Abstract
The chukar (Alectoris chukar, Galliformes) is a species hunted throughout its native range from the East Mediterranean to Manchuria and in the USA, which hosts the world’s largest introduced population. This study aims to investigate the genetic structure of Mediterranean chukar populations to aid management decisions. We genotyped 143 specimens at two regions of the mitochondrial DNA (mtDNA: cytochrome b, control region) and eight loci of the microsatellite DNA. Samples were collected in northern (Limnos, Lesvos, Chios) and southern (Crete) Aegean islands (Greece) and Cyprus. We also carried out mtDNA-based comparison with chukars (n = 124) from Asia (16 countries) and the USA (five states). We propose six management units for Mediterranean populations. Given their genetic integrity, Limnos and Cyprus, which host different subspecies, proved to be of primary conservation interest. We found exotic A. chukar mtDNA lineages in Lesvos, Chios and Crete and produced definitive genetic evidence for the Asian origin of the US chukars.



Similar content being viewed by others
References
Allendorf FW, Luikart G (2007) Conservation and the genetics of populations. Blackwell, Malden
Ballard JWO, Whitlock MC (2004) The incomplete natural history of mitochondria. Mol Ecol 13:729–744
Barbanera F, Negro JJ, Di Giuseppe G, Bertoncini F, Cappelli F, Dini F (2005) Analysis of the genetic structure of red-legged partridge (Alectoris rufa, Galliformes) populations by means of mitochondrial DNA and RAPD markers: a study from central Italy. Biol Conserv 122:275–287
Barbanera F, Guerrini M, Hadjigerou P, Panayides P, Sokos C, Wilkinson P, Khan AA, Khan BY, Cappelli F, Dini F (2007) Genetic insight into Mediterranean chukar (Alectoris chukar, Galliformes) populations inferred from mitochondrial DNA and RAPD markers. Genetica 131:287–298
Barbanera F, Guerrini M, Khan AA, Panayides P, Hadjigerou P, Sokos C, Gombobaatar S, Samadi S, Khan BY, Tofanelli S, Paoli G, Dini F (2009a) Human-mediated introgression of exotic chukar (Alectoris chukar, Galliformes) genes from East Asia into native Mediterranean partridges. Biol Invasions 11:333–348
Barbanera F, Zuffi MAL, Guerrini M, Gentilli A, Tofanelli S, Fasola M, Dini F (2009b) Molecular phylogeography of the asp viper Vipera aspis (Linnaeus, 1758) in Italy: evidence for introgressive hybridization and mitochondrial DNA capture. Mol Phylogenet Evol 53:103–114
Barilani M, Sfougaris A, Giannakopoulos A, Mucci A, Tabarroni C, Randi E (2007) Detecting introgressive hybridisation in rock partridge populations (Alectoris graeca) in Greece through Bayesian admixture analyses of multilocus genotypes. Conserv Genet 8:343–354
Belkhir K, Borsa P, Chikhi L, Raufaste N, Bonhomme F (2001) GENETIX 4.02, logiciel sous Windows™ pour la génétique des populations. Laboratoire Génome, Populations, Interactions, CNRS UMR 5000, Université de Montpellier II, Montpellier, France. Available via DIALOG. http://www.univ-montp2.fr/~genetix/genetix.htm
BirdLife International (2004) Birds in Europe: population estimates trends and conservation status. BirdLife conservation series, vol 12. Wageningen, BirdLife International
Byers SM, Burger GV (1979) Evaluation of three partridge species for put and take hunting. Wildl Soc Bull 7:17–20
Christensen GC (1970) The chukar partridge: its introduction life history and management. Biology bulletin, vol 4. Nevada Department of Fish and Game, Reno
Cottam C, Arnold LN, Saylor LW (1940) The chukar and Hungarian partridge in America. US Department Interior, Bio Survey, Wildlife Leaflets, BS-159, USA
Crandall KA, Bininda-Edmonds ORP, Mace GM (2000) Considering evolutionary processes in conservation biology. Trends Ecol Evol 15:290–295
Degner JF, Stout IJ, Roth JD, Parkinson CL (2007) Population genetics and conservation of the threatened southeastern beach mouse (Peromyscus polionotus niveiventris): subspecies and evolutionary units. Conserv Genet 8:1441–1452
Dragoev P (1974) On the population of the rock partridge (Alectoris graeca Meisner) in Bulgaria and methods of census. Acta Ornithologica 30:251–255
Evanno G, Reganut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Laval G, Schneider S (2005) ARLEQUIN ver 3.0: an integrated software package for population genetics data analysis. EBO 1:47–50
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fraser DJ, Bernatchez L (2001) Adaptive evolutionary conservation: towards a unified concept for defining conservation units. Mol Ecol 10:2741–2752
Godinho R, Crespo EG, Ferrand N (2008) The limits of mtDNA phylogeography: complex patterns of population history in a highly structured Iberian lizard are only revealed by the use of nuclear markers. Mol Ecol 17:4670–4683
Gonzalez EG, Castilla AM, Zardoya R (2005) Novel polymorphic microsatellites for the red-legged partridge (Alectoris rufa) and cross-species amplification in Alectoris graeca. Mol Ecol Notes 5:449–451
Goudet J (2001) FSTAT. Available via DIALOG. http://www2.unil.ch/popgen/softwares/ fstat.htm
Guerrini M, Panayides P, Hadjigerou P, Taglioli L, Dini F, Barbanera F (2007) Lack of genetic structure of Cypriot Alectoris chukar populations (Aves, Galliformes) as inferred from mtDNA sequencing data. Anim Biodivers Conserv 30:105–114
Hadjisterkotis E (1999) The survival of captive bred Alectoris chukar cypriotes released for restocking in Cyprus. Z Jagdwiss 45:238–249
Hastings A (1993) Complex interactions between dispersal and dynamics: lessons from couplet logistic equations. Ecology 74:1362–1372
Hochberg Y (1988) A sharper Bonferroni procedure for multiple tests of significance. Biometrika 75:800–802
Kizirian D, Donnelly MA (2004) The criterion of reciprocal monophyly and classification of nested diversity at the species level. Mol Phylogenet Evol 32:1072–1076
Liu N-F, L-Yi W, Huang Z-H, Hou P (2006) Introgressive hybridization between Alectoris magna and A. chukar in the Liu-pan Mountain Region. Acta Zool Sin 52:153–159
Lynch M, Crease J (1990) The analysis of population survey data on DNA sequence variation. Mol Biol Evol 7:377–394
Madge S, McGowan P (2002) Pheasants partridges and grouse. A and C Black, London
Masseti M (1997) Representations of birds in Minoan art. Int J Osteoarchaeol 7:354–363
Mills LS (2006) Conservation of wildlife populations demography genetics and management. Blackwell, Malden
Moritz C (1994) Defining evolutionarily significant units for conservation. Trends Ecol Evol 9:373–375
Palsbøll P, Bérubé M, Allendorf F (2007) Identification of management units using population genetic data. Trends Ecol Evol 22:11–16
Panayides P (2005) Six aspects of land use and development activity that result in adverse effects to Cypriot wildlife resources. In: Hadjisterkotis E (ed) Proceedings of the XXVth International Congress of the International Union of Game Biologists-IUGB and the IXth International Symposium Perdix 2. Government Printing Office, Nicosia, pp 182–196
Papaevangelou E, Thomaides C, Handrinos G, Haralambides A (2001) Status of partridges (Alectoris and Perdix) species in Greece. Game Wildl Sci 18:253–260
Posada D, Buckley TR (2004) Model selection and model averaging in phylogenetics: advantages of the AIC and Bayesian approaches over likelihood ratio tests. Systematic Biol 53:793–808
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818
Potts D (1988) The impact of releasing hybrid partridges on wild red-legged populations. Game Conservancy Rev 1988:81–85
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Pritchard J, Wen X, Falush D (2007) Documentation for structure software: version 2.2. Available via DIALOG. http://pritch.bsd.uchicago.edu/software/structure22/readme.pdf
Randi E (2008) Detecting hybridization between wild species and their domesticated relatives. Mol Ecol 17:285–293
Raymond M, Rousset F (1995) GENEPOP (version 3.1) is an update version of GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rozas J, Sanchez-Del Barrio JC, Messeguer X, Rozas R (2003) DNASP DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Ryder O (1986) Species conservation and systematics: the dilemma of subspecies. Trends Ecol Evol 1:9–10
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Slatkin M (1985) Rare alleles as indicators of gene flow. Evolution 39:53–65
Sokos CK, Birtsas PK, Tsachalidis EP (2008) The aims of galliforms release and choice of techniques. Wildlife Biol 14:414–422
Strimmer K, von Haeseler A (1996) Quartet puzzling: a quartet maximum likelihood method for reconstructing tree topologies. Mol Biol Evol 13:964–969
Swofford DL (2002) PAUP*: phylogenetic analysis using parsimony. Version 4.0b10. Sinauer, Sunderland
Swofford DL, Olsen GJ, Waddel PJ, Hillis DM (1996) Phylogenetic inference. In: Hillis DH, Moritz C, Bable BK (eds) Molecular systematics, 2nd edn. Sinauer, Sunderland, pp 407–514
Tejedor MT, Monteagudo LV, Hadjisterkotis E, Arruga MV (2005) Genetic variability and population structure in Cypriot chukar partridges (Alectoris chukar cypriotes) as determined by microsatellite analysis. Eur J Wildlife Res 51:232–236
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting positions-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
True GH (1937) The chukar partridge of Asia. Calif Fish Game 23:229–231
Vaha J-P, Primmer CR (2006) Efficiency of model-based Bayesian methods for detecting hybrid individuals under different hybridization scenarios and with different numbers of loci. Mol Ecol 15:63–72
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Waples RS (1991) Pacific Salmon Oncorhynchus spp. and the definition of “species” under the Endangered Species Act. Mar Fish Rev 53:11–22
Whitlock M, McCauley D (1999) Indirect measures of gene flow and migration: \( {{\rm F}_{ST}} \ne 1/\left( {4{{\rm N}_m} + 1} \right) \). Heredity 82:117–125
Wright S (1965) The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution 19:395–420
Acknowledgements
Chukar samples were provided by: P. Birtsas (Hunting Federation of Macedonia-Thrace), G. Arnellos and A. Sakoulis (Hunting Federation of Crete), P. Lemanis (Hunting Federation of Archipelago) and C. Barboutis (University of Crete) from Greece; L. Gilbertson (Nevada Department of Wildlife, Reno, Nevada) and R. Kaholoaa (Resources Management Division, Haleakala National Park, Maui, Hawaii) from USA; Natural History Museum of Crete (Heraklion, Greece, NHMC samples: from 80.4.59.11 to 80.4.59.23); University of Washington Burke Museum (Seattle, Washington, UWBM samples: 46402, 46516, 57853, 57857, 57859, 66692). We deeply thank G. Paoli and L. Taglioli (Department of Biology, University of Pisa) for their valuable support in the statistical analyses as well as Alan Crabtree (Cyprus) and Peter Wilkinson (UK) for their helpful linguistic revision of the paper and for their constructive comments. The Cypriot Game Fund Service, Ministry of the Interior, Nicosia (Cyprus) granted this research.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
S1
Chukar sampling localities are indicated for each island (a–e). a Limnos (477 km2): (1) Agios Dimitrios, (2) Papias Ormos, (3) Gomati, (4) Paradisi. b Lesvos (1,630 km2): (1) Mesotopus, (2) Lapsarna, (3) Tsichlioda. c Chios (840 km2): (1) Trypes, (2) Amades, (3) Kardamyla, (4) Agia Markella. d Cyprus (9,250 km2): (1) Paphos forest, (2) Karpasia, (3) Larnaka coastal area, (4) Stavrouvoni farm; dotted line marks out the border between the government-controlled area and the Turkish-occupied territory. e Crete (8,400 km2): (1) Lefka mountains, (2) Rethymno farm, (3) Psiloreitis, (4) Heraklion, (5) Dikti, (6) Ierapetra, (7) Sitia (DOC 4392 kb)
S2
Chukar sample size (n = 267). Geographic reference is reported together with population type, number of samples, type of tissue, number of haplotypes (Cyt-b + CR sequences) and literature record. *: with samples of Natural History Museum of Crete (acronym NHMC: from 80.4.59.11 to 80.4.59.23); **: with samples of University of Washington Burke Museum (acronym UWBM: 46402, 46516, 57853, 57857, 57859, 66692). One wild A. graeca specimen (Southern Apennines, Italy; type of tissue: liver; haplotype: H113) was used as out-group. Abbreviations: Prov., Province; NP, National Park (DOC 106 kb)
S3
The STR primers. Legend: F, forward; R, reverse; Ta (°C), first/second annealing temperature in touchdown (TD) PCR (DOC 33 kb)
S4
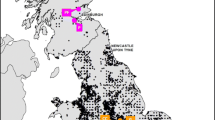
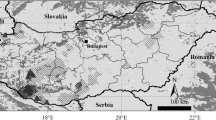
The frequency distribution of A. chukar specimens belonging to the A (white) or B (black) mtDNA clades of Fig. 1 is reported for all populations (S2). a Europe and Asia. For both Crete and Cyprus, the percentage was calculated by pooling together all wild and captive specimens. Triangles mark out the main mountain chains in Asia. b USA: the study areas are indicated (five states) (DOC 592 kb)
S5
The principal components analysis performed using the pairwise ϕST distances calculated for the ingroup of the mtDNA reconstructions. Specimens are grouped according to each population (S2) (DOC 164 kb)
S6
The outcome (t value, degrees of freedom or df, P value) of the Student t test for all pairs of populations (Pop. 1 vs. Pop. 2) as obtained for nucleotide diversity (π), haplotype diversity (h), mean number of pairwise differences (k) and index of Nei (I N). No significant differences were found after application of the Bonferroni correction (α = 0.05: for π, h and k, \( \alpha \prime = \alpha /28 = 0.001786 \); for I N, \( \alpha \prime = \alpha /36 = 0.001389 \)). Abbreviations: Cyprus-Paphos, Paphos forest; Cyprus-Larnaka, Larnaka coastal area; Cyprus-Karpasia, Karpasia; Cyprus-Farm, Stavrouvoni farm; Crete-Farm, Rethymno farm. −: Crete-Farm mtDNA data not computed, see “Material and methods” (DOC 84 kb)
S7
STR variability for each population: n, sample size; nA, number of alleles per locus; na, number of unique alleles; Ar, allelic richness; I n, Nei’s index with standard deviation (SD); H O, observed heterozygosity; H E, expected heterozygosity; P, probability value for the HWE test; χ 2 test with relative degrees of freedom (df; Fisher global test, all loci). Mono., monomorphic locus; *: significant departure from HWE after application of the Bonferroni correction (α = 0.05, \( \alpha \prime = \alpha /8 = 0.006 \)). Abbreviations: Cyprus-Paphos, Paphos forest; Cyprus-Larnaka, Larnaka coastal area; Cyprus-Karpasia, Karpasia; Cyprus-Farm, Stavrouvoni farm; Crete-Farm, Rethymno farm (DOC 139 kb)
S8
Fisher global test for departure from linkage disequilibrium (LE) with application of the Bonferroni correction (α = 0.05, \( \alpha \prime = \alpha /28 = 0.0018 \)). Significant departure was found only in the Lesvos population (see data in bold); –: comparison not possible as one locus was monomorphic (DOC 57 kb)
Rights and permissions
About this article
Cite this article
Barbanera, F., Marchi, C., Guerrini, M. et al. Genetic structure of Mediterranean chukar (Alectoris chukar, Galliformes) populations: conservation and management implications. Naturwissenschaften 96, 1203–1212 (2009). https://doi.org/10.1007/s00114-009-0586-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00114-009-0586-x