Abstract
Key message
A major QTL controlling early flowering in broccoli × cabbage was identified by marker analysis and next-generation sequencing, corresponding to GRF6 gene conditioning flowering time in Arabidopsis.
Abstract
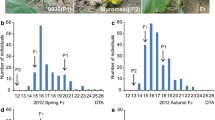
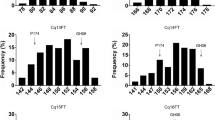
Flowering is an important agronomic trait for hybrid production in broccoli and cabbage, but the genetic mechanism underlying this process is unknown. In this study, segregation analysis with BC1P1, BC1P2, F2, and F2:3 populations derived from a cross between two inbred lines “195” (late-flowering) and “93219” (early flowering) suggested that flowering time is a quantitative trait. Next, employing a next-generation sequencing-based whole-genome QTL-seq strategy, we identified a major genomic region harboring a robust flowering time QTL using an F2 mapping population, designated Ef2.1 on cabbage chromosome 2 for early flowering. Ef2.1 was further validated by indel (insertion or deletion) marker-based classical QTL mapping, explaining 51.5% (LOD = 37.67) and 54.0% (LOD = 40.5) of the phenotypic variation in F2 and F2:3 populations, respectively. Combined QTL-seq and classical QTL analysis narrowed down Ef1.1 to a 228-kb genomic region containing 29 genes. A cabbage gene, Bol024659, was identified in this region, which is a homolog of GRF6, a major gene regulating flowering in Arabidopsis, and was designated BolGRF6. qRT-PCR study of the expression level of BolGRF6 revealed significantly higher expression in the early flowering genotypes. Taken together, our results provide support for BolGRF6 as a possible candidate gene for early flowering in the broccoli line 93219. The identified candidate genomic regions and genes may be useful for molecular breeding to improve broccoli and cabbage flowering times.





Similar content being viewed by others
References
Abe M, Abe M, Kobayashi Y, Yamamoto S, Daimon Y, Yamaguchi A, Ikeda Y, Ichinoki H, Notaguchi M, Goto K, Araki T (2005) FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science 309:1052–1056
Abe A, Kosugi S, Yoshida K, Natsume S, Takagi H, Kanzaki H, Matsumura H, Yoshida K, Mitsuoka C, Tamiru M (2012) Genome sequencing reveals agronomically important loci in rice using MutMap. Nat Biotechnol 30:174–178
Bergsma JG, Li C, Lesniewska K, Sivasithamparam K, Yang H (2008) Identification of quantitative trait loci (QTLs) influencing early vigour, height, flowering date, and seed size and their implications for breeding of narrow-leafed lupin (Lupinus angustifolius L.). Aust J Agric 59:527–535
Chen QJ, Zhang HY, Wang YJ, Li WY, Zhang F, Mao AJ, Cheng JH, Chen MY (2010) Mapping and analyzing QTLs of yield-associated agronomic traits of greenhouse cucumbers. Sci Agric Sin 43:112–122
Choi J, Hyun Y, Kang MJ, In Yun H, Yun JY, Lister C, Dean C, Amasino RM, Noh B, Noh YS, Choi Y (2009) Resetting and regulation of FLOWERING LOCUS C expression during Arabidopsis reproductive development. Plant J 57:918–931
Department for Environment, Food and Rural Affairs (DEFRA) (2010) Basic horticultural statistics for the United Kingdom. DEFRA UK. http://www.defra.gov.uk/
Fazio F, Staub JE, Stevens MR (2003) Genetic mapping and QTL analysis of horticultural traits in cucumber (Cucumis sativus L.) using recombinant inbred lines. Theor Appl Genet 107:864–874
Finley JW (2003) The antioxidant responsive element (ARE) may explain the protective effects of cruciferous vegetables on cancer. Nutr Rev 61:250–254
Fornara F, de Montaigu A, Coupland G (2010) SnapShot: control of flowering in Arabidopsis. Cell 141:550
Gómez-Lobato ME, Hasperuéb JH, Civelloa PM, Chaves AR, Martínez GA (2012) Effect of 1-MCP on the expression of chlorophyll degrading genes during senescence of broccoli (Brassica oleracea L.). Sci Hortic-Amsterdam 144:208–211
Ho WWH, Weigel D (2014) Structural features determining flower-promoting activity of Arabidopsis FLOWERING LOCUS T. Plant Cell 26:552–564
Jamalabadi JG, Saidi A, Karami E, Kharkesh M, Talebi R (2013) Molecular mapping and characterization of genes governing time to flowering, seed weight, and plant height in an intraspecific genetic linkage map of chickpea (Cicer arietinum). Biochem Genet 51:387–397
Ji XH, Yin L, Shen BY, Zhang L, Wang YG, Feng H (2013) Inheritance analysis of bolting correlated traits using mixed major gene plus polygene model in Brassica rapa. Chin Agric Sci Bull 29:76–82
Jung JH, Ju Y, Seo PJ, Lee JH, Park CM (2012) The SOC1-SPL module integrates photoperiod and gibberellic acid signals to control flowering time in Arabidopsis. Plant J 69:577–588
Kardailsky I, Shukla V, Ahn JH, Dagenais N, Christensen SK, Nguyen JT, Chory J, Harrison MJ, Weigel D (1999) Activation tagging of the floral inducer FT. Science 286:1962–1965
Kawamoto N, Endo M, Araki T (2015) Expression of a kinase-dead form of CPK33 involved in florigen complex formation causes delayed flowering. Plant Signal Behav 10:12
Kobayashi Y, Kaya H, Goto K, Iwabuchi M, Araki T (1999) A pair of related genes with antagonistic roles in mediating flowering signals. Science 286:1960–1962
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Lee H, Suh SS, Park E, Cho E, Ahn JH, Kim SG, Lee JS, Kwon YM, Lee I (2000) The AGAMOUS-LIKE 20 MADS domain protein integrates floral inductive pathways in Arabidopsis. Genes Dev 14:2366–2376
Lee JH, Yoo SJ, Park SH, Hwang I, Lee JS, Ahn JH (2007) Role of SVP in the control of flowering time by ambient temperature in Arabidopsis. Gene Dev 21:397–402
Lee J, Oh M, Park H, Lee I (2008) SOC1 translocated to the nucleus by interaction with AGL24 directly regulates leafy. Plant J 55:832–843
Li RQ, Yu C, Li YR, Lam TW, Yiu SM, Kristiansen K, Wang J (2009) SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25:1966–1967
Li CX, Lin HQ, Dubcovsky J (2015) Factorial combinations of protein interactions generate a multiplicity of florigen activation complexes in wheat and barley. Plant J 84:70–82
Lindhout P, Van Heusden S, Pet G, Van Ooijen JW, Sandbrink H, Verkerk R, Vrielink R, Zabel P (1994) Perspectives of molecular marker assisted breeding for earliness in tomato. Euphytica 79:279–286
Liu C, Chen H, Er HL, Soo HM, Kumar PP, Han JH, Liou YC, Yu H (2008) Direct interaction of AGL24 and SOC1 integrates flowering signals in Arabidopsis. Development 135:1481–1491
Liu SY, Liu YM, Yang XH, Tong CB, Edwards D, Parkin IAP, Zhao MX, Ma JX, Yu JY, Huang SM, Wang XY, Wang JY, Lu K, Fang ZY, Bancroft I, Yang TJ, Hu Q, Wang XF, Yue Z, Li HJ, Yang LF, Wu J, Zhou Q, Wang WX, King GJ, Pires JC, Lu CX, Wu ZY, Sampath P, Wang Z, Guo H, Pan SK, Yang LM, Min JM, Zhang D, Jin DC, Li WS, Belcram H, Tu JX, Guan M, Qi CK, Du DZ, Li JN, Jiang LC, Batley J, Sharpe AG, Park BS, Ruperao P, Cheng F, Waminal NE, Huang Y, Dong CH, Wang L, Li JP, Hu ZY, Zhuang M, Huang Y, Huang JY, Shi JQ, Mei DS, Liu J, Lee TH, Wang JP, Jin HZ, Li ZY, Li X, Zhang JF, Xiao L, Zhou YM, Liu ZS, Liu XQ, Qin R, Tang X, Liu WB, Wang YP, Zhang YY, Lee J, Kim HH, Denoeud F, Xu X, Liang XM, Hua W, Wang XW, Wang J, Chalhoub B, Paterson AH (2014) The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat Commun 5:3930
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods 25:402–408
Lu HF, Lin T, Klein J, Wang SH, Qi JJ, Zhou Q, Sun JJ, Zhang ZH, Weng YQ, Huang SW (2014) QTL-seq identifies an early flowering QTL located near Flowering Locus T in cucumber. Theor Appl Genet 127:1491–1499
Maheswaran M, Huang N, Sreerangasamy S, McCouch S (2000) Mapping quantitative trait loci associated with days to flowering and photoperiod sensitivity in rice (Oryza sativa L.). Mol Breed 6:145–155
Miao H, Gu XF, Zhang SP, Zhang ZH, Huang SW, Wang Y, Fang ZY (2012) Mapping QTLs for seedling-associated traits in cucumber. Acta Hortic Sin 39:879–887
Michaels SD, Amasino RM (2001) Loss of FLOWERING LOUCS C activity eliminates the late-flowering phenotype of FRIGIDA and autonomous pathway mutations but not responsiveness to vernalization. Plant Cell 13:935–936
Moon J, Suh SS, Lee H, Choi KR, Hong CB, Peak NC, Kim SG, Lee I (2003) The SOC1 MADS-box gene integrates vernalization and gibberellins signals for flowering in Arabidopsis. Plant J 35:613–623
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acid Res 8:4321–4325
O’Rawe J, Jiang T, Sun GQ, Wu YY, Wang W, Hu JC, Bodily P, Tian LF, Hakonarson H, Johnson WE, Wei Z, Wang K, Lyon GJ (2013) Low concordance of multiple variant-calling pipelines: practical implications for exome and genome sequencing. Genome Med 5:28
Panaud O, Chen X, McCouch S (1996) Development of microsatellite markers and characterization of simple sequence length polymorphism (SSLP) in rice (Oryza sativa L.). Mol Gen Genet 252:597–607
Pnueli L, Gutfinger T, Hareven D, Ben-Naim O, Ron N, Adir N, Lifschitz E (2001) Tomato SP-interacting proteins define a conserved signaling system that regulates shoot architecture and flowering. Plant Cell 13:2687–2702
Putterill J, Laurie R, Macknight R (2004) It’s time to flower: the genetic control of flowering time. BioEssays 26:363–373
Qin ZR, Wu JJ, Geng SF, Feng N, Chen FJ, Kong XC, Song GY, Chen K, Li AL, Mao L, Wu L (2016) Regulation of FT splicing by an endogenous cue in temperate grasses. Nat Commun 8:14320
Sheldon CC, Rouse DT, Finnegan EJ, Peacock WJ, Dennis ES (2000) The molecular basis of vernalization: the central role of FLOWERING LOCUS C (FLC). Proc Natl Acad Sci USA 97:3753–3758
Shu J, Liu Y, Fang Z, Yang L, Zhang L, Zhuang M, Zhang Y, Li Z, Sun P (2014) Study on the floral characteristics and structure in two types of male sterile lines of broccoli (Brassica oleracea var. italica). J Plant Genet Resour 15:113–119
Shu J, Liu Y, Li Z, Zhang L, Fang Z, Yang L, Zhuang M, Zhang Y, Sun P (2015a) Effect of different pruning methods on flowering and fruiting characteristics between different types of male sterile lines in broccoli seed plants. Acta Hortic Sin 42:689–696
Shu J, Liu Y, Li Z, Zhang L, Fang Z, Yang L, Zhuang M, Zhang Y, Lv H (2015b) Organelle simple sequence repeat markers help to distinguish carpelloid stamen and normal cytoplasmic male sterile sources in broccoli. PLoS One 10:e0138750
Suarez-Lopez P, Wheatley K, Robson F, Onouchi H, Valverde F, Coupland G (2001) CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature 410:1116–1120
Takagi H, Abe A, Yoshida K, Kosugi S, Natsume S, Mitsuoka C, Uemura A, Utsushi H, Tamiru M, Takuno S (2013) QTL-seq: rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations. Plant J 74:174–183
Taoka K, Ohki I, Tsuji H, Furuita K, Hayashi K, Yanase T, Yamaguchi M, Nakashima C, Purwestri YA, Tamaki S, Ogaki Y, Shimada C, Nakagawa A, Kojima C, Shimamoto K (2011) 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen. Nature 476:332-U97
Taoka K, Ohki I, Tsuji H, Kojima C, Shimamoto K (2013) Structure and function of florigen and the receptor complex. Trends Plant Sci 18:287–294
Van Ooijen JW (2006) Joinmap 4, Software for the calculation of genetic linkage maps in experimental populations. Kyazma BV, Wagenigen
Van Ooijen JW, Boer MP, Jansen RC, Maliepaard C (2002) Map QTL4.0, software for the calculation of QTL positions on genetic maps. Plant Research International, Wagenigen
Walley PG, Carder J, Skipper E, Mathas E, Lynn J, Pink D, Buchanan-Wollaston V (2012) A new broccoli × broccoli immortal mapping population and framework genetic map: tools for breeders and complex trait analysis. Theor Appl Genet 124:467–484
Wang JW, Czech B, Weigel D (2009) miR156-regulated SPL transcription factors define an endogenous flowering pathway in Arabidopsis thaliana. Cell 138:738–749
Wigge PA, Kim MC, Jaeger K-E, Busch W, Schmid M, Lohmann JU, Weigel D (2005) Integration of spatial and temporal information during floral induction in Arabidopsis. Science 309:1056–1059
Xu XW, Zeng L, Li Y, Luo SB, Wang HM, Tian YH (2012) Major gene plus poly-gene inheritance analysis of the flowing time in pepper (Capsicum annuum. L). J Biomath 27:755–757
Zhang F, Chen FD, Fang WM, Chen SM, Liu PS, Yin DM (2011) Heterosis and mixed genetic analysis for florescence-related traits of chrysanthemum. J Nanjing Agric Univ 34:31–36
Zou G, Zhai G, Feng Q, Yan S, Wang A, Zhao Q, Shao J, Zhang Z, Zou J, Han B, Tao Y (2012) Identification of QTLs for eight agronomically important traits using an ultra-high-density map based on SNPs generated from high-throughput sequencing in sorghum under contrasting photoperiods. J Exp Bot 63:5451–5462
Acknowledgments
This work was supported by the National Natural Science Foundation of China (Grant No. 31372067), the China Agriculture Research System (Grant No. CARS-25-A), the Key Projects in the National Science and Technology Pillar Program of China (Grant No. 2013BAD01B04), the Key Laboratory of Biology and Genetic Improvement of Horticultural Crops, Ministry of Agriculture, P.R. China, and the Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences (Grant No. CAAS-ASTIP-IVFCAAS). We thank Tom Buckle, MSc, from Liwen Bianji, Edanz Group China (http://www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
The experiments in this study comply with the current laws of China.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shu, J., Liu, Y., Zhang, L. et al. QTL-seq for rapid identification of candidate genes for flowering time in broccoli × cabbage. Theor Appl Genet 131, 917–928 (2018). https://doi.org/10.1007/s00122-017-3047-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-017-3047-5