Abstract
Key message
Genotyping data of a comprehensive Korean soybean collection obtained using a large SNP array were used to clarify global distribution patterns of soybean and address the evolutionary history of soybean.
Abstract
Understanding diversity and evolution of a crop is an essential step to implement a strategy to expand its germplasm base for crop improvement research. Accessions intensively collected from Korea, which is a small but central region in the distribution geography of soybean, were genotyped to provide sufficient data to underpin population genetic questions. After removing natural hybrids and duplicated or redundant accessions, we obtained a non-redundant set comprising 1957 domesticated and 1079 wild accessions to perform population structure analyses. Our analysis demonstrates that while wild soybean germplasm will require additional sampling from diverse indigenous areas to expand the germplasm base, the current domesticated soybean germplasm is saturated in terms of genetic diversity. We then showed that our genome-wide polymorphism map enabled us to detect genetic loci underlying flower color, seed-coat color, and domestication syndrome. A representative soybean set consisting of 194 accessions was divided into one domesticated subpopulation and four wild subpopulations that could be traced back to their geographic collection areas. Population genomics analyses suggested that the monophyletic group of domesticated soybeans was likely originated at a Japanese region. The results were further substantiated by a phylogenetic tree constructed from domestication-associated single nucleotide polymorphisms identified in this study.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Extensive genotyping and phenotyping can help harness favorable alleles underlying phenotypic variation in both wild and domesticated germplasm. Soybean [Glycine max (L.) Merr.] is a major crop for dietary protein and oil worldwide. Several hundred soybean genomes have been resequenced (Chung et al. 2014; Lam et al. 2010; Valliyodan et al. 2016; Zhou et al. 2015), and three genome-wide high-density SNP arrays have been developed and used to genotype thousands of soybean accessions (Lee et al. 2015; Song et al. 2013; Wang et al. 2016). These data have been primarily used to compare the patterns of genetic variation between G. max and its wild progenitor (G. soja Siebold & Zucc.) to understand the history of soybean domestication and identify selective sweeps related to the domestication and improvement of soybeans. The data have also been used to identify loci controlling important agronomic traits, such as protein-and-oil and seed-weight traits (Bandillo et al. 2015; Hwang et al. 2014; Zhou et al. 2015). However, further efforts will be required to implement genome-wide association studies (GWAS) (McCarthy et al. 2008) with higher statistical power and mapping resolution in soybean.
Soybean was domesticated ~ 5000 years ago from G. soja, its sympatric wild annual progenitor that is distributed throughout East Asia, including most of China, Korea, Japan, and part of Russia (Hymowitz 2004; Larson et al. 2014). Different regions of China have been proposed as a single center of soybean domestication on the basis of morphological, cytogenetic, and seed protein variation (Broich and Palmer 1981; Hymowitz 2004; Hymowitz and Kaizuma 1981). Multiple centers of domestication including the southern areas of Japan and China have also been proposed based on chloroplast sequence variation (Xu et al. 2002) and archeological records (Lee et al. 2011). However, recent phylogenetic studies using whole-genome resequencing data clearly indicated a monophyletic nature of domesticated soybean (Chung et al. 2014; Lam et al. 2010; Zhou et al. 2015). Of late, molecular studies that used hundreds of markers and accessions have proposed different areas, such as the Yellow River of China (Li et al. 2010) and southern China (Guo et al. 2010) as a center of soybean domestication. Yet, two other studies suggested the domestication center as northern and central China using high-density SNP array data (Wang et al. 2016), or central China surrounding the Yellow River using specific-locus amplified fragment sequencing data (Han et al. 2016).
In most of the previous studies, accessions collected from the Korean Peninsula were underrepresented, although this region is a central region of wild soybean distribution. For example, in the recent genome-wide analyses reported by Wang et al. (2016) and Han et al. (2016), accessions collected from China accounted for 91.8% and 100% of the total samples, respectively. Here, we present an analysis of SNP genotype data from 2824 domesticated, 1360 wild, and 50 putative hybrid accessions as part of an effort to characterize the entire Korean indigenous soybean collection deposited in the country’s National Agrobiodiversity Center. We genotyped soybean accessions using the 180 K Axiom® SoyaSNP array that was developed from soybean genome resequencing data (Lee et al. 2015). Our high-density SNP array data allowed us to evaluate levels of genetic diversity and patterns of population structure. We further attempted to detect genetic loci underlying soybean flower color, seed coat, and domestication traits, as well as provide a refined model of the evolutionary history of domesticated soybean.
Materials and methods
Plant materials and SNP genotyping
The majority of the accessions were from the National Agrobiodiversity Center in Jeonju, Korea, with a small number of accessions provided by individual laboratories (Table S1). The National Agrobiodiversity Center collection consists of approximately 14,000 accessions of improved and landrace cultivars (G. max) and approximately 3700 accessions of wild soybean (G. soja). The Korean germplasm collection substantially overlaps with those of other countries, particularly the United States. Most accessions collected from locations in other countries than Korea have been donated from the US National Genetic Resources Program. Notable exceptions were the 46 wild accessions from Japan, whose accession codes start with ‘B’. These were donated from National BioResource Project in Japan. As our primary goal was to characterize the indigenous soybean collection of Korea, we attempted as much as possible to genotype Korean accessions with even distribution across South Korea. At the same time, we analyzed representative sets of landrace accessions from China, North Korea, and Japan and approximately 400 improved lines, most of which are immediate descendants of ancestral lines of United States soybean cultivars (Gizlice et al. 1994), so that cultivated soybean from Korea could be assessed in the context of worldwide soybean germplasm pool (Table S2). Representative G. soja accessions from China, Russia, and Japan were also selected, allowing the even geographic distribution of wild soybean in each of these countries to be sampled. Initially, we planted approximately 5000 domesticated accessions and 2400 wild soybean accessions, of which approximately 90% were collected in Korea. In results, approximately 35% of the total number of domesticated accessions collected from each of provinces in South Korea were planted, and of the total number of wild accessions, approximately 65% that evenly represented across South Korea were planted. After pure line selection by single seed descent was performed at least two times, DNA samples from approximately 4400 diverse soybean germplasm lines were genotyped. However, because our SNP array data set ended up with smaller number of G. soja accessions from China than those from Korea and Japan, soybean population structure from the representative set was additionally assessed using genome resequencing data (downloaded from Figshare database, http://figshare.com/articles/Soybean_resequencing_project/1176133) from 45 G. soja accessions reported by Zhou et al. (2015).
DNA samples from the ~ 4400 diverse soybean accessions were extracted from a single plant of each accession and were genotyped with the Axiom® SoyaSNP array containing 180,961 SNP sites (Lee et al. 2015). Of the lines genotyped, 4234 with > 97% sample call rate were selected for further analysis. SNPs were scored following the Axiom® Genotyping Solution Data Analysis User Guide (http://www.affymetrix.com/) as described by Lee et al. (2015). Of the 180,961 SNPs, 170,223 were selected on the basis of the development and validation study. Missing data points in the 170,223 SNPs were imputed using BEAGLE 4.0 with default settings (Browning and Browning 2007). The 170,223 SNPs were then used to screen out duplicated and redundant accessions, leaving 3036 non-redundant accessions. After the initial filtration, SNPs with heterozygous rate > 0.02 and minor allele frequency < 0.02 were discarded from the genotype data of the non-redundant accessions, leaving a total of 117,095 high-quality SNPs for the further population analyses (Fig. S1).
Phenotypic data used for GWAS were obtained primarily from field evaluations in the field at National Institute of Crop Science, Suwon, Korea, in 2012 and 2013 (Table S1). The observed phenotype data were converted into binary data. The flower color phenotypes were divided into absence of color (white) or presence of colors ranging from light to dark purple. The seed-coat color phenotypes were divided into absence of colors (yellow or green) or presence of colors ranging from brown to black. Domestication phenotypes were divided into presence (G. max) or absence (G. soja) of domestication.
Population structure and genetic diversity pattern analyses
Principal component analysis (PCA) was conducted to summarize the genetic structure and variation present in the soybean collection using smartpca function in Eigensoft v7.2 (Patterson et al. 2006; Price et al. 2006). We plotted the first three PCs. Neighbor-joining trees were constructed by MEGA7 (Kumar et al. 2016) under the p-distances model. We used a model-based clustering method implemented in ADMIXTURE v1.23 (Alexander et al. 2009) to investigate the population structure of the soybean accessions. We determined the optimal K, the number of clusters based on the smallest cross-validation error calculated from v-fold cross-validation procedure. We plotted the membership coefficient using DISTRUCT (Rosenberg 2004). To investigate the level of genetic diversity maintained in soybean accessions, we calculated the nucleotide diversity (π) using VCFtools v 0.1.13 (Danecek et al. 2011). Genetic differentiation (Weir and Cockerham’s FST) between G. max and each of the G. soja subpopulations was calculated using the VCFtools V0.1.13 (Weir and Cockerham 1984). Hierarchical analyses of molecular variance (AMOVA) in the whole soybean set and the representative soybean set were performed using ARLEQUIN v.3.5.2.2 (Excoffier and Lischer 2010). The significance of the values for FCT (difference among groups), FSC (difference among populations within groups), and FST (difference among populations) was tested by 1023 permutations.
Genome-wide association studies
We conducted GWAS using PLINK 1.9 (Purcell et al. 2007). Flower color, seed-coat color, and domestication phenotype data were converted to binary data to perform a conditional logistic regression model analysis. Conditional specific SNPs were selected on the basis of minor allele frequencies of SNPs among the groups defined based on PCA analysis. A Bonferroni correction was used to control for the multiple testing problem by adjusting the alpha value from α = 0.01 to α = (0.01/117,095 SNPs) where 117,095 is the number of statistical tests conducted. Therefore, statistical significance of a SNP-trait association was set at 8.54e−8 (−log10P = 7.06). Manhattan plots were produced using the qqman package (Turner 2014). To define linkage disequilibrium (LD) patterns, correlation coefficient of alleles (r2) was calculated for SNPs under the peak regions that exhibited significant association using Haploview 4.2 (Barrett et al. 2005). The confidence interval (CI) method of Gabriel et al. (2002) was used to identify LD blocks.
Results
Overall structure of the genotyped soybean germplasm population
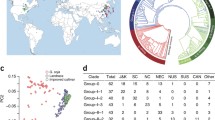
Of the approximately 4400 accessions genotyped, 4234 exhibited > 97% sample call rate. These were used as the total population for characterizing the Korean soybean germplasm (Table S1). The majority of this 4234 set contained accessions from Korea (78.7% G. max and 91.5% G. soja) (Fig. 1a). The rest were G. max landrace and G. soja accessions from China, Russia, North Korea, and Japan and improved lines mostly developed in the USA (Table S2). To eliminate potential confounding effects exerted by hybrids in the comparison of wild and domesticated soybean populations (Vaughan et al. 2008; Wang et al. 2017), we first removed 50 putative hybrid accessions from among the 4234 accessions (Fig. 1b; Table S2). In the field evaluation, most of these 50 accessions showed intermediate morphologies between domesticated soybean (G. max) and its wild relative (G. soja). Furthermore, principal component analysis (PCA) using 170,223 high-quality SNPs showed that the accessions were positioned between two large groups of G. max and G. soja (Fig. 1a). In further support of their suspected hybrid status, 50 accessions showed mixed wild or domesticated genome fractions ranging from 30 to 70% in K = 2 or 3 populations in the ADMIXTURE analysis (Fig. 1b).
Population structure of the genotyped 4234 soybean accessions. a Principal components of SNP variation. PC1 and PC2 indicate score of principal components 1 and 2, respectively. Each of PC1 and PC2 explained 15.6% and 2.7% of variance in the data. Glycine max, G. soja, and hybrids are shown by red, blue, and green dots, respectively. The majority of Korean accessions cluster together within dashed eclipses. b ADMIXTURE plots. The accessions were divided into three groups: G. max, G. soja, and their hybrids. c Distribution of number of redundant accession groups that showed < 1.25% inconsistencies between the SNP calls. G. max and G. soja are shown by white and gray boxes. d Geographic distribution of the collection sites for G. soja accessions
In our previous development and validation study (Lee et al. 2015), the SNP calls genotyped by the SoyaSNP array were highly reproducible, with inconsistencies of ≤ 1.17% observed within pairs of 27 duplicated samples after excluding missing genotypes in either sample. Several sets of near-isogenic isolines were genotyped (Table S3). Single-gene isolines (backcross-derived isolines for single genes) showed approximately 1.0% inconsistencies after excluding missing genotypes in either sample (e.g., 1.16% between Harosoy and L67-153 [Harosoy(6) × Higan]). As expected, a slightly higher level of inconsistency (up to 1.5%) was observed for samples from multiple-gene isolines (e.g., 1.48% between L62-667 [Harosoy(6) × T204] and OT94-51 [OT89-5/L71-802//OT89-6]). However, we occasionally observed that some soybean accessions that had no known pedigree relationship showed < 1.50% inconsistencies (e.g., 0.87% between Williams 82 K and KLS85102). Therefore, we used a 1.25% inconsistency value as the cutoff to remove redundant or highly similar accessions from groups of duplicates or near-isogenic lines. The same cutoff value was applied to filtration of wild soybean accessions. In each of the genotype duplicate sets, an accession with a sample call rate ≥ 99% was preferentially retained. For each of the near-isogenic line groups, the recurrent parent or representative single-gene isoline in case of the absence of parents was retained. Of the 4184 accessions genotyped in this study, 1148 (867 domesticated and 281 wild) were removed (Fig. 1c). The high rate of redundant and highly similar accessions has been frequently reported in worldwide germplasm collections (Food and Agriculture Organization of the United Nations 2010; McCouch et al. 2012). For domesticated soybeans, the major cause in the National Agrobiodiversity Center in Korea is probably the unknowing submission of the same accession with different collection sites and designators because there are many accessions with the same common name but with different collection sites or collectors. For wild soybeans, multiple accessions were collected from a narrow habitat area.
After the filtration, a final set of 3036 genotyped accessions was available for population structure analysis (Table S1 and S3). Only a few accessions were removed from countries other than South Korea. As a result, overall proportion of soybean accessions among countries in this 3036 set was similar to that of the 4234 set (Table S2). In the 3036 set, 1957 were G. max accessions and 1079 were G. soja accessions. Representative G. soja accessions from China, Russia, and Japan remained to evenly reflect the geographic distribution of native G. soja in each of these country regions (Fig. 1d).
Population structure
ADMIXTURE (Alexander et al. 2009) and PCA (Patterson et al. 2006) were used to infer population structure of the 3036 non-redundant soybean set using 117,095 SNPs with heterozygous rate < 0.02 and minor allele frequency > 0.02 (Fig. S1). As observed in the analysis of our total population of 4234 accessions, the 3036 accessions were clearly divided into two large groups, representing G. max and G. soja (Fig. S2). Both the estimated cross-validation (CV) error plot from ADMIXTURE and scree plot from the PCA supported the presence of two large groups (Fig. S2), although the slopes did not level off, which is likely because of subgroupings within the two large groups. The clear separation of G. max and G. soja groups might be expected by the ascertainment bias, which favored selection of G. max SNPs (Lee et al. 2015), and the sampling bias. Unlike the previous observations that the genome diversity level was > twofold lower in the domesticated soybeans relative to that in the wild soybeans, the diversity level of the domesticated soybeans (mean per-site nucleotide diversity (π = 0.189) estimated from the 117,095 SNPs was ~ 1.58-fold lower than that of the wild soybeans (π = 0.298) and nearly two times more domesticated soybean accessions were used for the population structure analyses. The 3036 set also contained excessive numbers of accessions collected in South Korea in both the domesticated and wild soybean groups. Interestingly, in the PCA space constructed with the first two PCs, G. soja accessions from Japan and China that were located at both ends of the G. soja cluster were almost equally close to the G. max cluster. However, in the PC1 and PC3 plot, accessions from Japan were closest to the G. max cluster and accessions from China were the most distantly related to the G. max cluster (Fig. S2).
When we analyzed G. max and G. soja separately, a somewhat distinct subpopulation structure was revealed. Within each of G. max and G. soja populations, CV errors of the ADMIXTURE runs decreased gradually without a steep drop (Fig. S3), whereas eigenvalues of the PCA runs showed steep decrease up to K = 5 (Fig. 2), indicating that there were at least four distinct subpopulations in each of the G. max and G. soja populations. Both the ADMIXTURE and PCA plots from the 1957 domesticated soybeans did not show distinctive grouping (Figs. 2 and S3). The majority of South Korean domesticated accessions (~ 80%) formed a dense subpopulation likely because of recent overcollection. This notion is supported by that the rest of the Korea accessions were well mixed with Chinese, North Korean, and Japanese accessions, which did not show distinct subgrouping on a geographic basis. Notably, the North Korea accessions, which could be considered true landraces because of their collections during the first half of the twentieth century before modern breeding research, were evenly distributed across subpopulations. The improved cultivars were narrowly clustered in the PCA plot, indicating much lower diversity relative to that of the entire domesticated soybeans. The 1079 wild soybean population showed distinctive subpopulations (Fig. 2 and S3), as shown in the analyses of the entire 4234 population. The groupings were consistent with geographic distributions of the collection sites. Korean accessions and Japanese accessions formed a unique subpopulation, respectively. Chinese formed two subpopulations. Five accessions from the Russian border clustered together with those from northeast China. A strong relationship between subpopulations and their geographic distribution was notably exemplified by accessions from Jeju Island located 130 km off the southern coast of the Korean Peninsula; although they belong to the Korean subpopulation, all accessions from Jeju Island formed a subgroup that was the closest to the Japanese subpopulation (Fig. 2c).
Population structures of 1957 domesticated and 1079 wild soybean accessions in the 3036 non-redundant soybean accession set. a Principal components (PC) of SNP variation in the domesticated population. The plots show the first three principal components. The countries of collection or improvement status of the soybean accessions in a and c are represented by two-letter codes—CN, China; IP, improved breeding line; JP, Japan; ND, not determined; NK, North Korea; RS, Russia; and SK (KR), South Korea. b Scree plot of the PC number and their contribution to variance from principal component analysis of the domesticated accessions. c Principal components of SNP variation in the wild population. The plots show the first three principal components. A cluster of accessions from Jeju Island is indicated by a dashed eclipse. d Scree plot of the PC number and their contribution to variance from principal component analysis of the wild accessions
Detection of SNPs associated with domestication history
Domestication is a process of continuous artificial selection of a group of traits, collectively called domestication syndrome. The domestication process has produced selective sweeps with significant reductions in nucleotide diversity (Chung et al. 2014; Doebley et al. 2006; Hufford et al. 2012) on limited regions of the genome (approximately 5–10% of the genome). Numerous recent whole-genome resequencing studies have effectively detected the selective sweeps, which are associated with domesticated genes (Meyer and Purugganan 2013) by examining reduction of diversity (ROD) in windows along chromosomes. However, our SoyaSNP array data are not dense enough to detect the reduction of diversity. Thus, we attempted to detect SNPs associated with domestication using a case–control GWAS method that analyzed binary domestication phenotypes, which were determined by presence (G. max) or absence (G. soja) of domestication.
To test whether our case–control GWAS method enabled to find genes or chromosomal regions underlining binary phenotypes in our 3036 non-redundant population, we chose two highly studied phenotypes—flower and seed-coat colors—which are monogenic and multigenic, respectively. Because our population was highly structured, we performed logistic regression model analysis conditional on a list of subpopulation-specific SNPs. The selected specific SNPs included one perfect domestication-specific SNP with each allele being perfectly correlated with G. max or G. soja membership of soybean accessions, and ten subpopulation-specific SNP within each of the G. max and G. soja populations (Table S4). The flower color phenotypes were divided into absence or presence of anthocyanin deposition colors. The seed-coat color phenotypes were divided into absence or presence of anthocyanin deposition colors. Using the conditional logistic regression model, we detected a broad and strong peak for flower color with the most significant SNP (max − log10P = 80.6) located at 17,877,234 on chromosome 13 (Figs. 3a, S4, and S5c). This peak area contained the W1 locus, which is the major locus determining flower color (Zabala and Vodkin 2007). However, the most significant SNPs were located ~ 500 kb off the position of the flavonoid 3′5′-hydroxylase gene, which is the causal gene of the W1 locus. To understand this region, we estimated pairwise LD for SNPs from 16.8 to 18.8 Mb. A strong LD pattern was observed between all the SNPs under the most significant SNPs; however, no clear LD pattern was observed near the flavonoid 3′5′-hydroxylase gene. Although the result might be due to a SNP density insufficient for a long LD block frequently observed in soybean whose LD decayed to half of its maximum at > 100 kb with genome resequencing data (Chung et al. 2014; Lam et al. 2010; Valliyodan et al. 2016), it has been often observed that a causal gene for a strong peak are not always corresponding with the highest −log10P value (Segura et al. 2012; Yano et al. 2016).
Genome-wide association scans for 3036 soybean accessions for flower color, seed-coat color, and domestication. a Manhattan plot for flower color. The solid horizontal line denotes the Bonferroni-adjusted significance threshold. Chromosomal regions of known genes (T, I, R, W1) or loci (PD05 and qSW) are indicated by dashed vertical lines. b Manhattan plot for seed-coat color. c Manhattan plot for domestication. d Local Manhattan plot (top) and LD heatmap (bottom) surrounding the T locus on chromosome 6. Dashed lines indicate the region of the T locus. Physical locations (kb) are indicated under the Manhattan plot. e Local Manhattan plot (top) and LD heatmap (bottom) surrounding the qSW locus on chromosome 17. A bar indicates the region of the qSW locus
We detected more than 30 peaks exceeding a significant threshold (-log10P ≥ 7.07) for seed-coat color (Figs. 3b, S4, and S5). The top three peaks were correlated with three known major loci (I, R, and T on chromosomes 8, 9, and 6, respectively) that control the deposition of various anthocyanin pigments in seed coat (Yang et al. 2010). The highest peak was generated from a chromosomal region surrounding the inverted CHS gene repeats, which is the causal region of the I locus (Clough et al. 2004). A strong LD pattern was observed at the I locus region. An SNP AX-90432942 on chromosome 6 with the second highest − log10P value = 69.8 was generated from flavonoid 3′ hydroxylase, which is the causal gene of the T locus. Finally, the SNPs near the R2R3 MYB transcription factor gene, a strong candidate gene for the R locus reported by Gillman et al. (2011), were not significantly associated with seed-coat colors. However, the gene is one of the R2R3 MYB transcription factor genes tandemly repeated in this chromosomal region and numerous highly significant SNPs were observed 100-kb away from the proposed R gene. Interestingly, the peak on chromosome 13 is located on the W1 locus that influences seed-coat color in a case of the homozygous recessive it genotypes (Palmer et al. 2004). Considering such a wide range of soybean seed-coat color variations, the detection of numerous minor peaks is not surprising, as reported by Song et al. (2016), although it is still surprising to detect this large number of significant peaks using binary phenotyping data. Nevertheless, we think that some of those minor peaks are inevitably false because the limited number of conditional SNPs could not correct for all inflation of the statistic caused by population substructure.
Since our logistic regression model could readily detect loci associated with flower and seed-coat colors using binary phenotypes, we performed GWAS for domestication syndrome using the binary domestication phenotypes. For this analysis, we excluded the perfect domestication-specific SNP in the list of subpopulation-specific SNPs (Table S4). We detected numerous peaks for domestication syndrome over the genome, as expected from the previous studies (Chung et al. 2014; Hufford et al. 2012; Meyer and Purugganan 2013) that showed that domestication features covered approximately ~ 7% of the crop genome. To examine whether previously detected domestication regions were also detected in the current study, we compared peak locations from our study with selective signals previously detected for two domestication traits, pod dehiscence and seed weight (Figs. 3 and S5d), which are two of the most critical domestication traits among the traits assayed by Zhou et al. (2015). The 190-kb region (PD05) responsible for pod dehiscence was also detected in our study with three highly significant SNPs (-log10P ≥ 20), and the qSW locus for seed weight was detected with > 30 highly significant SNPs (-log10P ≥ 20). Lengths of strong LD blocks under the peaks corresponding to the PD05 and qSW loci were not sufficient to define the selective sweeps that have been known to be > 100 kb, as observed in our LD analyses for the flower and seed-coat color loci. Flower and seed-coat colors analyzed in this study are considered domestication- or diversification-related morphological features because nearly all wild soybean accessions have purple flowers and black seed coats. As expected, their major loci, W1, I, R, and T were also detected with highly significant SNPs (-log10P ≥ 20) in this GWAS for domestication syndrome (Fig. 3).
Extraction of a representative set
Although we used excessive number of Korean wild accessions in the above analysis, Korean accessions formed a unique group from those of neighbor countries. However, since a fundamental assumption of model-based methods, such as ADMIXTRUE and PCA, is that the sample available for analysis is representative of the entire population distribution, sample sizes of subpopulations can substantially affect population stratification and ancestral population inference (McVean 2009; Shringarpure and Xing 2014). To investigate the possibility that excessive numbers of domesticated or Korean soybean accessions might have caused bias in inference of population structure of wild and domesticated soybeans, we obtained representative domesticated and wild soybean sets by selecting one from each of tightly distributed soybean miniclusters in the PCA plots, with a caution that overall distribution patterns are maintained (Fig. S6). For the representative set of wild soybeans, we filtered the population of the tightly distributed Korean accessions and selected 50 diverse Korean wild accessions (Table S2). In addition, four wild soybean anomalies misplaced to subpopulations different from subpopulations predicted by their collection site records were excluded: three Korean and one Chinese accessions (Fig. S7). For the representative set of domesticated soybeans, we selected 50 diverse G. max accessions that represent diversity of 1957 G. max accessions. The results of the AMOVA indicated that the overall genetic structure observed in the 3036 non-redundant soybean population was well represented by the extracted representative set with some decrease in genetic diversity in the G. max population (Table S5). PCA plots from the resultant representative set of 194 soybean accessions showed distribution patterns similar to those from the 3036 non-redundant soybean accessions, although relative sizes of G. max and G. soja distributions in the PCA spaces constructed with the first and third PCs were reversed. The last drop of eigenvalues from the PCA runs occurred between K = 5 and K = 6 (Fig. S8), indicating that there were five distinct subpopulations.
Center of soybean domestication
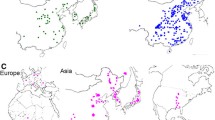
Because G. soja can be found in situ across most of the East Asia, it is important to establish the population structure, if any, of a diverse collection of G. soja accessions and to associate one or more of these populations with a collection of domesticated G. max varieties. To perform these experiments, we analyzed the population structure of the representative set of 194 soybean accessions with ADMIXTURE and found K = 5 populations based on the estimated CV error plot (Fig. S8). Thus, the soybean accessions were partitioned into one G. max (Gm) and four G. soja (Gs-I, Gs-II, Gs-III, and Gs-IV) subgroups (Fig. 4a–c). Wild accessions from China were divided into two subgroups, Gs-I and Gs-II (Fig. 4d). The Gs-I group showed the least diversity (Table 1), and most of them distributed in the middle region of the Yellow River basin. The Gs-II group was dispersed across northeast China, south China, and the Russian border of northeast China. Interestingly, this grouping result is remarkably similar to that obtained by a recent comprehensive study that showed that, by analyzing a total of 712 G. soja individuals from 40 natural populations in China, Chinese wild soybeans were grouped into two main subgroups, which were one from the Yellow River basin and the other from northeast China and south China (Guo et al. 2012). The Gs-IV distributed in Japan. The Gs-III showed the greatest diversity and distributed in South Korea and most of them appeared to be ancient admixture between Gs-II and Gs-IV. Interestingly, despite clear separation of the Chinese G. soja, diversity level of the combined population of Gs-I and Gs-II was similar to that of Korean or Japanese G. soja. An independent Gm group appeared from K = 2 to K = 5 (Fig. 4c). Interestingly, major genomic fractions of the Gm subgroup consistently appeared as minor genomic fractions of the Gs-IV and minor genomic fractions of the Gm group were a genomic fraction of Gs-I ancestry. The results suggested that after domestication of the Gm subgroup from the Gs-IV subgroup, the Gm subgroup was substantially diversified by introgression of the Gs-I genomic fractions. One of the Gs-I accessions, PI 549046, appeared to be a G. max × G. soja hybrid, although its genomic fraction (~ 22%) from G. max was lower than our hybrid filtration criteria (30% domesticated ancestry), which was less stringent than 20% domesticated ancestry used in other admixture studies (e.g., Wang et al. (2017)).
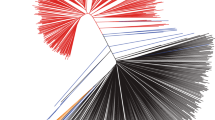
Identification of the domestication center of G. max. a, b Principal components plots of SNP variation. PC1, PC2, and PC3 indicate score of principal components 1, 2, and 3, respectively. Each of PC1, PC2, and PC3 explained 12.0, 5.2, and 2.6% of variance in the data. Countries of collection of the soybean accessions and species names are represented by two-letter codes—CN, China; JP, Japan; NK, North Korea; RS, Russia; SK, South Korea; Gm, G. max; and Gs, G. soja. A putative hybrid PI 549046 is labeled. c Population structure of 50 G. max (Gm) and 144 G. soja (Gs-I, Gs-II, Gs-III, and Gs-IV) accessions inferred using ADMIXTURE. Each color represents one population. PI 549046 showed ~ 20% of ancestral genomic fractions from G. max. d Geographic distribution of the four G. soja subgroups. Gs-I is red, Gs-II green, Gs-III orange, and Gs-IV blue. e–g Neighbor-joining phylogenetic tree of 194 soybean accessions based on the SNPs genotyped by the 180 K AXIOM SoyaSNP array, with evolutionary distances measured by the p distance. The taxa used in the neighbor-joining tree and bootstrap values from 1000 bootstrap replications at branches are described in Fig. S9. e Phylogenetic tree based on 117,095 SNPs. f Phylogenetic tree based on 108,899 SNPs, which are weakly or not significantly associated with domestication traits. g Phylogenetic tree based on 8196, which are very significantly associated with domestication traits. PI 549046 from group Gs-I clusters between Gm and Gs-IV likely because of contribution of ancestral genomic fraction from Gm
We constructed a neighbor-joining tree for the representative soybean set (Figs. 4e and S9). The tree showed that all G. max accessions formed a monophyletic cluster. Although G. max was artificially selected recently, terminal branch lengths were similar to those of G. soja likely because of ascertainment bias that more SNPs were selected from G. max than from G. soja (Lee et al. 2015). The population of the nearest branches, which were basal to the G. max soybean lineage, was G. soja subgroup Gs-IV. Within the Gs-IV that contains wild soybeans from Japan, those from eastern Japan area were closer to the G. max soybean lineage. To measure population differences and similarities, we calculated the fixation index values (FST) (Holsinger and Weir 2009) between G. max and each G. soja population (Table 2). The pairwise FST values ranged from 0.201 to 0.334. The value of FST for the G. max and Gs-IV populations was the smallest, suggesting that G. max was domesticated directly from G. soja subpopulation Gs-IV. The level of population differentiation among G. soja subpopulation was much lower than that between G. soja and G. max, similar to the case of rice (Huang et al. 2012). However, FST values between G. soja subpopulations corresponded with their geographic distances. The regions containing those wild populations that are phylogenetically close with cultivars could be proposed as the domestication region of crops (Matsuoka et al. 2002; Spooner et al. 2005). Thus, our results suggested that soybeans had been most likely domesticated only once in eastern Japan.
Because the number of G. soja accessions from China in our representative set was smaller than those from Korea and Japan, the population structure revealed by our representative set was further resolved by incorporating the SNP data from 62 G. soja accession genomes resequenced by Zhou et al. (2015) By intersecting these SNPs with the set of 117,095 high-quality SNPs selected in this study, we extracted 103,801 SNPs, which were shared between the genome resequencing data and our representative set. Of the 62 accessions, only 45 were incorporated into the representative set because of the high level (eleven accessions, > 20%) of heterozygous SNPs, hybrid (one), and overlapping (five) (Fig. S10 and Table S6). The resultant expanded set contained twelve diverse accessions from Zhejiang, China, and one accession from Taiwan, thus increasing geographic coverage of this study further down to southern China. In results, the diversity level of G. soja accessions from China and its Russian border was similar to that from Korea or Japan (Table 1). Population structure of the expanded set inferred from ADMIXTURE and PCA was quite similar to that of our representative set, although the G. max accessions appeared to be divided into two groups likely because of the high level of heterozygous SNPs from the genome resequencing data (Fig. S11, S12, and Table 2). Phylogenetic analysis and estimated FST values between subpopulations indicated that G. soja accessions collected from eastern Japan were closest to the G. max soybean lineage.
To evaluate the distribution of SNPs associated with domestication syndrome across soybean subpopulations, we divided the 117,095 SNPs into 8196 SNPs highly significantly associated with domestication traits and 108,899 SNPs weakly or not significantly associated with domestication traits. The 8196 SNPs (-log10P > 17) were selected because the previous studies have shown that ~ 7% of the crop genome is domestication related (Chung et al. 2014; Hufford et al. 2012; Xu et al. 2012). Our GWAS also indicated that, in our GWAS population, strong LD extent under the highly significant SNPs in a peak tended to be much shorter than chromosomal extent under all the significant SNPs of the peak. The tree constructed from the 108,899 SNPs (Fig. 4f) was similar to that constructed from the 117,095 SNPs in their overall grouping and branch patterns, except that the branch length and grouping of the G. max clade were slightly different to each other. Grouping patterns in the tree constructed from the 8196 SNPs (Fig. 4g) were similar to those in the trees constructed from both 108,899 SNPs and 117,095 SNPs, except that the putative hybrid PI 549046 moved the closest to the G. max clade. Interestingly, the lengths of the basal and terminal branches for the G. max clade and the G. soja Gs-IV clade in the tree from the 8196 SNPs became distinctively longer than those in the other G. soja clades. The results indicated that initial major artificial selection for soybean domestication was limited to a G. soja group from Japan.
Discussion
The present study analyzed genome-wide SNP variations obtained from thousands of soybean accessions, the majority of which were collected from the Korean peninsula. The results provide insight into the development of strategies for efficient and directed germplasm use as well as for collection of novel landraces and wild relatives. Population structure and grouping analyses revealed strong correlations between genetic distance and geographic distance in G. soja (wild soybean) populations and weak correlations in G. max (domesticated soybean) populations. G. soja accessions were divided into four distinct subgroups; Gs-I and Gs-II from China and its Russian border, Gs-III from Korea, and Gs-IV from Japan. The results suggest that although the Korean territory is much smaller than Chinese and Japanese territories, the sea-imposed geographic separation among these countries has been a major contributor to the evolutionary divergence of G. soja. Most of the Gs-III group from Korea appeared to be ancient admixtures between Gs-II and Gs-IV, suggesting that G. soja spread from each of China and Japan might be mixed in Korea. Thus, Korean wild soybeans are expected to be rich in terms of diversity than the other countries’ wild soybeans because they alone provide variations from two large subgroups. Interestingly, accessions from Jeju Island off the southern coast of the Korean Peninsula are the closest grouped to the Gs-IV group among members of the Gs-III group, indicating that although our estimated FST values between G. soja groups denied appearance of a new distinct group, more extensive sampling from diverse areas will likely reveal better correlations between geographic and genetic distances among G. soja subpopulations. The G. max population was divided into four subgroups. However, it was apparent that the subgrouping did not reflect geographic origins. Particularly, landraces from North Korea that would be considered true landraces based on their collection time appeared in every subgroup. The majority of South Korean landraces that had been collected recently were grouped together. The majority of improved accessions from the USA were clustered closely together, supporting a previous observation (Hyten et al. 2006). Taken together, our results suggest that while G. soja germplasm will require additional sampling from diverse indigenous areas to expand the germplasm base, G. max germplasm is saturated in terms of genetic diversity. Thus, extensive genotyping and phenotyping of extant G. max germplasm would be the next step to expand the germplasm base of G. max.
Our results provided strong support for a single origin of G. max from eastern region in Japan, although pointing to a specific region in Japan likely requires analysis of more extensive wild and landrace soybean accessions from Japan. Whether a crop species stems from a single domestication event or from multiple independent domestications has been consistent with whether the domesticated species are monophyletic or polyphyletic, respectively, in the phylogenetic trees constructed from both the domesticated and wild progenitor species. Although diversity of chloroplast DNA, which represents maternal lineage of soybean, revealed multiple lineages of domesticated soybeans, analyses of recent genome-wide soybean variation data (Chung et al. 2014; Guo et al. 2010; Lam et al. 2010; Zhou et al. 2015) consistently showed the monophyletic nature of G. max, as observed in this study. In other words, recent soybean phylogenetic studies collectively indicated a single origin of G. max. The best examples of monophyletic grouping are wheat and barley, which appear to have been domesticated once from their wild ancestors in the Fertile Crescent (Badr et al. 2000; Ozkan et al. 2002). The origin of barley was further supported by the genome sequences of five 6000-year-old barley grains (Mascher et al. 2016). In cases of rice and common bean that showed polyphyletic groupings, single or multiple regions of origin of these crop species are still contentious (Bitocchi et al. 2012; Huang et al. 2012; Molina et al. 2011). This controversy may have arisen because most modern wild accessions studied represent descendants of ancient feralization of admixed accessions that resulted from hybridization events between domesticated species and wild species populations after domestication (Wang et al. 2017), indicating that one of the previously thought independent origin regions might be a secondary origin region. The grouping of PI 459046 in this study is a good example that shows how hybrids could mislead inference of relationship between wild and domesticated crop species.
A recent comprehensive study of the archeological records for soybean from Japan, China, and Korea indicated that Japan could have been a source of a large-seeded landrace of domesticated soybean that spread to Korea and subsequently to China (Lee et al. 2011). The archeological records suggest that selection of large seed sizes occurred in Japan (Lee et al. 2011; Nakayama 2015) by 5500 calibrated years (cal) before present (BP) and in Korea (Lee et al. 2011) by 3500 cal BP. Seed size is clearly a domestication trait because the seed size of G. soja is much smaller than that of G. max landraces (Broich and Palmer 1980). However, the archeological data were interpreted to suggest the multiple origins hypothesis of soybean. One particular reason is that the excavated tiny seeds were as old as 9000–8600 cal BP in northern China and 7000 cal BP in Japan. However, the size of the seeds is similar to that of the seeds of present-day wild soybeans, and so would have been quite different from the landraces already grown in China by 2500 BP. Another reason is that the interpretation was greatly influenced by a previous report that diversity of chloroplast DNA SSRs in wild and domesticated soybeans showed evidence for multiple origins of domesticated soybean (Xu et al. 2002). However, as mentioned above, numerous recent genome-wide soybean genome variation studies consistently show a single origin of G. max.
One of the main reasons that the previous studies pointed different regions in China as the center of soybean domestication is likely sampling bias. Our results suggested that wild accessions from China had genetic diversity level almost equal to those from Korea or Japan. However, most previous studies tended to neglect this fact. In an extreme case (Han et al. 2016), no accession from Korea and Japan was used, with the conclusion that central China is the initial domestication region. Another confounding factor is the inclusion of hybrid soybeans from natural mating between G. soja and G. max. Hybrid soybeans were not recognized in many previous soybean population studies, although hybrids between wild and domesticated species have been increasingly regarded as a major problem in studies of crop domestication history (Bitocchi et al. 2012; Wang et al. 2017). Furthermore, it was often assumed that a region in China is a center of soybean domestication because hybrid soybeans are frequently found in China (Han et al. 2016). However, of the 50 hybrids that we removed to avoid their potential confounding effects in this analysis, the majority (36 of 50) were accessions from Korea. The domestication of domesticated plant species from their wild ancestors arose from rapid evolutionary changes in the past 13,000 years of Holocene human history (Diamond 2002; Larson et al. 2014). The list of origins and the list of the most productive areas of most of major crops in the modern world are almost mutually exclusive. This could be explained by that the domestication origin of a crop was merely a region to which the most numerous and most valuable domesticable wild plant species were native. In this respect, our result that shows Japan as the domestication origin of soybean is not totally unexpected one.
Expanding on previous studies that reported genome-wide polymorphism data of soybean germplasm (Bandillo et al. 2015; Chung et al. 2014; Lam et al. 2010; Valliyodan et al. 2016; Wang et al. 2016; Zhou et al. 2015), our results show that accessions intensively collected from Korea, which is a small area of the entire soybean distribution, provide sufficient amounts of data to underpin genome-wide population genetic questions that have been neglected or misled in the context of diversity and domestication panels of extant individuals. Our analysis demonstrates the value of current germplasm collections and how to expand the germplasm base. Furthermore, the findings show that a single major domestication event had occurred in a region of Japan. In addition, the high-density SNP array data enabled detection of domestication-associated SNPs and regions controlling important agronomic traits in a highly accurate manner. This suggests that our results will likely be useful for marker-assisted selection and genomic prediction to utilize unexplored genetic diversity in the soybean germplasm.
Author contribution statement
SCJ and JKM designed the study; SCJ, JKM, SKP, NJ, MSC, STK, and EP collected data; SCJ, JKM, MSK, KL, SRL, and NK performed the analyses; SCJ and JKM wrote the paper.
References
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19:1655–1664
Badr A, Muller K, Schafer-Pregl R, El Rabey H, Effgen S, Ibrahim HH, Pozzi C, Rohde W, Salamini F (2000) On the origin and domestication history of Barley (Hordeum vulgare). Mol Biol Evol 17:499–510
Bandillo N, Jarquin D, Song Q, Nelson RL, Cregan P, Specht J, Lorenz A (2015) A population structure and genome-wide association analysis on the USDA soybean germplasm collection. Plant Genome. https://doi.org/10.3835/plantgenome2015.04.0024
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21:263–265
Bitocchi E, Nanni L, Bellucci E, Rossi M, Giardini A, Zeuli PS, Logozzo G, Stougaard J, McClean P, Attene G, Papa R (2012) Mesoamerican origin of the common bean (Phaseolus vulgaris L.) is revealed by sequence data. Proc Natl Acad Sci USA 109:E788–E796
Broich SL, Palmer RG (1980) A cluster analysis of wild and domesticated soybean phenotypes. Euphytica 29:23–32
Broich SL, Palmer RG (1981) Evolutionary studies of the soybean: The frequency and distribution of alleles among collections of Glycine max and G. soja of various origin. Euphytica 30:55–64
Browning SR, Browning BL (2007) Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am J Hum Genet 81:1084–1097
Chung WH, Jeong N, Kim J, Lee WK, Lee YG, Lee SH, Yoon W, Kim JH, Choi IY, Choi HK, Moon JK, Kim N, Jeong SC (2014) Population structure and domestication revealed by high-depth resequencing of Korean cultivated and wild soybean genomes. DNA Res 21:153–167
Clough SJ, Tuteja JH, Li M, Marek LF, Shoemaker RC, Vodkin LO (2004) Features of a 103-kb gene-rich region in soybean include an inverted perfect repeat cluster of CHS genes comprising the I locus. Genome 47:819–831
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, Sherry ST, McVean G, Durbin R, Genomes Project Analysis G (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158
Diamond J (2002) Evolution, consequences and future of plant and animal domestication. Nature 418:700–707
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127:1309–1321
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Food and Agriculture Organization of the United Nations (2010) The second report on the state of the world’s plant genetic resources for food and agriculture. FAO, Rome
Gabriel SB, Schaffner SF, Nguyen H, Moore JM, Roy J, Blumenstiel B, Higgins J, DeFelice M, Lochner A, Faggart M, Liu-Cordero SN, Rotimi C, Adeyemo A, Cooper R, Ward R, Lander ES, Daly MJ, Altshuler D (2002) The structure of haplotype blocks in the human genome. Science 296:2225–2229
Gillman JD, Tetlow A, Lee JD, Shannon JG, Bilyeu K (2011) Loss-of-function mutations affecting a specific Glycine max R2R3 MYB transcription factor result in brown hilum and brown seed coats. BMC Plant Biol 11:155
Gizlice Z, Carter TE, Burton J (1994) Genetic base for North American public soybean cultivars released between 1947 and 1988. Crop Sci 34:1143–1151
Guo J, Wang Y, Song C, Zhou J, Qiu L, Huang H, Wang Y (2010) A single origin and moderate bottleneck during domestication of soybean (Glycine max): implications from microsatellites and nucleotide sequences. Ann Bot 106:505–514
Guo J, Liu Y, Wang Y, Chen J, Li Y, Huang H, Qiu L, Wang Y (2012) Population structure of the wild soybean (Glycine soja) in China: implications from microsatellite analyses. Ann Bot 110:777–785
Han Y, Zhao X, Liu D, Li Y, Lightfoot DA, Yang Z, Zhao L, Zhou G, Wang Z, Huang L, Zhang Z, Qiu L, Zheng H, Li W (2016) Domestication footprints anchor genomic regions of agronomic importance in soybeans. New Phytol 209:871–884
Holsinger KE, Weir BS (2009) Genetics in geographically structured populations: defining, estimating and interpreting F ST. Nat Rev Genet 10:639–650
Huang X, Kurata N, Wei X, Wang ZX, Wang A, Zhao Q, Zhao Y, Liu K, Lu H, Li W, Guo Y, Lu Y, Zhou C, Fan D, Weng Q, Zhu C, Huang T, Zhang L, Wang Y, Feng L, Furuumi H, Kubo T, Miyabayashi T, Yuan X, Xu Q, Dong G, Zhan Q, Li C, Fujiyama A, Toyoda A, Lu T, Feng Q, Qian Q, Li J, Han B (2012) A map of rice genome variation reveals the origin of cultivated rice. Nature 490:497–501
Hufford MB, Xu X, van Heerwaarden J, Pyhajarvi T, Chia JM, Cartwright RA, Elshire RJ, Glaubitz JC, Guill KE, Kaeppler SM, Lai J, Morrell PL, Shannon LM, Song C, Springer NM, Swanson-Wagner RA, Tiffin P, Wang J, Zhang G, Doebley J, McMullen MD, Ware D, Buckler ES, Yang S, Ross-Ibarra J (2012) Comparative population genomics of maize domestication and improvement. Nat Genet 44:808–811
Hwang EY, Song Q, Jia G, Specht JE, Hyten DL, Costa J, Cregan PB (2014) A genome-wide association study of seed protein and oil content in soybean. BMC Genom 15:1
Hymowitz T (2004) Speciation and cytogenetics. In: Boerma HR, Specht JE (eds) Soybeans: improvement, production, and uses. American Society of Agronomy, Madison, pp 97–136
Hymowitz T, Kaizuma N (1981) Soybean seed protein electrophoresis profiles from 15 Asian countries or regions: hypotheses on paths of dissemination of soybeans from China. Econ Bot 13:10–23
Hyten DL, Song Q, Zhu Y, Choi IY, Nelson RL, Costa JM, Specht JE, Shoemaker RC, Cregan PB (2006) Impacts of genetic bottlenecks on soybean genome diversity. Proc Natl Acad Sci USA 103:16666–16671
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lam HM, Xu X, Liu X, Chen W, Yang G, Wong FL, Li MW, He W, Qin N, Wang B, Li J, Jian M, Wang J, Shao G, Wang J, Sun SS, Zhang G (2010) Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat Genet 42:1053–1059
Larson G, Piperno DR, Allaby RG, Purugganan MD, Andersson L, Arroyo-Kalin M, Barton L, Climer Vigueira C, Denham T, Dobney K, Doust AN, Gepts P, Gilbert MT, Gremillion KJ, Lucas L, Lukens L, Marshall FB, Olsen KM, Pires JC, Richerson PJ, Rubio de Casas R, Sanjur OI, Thomas MG, Fuller DQ (2014) Current perspectives and the future of domestication studies. Proc Natl Acad Sci USA 111:6139–6146
Lee GA, Crawford GW, Liu L, Sasaki Y, Chen X (2011) Archaeological soybean (Glycine max) in East Asia: Does size matter? PLoS ONE 6:e26720
Lee YG, Jeong N, Kim JH, Lee K, Kim KH, Pirani A, Ha BK, Kang ST, Park BS, Moon JK, Kim N, Jeong SC (2015) Development, validation and genetic analysis of a large soybean SNP genotyping array. Plant J 81:625–636
Li YH, Li W, Zhang C, Yang L, Chang RZ, Gaut BS, Qiu LJ (2010) Genetic diversity in domesticated soybean (Glycine max) and its wild progenitor (Glycine soja) for simple sequence repeat and single-nucleotide polymorphism loci. New Phytol 188:242–253
Mascher M, Schuenemann VJ, Davidovich U, Marom N, Himmelbach A, Hubner S, Korol A, David M, Reiter E, Riehl S, Schreiber M, Vohr SH, Green RE, Dawson IK, Russell J, Kilian B, Muehlbauer GJ, Waugh R, Fahima T, Krause J, Weiss E, Stein N (2016) Genomic analysis of 6000-year-old cultivated grain illuminates the domestication history of barley. Nat Genet 48:1089–1093
Matsuoka Y, Vigouroux Y, Goodman MM, Sanchez GJ, Buckler E, Doebley J (2002) A single domestication for maize shown by multilocus microsatellite genotyping. Proc Natl Acad Sci USA 99:6080–6084
McCarthy MI, Abecasis GR, Cardon LR, Goldstein DB, Little J, Ioannidis JP, Hirschhorn JN (2008) Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat Rev Genet 9:356–369
McCouch SR, McNally KL, Wang W, Sackville Hamilton R (2012) Genomics of gene banks: a case study in rice. Am J Bot 99:407–423
McVean G (2009) A genealogical interpretation of principal components analysis. PLoS Genet 5:e1000686
Meyer RS, Purugganan MD (2013) Evolution of crop species: genetics of domestication and diversification. Nat Rev Genet 14:840–852
Molina J, Sikora M, Garud N, Flowers JM, Rubinstein S, Reynolds A, Huang P, Jackson S, Schaal BA, Bustamante CD, Boyko AR, Purugganan MD (2011) Molecular evidence for a single evolutionary origin of domesticated rice. Proc Natl Acad Sci USA 108:8351–8356
Nakayama S (2015) Domestication of the soybean (Glycine max) and morphological differentiation of seeds in the Jomon period. Jpn J Histor Bot 23:33–42
Ozkan H, Brandolini A, Schafer-Pregl R, Salamini F (2002) AFLP analysis of a collection of tetraploid wheats indicates the origin of emmer and hard wheat domestication in southeast Turkey. Mol Biol Evol 19:1797–1801
Palmer RG, Pfeiffer TW, Buss GR, Kilen TC (2004) Qualitative genetics. In: Boerma HR, Specht HE (eds) Soybeans: improvement, production, and uses, 3rd edn. ASA, CSSA, and SSSA, Madison, pp 137–214
Patterson N, Price AL, Reich D (2006) Population structure and eigenanalysis. PLoS Genet 2:e190
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Segura V, Vilhjalmsson BJ, Platt A, Korte A, Seren U, Long Q, Nordborg M (2012) An efficient multi-locus mixed-model approach for genome-wide association studies in structured populations. Nat Genet 44:825–830
Shringarpure S, Xing EP (2014) Effects of sample selection bias on the accuracy of population structure and ancestry inference. G3 4:901–911
Song Q, Hyten DL, Jia G, Quigley CV, Fickus EW, Nelson RL, Cregan PB (2013) Development and evaluation of SoySNP50K, a high-density genotyping array for soybean. PLoS ONE 8:e54985
Song J, Liu Z, Hong H, Ma Y, Tian L, Li X, Li YH, Guan R, Guo Y, Qiu LJ (2016) Identification and validation of loci governing seed coat color by combining association mapping and bulk segregation analysis in soybean. PLoS ONE 11:e0159064
Spooner DM, McLean K, Ramsay G, Waugh R, Bryan GJ (2005) A single domestication for potato based on multilocus amplified fragment length polymorphism genotyping. Proc Natl Acad Sci USA 102:14694–14699
Turner SD (2014) qqman: an R package for visualizing GWAS results using Q–Q and Manhattan plots. biorXiv
Valliyodan B, Dan Q, Patil G, Zeng P, Huang J, Dai L, Chen C, Li Y, Joshi T, Song L, Vuong TD, Musket TA, Xu D, Shannon JG, Shifeng C, Liu X, Nguyen HT (2016) Landscape of genomic diversity and trait discovery in soybean. Sci Rep 6:23598
Vaughan DA, Lu BR, Tomooka N (2008) The evolving story of rice evolution. Plant Sci 174:394–408
Wang J, Chu S, Zhang H, Zhu Y, Cheng H, Yu D (2016) Development and application of a novel genome-wide SNP array reveals domestication history in soybean. Sci Rep 6:20728
Wang H, Vieira FG, Crawford JE, Chu C, Nielsen R (2017) Asian wild rice is a hybrid swarm with extensive gene flow and feralization from domesticated rice. Genome Res 27:1029–1038
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Xu H, Abe J, Gai Y, Shimamoto Y (2002) Diversity of chloroplast DNA SSRs in wild and cultivated soybeans: evidence for multiple origins of cultivated soybean. Theor Appl Genet 105:645–653
Xu X, Liu X, Ge S, Jensen JD, Hu F, Li X, Dong Y, Gutenkunst RN, Fang L, Huang L, Li J, He W, Zhang G, Zheng X, Zhang F, Li Y, Yu C, Kristiansen K, Zhang X, Wang J, Wright M, McCouch S, Nielsen R, Wang J, Wang W (2012) Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat Biotechnol 30:105–111
Yang K, Jeong N, Moon JK, Lee YH, Lee SH, Kim HM, Hwang CH, Back K, Palmer RG, Jeong SC (2010) Genetic analysis of genes controlling natural variation of seed coat and flower colors in soybean. J Hered 101:757–768
Yano K, Yamamoto E, Aya K, Takeuchi H, Lo PC, Hu L, Yamasaki M, Yoshida S, Kitano H, Hirano K, Matsuoka M (2016) Genome-wide association study using whole-genome sequencing rapidly identifies new genes influencing agronomic traits in rice. Nat Genet 48:927–934
Zabala G, Vodkin LO (2007) A rearrangement resulting in small tandem repeats in the F3′5′H gene of white flower genotypes is associated with the soybean W1 locus. Crop Sci 47:S113–S124
Zhou Z, Jiang Y, Wang Z, Gou Z, Lyu J, Li W, Yu Y, Shu L, Zhao Y, Ma Y, Fang C, Shen Y, Liu T, Li C, Li Q, Wu M, Wang M, Wu Y, Dong Y, Wan W, Wang X, Ding Z, Gao Y, Xiang H, Zhu B, Lee SH, Wang W, Tian Z (2015) Resequencing 302 wild and cultivated accessions identifies genes related to domestication and improvement in soybean. Nat Biotechnol 33:408–414
Acknowledgements
We thank Dr. Changyong Lee at Kongju National University for helpful comments in statistical analysis. We are grateful to Prof. Michael Gore at Cornell University for his critical reading of the revised version of this paper. S.C.J. was supported by the Next-Generation BioGreen 21 Program (PJ01321304), Rural Development Administration, and partly by the National Research Foundation grant (NRF-2018R1A2A2A05021904) funded by the Korea government and by the Korea Research Institute of Bioscience and Biotechnology Research Initiative Program. The work at Rural Development Administration was funded in part by Rural Development Administration Project No. PJ01155401.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Data accessibility
SNP genotype data are listed in Table S1 and are publicly available at Korean Soya Base (http://koreansoyabase.org/Data_Resource/).
Additional information
Communicated by Henry T. Nguyen.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Jeong, SC., Moon, JK., Park, SK. et al. Genetic diversity patterns and domestication origin of soybean. Theor Appl Genet 132, 1179–1193 (2019). https://doi.org/10.1007/s00122-018-3271-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-018-3271-7