Abstract
Key message
The HaOr5 resistance gene is located in a large genomic insertion containing putative resistance genes and provides resistance to O. cumana, preventing successful connection to the sunflower root vascular system.
Abstract
Orobanche cumana (sunflower broomrape) is a parasitic plant that is part of the Orobanchaceae family and specifically infests sunflower crops. This weed is an obligate parasitic plant that does not carry out photosynthetic activity or develop roots and is fully dependent on its host for its development. It produces thousands of dust-like seeds per plant. It possesses a high spreading ability and has been shown to quickly overcome resistance genes successively introduced by selection in cultivated sunflower varieties. The first part of its life cycle occurs underground. The connection to the sunflower vascular system is essential for parasitic plant survival and development. The HaOr5 gene provides resistance to sunflower broomrape race E by preventing the connection of O. cumana to the root vascular system. We mapped a single position of the HaOr5 gene by quantitative trait locus mapping using two segregating populations. The same location of the HaOr5 gene was identified by genome-wide association. Using a large population of thousands of F2 plants, we restricted the location of the HaOr5 gene to a genomic region of 193 kb. By sequencing the whole genome of the resistant line harboring the major resistance gene HaOr5, we identified a large insertion of a complex genomic region containing a cluster of putative resistance genes.





Similar content being viewed by others
Data availability
The GenBank accession numbers of the HaOr5 genomic regions are OR715020 and OR715021. Thewhole-genome sequence data of the LC1093 sunflower line reported in this paper have been deposited in the Genome Warehouse of the National Genomics Data Center, Beijing Institute of Genomics, Chinese Academy of Sciences/China National Center for Bioinformation, under accession number GWHAAEK00000000, which is publicly accessible at https://ngdc.cncb.ac.cn/gwh.
References
Abd-Hamid N-A, Ahmad-Fauzi M-I, Zainal Z, Ismail I (2020) Diverse and dynamic roles of F-box proteins in plant biology. Planta 251:68. https://doi.org/10.1007/s00425-020-03356-8
Albert M, Axtell MJ, Timko MP (2021) Mechanisms of resistance and virulence in parasitic plant–host interactions. Plant Physiol 185:1282–1291. https://doi.org/10.1093/plphys/kiaa064
Antonova TS (2014) The history of interconnected evolution of Orobanche cumana Wallr. and sunflower in the Russian Federation and Kazakhstan. Helia 37:215–225. https://doi.org/10.1515/helia-2014-0017
Badouin H, Gouzy J, Grassa CJ et al (2017) The sunflower genome provides insights into oil metabolism, flowering and Asterid evolution. Nature 546:148–152. https://doi.org/10.1038/nature22380
Broman KW, Wu H, Sen S, Churchill GA (2003) R/qtl: QTL mapping in experimental crosses. Bioinforma Oxf Engl 19:889–890. https://doi.org/10.1093/bioinformatics/btg112
Cabanettes F, Klopp C (2018) D-GENIES: dot plot large genomes in an interactive, efficient and simple way. PeerJ 6:e4958. https://doi.org/10.7717/peerj.4958
Calderón-González Á, Pouilly N, Muños S et al (2019) An SSR-SNP linkage map of the parasitic weed Orobanche cumana Wallr. Including a gene for plant pigmentation. Front Plant Sci. https://doi.org/10.3389/fpls.2019.00797
Calderón-González Á, Pérez-Vich B, Pouilly N et al (2022) Association mapping for broomrape resistance in sunflower. Front Plant Sci 13:1056231. https://doi.org/10.3389/fpls.2022.1056231
Carrere S, Gouzy J (2023a) Eukaryote EuGene pipeline Version 2
Carrere S, Gouzy J (2023b) LIPME functional annotation pipeline
Chabaud M, Auriac M-C, Boniface M-C et al (2022) Wild Helianthus species: a reservoir of resistance genes for sustainable pyramidal resistance to broomrape in sunflower. Front Plant Sci 13:1038684. https://doi.org/10.3389/fpls.2022.1038684
Cheng H, Concepcion GT, Feng X et al (2021) Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat Methods 18:170–175. https://doi.org/10.1038/s41592-020-01056-5
Cheung WY, Hubert N, Landry BS (1993) A simple and rapid DNA Microextraction method for plant, animal, and insect suitable for RAPD and other PCR analyzes. PCR Meth Applics 3:69–70
de Givry S, Bouchez M, Chabrier P et al (2005) Carhta Gene: multipopulation integrated genetic and radiation hybrid mapping. Bioinformatics 21:1703–1704. https://doi.org/10.1093/bioinformatics/bti222
Duriez P, Vautrin S, Auriac M-C et al (2019) A receptor-like kinase enhances sunflower resistance to Orobanche cumana. Nat Plants 5:1211–1215. https://doi.org/10.1038/s41477-019-0556-z
Eshed Y, Zamir D (1995) An introgression line population of Lycopersicon pennellii in the cultivated tomato enables the identification and fine mapping of yield-associated QTL. Genetics 141:1147–1162. https://doi.org/10.1093/genetics/141.3.1147
Fernández-Aparicio M, del Moral L, Muños S et al (2022) Genetic and physiological characterization of sunflower resistance provided by the wild-derived OrDeb2 gene against highly virulent races of Orobanche cumana Wallr. Theor Appl Genet 135:501–525. https://doi.org/10.1007/s00122-021-03979-9
Galindo-González L, Mhiri C, Deyholos MK, Grandbastien M-A (2017) LTR-retrotransposons in plants: engines of evolution. Gene 626:14–25. https://doi.org/10.1016/j.gene.2017.04.051
Gobena D, Shimels M, Rich PJ et al (2017) Mutation in sorghum LOW GERMINATION STIMULANT 1 alters strigolactones and causes striga resistance. Proc Natl Acad Sci 114:4471–4476. https://doi.org/10.1073/pnas.1618965114
Guo X, Zhang Y, Tu Y et al (2018) Overexpression of an EIN3-binding F-box protein2-like gene caused elongated fruit shape and delayed fruit development and ripening in tomato. Plant Sci 272:131–141. https://doi.org/10.1016/j.plantsci.2018.04.016
Hegenauer V, Fürst U, Kaiser B et al (2016) Detection of the plant parasite Cuscuta reflexa by a tomato cell surface receptor. Science 353:478–481. https://doi.org/10.1126/science.aaf3919
Hegenauer V, Slaby P, Körner M et al (2020) The tomato receptor CuRe1 senses a cell wall protein to identify Cuscuta as a pathogen. Nat Commun 11:5299. https://doi.org/10.1038/s41467-020-19147-4
Huang K, Jahani M, Gouzy J et al (2023) The genomics of linkage drag in inbred lines of sunflower. Proc Natl Acad Sci 120:e2205783119. https://doi.org/10.1073/pnas.2205783119
Imerovski I, Dedić B, Cvejić S et al (2019) BSA-seq mapping reveals major QTL for broomrape resistance in four sunflower lines. Mol Breed 39:41. https://doi.org/10.1007/s11032-019-0948-9
Joel DM, Chaudhuri SK, Plakhine D et al (2011) Dehydrocostus lactone is exuded from sunflower roots and stimulates germination of the root parasite Orobanche cumana. Phytochemistry 72:624–634. https://doi.org/10.1016/j.phytochem.2011.01.037
Koressaar T, Remm M (2007) Enhancements and modifications of primer design program Primer3. Bioinformatics 23:1289–1291. https://doi.org/10.1093/bioinformatics/btm091
Krupp A, Heller A, Spring O (2019) Development of phloem connection between the parasitic plant Orobanche cumana and its host sunflower. Protoplasma 256:1385–1397. https://doi.org/10.1007/s00709-019-01393-z
Le Ru A, Ibarcq G, Boniface M-C et al (2021) Image analysis for the automatic phenotyping of Orobanche cumana tubercles on sunflower roots. Plant Methods 17:80. https://doi.org/10.1186/s13007-021-00779-6
Lechner E, Achard P, Vansiri A et al (2006) F-box proteins everywhere. Curr Opin Plant Biol 9:631–638. https://doi.org/10.1016/j.pbi.2006.09.003
Li H (2018) Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34:3094–3100. https://doi.org/10.1093/bioinformatics/bty191
Li J, Timko MP (2009) Gene-for-Gene resistance in Striga-cowpea associations. Science 325:1094–1094. https://doi.org/10.1126/science.1174754
Limpert E, Müller K (1994) Designation of pathotypes of plant pathogens. J Phytopathol 140:346–358. https://doi.org/10.1111/j.1439-0434.1994.tb00617.x
Liu Y, Zhang Y-M, Tang Y et al (2023) The evolution of plant NLR immune receptors and downstream signal components. Curr Opin Plant Biol 73:102363. https://doi.org/10.1016/j.pbi.2023.102363
Louarn J, Boniface M-C, Pouilly N et al (2016) Sunflower resistance to broomrape (Orobanche cumana) is controlled by specific QTLs for different parasitism stages. Front Plant Sci. https://doi.org/10.3389/fpls.2016.00590
Martín-Sanz A, Pérez-Vich B, Rueda S et al (2020) Characterization of post-haustorial resistance to sunflower broomrape. Crop Sci 60:1188–1198. https://doi.org/10.1002/csc2.20002
Mayjonade B, Gouzy J, Donnadieu C et al (2016) Extraction of high-molecular-weight genomic DNA for long-read sequencing of single molecules. Biotechniques 61:203–205. https://doi.org/10.2144/000114460
Molinero-Ruiz L, Delavault P, Pérez-Vich B et al (2015) History of the race structure of Orobanche cumana and the breeding of sunflower for resistance to this parasitic weed: a review. Span J Agric Res. https://doi.org/10.13039/501100000780
Nelson DC (2021) The mechanism of host-induced germination in root parasitic plants. Plant Physiol 185:1353–1373. https://doi.org/10.1093/plphys/kiab043
Pacureanu-Joita M, Veronesi C, Raranciuc S, Stanciu D (2004) Parasite-host plant interaction of Orobanche cumana Wallr. (Orobanche cernua Loefl) with Helianthus annuus. In: Proc. 16th Int. Sunflower Conf., Fargo, ND, USA. pp 171–177
Papadakis JS (1937) Méthode statistique pour des expériences sur champ. Thessalonike: Institut d'Amélioration des Plantes à Salonique. (Bullettin, 23). p 30
Parks T, Yordanov YS (2020) Composite plants for a composite plant: an efficient protocol for root studies in the sunflower using composite plants approach. Plant Cell Tissue Organ Cult PCTOC 140:647–659. https://doi.org/10.1007/s11240-019-01760-x
Pecrix Y, Staton SE, Sallet E et al (2018) Whole-genome landscape of Medicago truncatula symbiotic genes. Nat Plants 4:1017–1025. https://doi.org/10.1038/s41477-018-0286-7
Pérez-Vich B, Akhtouch B, Muñoz-Ruz J et al (2002) Inheritance of resistance to a highly virulent race f of Orobanche cumana wallr. in a sunflower line derived from interspecific amphiploid. Helia 25:137–144. https://doi.org/10.2298/hel0236137p
Pérez-Vich B, Akhtouch B, Knapp SJ et al (2004) Quantitative trait loci for broomrape (Orobanche cumana Wallr.) resistance in sunflower. Theor Appl Genet 109:92–102. https://doi.org/10.1007/s00122-004-1599-7
Pujadas-salvà AJ, Velasco L (2000) Comparative studies on Orobanche cernua L. and O. cumana Wallr. (Orobanchaceae) in the Iberian Peninsula. Bot J Linn Soc 134:513–527. https://doi.org/10.1111/j.1095-8339.2000.tb00548.x
Qiu S, Bradley JM, Zhang P et al (2022) Genome-enabled discovery of candidate virulence loci in Striga hermonthica, a devastating parasite of African cereal crops. New Phytol 236:622–638. https://doi.org/10.1111/nph.18305
Rahimi M, Bouwmeester H (2021) Are sesquiterpene lactones the elusive KARRIKIN-INSENSITIVE2 ligand? Planta 253:54. https://doi.org/10.1007/s00425-021-03571-x
Raupp FM, Spring O (2013) New sesquiterpene lactones from sunflower root exudate as germination stimulants for Orobanche cumana. J Agric Food Chem 61:10481–10487. https://doi.org/10.1021/jf402392e
Rodríguez-Ojeda MI, Pineda-Martos R, Alonso LC et al (2013) A dominant avirulence gene in Orobanche cumana triggers Or5 resistance in sunflower. Weed Res 53:322–327. https://doi.org/10.1111/wre.12034
Sallet E, Gouzy J, Schiex T (2019) EuGene: an automated integrative gene finder for eukaryotes and prokaryotes. In: Kollmar M (ed) Gene prediction: methods and protocols. Springer, New York, NY, pp 97–120
Segura V, Vilhjálmsson BJ, Platt A et al (2012) An efficient multi-locus mixed-model approach for genome-wide association studies in structured populations. Nat Genet 44:825–830. https://doi.org/10.1038/ng.2314
Shahid S, Kim G, Johnson NR et al (2018) MicroRNAs from the parasitic plant Cuscuta campestris target host messenger RNAs. Nature 553:82–85. https://doi.org/10.1038/nature25027
Sisou D, Tadmor Y, Plakhine D et al (2021) Biological and transcriptomic characterization of pre-haustorial resistance to sunflower broomrape (Orobanche cumana W.) in sunflowers (Helianthus annuus). Plants Basel 10:1810
Subrahmaniam HJ, Libourel C, Journet E-P et al (2018) The genetics underlying natural variation of plant-plant interactions, a beloved but forgotten member of the family of biotic interactions. Plant J Cell Mol Biol 93:747–770. https://doi.org/10.1111/tpj.13799
Tang S, Heesacker A, Kishore V et al (2003) Genetic mapping of the Or5 gene for resistance to Orobanche race E in sunflower. Crop Sci. https://doi.org/10.2135/cropsci2003.1021
Terzić S, Boniface M-C, Marek L et al (2020) Gene banks for wild and cultivated sunflower genetic resources. OCL 27:9. https://doi.org/10.1051/ocl/2020004
Ueno K, Furumoto T, Umeda S et al (2014) Heliolactone, a non-sesquiterpene lactone germination stimulant for root parasitic weeds from sunflower. Phytochemistry 108:122–128. https://doi.org/10.1016/j.phytochem.2014.09.018
Untergasser A, Cutcutache I, Koressaar T et al (2012) Primer3—new capabilities and interfaces. Nucleic Acids Res 40:e115. https://doi.org/10.1093/nar/gks596
van den Burg HA, Tsitsigiannis DI, Rowland O et al (2008) The F-box protein ACRE189/ACIF1 regulates cell death and defense responses activated during pathogen recognition in tobacco and tomato. Plant Cell 20:697–719. https://doi.org/10.1105/tpc.107.056978
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78. https://doi.org/10.1093/jhered/93.1.77
Vranceanu AV, Tudor VA, Stoenescu FM, Pirvu N (1981) Virulence groups of Orobanche cumana Wallr.[root parasite], differential hosts and resistance sources and genes in sunflower. In 9. international conference of sunflower, Torremolinos, Malaga (Spain), 8–13 Jun 1980. Ministerio de Agricultura, Servicio de Publicaciones Agrarias
Westwood JH, Yoder JI, Timko MP, dePamphilis CW (2010) The evolution of parasitism in plants. Trends Plant Sci 15:227–235. https://doi.org/10.1016/j.tplants.2010.01.004
Wimmer V, Albrecht T, Auinger H-J, Schön C-C (2012) synbreed: a framework for the analysis of genomic prediction data using R. Bioinformatics 28:2086–2087. https://doi.org/10.1093/bioinformatics/bts335
Yang C, Fu F, Zhang N et al (2020) Transcriptional profiling of underground interaction of two contrasting sunflower cultivars with the root parasitic weed Orobanche cumana. Plant Soil 450:303–321. https://doi.org/10.1007/s11104-020-04495-3
You FM, Huo N, Gu YQ et al (2008) BatchPrimer3: A high throughput web application for PCR and sequencing primer design. BMC Bioinformatics 9:253. https://doi.org/10.1186/1471-2105-9-253
Acknowledgements
We thank all the students and people who worked in the nurseries for selfing and harvesting the capitula of the sunflower plants. We thank Ming Denaes for harvesting the seeds. The Genotoul bioinformatics platform Toulouse Midi provided computing resources. This work was supported by the French Laboratory of Excellence project TULIP (ANR-10-LABX-41; ANR-11-IDEX-0002-02). This study was performed within the framework of the École Universitaire de Recherche TULIP-GS (ANR-18-EURE-0019).
Funding
This work was funded by MAS SEEDS, INRAE, ANRT and the Region Occitanie (Genosol project).
Author information
Authors and Affiliations
Contributions
Camille Pubert coordinated the selection of the recombinant plants and participated in the fine mapping, genomic analysis, marker design, molecular experiments and phenotyping. Marie-Claude Boniface coordinated the nursery, phenotyping and harvesting of the plants and the seeds. Alexandra Legendre performed the gDNA extractions and the genotyping experiments. Mireille Chabaud performed the cytological experiments. Sébastien Carrère annotated the genome sequences. Caroline Callot prepared the sequencing libraries. Isabelle Dufau performed the optical map experiments. Charlotte Cravero assembled the genome sequences. Sonia Vautrin coordinated the Genosol project for genome sequence assembly. Mihaela Patrascoiu supervised the production of the F2 population. Aurélie Baussart participated in the phenotyping of the parental lines and phenotyped the plants for QTL mapping and GWA analyzes. Véronique Gautier and Elodie Belmonte performed the sequencing. Charles Poncet coordinated the sequencing. Jun Zhao collected the samples of the Chinese O. cumana population. Luyang Hu characterized the Chinese O. cumana population. Weijun Zhou supervised the characterization of the Chinese O. cumana population. Claire Coussy performed the QTL mapping and GWA analyzes. Nicolas Langlade, Claire Coussy and Stéphane Muños supervised the study. Claire Coussy and Stéphane Muños designed the study. Stéphane Muños coordinated the study. Camille Pubert and Stéphane Muños contributed to the data analysis and wrote the manuscript. All the authors contributed to and revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
We declare no conflicts of interest in regard to this manuscript.
Additional information
Communicated by Brent Hulke.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
122_2024_4594_MOESM1_ESM.pptx
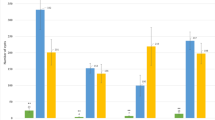
Figure S1 Evaluation of the resistance for various broomrape populations. The parental lines LSMS1 and LC1093 were phenotyped using six broomrape populations (E-DDP, E-BOU, E/F-EEC, F-IN23, G-KZP and G-RO) in 3L pots in greenhouse. The figure represents the results for E-DDP, E/F-EEC and G-RO populations (see Fig. 1 for the three others). We added the susceptible line XRQ and two resistant lines LRMS1 and DEB2 carrying the resistance gene HaOr7 and OrDeb2 respectively as controls. Broomrape emergences were counted 12 weeks after inoculation. Significant pair-wise t-tests between lines are grouped for clarity and illustrated as ***p < 0.01 and **p < 0.05 (PPTX 109 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pubert, C., Boniface, MC., Legendre, A. et al. A cluster of putative resistance genes is associated with a dominant resistance to sunflower broomrape. Theor Appl Genet 137, 103 (2024). https://doi.org/10.1007/s00122-024-04594-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00122-024-04594-0