Abstract
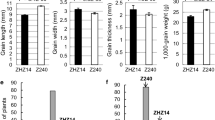
The GS3 locus located in the pericentromeric region of rice chromosome 3 has been frequently identified as a major QTL for both grain weight (a yield trait) and grain length (a quality trait) in the literature. Near isogenic lines of GS3 were developed by successive crossing and backcrossing Minghui 63 (large grain) with Chuan 7 (small grain), using Minghui 63 as the recurrent parent. Analysis of a random subpopulation of 201 individuals from the BC3F2 progeny confirmed that the GS3 locus explained 80–90% of the variation for grain weight and length in this population. In addition, this locus was resolved as a minor QTL for grain width and thickness. Using 1,384 individuals with recessive phenotype (large grain) from a total of 5,740 BC3F2 plants and 11 molecular markers based on sequence information, GS3 was mapped to a DNA fragment approximately 7.9 kb in length. A full-length cDNA corresponding to the target region was identified, which provided complete sequence information for the GS3 candidate. This gene consists of five exons and encodes 232 amino acids with a putative PEBP-like domain, a transmembrane region, a putative TNFR/NGFR family cysteine-rich domain and a VWFC module. Comparative sequencing analysis identified a nonsense mutation, shared among all the large-grain varieties tested in comparison with the small grain varieties, in the second exon of the putative GS3 gene. This mutation causes a 178-aa truncation in the C-terminus of the predicted protein, suggesting that GS3 may function as a negative regulator for grain size. Cloning of such a gene provided the opportunity for fully characterizing the regulatory mechanism and related processes during grain development.





Similar content being viewed by others
References
Abreu JG, Coffinier C, Larraín J, Oelgeschläger M, Robertis EMD (2002) Chordin-like CR domains and the regulation of evolutionarily conserved extracellular signaling systems. Gene (Amst) 287:39–47
Aluko G, Martinez C, Tohme J, Castano C, Bergman C, Oard HJ (2004) QTL mapping of grain quality traits from the interspecific cross Oryza sativa × O. glaberrima. Theor Appl Genet 109:630–639
Brondani C, Rangel PHN, Brondani RPV, Ferreira ME (2002) QTL mapping and introgression of yield-related traits from Oryza glumaepatula to cultivated rice (Oryza sativa) using microsatellite markers. Theor Appl Genet 104:1192–1203
Evans LT (1972) Storage capacity as a limitation on grain yield. In: Banos Los (ed) Rice breeding. International Rice Research Institute, Manila, pp 499–511
Fan CC, Yu XQ, Xing YZ, Xu CG, Luo LJ, Zhang Q (2005) The main effects, epistatic effects and environmental interactions of QTLs on cooking and eating quality of rice in a doubled haploid line population. Theor Appl Genet 110:1445–14452
Hua JP, Xing YZ, Xu CG, Sun XL, Yu SB, Zhang Q (2002) Genetic dissection of an elite rice hybrid revealed that heterozygotes are not always advantageous for performance. Genetics 162:1885–1895
Huang N, Parco A, Mew T, Magpantay G, McCouch S, Guiderdoni E, Xu JC, Subudhi P, Angeles ER, Khush GS (1997) RFLP mapping of isozymes, RAPD, and QTLs for grain shape, brown planthopper resistance in a doubled-haploid rice population. Mol Breed 3:105–113
Juliano BO, Villareal CP (1993) Grain quality evaluation of world rices. International Rice Research Institute, Manila
Kubo T, Takano-kai N, Yoshimura A (2001) RFLP mapping of genes for long kernel and awn on chromosome 3 in rice. Rice Genet Newsl 18:26–28
Li JX, Yu SB, Xu CG, Tan YF, Gao YJ, Li XH, Zhang Q (2000) Analyzing quantitative trait loci for yield using a vegetatively replicated F2 population from a cross between the parents of an elite rice hybrid. Theor Appl Genet 101:248–254
Li J, Thomson M, McCouch SR (2004a) Fine mapping of a grain-weight quantitative trait locus in the pericentromeric region of rice chromosome 3. Genetics 168:2187–2195
Li J, Xiao J, Grandillo S, Jiang L, Wan Y, Deng Q, Yuan L, McCouch SR (2004b) QTL detection for rice grain quality traits using an interspecific backcross population derived from cultivated Asian (O. sativa L.) and African (O. glaberrima S.) rice. Genome 47:697–704
Lincoln S, Daly M, Lander E (1992) Constructing genetics maps with MAPMAKER/EXP 3.0. Whitehead Institute Technical Report, Whitehead Institute, Cambridge
Liu JP, Eck JV, Cong B, Tanksley SD (2002) A new class of regulatory genes underlying the cause of pear-shaped tomato fruit. Proc Natl Acad Sci USA 99:13302–13306
McCouch SR, Teytelman L, Xu Y, Lobos KB, Clare K, Walton M, Fu B, Maghirang R, Li Z, Xing Y, Zhang Q, Kono I, Yano M, Fjellstrom R, DeClerck G, Schneider D, Cartinhour S, Ware D, Stein L (2002) Development and mapping of 2,240 new SSR markers for rice (Oryza sativa L.). DNA Res 9:199–207
O’Leary JM, Hamilton JM, Deane CM, Valeyev NV, Sandell LJ, Downing AK (2004) Solution structure and dynamics of a prototypical Chordin-like cysteine-rich repeat (von Willebrand factor type C module) from collagen IIA. J Biol Chem 279:53857–53866
Redoña ED, Mackill (1998) Quantitative trait locus analysis for rice panicle and grain characteristics. Theor Appl Genet 96:957–963
Tan YF, Xing YZ, Li JX, Yu SB, Xu CG, Zhang Q (2000) Genetic bases of appearance quality of rice grains in Shanyou 63, an elite rice hybrid. Theor Appl Genet 101:823–829
Temnykh S, Park WD, Ayres N, Cartihour S, Hauck N, Lipovich L, Cho YG, Ishii T, McCouch SR (2000) Mapping and genome organization of microsatellite sequences in rice (Oryza sativa L.). Theor Appl Genet 100:697–712
Temnykh S, Declerck G, Luashova A, Lipovich L, Cartinhour S, McCouch S (2001) Computational and experimental analysis of microsatellites in rice (Oryza sativa L.): frequency, length variation, transposon associations, and genetic marker potential. Genome Res 11:1441–1452
Thomson M, Tai T, McClung A, Xai XH, Hinga M, Lobos K, Xu Y, Martinez P, McCouch S (2003) Mapping quantitative trait loci for yield, yield components and morphological traits in an advanced backcross population between Oryza rufipogon and the Oryza sativa cultivar Jefferson. Theor Appl Genet 107:479–493
Unnevehr LJ, Duff B, Juliano BO (1992) Consumer demand for rice grain quality. International Rice Research Institute, Manila, and International Development Research Center, Ottawa
Xiao JH, Li JM, Grandillo S, Ahn SN, Yuan LP, Tanksley SD, McCouch SR (1998) Identification of trait-improving quantitative trait loci alleles from a wild rice relative, Oryza rufipogon. Genetics 150:899–909
Xing YZ, Tan YF, Xu CG, Hua JP, Sun XL (2001) Mapping quantitative trait loci for grain appearance traits of rice using a recombinant inbred line population. Acta Bot Sin 43:721–726
Xing YZ, Tan YF, Hua JP, Sun XL, Xu CG, Zhang Q (2002) Characterization of the main effects, epistatic effects and their environmental interactions of QTLs on the genetic basis of yield traits in rice. Theor Appl Genet 105:248–257
Xu JY, Xue QZ, Luo LJ, Li ZK (2002) Genetic dissection of grain weight and its related traits in rice (Oryza sativa L.). Chin J Rice Sci 16:6–10 (in Chinese with an English abstract)
Yu SB, Li JX, Xu CG, Tan YF, Gao YJ, Li XH, Zhang Q, Saghai Maroof MA (1997) Importance of epistasis as the genetic basis of heterosis in an elite rice hybrid. Proc Natl Acad Sci USA 94:9226–9231
Acknowledgements
This work was supported in part by grants from the National Program on the Development of Basic Research, the National Special Key Project of Functional Genomics and Biochips, and the National Natural Science Foundation of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Y. Xue
Rights and permissions
About this article
Cite this article
Fan, C., Xing, Y., Mao, H. et al. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor Appl Genet 112, 1164–1171 (2006). https://doi.org/10.1007/s00122-006-0218-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-006-0218-1