] in the FP7 of the European Community) and the Belgium Program on Interuniversity Poles of Attraction initiated by the Belgium State (IUAP P6/40). A. K. Cardozo is a Research Associate of the FNRS. We thank M. A. Neef, G. Vandenbroeck, M. Urbain, R. Leeman, A. M. Musuaya, R. Makhnas, J. Schoonheydt and S. Mertens from the Laboratory of Experimental Medicine, ULB, for excellent technical support.
Duality of interest
The authors declare that there is no duality of interest associated with this manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM Table 1
Primer sequences for quantitative RT-PCR and XBP siRNA sequences (PDF 45 kb)
ESM Fig. 1
Schematic representation of unspliced, spliced and truncated forms of XBP1 (XBP1u, XBP1s, tXBP1s, respectively) mRNA and protein-coding regions. Diagram shows the translation portions of the two open reading frames (ORF) 1 for unspliced and ORF 2 for spliced XBP1, respectively, the 26-bp intron and the primers F (forward), R1 (reverse 1) and R2 (reverse 2), which were used to amplify the mRNA inserts cloned in the adenoviral plasmid (pcDNA3). For more details, see main text, “Methods”. Adapted with kind permission from Lee et al. [1] (PDF 634 kb)
ESM Fig. 2
Dispersed islet cells were treated with CPA or infected with Ad-LUC or Ad-XBP1s at MOI 15, as indicated. Between 48 and 72 h after infection or 24 h after CPA treatment, mRNA was extracted and real-time PCR experiments performed on rat (r)Xbp1s (a), Chop (b) and Bip (c), normalised per Gapdh expression. Results are means ± SEM of four independent experiments.*p < 0.05 vs respective control (PDF 1918 kb)
ESM Fig. 3
208F rat fibroblasts were infected with Ad-LUC or Ad-XBP1s at MOI 15 or 30, as indicated. At 48 h after infection, cells were treated or not for 48 h with CPA (50 µmol/l). a XBP1s and β-actin Western blot, representative of five independent experiments. b Fibroblasts were fixed and subjected to immunofluorescence anti- cytochrome c and counterstained with Hoechst 33342 as indicated. Arrows indicate: cells that have lost mitochondrial cytochrome c staining; and cells with typical apoptotic nuclei or cells that have lost their DNA staining. Merged image shows cytochrome c and Hoechst staining. Fields are representative of five independent experiments. c Quantitative assessment of cytochrome c release in fibroblasts treated with CPA. Data are expressed as percentage of cells showing released cytochrome c over the total number of cells counted ± SEM. Results are mean of five independent experiments. *p < 0.05 vs non-infected condition (PDF 4427 kb)
Rights and permissions
About this article
Cite this article
Allagnat, F., Christulia, F., Ortis, F. et al. Sustained production of spliced X-box binding protein 1 (XBP1) induces pancreatic beta cell dysfunction and apoptosis. Diabetologia 53, 1120–1130 (2010). https://doi.org/10.1007/s00125-010-1699-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-010-1699-7