Abstract
The deubiquitinating enzyme CYLD negatively regulates NF-κB signaling by removing activating ubiquitin chains from several members of the NF-κB pathway. Thereby, CYLD is critical for the maintenance and differentiation of various immune cells. Despite the importance of the NF-κB pathway in microglia regulation, the role of CYLD in microglia has not been investigated so far. In this study, we investigated whether CYLD in microglia can protect against neuroinflammation using a newly generated conditional mouse strain (Rosa26-Cyld-tdTomato) that allows cell type-specific CYLD overexpression. Here, we show that overexpression of CYLD in microglia did not alter microglia numbers or microglia morphology in different brain regions. Additionally, CYLD overexpression did not modify the microglial response to LPS-induced neuroinflammation or the disease severity in experimental autoimmune encephalomyelitis (EAE). Finally, also immune cell infiltration into the CNS during EAE and under steady state conditions remained unaffected by microglial CYLD overexpression. Our findings suggest that CYLD overexpression does not alter microglial function, and thus does not represent a viable therapeutic strategy in neuroinflammatory conditions. This study highlights the complexity of ubiquitin-mediated signaling in neuroinflammation and the need for cell-type-specific investigations. The Rosa26-Cyld-tdTomato mouse model offers a valuable tool for studying CYLD’s role across various tissues and cell types.
Key messages
-
Novel mouse strain for cell type-specific overexpression of the deubiquitinating enzyme CYLD.
-
CYLD overexpression in microglia did not alter microglia numbers or morphology in the steady state.
-
CYLD overexpression in microglia did not protect mice from LPS-induced neuroinflammation or EAE.
-
CYLD overexpression in microglia did not influence their gene expression during neuroinflammation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The deubiquitinating enzyme CYLD was first identified as a tumor suppressor gene mutated in familial cylindromatosis, a genetic condition predisposing individuals to develop multiple skin tumors [1, 2]. Located in the cytoplasm, CYLD removes Lysine 63 (K63)- or Met1-linked polyubiquitin chains from several substrates [3]. Among these substrates are molecules of the proinflammatory NF-κB pathway such as TRAF2, TRAF6, and NEMO, and CYLD acts as a negative regulator of NF-κB signaling by removing activating K63-linked ubiquitin chains from these proteins [4,5,6]. By controlling NF-κB pathway activity, CYLD regulates the maintenance and differentiation of various immune cell types. Studies with CYLD-deficient mice showed that CYLD is required for the proper development and activation of T and B cells [7,8,9]. Previously, we have generated mice deficient in full-length CYLD but overexpressing a naturally occurring splice variant of CYLD that lacks the binding sites for TRAF2 and NEMO [10]. These CYLDex7/8 mice show defects in B cell homeostasis [10] and regulatory T cell function [11], and exhibit a hyperactive phenotype of dendritic cells and T cells [11,12,13].
CYLD is expressed in the brain [2] and is required for maintaining neuronal activity and synaptic transmission [14, 15]. Recent studies showed that CYLD deficiency leads to an autistic phenotype characterized by impaired sociability and other behavioral abnormalities, likely due to the loss of CYLD’s regulation of mTOR signaling and autophagy [16, 17].
Microglia are innate immune cells residing in the central nervous system (CNS) parenchyma. In the steady state, microglia have a ramified morphology with finely branched processes that scan the CNS environment for potential threats [18]. Upon infection or disease, microglia adopt an amoeboid shape, become reactive and phagocytose cellular debris, dead cells and foreign material. In studies with Cyld−/− mice, microglia seem to exhibit a more activated phenotype in certain brain regions [14, 19]. A similar phenotype was observed in mice lacking another deubiquitinating enzyme that negatively regulates the NF-κB pathway, namely A20, specifically in microglia [20, 21]. Mice with A20 deletion in microglia exhibited increased microglia numbers and altered microglia morphology, suffered from spontaneous neuroinflammation and showed exacerbated responses to different mouse models of neuroinflammation [20, 21]. Interestingly, while the protein substrates of A20 and CYLD partially overlap, mice deficient in one of the enzymes nevertheless display different phenotypes [22].
We hypothesized that mice with CYLD-deficient microglia could also be more prone to develop neuroinflammation, while constitutive overexpression of CYLD in microglia could protect against CNS inflammation. To study the cell-type specific role of CYLD in microglia, we generated a novel mouse strain allowing the conditional overexpression of CYLD in a cell type of interest (Rosa26-Cyld-tdTomato mice) and used these mice to study the effects of constitutive CYLD expression in microglia. Surprisingly, microglia numbers and microglia morphology were not altered in several brain regions of mice overexpressing CYLD in microglia. Additionally, constitutive CYLD expression in microglia did not affect CNS immune cell infiltration either in the steady state or in experimental autoimmune encephalomyelitis (EAE), a mouse model for Multiple Sclerosis (MS). Even when directly stimulating the NF-κB pathway in microglia in a model of LPS-induced neuroinflammation, CYLD overexpression had no effect on microglia gene expression and function. Taken together, our data strongly suggest that constitutive expression of CYLD in microglia does not impact microglia function and is not protective in the context of neuroinflammation.
Results
Overexpression of Cyld in microglia does not influence microglia numbers and microglia morphology in the steady state
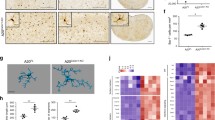
To assess the role and function of CYLD in microglia, a new conditional mouse strain was generated in which CYLD can be overexpressed in specific cell types in a Cre-dependent manner. These Rosa26-Cyld-tdTomato mice were generated by insertion of the Cyld and tdTomato genes into the Rosa26 locus behind a loxP-flanked STOP cassette (Fig. 1A). After Cre-mediated removal of the STOP cassette, a chimeric transcript of Cyld and tdTomato fused by a T2A site is expressed. The T2A sequence leads to co-translational cleavage between the CYLD and tdTomato proteins allowing the co-expression of CYLD and tdTomato in the cell type of interest. We crossed these Rosa26-Cyld-tdTomato mice to the tamoxifen-inducible Cx3cr1CreER strain [23] to obtain R26-Cyld-OECx3 mice. In the CNS of R26-Cyld-OECx3 mice in the steady state, the expression of CYLD and tdTomato is limited to microglia and other CNS-resident macrophage populations (for simplicity, we will only use the term "microglia" in the following sections). We used littermate control animals for all experiments, which harbored an allele of the tdRFP reporter gene in the Rosa26 locus [24] but no Cyld (R26RFP/WT), and also expressed the tamoxifen-inducible Cre recombinase under control of the Cx3Cr1 promotor. We verified the expression of Cyld in isolated primary microglia of R26-Cyld-OECx3 mice where we observed on average 5.95-fold higher expression levels of Cyld than in R26RFP/WT controls (Fig. 1B). Next, we quantified microglia numbers in the motor cortex, hippocampus, striatum and cerebellum of R26-Cyld-OECx3 mice and controls two weeks (Fig. 1C, D) and twelve weeks (Fig. 1E, F) after tamoxifen administration. CYLD overexpression in R26-Cyld-OECx3 mice had no effect on microglia numbers in the brain regions assessed. We also performed a Sholl analysis in the motor cortex and hippocampus to assess the morphology of the microglia. Microglia branching complexity was not altered in R26-Cyld-OECx3 mice in the motor cortex and hippocampus two weeks after tamoxifen administration (Fig. 1G), as well as in the hippocampus twelve weeks after tamoxifen administration (Fig. 1H). Surprisingly, microglia of R26-Cyld-OECx3 mice showed a slightly decreased branching complexity with less intersections in the motor cortex twelve weeks after tamoxifen administration (Fig. 1H, J). To further investigate this phenotype, we also performed a Sholl analysis in the motor cortex six months after tamoxifen administration. At this timepoint, microglia branching complexity was not altered in R26-Cyld-OECx3 mice compared to R26RFP/WT littermate controls (Fig. 1I).
Overexpression of Cyld in microglia does not influence microglia numbers and microglia morphology in the steady state. A Rosa26 Cyld conditional knock-in mice were generated in which the Cyld gene and the tdTomato gene were inserted into the Rosa26 locus behind a STOP cassette flanked by loxP sites (top). After Cre-mediated removal of the STOP cassette, the constitutive knock-in allele is obtained (bottom). B Expression of Cyld in primary murine microglia isolated by magnetic-activated cell sorting (MACS) for CD11b was validated by RT-PCR. Cyld mRNA levels were normalized to Hprt and plotted relative to control. C Representative images of tdTomato/tdRFP-labelled Iba-1+ (green) microglia in different brain regions of R26-Cyld-OECx3 mice and littermate controls 2 weeks after tamoxifen administration. Scale bar 100 μm. D Quantification of Iba-1+ cells per mm2 2 weeks after tamoxifen administration. E Representative images of tdTomato/tdRFP-labelled Iba-1+ (green) microglia in different brain regions of R26-Cyld-OECx3 mice and littermate controls 12 weeks after tamoxifen administration. Scale bar 100 μm. F Quantification of Iba-1+ cells per mm2 12 weeks after tamoxifen administration. G Sholl analysis of microglia in the cortex and hippocampus of R26-Cyld-OECx3 mice and littermate controls 2 weeks after tamoxifen administration. N = 6 mice for R26RFP/WT controls; N = 4 mice for R26-Cyld-OECx3; n = 9–17 cells. H Sholl analysis of microglia in the cortex and hippocampus of R26-Cyld-OECx3 mice and littermate controls 12 weeks after tamoxifen administration. N = 3 mice for R26RFP/WT controls; N = 3 mice for R26-Cyld-OECx3; n = 4–9 cells. I Sholl analysis of microglia in the cortex of R26-Cyld-OECx3 mice and littermate controls 6 months after tamoxifen administration. N = 4 mice for R26RFP/WT controls; N = 4 mice for R26-Cyld-OECx3; n = 36 cells. J Area under curve of the Sholl analysis of the cortex 12 weeks after tamoxifen administration shown in (H). In B, D, F, and J, every circle represents a single mouse. Data is represented as mean ± SEM. Statistical significance was determined by two-tailed unpaired Student’s t-test (B, D, F, J) or two-way ANOVA with Bonferroni Multiple comparisons correction (G, H, I). * p < 0.05, ** p < 0.01, **** p < 0.0001, ns = non-significant
Overexpression of Cyld in microglia does not influence immune cell infiltration and microglia phenotype in the steady state
Since mice lacking A20 in microglia developed spontaneous neuroinflammation with an increased infiltration of CD8+ T cells into the CNS of these mice [21], we hypothesized that mice overexpressing CYLD in microglia might show an opposite phenotype. Therefore, we analyzed immune cell infiltrates in the brain and spinal cord at steady state by flow cytometry (Fig. 2A). No alterations in the number of CD45hi immune cells in the brains of R26-Cyld-OECx3 mice compared to controls were detected two weeks after tamoxifen administration. Similarly, also no changes in the percentages of CD45hi CD11b− infiltrates, CD45hi CD11b+ infiltrates, and TCR-β+ T cells among all brain cells could be observed (Fig. 2B). Of note, at this early time point after tamoxifen administration, also short-lived Cx3Cr1+ cells such as peripheral monocytes could harbor the Rosa26-Cyld-tdTomato allele and therefore might overexpress CYLD. Next, we analyzed the brain and spinal cord twelve weeks after tamoxifen administration. Also here, no differences in the numbers and percentages of immune cells were observed in R26-Cyld-OECx3 mice compared to littermate controls (Fig. 2D, F). Finally, we also examined expression levels of CD68, a lysosomal marker upregulated in phagocytically active microglia. No alterations in the mean fluorescence intensity (MFI) of CD68 in microglia could be observed in R26-Cyld-OECx3 mice compared to controls in the brain and spinal cord at the respective time points (Fig. 2 C, E, G). Therefore, we concluded that CYLD overexpression in microglia does not affect immune cell infiltration in the steady state.
Overexpression of Cyld in microglia does not influence immune cell infiltration and microglia phenotype in the steady state. A Representative gating strategy for the flow cytometry analysis of immune cells and microglia isolated from the CNS. B Quantification of the absolute cell number of CD45hi immune cells in the brain (left) and percentages of CD45hi CD11b−, CD45hi CD11b+ and TCR-β+ cells among all brain cells (right) 2 weeks after tamoxifen administration. C Mean fluorescence intensity (MFI) as geometric mean of CD68 on brain microglia 2 weeks after tamoxifen administration. D Quantification of the absolute cell number of CD45hi immune cells in the brain (left) and percentages of CD45hi CD11b−, CD45hi CD11b+ and TCR-β+ cells among all brain cells (right) 12 weeks after tamoxifen administration. E Mean fluorescence intensity (MFI) as geometric mean of CD68 on brain microglia 12 weeks after tamoxifen administration. F Quantification of the absolute cell number of CD45hi immune cells in the spinal cord (left) and percentages of CD45hi CD11b−, CD45hi CD11b+ and TCR-β+ cells among all spinal cord cells (right) 12 weeks after tamoxifen administration. G Mean fluorescence intensity (MFI) as geometric mean of CD68 on spinal cord microglia 12 weeks after tamoxifen administration. Every circle represents a single mouse. Data is represented as mean ± SEM. Statistical significance was determined by two-tailed unpaired Student’s t-test. ns = non-significant
Conditional overexpression of Cyld in microglia and Cyld deletion do not affect EAE pathology
Since immune cell infiltration into the CNS in the steady state is very limited, the potential role of CYLD in microglia as a negative regulator of neuroinflammation may only be required during neuroinflammatory challenges. Therefore, we immunized R26-Cyld-OECx3 mice and controls with the MOG35-55 peptide to induce experimental autoimmune encephalomyelitis (EAE) and monitored clinical signs of disease. We observed no effect of CYLD overexpression in microglia on EAE disease progression (Fig. 3A). We also isolated spinal cord cells of the EAE mice at day 23 after induction of EAE for flow cytometry analysis (Fig. 3B). As was already expected with regard to the similar EAE scores, no differences in immune cell infiltrates could be observed between R26-Cyld-OECx3 mice and littermate controls (Fig. 3C). Also the expression levels of CD68 on microglia did not differ between R26-Cyld-OECx3 mice and controls, indicating no influence of CYLD overexpression on microglia activation during EAE (Fig. 3D). We also wanted to analyze the effect of Cyld deletion on the development of EAE and immunized Cyld−/− mice [25] and their heterozygous (Cyld+/-) and homozygous (Cyld+/+) littermate controls with the MOG35-55 peptide to induce EAE. Also the full knock-out of Cyld had no effect on EAE disease progression (Fig. 3E) questioning the importance of CYLD in microglia as a regulator of autoimmune neuroinflammation.
Conditional overexpression of Cyld in microglia and Cyld deletion do not affect EAE pathology. A EAE was induced in R26-Cyld-OECx3 mice and littermate controls and clinical signs of EAE were monitored daily. Data is pooled from two independent experiments with a total of n = 8 mice per group. B Representative gating strategy for the flow cytometry analysis of immune cells and microglia isolated from the spinal cord of EAE mice at day 23 after EAE induction. C Quantification of the absolute cell number of CD45hi immune cells in the spinal cord (left) and percentages of CD45hi CD11b−, CD45hi CD11b+ and TCR-β+ cells among all spinal cord cells (right). D Mean fluorescence intensity (MFI) as geometric mean of CD68 on spinal cord microglia. E EAE was induced in Cyld−/− mice (n = 7) and Cyld+/- (n = 6) and Cyld+/+ (n = 6) littermate controls. Clinical signs of EAE were monitored daily. In C and D, every circle represents a single mouse. Data is represented as mean ± SEM. Statistical significance was determined by two-tailed unpaired Student’s t-test (C, D). ns = non-significant
Overexpression of Cyld in microglia does not alter the microglial response to lipopolysaccharide (LPS)
In addition to the EAE model in which CYLD overexpression in microglia did not confer protection against autoimmune neuroinflammation in R26-Cyld-OECx3 mice, we used a model of LPS-induced neuroinflammation to directly stimulate microglial inflammatory responses. LPS activates microglia and induces the production of proinflammatory cytokines in the brain [26] by binding to toll-like receptor 4 (TLR4) on microglia [27], which activates the NF-κB pathway. We injected R26-Cyld-OECx3 mice and controls intraperitoneally with 1 mg/kg LPS and isolated their microglia for flow cytometry and RNA sequencing 24 h later. All mice showed a significant decrease in body weight 24 h after LPS injection confirming that an inflammatory response was triggered (Fig. 4A). Microglia were isolated by MACS for CD11b+ cells and purity was confirmed by flow cytometry (Fig. 4B), with microglia accounting for 87–91% of all live cells in the samples isolated for RNA sequencing (Fig. 4C). No differences in microglia activation could be observed in flow cytometry with similar expression levels of CD68 and MHC-II on brain microglia (Fig. 4D) and spinal cord microglia (Fig. 4E) of R26-Cyld-OECx3 mice and littermate controls. The RNA sequencing data were analyzed for signature genes of different CNS and immune cell types to further confirm the purity of the samples. Microglia signature genes Tmem119, Hexb and C1qa were highly expressed, while marker genes of neurons, astrocytes, endothelial cells, pericytes and oligodendrocytes were barely detectable (Fig. 4F). Also marker genes of other immune cell types such as monocytes, dendritic cells and neutrophils, as well as marker genes of CNS-associated macrophages such as Mrc1 and Lyve1 were absent. Interestingly, genes known to be expressed by “disease-associated microglia” (DAMs) [28], namely Apoe, Lgals3 and Trem2, were found to be expressed by the microglia. We also confirmed that LPS treatment efficiently activated the microglia by comparing the gene expression profile of naïve versus LPS-treated microglia, where we observed 3001 differentially expressed genes (1471 upregulated; 1530 downregulated; threshold: adjusted p-value < 0.05, log2 fold change > 1; data not shown). Next, we performed a DESeq2 analysis for differentially expressed genes in LPS-treated R26-Cyld-OECx3 versus LPS-treated R26RFP/WT microglia samples (threshold: adjusted p-value < 0.05, log2 fold change > 1). Figure 4G shows the 10 genes with the greatest difference in expression between the two groups. However, surprisingly, we could not detect any statistically significant DEGs. This rather unexpected finding strongly supports our previous results and suggests the conclusion that CYLD overexpression in microglia has no effect on microglia gene expression and function in the context of neuroinflammation.
Overexpression of Cyld in microglia does not alter the microglial response to LPS. R26-Cyld-OECx3 mice and littermate controls were injected intraperitoneally with 1 mg/kg LPS and microglia were isolated for flow cytometry and RNA sequencing 24 h later. A Change in body weight 24 h after LPS injection (shown as % of starting body weight). B Representative gating strategy for the flow cytometry analysis of brain and spinal cord microglia (brain is shown). From brain CNS samples, microglia were first isolated by MACS for CD11b+ cells. For spinal cord samples, the whole CNS cell suspension was used for flow cytometry. After gating on live, single cells, microglia were selected as CD45intermediate, CD68+ and Ly6C−. C Microglia purity was analyzed by flow cytometry in the MACS-isolated brain samples used for RNA sequencing. D Mean fluorescence intensity (MFI) as geometric mean of CD68 and MHC-II on brain microglia. E Mean fluorescence intensity (MFI) as geometric mean of CD68 and MHC-II on spinal cord microglia. F Purity of MACS-isolated microglia was confirmed by plotting reads per kilobase of transcript per million mapped reads (RPKM) for marker genes of CNS and immune cell types. G Normalized counts of the top 10 genes with the greatest difference in expression between R26-Cyld-OECx3 and control microglia (not differentially expressed). In A, C, D and E, every circle represents a single mouse. Data in A, C, D and E is represented as mean ± SEM. Statistical significance was determined by two-tailed unpaired Student’s t-test (A, C, D, E). ***** p < 0.0001, ns = non-significant
Discussion
CYLD inhibits NF-κB activation, which is a key pathway in regulating microglia during inflammation and in various CNS diseases [29]. However, the specific mechanisms by which NF-κB is regulated in microglia and the potential pathways targeted by CYLD beyond NF-κB remain incompletely understood. Given the detrimental effects of A20 deletion in microglia [20, 21], an enzyme that functions similar to CYLD, we hypothesized that CYLD overexpression in microglia could have the opposite effect and might protect mice against CNS inflammation. Contrary to our expectations, overexpression of CYLD in microglia did not affect microglial gene expression or function in LPS-induced neuroinflammation and did not influence disease severity in the EAE model. Additionally, in the steady state, CYLD overexpression had no effect on immune cell infiltration into the CNS or on microglia numbers in different brain regions. Thus, we conclude that constitutive overexpression of CYLD in microglia cannot serve as a strategy to protect mice from neuroinflammation.
Our most compelling evidence for this conclusion comes from the model of LPS-induced neuroinflammation. LPS is the most common stimulus for microglia activation in vivo and in vitro [30] and acts by activating the NF-κB pathway in microglia via the activation of TLR4. We induced the overexpression of CYLD in microglia by tamoxifen injection and analyzed the microglial response to LPS 12 weeks later. To our surprise, no differences between R26-Cyld-OECx3 mice and R26RFP/WT littermate controls could be observed on the protein and gene expression levels, despite clear expression changes between naïve and LPS-treated mice, which confirmed that our analysis approach in general worked. This was further corroborated in the EAE model, where CYLD overexpression in microglia did not influence disease severity or immune cell infiltration into the CNS.
In contrast, deletion of the deubiquitinating enzyme A20 in microglia significantly enhanced microglia activation and neuroinflammation. Mice with A20 deletion were more susceptible to LPS and EAE and exhibited a hyperactive microglial phenotype with increased proinflammatory cytokine production [20]. One possible explanation for our observation that microglial CYLD overexpression had no effect, while microglial A20 deletion was detrimental, is that even though CYLD and A20 share a number of substrates, they also have distinct substrate specificities and A20 has non-catalytic functions that CYLD lacks [22]. Furthermore, while mice with full deletion of A20 suffer from multiorgan inflammation and die at a young age [31], the loss of CYLD is not problematic [25] indicating that A20 is more critical in preventing excessive immune responses. This could be due to the fact that A20 is upregulated in a NF-κB-dependent manner which serves as a negative feedback mechanism to terminate NF-κB signaling [32]. Another factor could be that CYLD is expressed at much lower levels in microglia compared to A20 [33], potentially limiting its role in regulating microglia function under physiological conditions. However, even if CYLD is physiologically not required for microglia function, we hypothesized that its overexpression could nonetheless protect against neuroinflammation by restraining NF-κB signaling. One hypothesis for the lack of this protective effect is that the artificially expressed CYLD could be functionally inactivated, possibly through phosphorylation or cleavage [22]. In conclusion, we are convinced that CYLD overexpression in microglia is not effective to protect mice from neuroinflammation. However, we cannot rule out the possibility that endogenous CYLD in microglia could play a role in the regulation of neuroinflammation. Further studies with conditional CYLD deletion in microglia are needed to explore this possibility.
Also in the steady state, CYLD overexpression in microglia did not affect microglia numbers across different brain regions or the infiltration of peripheral immune cells into the CNS. In contrast, mice with A20 deletion in microglia showed increased microglia numbers [20, 21] and spontaneous infiltration of CD8+ T cells in the CNS [21], again suggesting that A20 is more critical for the regulation of microglia physiology. We observed a slight change in microglia morphology in the cortex of R26-Cyld-OECx3 mice twelve weeks post-tamoxifen injection, but this was not seen at two weeks or six months, questioning the significance of this finding. While no studies with a microglia-specific Cyld deletion have been conducted so far, global Cyld−/− mice displayed altered microglial numbers and morphology, along with elevated levels of the proinflammatory cytokines IL-1β and TNF-α in the dorsal striatum, indicating a more activated microglia phenotype [14, 19]. This may result from the loss of CYLD in the microglia themselves, which might remove a regulatory brake on microglia proliferation. The additional overexpression of CYLD in our model does not necessarily need to have the opposite effect, it rather seems likely that it would not have any additional benefit in this context. Furthermore, CYLD is crucial for maintaining neuronal activity, synaptic transmission and normal behavior of mice [14,15,16,17]. The loss of CYLD in microglia could contribute to these neurological phenotypes in Cyld−/− mice, for example due to alterations in synaptic pruning conducted by the microglia. Nevertheless, it seems more likely that the absence of CYLD in neurons is the primary cause for these phenotypes, since CYLD highly accumulates in the postsynaptic density in a neuronal activity-dependent manner [34] and therefore can directly regulate synaptic signaling pathways.
In conclusion, overexpression of CYLD in microglia does not protect mice against neuroinflammation and thus is not a viable therapeutic strategy. Further studies using conditional CYLD knock-out mice are necessary to fully elucidate CYLD’s role in microglial regulation. Our novel Rosa26-Cyld-tdTomato mice provide a valuable tool for exploring CYLD’s function across various cell types, potentially advancing our understanding of inflammatory disorders and cancer.
Methods
Mice
All mice used were on a C57BL/6 background and housed under specific pathogen-free conditions. Rosa26-Cyld-tdTomato mice were generated by Taconic Biosciences. The genotypes of the generated Rosa26-Cyld-tdTomato mice were assessed with the following primers: forward_1 5’-TTGGGTCCACTCAGTAGATGC-3’; forward_2 5’- CTCTTCCCTCGTGATCTGCAACTCC-3’; reverse 5’-CATGTCTTTAATCTACCTCGATGG-3’. A PCR with all 3 primers detects the Rosa26 wildtype allele (299 bp; forward_2 and reverse primer) and the Rosa26-Cyld-tdTomato knock-in allele (744 bp; forward_1 and reverse primer). Rosa26-Cyld-tdTomato mice were crossed to Cx3cr1CreER mice [23] to obtain R26-Cyld-OECx3 mice with the genotype Cx3Cr1CreER/WT; Rosa26Cyld−OE−tdTom/WT. Rosa26-tdRFP reporter mice [24] were crossed to R26-Cyld-OECx3 mice to obtain littermate control animals (R26RFP/WT) with the genotype Cx3Cr1CreER/WT; Rosa26RFP/WT. For the EAE experiment, heterozygous Cyld+/- mice [25] were crossed to obtain Cyld−/− mice and heterozygous (Cyld+/-) and homozygous (Cyld+/+) littermate controls.
Tamoxifen treatment
Tamoxifen (Sigma-Aldrich) at a concentration of 20 mg/ml was dissolved in olive oil (Sigma-Aldrich) containing 5% ethanol by rotation overnight at 4 °C. Mice were injected subcutaneously with 100 μl (= 2 mg tamoxifen) at the age of 3–4 weeks on two days with a one-day break in between. R26RFP/WT littermate controls were treated equally.
Immunofluorescence stainings and Sholl analysis
Brains were isolated from mice perfused with 0.9% NaCl (Sigma-Aldrich), were fixed in 4% PFA for 6 h at 4 °C and cryoprotected in 30% sucrose. The tissue was embedded in TissueTek® O.C.T™ compound (Sakura Finetek) and 50 μm free-floating sections were cut at a cryostat.
Endogenous peroxidase was blocked in methanol containing 0.6% H2O2 for 20 min. at room temperature. Endogenous biotin and avidin binding sites were blocked with the Avidin/Biotin Blocking Kit (Vector Laboratories) for 15 min. each at room temperature. Sections were blocked with 10% ROTI®ImmunoBlock (Carl Roth) and 2% bovine serum albumin (BSA; PAN-Biotech) diluted in TBST (Tris Buffer Saline containing 1% TWEEN® 20) for 15 min. at room temperature. Sections were incubated with primary antibodies against Iba-1 (goat polyclonal, 1:500, FUJIFILM Wako Chemicals) and RFP (rabbit polyclonal, 1:1000, Rockland Immunochemicals) diluted in 2% BSA/TBST overnight at 4 °C. Sections were washed and incubated with secondary antibodies against goat IgG (donkey polyclonal, Alexa Fluor 647, 1:400, Invitrogen) and against rabbit IgG (donkey polyclonal, Biotin, 1:500, Invitrogen) in TBST for 1.5 h at room temperature. After washing, sections were treated with the TSA® Cyanine 3 detection kit (Akoya Biosciences) following manufacturer’s instructions. Sections were washed and mounted with Vectashield® Antifade Mounting Medium with DAPI (Vector Laboratories). For microglia quantification, images were acquired at a DMi8 widefield microscope (Leica) with an objective magnification of 20 × and only microglia with a clear cell body in-focus were manually counted. For Sholl analysis, images were acquired at a TCS SP8 multiphoton microscope (Leica) with an objective magnification of 20 × with 3 × optical zoom and 1.0 µm increments. Sholl analysis was performed in Fiji (ImageJ). First, Iba1 stacks were collapsed using maximum projection. The threshold was set to the same level for all images and only microglia with the cell body at the center of the stack analyzed. The cell body was defined and Sholl analysis was performed with a starting radius of 1 μm from the center and in increments of 1 μm.
Cell isolation from the CNS
Brains were dissected from mice transcardially perfused with 0.9% NaCl solution (Sigma-Aldrich) and digested with 1 mg/ml Papain (Sigma-Aldrich) and 100 µg/ml DNase I (Roche) in HBSS with calcium and magnesium with mechanical dissociation with the GentleMACS homogenizer (Miltenyi, Bergisch Gladbach Germany) at 37 °C for 30 min. CNS homogenates were passed through a 70 μm cell strainer and cells were separated using a 70/30% Percoll (Sigma-Aldrich) gradient centrifugation for 45 min, 500 × g at 18 °C without brakes. Myelin was discarded and cells at the 70/30% interphase were carefully collected for further processing.
Microglia isolation by magnetic-activated cell sorting (MACS)
Cells obtained from the CNS cell isolation were resuspended in 90 µl MACS buffer and 10 µl anti-CD11b MicroBeads (Miltenyi Biotec) and incubated for 15 min. at 4 °C. After washing, cells were applied to a MS column in an OctoMACS separator (Miltenyi Biotec), washed 3 × with MACS buffer and CD11b+ cells were eluted.
Flow cytometry
Cells obtained from the CNS cell isolation were resuspended in anti-mouse CD16/CD32 (5 µg/ml, BioXCell,) in FACS buffer (PBS containing 2% fetal calf serum) for 10 min. at 4 °C to block Fc receptors. Cells were stained for 30 min at 4 °C on the cell surface with antibodies against CD4 BV421 (GK1.5, rat monoclonal, 1:400, BioLegend), CD45 BV510 (30-F11, rat monoclonal, 1:300, BioLegend), TCR-β FITC (H57-597, hamster monoclonal, 1:300, BioLegend), CD11b PECy7 (M1/70, rat monoclonal, 1:1000, eBioscience™, Thermo Fisher Scientific), CD8a PerCP (53–6.7, rat monoclonal, 1:500, BioLegend), MHC-II eFluor™450 (M5/114.15.2, rat monoclonal, 1:4000, eBioscience™, Thermo Fisher Scientific). For dead cell exclusion, fixable viability dye eFluor™780 (eBioscience™, Thermo Fisher Scientific) was added to the surface staining. Afterwards, cells were fixed and permeabilized with Cytofix/Cytoperm (BD Bioscience) and stained overnight at 4 °C with anti-CD68 APC antibody (FA-11, 1:500, rat monoclonal, BioLegend). Samples were acquired at a FACSCanto II using FACS Diva software (BD Bioscience) and analyzed with FlowJoTM v10 software (BD Bioscience). For all analyses, doublets (FSC and SSC properties) and dead cells were excluded.
RNA isolation, reverse transcription and real-time PCR
RNA was isolated with the ReliaPrep™ RNA Cell Miniprep System (Promega) following manufacturer’s guidelines. RNA concentrations were determined by measuring absorbance at 260 nm and 280 nm using the NanoQuant Plate™ (Tecan) at an Infinite M200 pro plate reader (Tecan). cDNA was synthesized with the QuantiTect® Reverse Transcription Kit (Qiagen) using all RNA obtained following manufacturer’s guidelines. Real-time PCR (RT-PCR) was performed with the StepOnePlus™ Real-Time PCR System (Life Technologies) using SYBR Green reagent (Promega). Fold enrichment was calculated using the Delta-Delta CT method normalized to hypoxanthin-guanin-phosphoribosyltransferase (Hprt) as house-keeping gene reference. Hprt primers were self-designed: forward 5’-CGTCGTGATTAGCGATGATG-3’, reverse 5’- TCCAAATCCTCGGCATAATG-3’. Cyld primers were ordered as QuantiTect Primer Assay (QT00103768, Qiagen).
RNA isolation and RNA sequencing
For RNA sequencing, RNA was isolated with the RNeasy Micro Kit (Qiagen). RNA quality was assessed with the Qubit RNA High Sensitivity Assay Kit on a Qubit 3 Fluorometer (Invitrogen) and with the High Sensitivity RNA ScreenTape on a 4200 TapeStation system (Agilent). 100 ng RNA were used for library preparation with the NEBNext Single Cell/Low Input RNA Library Prep Kit for Illumina using NEBNext Multiplex Oligos for Illumina (New England Biolabs). cDNA quality was assessed with Agilent’s High Sensitivity DNA kit on an Agilent 2100 Bioanalyzer. Quantity of the library was assessed with the Qubit 1X dsDNA assay kit on a Qubit 4 Fluorometer (Invitrogen) and library size was determined with a D1000 ScreenTape analysis on a 4200 TapeStation system (Agilent). All libraries were sequenced in paired-end mode (2 × 50 nt) on an Illumina NovaSeq 6000 instrument resulting in an average of 20 million distinct sequencing reads per library.
RNA sequencing data analysis
Quality control of the sequencing data was performed with the FastQC tool (v0.12.1). RNA sequencing reads were mapped to the Mus_musculus.GRCm38 reference genome (ENSEMBL v84) using kallisto (v0.46.0). Alignments were processed with kallisto to obtain gene transcripts per million (TPM) values and gene counts. Differential gene expression analysis was performed with DESeq2 (v1.34.0) and apeglm (v1.16.0) was applied for shrinkage of log-fold changes.
Experimental autoimmune encephalomyelitis (EAE)
Mice were immunized 12 weeks after tamoxifen administration at the age of 15–16 weeks. MOG35-55/CFA was prepared by mixing 1 mg/ml MOG35-55 (GenScript) in PBS with Complete Freund’s Adjuvants (CFA, BD Biosciences) containing mycobacterium tuberculosis H37RA (BD Biosciences). 50 μg of the MOG35-55/CFA emulsion were injected subcutaneously into the tail base. Mice were also injected intraperitoneally with 150 ng pertussis toxin (List Biological Laboratories) in PBS at the day of immunization and two days later.
Mice were weighed daily and clinical scores were documented as follows: 0 - no disease; 0.5 - limb tail; 1 - paralyzed tail; 1.5 - weakened righting reflex; 2 - no righting reflex; 3 - partial paralysis of hind legs; 3.5 - paralysis of one hind leg; 4 - paralysis of both hind legs.
LPS-induced neuroinflammation
LPS (from Salmonella typhimurium, Sigma-Aldrich) was solved in PBS at a concentration of 0.2 mg/ml. Mice were injected 12 weeks after tamoxifen administration at the age of 15–16 weeks with 1 mg/kg LPS intraperitoneally and were sacrificed 24 h later.
Statistics
Statistical analyses were performed with Prism v10 software (GraphPad). Values are shown as mean ± standard error of the mean (SEM) unless indicated otherwise. P values were considered significant with * p < 0.05, ** p < 0.01, *** p < 0.001 and **** p < 0.0001.
Data availability
The RNA sequencing data generated in this study will be made available on request.
References
Biggs PJ, Wooster R, Ford D, Chapman P, Mangion J, Quirk Y, Easton DF, Burn J, Stratton MR (1995) Familial cylindromatosis (turban tumour syndrome) gene localised to chromosome 16q12-q13: evidence for its role as a tumour suppressor gene. Nat Genet 11:441–443. https://doi.org/10.1038/ng1295-441
Bignell GR, Warren W, Seal S, Takahashi M, Rapley E, Barfoot R, Green H, Brown C, Biggs PJ, Lakhani SR et al (2000) Identification of the familial cylindromatosis tumour-suppressor gene. Nat Genet 25:160–165. https://doi.org/10.1038/76006
Sato Y, Goto E, Shibata Y, Kubota Y, Yamagata A, Goto-Ito S, Kubota K, Inoue J, Takekawa M, Tokunaga F et al (2015) Structures of CYLD USP with Met1- or Lys63-linked diubiquitin reveal mechanisms for dual specificity. Nat Struct Mol Biol 22:222–229. https://doi.org/10.1038/nsmb.2970
Trompouki E, Hatzivassiliou E, Tsichritzis T, Farmer H, Ashworth A, Mosialos G (2003) CYLD is a deubiquitinating enzyme that negatively regulates NF-kappaB activation by TNFR family members. Nature 424:793–796. https://doi.org/10.1038/nature01803
Kovalenko A, Chable-Bessia C, Cantarella G, Israël A, Wallach D, Courtois G (2003) The tumour suppressor CYLD negatively regulates NF-kappaB signalling by deubiquitination. Nature 424:801–805. https://doi.org/10.1038/nature01802
Brummelkamp TR, Nijman SM, Dirac AM, Bernards R (2003) Loss of the cylindromatosis tumour suppressor inhibits apoptosis by activating NF-kappaB. Nature 424:797–801. https://doi.org/10.1038/nature01811
Reiley WW, Zhang M, Jin W, Losiewicz M, Donohue KB, Norbury CC, Sun SC (2006) Regulation of T cell development by the deubiquitinating enzyme CYLD. Nat Immunol 7:411–417. https://doi.org/10.1038/ni1315
Reiley WW, Jin W, Lee AJ, Wright A, Wu X, Tewalt EF, Leonard TO, Norbury CC, Fitzpatrick L, Zhang M et al (2007) Deubiquitinating enzyme CYLD negatively regulates the ubiquitin-dependent kinase Tak1 and prevents abnormal T cell responses. J Exp Med 204:1475–1485. https://doi.org/10.1084/jem.20062694
Jin W, Reiley WR, Lee AJ, Wright A, Wu X, Zhang M, Sun SC (2007) Deubiquitinating enzyme CYLD regulates the peripheral development and naive phenotype maintenance of B cells. J Biol Chem 282:15884–15893. https://doi.org/10.1074/jbc.M609952200
Hövelmeyer N, Wunderlich FT, Massoumi R, Jakobsen CG, Song J, Wörns MA, Merkwirth C, Kovalenko A, Aumailley M, Strand D et al (2007) Regulation of B cell homeostasis and activation by the tumor suppressor gene CYLD. J Exp Med 204:2615–2627. https://doi.org/10.1084/jem.20070318
Reissig S, Hovelmeyer N, Weigmann B, Nikolaev A, Kalt B, Wunderlich TF, Hahn M, Neurath MF, Waisman A (2012) The tumor suppressor CYLD controls the function of murine regulatory T cells. J Immunol 189:4770–4776. https://doi.org/10.4049/jimmunol.1201993
Srokowski CC, Masri J, Hövelmeyer N, Krembel AK, Tertilt C, Strand D, Mahnke K, Massoumi R, Waisman A, Schild H (2009) Naturally occurring short splice variant of CYLD positively regulates dendritic cell function. Blood 113:5891–5895. https://doi.org/10.1182/blood-2008-08-175489
Bros M, Dexheimer N, Besche V, Masri J, Trojandt S, Hövelmeyer N, Reissig S, Massoumi R, Grabbe S, Waisman A et al (2010) Mutated cylindromatosis gene affects the functional state of dendritic cells. Eur J Immunol 40:2848–2857. https://doi.org/10.1002/eji.200939285
Chen SY, Liu KF, Tan SY, Chen XS, Li HD, Li JJ, Zhou JW, Yang L, Long C (2023) Deubiquitinase CYLD regulates excitatory synaptic transmission and short-term plasticity in the hippocampus. Brain Res 1806:148313. https://doi.org/10.1016/j.brainres.2023.148313
Li HD, Li DN, Yang L, Long C (2021) Deficiency of the CYLD Impairs Fear Memory of Mice and Disrupts Neuronal Activity and Synaptic Transmission in the Basolateral Amygdala. Front Cell Neurosci 15:740165. https://doi.org/10.3389/fncel.2021.740165
Colombo E, Horta G, Roesler MK, Ihbe N, Chhabra S, Radyushkin K, Di Liberto G, Kreutzfeldt M, Schumann S, von Engelhardt J et al (2021) The K63 deubiquitinase CYLD modulates autism-like behaviors and hippocampal plasticity by regulating autophagy and mTOR signaling. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.2110755118
Zajicek AS, Ruan H, Dai H, Skolfield MC, Phillips HL, Burnette WJ, Javidfar B, Sun SC, Akbarian S, Yao WD (2022) Cylindromatosis drives synapse pruning and weakening by promoting macroautophagy through Akt-mTOR signaling. Mol Psychiatry 27:2414–2424. https://doi.org/10.1038/s41380-022-01571-1
Nimmerjahn A, Kirchhoff F, Helmchen F (2005) Resting microglial cells are highly dynamic surveillants of brain parenchyma in vivo. Science 308:1314–1318. https://doi.org/10.1126/science.1110647
Han YY, Jin K, Pan QS, Li B, Wu ZQ, Gan L, Yang L, Long C (2020) Microglial activation in the dorsal striatum participates in anxiety-like behavior in Cyld knockout mice. Brain Behav Immun 89:326–338. https://doi.org/10.1016/j.bbi.2020.07.011
Voet S, Mc Guire C, Hagemeyer N, Martens A, Schroeder A, Wieghofer P, Daems C, Staszewski O, Vande Walle L, Jordao MJC et al (2018) A20 critically controls microglia activation and inhibits inflammasome-dependent neuroinflammation. Nat Commun 9:2036. https://doi.org/10.1038/s41467-018-04376-5
Mohebiany AN, Ramphal NS, Karram K, Di Liberto G, Novkovic T, Klein M, Marini F, Kreutzfeldt M, Härtner F, Lacher SM et al (2020) Microglial A20 Protects the Brain from CD8 T-Cell-Mediated Immunopathology. Cell Rep 30:1585-1597.e1586. https://doi.org/10.1016/j.celrep.2019.12.097
Lork M, Verhelst K, Beyaert R (2017) CYLD, A20 and OTULIN deubiquitinases in NF-κB signaling and cell death: so similar, yet so different. Cell Death Differ 24:1172–1183. https://doi.org/10.1038/cdd.2017.46
Yona S, Kim KW, Wolf Y, Mildner A, Varol D, Breker M, Strauss-Ayali D, Viukov S, Guilliams M, Misharin A et al (2013) Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38:79–91. https://doi.org/10.1016/j.immuni.2012.12.001
Luche H, Weber O, Nageswara Rao T, Blum C, Fehling HJ (2007) Faithful activation of an extra-bright red fluorescent protein in “knock-in” Cre-reporter mice ideally suited for lineage tracing studies. Eur J Immunol 37:43–53. https://doi.org/10.1002/eji.200636745
Massoumi R, Chmielarska K, Hennecke K, Pfeifer A, Fässler R (2006) Cyld inhibits tumor cell proliferation by blocking Bcl-3-dependent NF-kappaB signaling. Cell 125:665–677. https://doi.org/10.1016/j.cell.2006.03.041
Qin L, Wu X, Block ML, Liu Y, Breese GR, Hong JS, Knapp DJ, Crews FT (2007) Systemic LPS causes chronic neuroinflammation and progressive neurodegeneration. Glia 55:453–462. https://doi.org/10.1002/glia.20467
Hines DJ, Choi HB, Hines RM, Phillips AG, MacVicar BA (2013) Prevention of LPS-induced microglia activation, cytokine production and sickness behavior with TLR4 receptor interfering peptides. PLoS ONE 8:e60388. https://doi.org/10.1371/journal.pone.0060388
Keren-Shaul H, Spinrad A, Weiner A, Matcovitch-Natan O, Dvir-Szternfeld R, Ulland TK, David E, Baruch K, Lara-Astaiso D, Toth B et al (2017) A Unique Microglia Type Associated with Restricting Development of Alzheimer’s Disease. Cell 169:1276-1290.e1217. https://doi.org/10.1016/j.cell.2017.05.018
Anilkumar S, Wright-Jin E (2024) NF-κB as an Inducible Regulator of Inflammation in the Central Nervous System. Cells. https://doi.org/10.3390/cells13060485
Lively S, Schlichter LC (2018) Microglia Responses to Pro-inflammatory Stimuli (LPS, IFNγ+TNFα) and Reprogramming by Resolving Cytokines (IL-4, IL-10). Front Cell Neurosci 12:215. https://doi.org/10.3389/fncel.2018.00215
Lee EG, Boone DL, Chai S, Libby SL, Chien M, Lodolce JP, Ma A (2000) Failure to regulate TNF-induced NF-kappaB and cell death responses in A20-deficient mice. Science 289:2350–2354. https://doi.org/10.1126/science.289.5488.2350
Catrysse L, Vereecke L, Beyaert R, van Loo G (2014) A20 in inflammation and autoimmunity. Trends Immunol 35:22–31. https://doi.org/10.1016/j.it.2013.10.005
Zhang Y, Chen K, Sloan SA, Bennett ML, Scholze AR, O’Keeffe S, Phatnani HP, Guarnieri P, Caneda C, Ruderisch N et al (2014) An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J Neurosci 34:11929–11947. https://doi.org/10.1523/jneurosci.1860-14.2014
Dosemeci A, Thein S, Yang Y, Reese TS, Tao-Cheng JH (2013) CYLD, a deubiquitinase specific for lysine63-linked polyubiquitins, accumulates at the postsynaptic density in an activity-dependent manner. Biochem Biophys Res Commun 430:245–249. https://doi.org/10.1016/j.bbrc.2012.10.131
Acknowledgements
We would like to thank Marity Dessolain and Michaela Blanfeld for their help with the experiments.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under project-ID 318346496 - SFB 1292 to EvS and AW, project-ID 500247271 - SPP2395 to AW and project-ID 221828878 - SFB 1080 to MJS.
Author information
Authors and Affiliations
Contributions
Experiment preparation, data collection and data analysis were performed by Eva Schramm, Vanessa Becker, Ilaria Palagi, Melanie Müller, Thomas Rösler, Feyza Durak, Anna Ebering and Khalad Karram. Ari Waisman designed the study together with Eva Schramm, supervised the project and acquired funding. Michael J. Schmeisser and Esther von Stebut contributed to the conception of the study; Michael J. Schmeisser provided the Rosa26-Cyld-tdTomato mouse strain. Figures were prepared and the manuscript draft was written by Eva Schramm and revised by Ari Waisman. All authors commented on previous versions of the manuscript and read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
All animal experiments in this study were approved by the local administrations (Landesuntersuchungsamt Koblenz, Germany; approval numbers G22-1-090, G13-1-099). Experiments were performed in accordance with the guidelines from the translational animal research center (TARC) Mainz, Germany. All efforts were made to minimize suffering of the mice.
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Disclosure
The data in Fig. 1 was generated by Vanessa Becker as part of her medical doctoral thesis at the Johannes Gutenberg University Mainz, Germany. The dissertation has not yet been published.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schramm, E., Becker, V., Palagi, I. et al. Constitutive expression of the deubiquitinating enzyme CYLD does not affect microglia phenotype or function in homeostasis and neuroinflammation. J Mol Med (2024). https://doi.org/10.1007/s00109-024-02489-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00109-024-02489-7