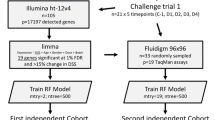
Abstract
Despite increases in vaccination coverage, reductions in influenza-related mortality have not been observed. Better vaccines are therefore required and influenza challenge studies can be used to test the efficacy of new vaccines. However, this requires the accurate post-challenge classification of subjects by outcome, which is limited in current methods that use artificial thresholds to assign ‘symptomatic’ and ‘asymptomatic’ phenotypes. We present data from an influenza challenge study in which 22 healthy adults (11 vaccinated) were inoculated with H3N2 influenza (A/Wisconsin/67/2005). We generated genome-wide gene expression data from peripheral blood taken immediately before the challenge and at 12, 24 and 48 h post-challenge. Variation in symptomatic scoring was found amongst those with laboratory confirmed influenza. By combining the dynamic transcriptomic data with the clinical parameters this variability can be reduced. We identified four subjects with severe laboratory confirmed influenza that show differential gene expression in 1103 probes 48 h post-challenge compared to the remaining subjects. We have further reduced this profile to six genes (CCL2, SEPT4, LAMP3, RTP4, MT1G and OAS3) that can be used to define these subjects. We have used this gene set to predict symptomatic infection from an independent study. This analysis gives further insight into host-pathogen interactions during influenza infection. However, the major potential value is in the clinical trial setting by providing a more quantitative method to better classify symptomatic individuals post influenza challenge.
Key message
-
Differential gene expression signatures are seen following influenza challenge.
-
Expression of six predictive genes can classify response to influenza challenge.
-
The genomic influenza response classification replicates in an independent dataset.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Seasonal influenza is common, affecting 5–10 % of the 900 million people in the European Region [1] with substantial associated mortality, morbidity and socioeconomic costs. Despite increases in vaccination coverage, reductions in influenza-related mortality have not been observed [2]. In order to develop better vaccines for seasonal and pandemic influenza, influenza challenge studies are central to testing vaccine efficacy. Consenting subjects are vaccinated, quarantined and then inoculated with influenza via intranasal administration. This approach has the advantage that potentially susceptible individuals can be positively selected based on low pre-existing immunity to the challenge virus. The sample size is restricted by the capacity of quarantine facilities and the high costs associated with performing a challenge experiment. Accurate post-challenge classification of subjects by outcome is then critical to determining vaccine efficacy. However these outcomes are based on scoring systems, which have artificial thresholds for ‘symptomatic’ and ‘asymptomatic’ phenotypes and include subjective self-reported observations and on biological evidence for infection (i.e. virus shedding and post-challenge serology) that are often discordant.
We sought to investigate the utility of genome-wide gene expression profiling in classification of volunteers following challenge. Application of a transcriptomic approach in the context of a challenge study is also a powerful approach to gain new insights into the biology of disease, as concordance in the exposure history and detailed clinical phenotyping before and after infection reduce the heterogeneity inherent to study of naturally acquired infection. Gene expression microarrays have been used to characterise the aberrant nature of the immune response to the 1918 pandemic influenza virus in macaques [3]. A number of studies have applied blood transcriptomics to understand host response to influenza in the context of lower respiratory tract infection and the H1N1 pandemic [4–8]. These approaches have also been applied to the field of influenza vaccinology [9], giving insights into the mechanisms of protective immunity. There has been one previous challenge study involving H3N2 with application of transcriptomic approaches [10, 11].
Here, we describe application of gene expression profiling to an influenza challenge study involving a vaccine called MVA-NP+M1. This viral-vectored vaccine has the potential to induce heterosubtypic immunity through the induction of cytotoxic T cells with specificity for epitopes within conserved influenza proteins. We have found MVA-NP+M1 to be safe and immunogenic [12, 13] with evidence that it may increase antibody responses to inactivated influenza vaccine when both vaccines are administered together [14]. The influenza challenge study (FLU002) to test the efficacy of MVA-NP+M1 [15] showed that vaccinees with laboratory confirmed influenza (LCI) shed virus for fewer days than control subjects with LCI. In this paper, we report the nature of the transcriptional response to influenza based on genome-wide gene expression profiling of participants during the FLU002 study. We describe how this information could be used to improve the biological classification of influenza challenge subjects. These results have implications for our understanding of the host-pathogen interactions for influenza and have the potential to inform clinical diagnostic testing.
Materials and methods
Study subjects and clinical trial procedures
Detailed description of the subjects and methods used in the clinical trial are provided in Lillie et al. [15]. The trial protocol was approved by the local Research Ethics Committee and conducted in accordance with the Declaration of Helsinki. All participants provided written informed consent. Briefly, 22 healthy volunteers, aged 18–45 years, were enrolled and 11 were vaccinated with 1.5×108 plaque forming units (pfu) of MVA-NP+M1. Thirty days after vaccination, all 22 volunteers underwent intranasal challenge with H3N2 influenza (A/Wisconsin/67/2005) at a dose of 1 ml of 105.25 TCID50/ml in a quarantine facility. Volunteers were all challenged within a 2-h period. All subjects had haemagglutination inhibition (HI) titres of <1:10 to the challenge virus on admission to the quarantine unit.
After virus challenge, volunteers had a physical examination by a trial physician daily and self-reported their symptoms using a modified Jackson scoring system [16, 17] twice daily, from 12-h post-challenge, for 6 days. This system lists upper respiratory and systemic symptoms on a scale of 0–3: ‘no symptoms’, ‘just noticeable’, ‘bothersome but can still do activities’ and ‘bothersome and cannot do daily activities’. Scores were summed over the duration of the challenge period to give a total symptom score. Those volunteers with scores of ≥4 and positive viral culture from nasal wash samples were categorised as having LCI. Severity of infection was graded as mild if the summed symptom score was 4–28, with scores of ≥29 classified as moderate/severe. HI titres for each individual at 26 days after challenge are provided (Online Resource, Table S1). Volunteers were kept in quarantine until day 6 post-challenge, and all had a negative rapid antigen test on nasal wash sample prior to discharge.
Whole blood was collected in PAXgene tubes prior to influenza challenge (day 30) and then at three further time points (12, 24 and 48 h post-challenge). All nasal washing and blood collections for viral shedding and gene expression analysis were taken in the same order that the volunteers were challenged in.
RNA extraction and globin mRNA depletion
Total RNA was extracted from whole blood using the PAXgene Blood RNA Kit (Qiagen) according to the manufacturer’s instructions. The abundance of globin mRNA present in whole blood reduces the sensitivity of expression array data, and therefore, globin mRNA depletion was carried out using the GLOBINclearTM-Human kit (Ambion) following the manufacturer’s instructions. Briefly, total RNA from whole blood is mixed with a biotinylated capture oligo mix specific for human globin mRNA. Streptavidin magnetic beads capture the globin mRNA. Placing the tube against a magnet then allows the globin mRNA depleted RNA to be transferred to a fresh tube. Globin-depleted RNA is further purified using a rapid, magnetic bead-based clean up procedure. Globin mRNA depleted samples were quantified using a NanoDrop spectrophotometer, and the quality for a subset was verified using a Bioanalyzer following the manufacturer’s instructions (Bioanalyzer RNA 6000 Nano kit, Agilent).
Microarray data processing and analysis
Genome-wide gene expression analysis was carried out on 500-ng globin depleted RNA using the Illumina HumanHT-12 v4 Expression BeadChip gene expression platform comprising 47,231 probes (Core Genomics, WTCHG). The report for analysis was generated by Illumina’s GenomeStudio version 1.6.0. Background signals were subtracted and 11,158 low expression probes (detection value <0.95) were removed. The raw data were transformed and normalised using the Variance Stabilization and Normalization method [18]. Quality control checks including principal component analysis to check for batch and array effects were carried out using R [19]. Two samples were excluded as outliers. The raw and normalised data are accessible through GEO (GSE61754).
Statistical analysis
Differential gene expression analysis was carried out on 36,073 probes using three different R packages: limma [20], maSigPro [21] and timecourse [22]. Benjamini-Hochberg corrected values are used to control for multiple testing in differential gene expression analysis and subsequent pathway analysis. Pathway and network analysis was performed using IPA (Ingenuity® Systems, www.ingenuity.com) analysing probes differentially expressed between the four moderate/severe LCI and the remaining samples, 48 h post-challenge. The pamr R package [23] was used to identify genes whose expression could accurately categorise moderate/severe LCI compared to the remaining samples. The pamr package employs a ‘nearest shrunken centroids’ method to identify subsets of genes that are best able to classify samples into the specified groups of interest. The Huang dataset [11] was normalised using the robust multi-array (RMA) method (data accessible at NCBI GEO database [24], accession GSE30550).
Results
Following influenza challenge, two individuals in the vaccinated group and five individuals in the unvaccinated group developed LCI as defined in the clinical trial protocol [15] (Fig. 1). Within the LCI group, there was considerable variation in the summed modified Jackson scores (ranging from 4 to 38) that describe the extent to which volunteers were symptomatic. We therefore hypothesised that these differences in clinical phenotype might be reflected in patterns of gene expression detected through microarray gene expression analysis.
Clinical observations and classification of study subjects. Each individual is assigned a unique letter code (A-V). Vaccinated individuals are shown in red. Individuals are defined as having no symptoms, mild or moderate/severe based on the summed self-reported symptoms collected over 6 days. Positive shedding indicating viral shedding was detected on at least 1 day after challenge. Seroconversion was determined by HI titre using serum samples obtained at 26-days post-challenge (Online Resource, Table S1). LCI is defined as mild or moderate/severe symptoms and viral shedding (shown by grey shaded area)
Principal component analysis (PCA) and hierarchical clustering was performed to visualise variation in the dataset (Fig. 2). This demonstrated that four subjects shared a similar profile of variance in gene expression. These four subjects were the only cases of LCI whose symptom score was moderate/severe rather than mild. They also shared other virology outcomes as shown in Fig. 1.
Principal components and hierarchical clustering plot. a The first three principal components are plotted with the proportion of variance explained by each component. The yellow oval highlights the samples taken from the four subjects with moderate/severe LCI 48 h post-challenge. N none or mild LCI, S moderate/severe LCI, V vaccinee, C control. b The hierarchical tree illustrates the relationship between clusters of samples. The height of the branches indicates the strength of the separation. The red oval highlights the samples taken from the four subjects with moderate/severe LCI 48 h post-challenge
We proceeded to analyse differential gene expression using a linear regression model. The likelihood of observed differences in gene expression was defined using an empirical Bayes approach in which prior distribution was estimated from the overall variance in the dataset, allowing a more stable inference given small numbers of arrays [25]. Implementing this approach using the Limma package [15], a total of 1103 probes were differentially expressed between these four volunteers and the remaining samples at 48 h post-challenge (q < 0.05 controlling for multiple testing using Benjamini and Hochberg’s method, Online Resource, Table S2 and Table S3). Using the same approach, no probes were differentially expressed at 12 or 24-h post-challenge (Online Resource, Table S2). This specific effect on differential expression at 48 h post-challenge is consistent with the observed temporal onset of symptoms [15].
In further analyses, the most significant 100 of these differentially expressed probes (9 %) were selected and almost all were found to be upregulated at 48 h post-challenge in the moderate/severe LCI individuals (Fig. 3a). In contrast, subjects with LCI and mild symptoms did not show substantial changes in expression of the same probes (Fig. 3b). To further validate these findings, two alternative methods based on a two-step regression approach (maSigPro package [21]) and on multivariate empirical Bayes statistics (timecourse package [22]) were employed to generate an equivalent list of the 200 most significant differentially expressed probes. The overlap with the original probe list was 81 and 74 %, respectively (Online Resource Fig. S1).
Heatmaps for differentially expressed probes. The top 100 most significant differentially expressed probes between the four moderate/severe LCI, and the remaining samples at 48 h post-challenge were used to generate heatmaps of gene expression. a Moderate/severe LCI subjects. b Mild LCI subjects. Individuals D and G were vaccinated; others are unvaccinated
To assess the functional significance of the differentially expressed genes in subjects with moderate/severe LCI at 48 h post-challenge, we performed pathway and network analysis (Fig. 4, Online Resource Figure S2). This showed significant enrichment for genes involved in a number of pathways notably interferon signalling (Benjamini-Hochberg corrected p value = 6.4 × 10−10), role of pattern recognition receptors (PRR) in recognition of bacteria and viruses (p = 4.0 × 10−6), activation of IRF by cytosolic PRR (p = 7.7 × 10−4), eIF2 signalling (p = 8.1 × 10−3) and death receptor signalling (p = 8.1 × 10−3) (Fig. 4). The most significant upstream regulators identified were type III, II and I interferons (IFNL1 p = 2.2 × 10−59, IFNA2 2.7 × 10−53, IFNG 3.4 × 10−42). Network analysis further highlighted the significant upregulation of genes involved in the IFN response seen in these patients (Fig. S2). IFNs play a critical role in host defence to viral infection with the significance of IFNλ recently recognised [26–28]. The eIF2 signalling pathway plays a role in host defence by reducing translation in the host cell thereby reducing the rate at which new viral particles are generated [29] while death receptor signalling involving apoptosis has been related to disease severity in influenza A infection [30]. In terms of diseases and functions related to the differentially expressed genes at 48 h post-challenge, infectious disease was most significant, specifically replication of virus (P = 9.3 × 10−17).
Pathway analysis of differential expression in LCI. The differentially expressed probes between the four moderate/severe LCI samples 48 h post-challenge and the remaining samples were used to determine pathway enrichment with IPA. a Pathways with significant enrichment based on Benjamini-Hochberg (B-H) multiple testing corrected p value <0.01 are shown. The blue bars show the −log(p value). Differentially expressed genes are plotted for each enriched pathway (red upregulated in moderate/severe LCI individuals, green downregulated) with numbers of enriched genes shown in brackets below x-axis labels. b The most significantly enriched pathway, interferon signalling, is shown with enriched genes shaded in colour
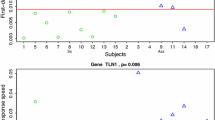
Although the analysis of over 1000 differentially expressed probes can provide insight into host-pathogen interactions, this magnitude of variables does not lend itself well to use in diagnostics. We therefore sought to find the minimum number of genes that could be used to successfully classify our data. Using the tool ‘prediction analysis for microarrays’ (pamr) [23], we identified six genes (CCL2, SEPT4, LAMP3, RTP4, MT1G and OAS3) which alone classified our samples into two groups with 100 % accuracy. These two groups comprised those from subjects with moderate/severe LCI taken at 48 h post-challenge in one group and the remaining 18 samples (with either mild LCI or without LCI) in the second group. The six selected genes are strongly upregulated in individuals with moderate/severe LCI with network analysis showing relationships involving interferon activation (Online Resource Fig. S3).
In order to assess the validity of this six gene signal, we identified a data set from an independent but similarly conducted influenza challenge study, published by Huang et al. [11]. This study used similar methods for RNA collection and extraction, and the challenge was performed with the same strain of influenza (A/Wisconsin/67/2005). Of the 17 subjects in this study, eight were described as having asymptomatic infection and nine as having symptomatic infection. Those with symptomatic infection had a median summed Jackson score of 39.
For each of the subjects in the Huang study, we included six samples taken at different time points: two samples prior to challenge were included, and the remaining four samples were from consecutive post-challenge time points taken at 45, 53, 60 and 69 h. For all the post-challenge samples in the eight asymptomatic volunteers, the six gene set classified the samples with 100 % accuracy. For 89 % (8/9) of the symptomatic volunteers, pamr correctly classified at least one of these six post-challenge timepoints (Table 1).
In addition, we investigated whether any differences were observed between other subgroups in the study, for instance, individuals who do not shed virus. We found no significant difference between the symptomatic and asymptomatic individuals in this subgroup. Finally, we attempted to identify whether vaccination modulated the host responses to influenza challenge. No significant difference in the temporal response to influenza was seen between vaccinees and controls on limma analysis.
Discussion
The results of this study highlight the value of combining genome-wide gene expression profiling with clinical observations in order to improve the biological classification of influenza challenge studies and assist with clinical diagnostic testing.
Using the principal components generated from the genome-wide gene expression profiles, we were able to identify four LCI samples taken 48 h post-challenge that clustered separately from the other samples. Upon further investigation, we discovered a pattern of predominately upregulated gene expression that is not observed in the remaining samples, including LCI cases with mild symptoms according to the clinical trial protocol. The signature correlates with the clinical observations, and therefore, setting an arbitrary symptom score threshold to classify LCI may not be the most informative end result of an influenza challenge study. Instead, this gene signature may assist with redefining the symptom score threshold or could be used in combination with the symptom score to better classify moderate/severe LCI.
The informativeness of differentially expressed genes at 48 h post-challenge are likely to reflect the observed peak in influenza A viral titres that have been reported at this time for experimentally infected adults which has been shown to be closely mirrored by changes in innate immune response notably type 1 interferon levels [31, 32]. From the list of probes that we found are differentially expressed in the four moderate/severe LCI cases 48 h post-challenge, pathway analysis identified a number of pathways which are enriched within the dataset. Many of these pathways are involved in the anti-viral response illustrating that these four individuals show a different immune response to those with mild LCI. Similar pathways, and in particular, those regulated by interferon, have previously been identified in a number of other studies; however, few have focused on the human in vivo response [33–35].
We were able to reduce the 48 h time point gene expression signature to just six genes (CCL2, SEPT4, LAMP3, RTP4, MT1G and OAS3). This enabled us to classify our samples into two distinct groups (symptomatic and asymptomatic) with 100 % accuracy providing a potentially convenient, cost effective and quantitative method for determining post-challenge outcome during vaccine and challenge trials. The reproducibility of this gene set has been demonstrated using samples from an independent vaccine study [17] with a high level of accuracy (100 % for asymptomatic and 89 % for symptomatic volunteers). To our knowledge, the only other transcriptomics analysis involving an H3N2 influenza challenge study was for 17 adults, described by Huang, Zaas, Woods and colleagues [10, 11, 17]. Woods et al. have also published data on H1N1 challenge [10]. They found the gene expression signatures induced by these two strains to be highly similar, with 88 % overlap between the most differentially expressed genes. Four of our six genes (CCL2, LAMP3, RTP4 and OAS3) are represented in the list of the top 50 genes from this H1N1 study. Two studies in 2013 examined genome-wide gene expression in patients with influenza acquired in the community. While none of these six genes were amongst the best discriminators of respiratory syncytial virus (RSV) from influenza, four of the six genes (OAS3, SEPT4, LAMP3 and RPT4) were represented in a gene list of 161 interferon-related differentially expressed genes [6]. In a separate study, Herberg et al. [5] reported genome-wide gene expression in 19 children with H1N1 infection. Of the 50 probes which were most significantly upregulated, four of our six genes were represented (OAS3, LAMP3, SEPT4 and MT1G).
The OAS3 gene product is an enzyme that functions to resist viral infection via the destruction of intracellular RNA [29]. CCL2 encodes a chemokine that attracts monocytes. The clinical significance of CCL2/MCP1 in influenza disease has been suggested by studies that have associated elevated CCL2 levels with a poor prognosis in H1N1 infected Chinese patients [36] and a small interfering RNA (siRNA) screen showing effects on vaccinia virus infection [37]. While CCL2 attracts monocytes, receptor transporter protein 4 (RTP4) and the metallothionein isoform MT1G can be expressed by monocytes [38, 39]. RTP4 is known to be induced by type I interferons and have targeted anti-viral actions [40]. A high-throughput siRNA screen in primary lung cells showed that RPT4 was a key determinant of influenza virus replication [35]. SEPT4 encodes a member of the septin family of nucleotide binding proteins, and Septin 5 was reported to be upregulated in chicken brain following H5N1 infection [41]. The lysosomal-associated membrane protein 3 (LAMP3) is expressed by activated DCs [42] and is significantly induced in human lung epithelial cells (A549 cells) following influenza infection in vitro [43]. Our network analysis and the published evidence described here links these genes to the immune and specifically anti-viral response but shows that they are not involved in the same biological pathways. By representing different biological pathways, these six genes may succinctly describe a complex pattern of gene expression.
The Huang dataset illustrates the variability that can be seen between individuals in response to influenza challenge. For the eight samples that we successfully predicted as being symptomatic using the panel of six genes, there is no one single time point at which all eight samples are predicted as symptomatic. This suggests that the window for symptomatic classification can vary from individual to individual over a number of hours. Further work may be required to optimise the panel of genes used dependent on the specific time window post-challenge used to measure gene expression.
Fewer vaccinated volunteers were expected to develop LCI and experience moderate/severe symptoms less frequently compared to unvaccinated volunteers. We did not detect a difference in the gene expression profile between vaccinated and unvaccinated individuals in this study. This is perhaps not surprising as initial samples were taken 30 days after vaccination, and previous studies [44] with viral-vectored vaccines have shown differential gene expression to be undetectable at 7 days post-vaccination. Further studies will be needed to confirm the preliminary evidence of vaccine efficacy, and these will be facilitated by the use of gene expression data to allow objective classification of outcomes.
This analysis gives further insight into host-pathogen interactions which could be beneficial for early detection of influenza. However, the major value is in the clinical trial setting. Current biological phenotyping methods often show inconsistency, but this work illustrates the use of a more quantitative method. The ability to better classify symptomatic individuals post influenza challenge will help accelerate future influenza vaccine efficacy studies.
References
World Health Organization. Influenza. Available via: http://www.euro.who.int/en/what-we-do/health-topics/communicable-diseases/influenza/facts-and-figures. Accessed: 11/06/2013
Simonsen L, Taylor RJ, Viboud C, Miller MA, Jackson LA (2007) Mortality benefits of influenza vaccination in elderly people: an ongoing controversy. Lancet Infect Dis 7:658–666
Kobasa D, Jones SM, Shinya K, Kash JC, Copps J, Ebihara H, Hatta Y, Kim JH, Halfmann P, Hatta M et al (2007) Aberrant innate immune response in lethal infection of macaques with the 1918 influenza virus. Nature 445:319–323
Bermejo-Martin JF, Martin-Loeches I, Rello J, Anton A, Almansa R, Xu L, Lopez-Campos G, Pumarola T, Ran L, Ramirez P et al (2010) Host adaptive immunity deficiency in severe pandemic influenza. Crit Care 14:R167
Herberg JA, Kaforou M, Gormley S, Sumner ER, Patel S, Jones KD, Paulus S, Fink C, Martinon-Torres F, Montana G et al (2013) Transcriptomic profiling in childhood H1N1/09 influenza reveals reduced expression of protein synthesis genes. J Infect Dis 208:1664–1668
Mejias A, Dimo B, Suarez NM, Garcia C, Suarez-Arrabal MC, Jartti T, Blankenship D, Jordan-Villegas A, Ardura MI, Xu Z et al (2013) Whole blood gene expression profiles to assess pathogenesis and disease severity in infants with respiratory syncytial virus infection. PLoS Med 10:e1001549
Parnell GP, McLean AS, Booth DR, Armstrong NJ, Nalos M, Huang SJ, Manak J, Tang W, Tam OY, Chan S et al (2012) A distinct influenza infection signature in the blood transcriptome of patients with severe community-acquired pneumonia. Crit Care 16:R157
Song H, Wang Q, Guo Y, Liu S, Song R, Gao X, Dai L, Li B, Zhang D, Cheng J (2013) Microarray analysis of microRNA expression in peripheral blood mononuclear cells of critically ill patients with influenza A (H1N1). BMC Infect Dis 13:257
Bucasas KL, Franco LM, Shaw CA, Bray MS, Wells JM, Nino D, Arden N, Quarles JM, Couch RB, Belmont JW (2011) Early patterns of gene expression correlate with the humoral immune response to influenza vaccination in humans. J Infect Dis 203:921–929
Woods CW, McClain MT, Chen M, Zaas AK, Nicholson BP, Varkey J, Veldman T, Kingsmore SF, Huang Y, Lambkin-Williams R et al (2013) A host transcriptional signature for presymptomatic detection of infection in humans exposed to influenza H1N1 or H3N2. PLoS One 8:e52198
Huang Y, Zaas AK, Rao A, Dobigeon N, Woolf PJ, Veldman T, Oien NC, McClain MT, Varkey JB, Nicholson B et al (2011) Temporal dynamics of host molecular responses differentiate symptomatic and asymptomatic influenza A infection. PLoS Genet 7:e1002234
Berthoud TK, Hamill M, Lillie PJ, Hwenda L, Collins KA, Ewer KJ, Milicic A, Poyntz HC, Lambe T, Fletcher HA et al (2011) Potent CD8+ T-cell immunogenicity in humans of a novel heterosubtypic influenza A vaccine, MVA-NP M1. Clin Infect Dis 52:1–7
Antrobus RD, Lillie PJ, Berthoud TK, Spencer AJ, McLaren JE, Ladell K, Lambe T, Milicic A, Price DA, Hill AV et al (2012) A T cell-inducing influenza vaccine for the elderly: safety and immunogenicity of MVA-NP+M1 in adults aged over 50 years. PLoS One 7:e48322
Antrobus RD, Berthoud TK, Mullarkey CE, Hoschler K, Coughlan L, Zambon M, Hill AV, Gilbert SC (2014) Coadministration of seasonal influenza vaccine and MVA-NP+M1 simultaneously achieves potent humoral and cell-mediated responses. Mol Ther 22:233–238
Lillie PJ, Berthoud TK, Powell TJ, Lambe T, Mullarkey C, Spencer AJ, Hamill M, Peng Y, Blais ME, Duncan CJ et al (2012) Preliminary assessment of the efficacy of a T-cell-based influenza vaccine, MVA-NP+M1, in humans. Clin Infect Dis 55:19–25
Jackson GG, Dowling HF, Spiesman IG & Boand AV. (1958) Transmission of the common cold to volunteers under controlled conditions. I. The common cold as a clinical entity. A.M.A. archives of internal medicine 101:267–278
Zaas AK, Chen M, Varkey J, Veldman T, Hero AO 3rd, Lucas J, Huang Y, Turner R, Gilbert A, Lambkin-Williams R et al (2009) Gene expression signatures diagnose influenza and other symptomatic respiratory viral infections in humans. Cell Host Microbe 6:207–217
Huber W, von Heydebreck A, Sultmann H, Poustka A, Vingron M (2002) Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 18(Suppl 1):S96–S104
R: A language and environment for statistical computing (R Foundation for Statistical Computing, Vienna, Austria, 2013)
Smyth GK. in Bioinformatics and computational biology solutions using R and Bioconductor (ed Robert Gentleman et al.) 397–420 (Springer, 2005)
Conesa A, Nueda MJ, Ferrer A, Talon M (2006) maSigPro: a method to identify significantly differential expression profiles in time-course microarray experiments. Bioinformatics 22:1096–1102
Tai YC. timecourse: Statistical Analysis for Developmental Microarray Time Course Data. Available via: http://www.bioconductor.org . Accessed: version 1.28.0. 2007
Hastie TT, Narasimhan B & Chu G. pamr: Pam: prediction analysis for microarrays. Available via : http://CRAN.R-project.org/package=pamr . Accessed: version 1.54.1. 2013
Edgar R, Domrachev M, Lash AE (2002) Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–210
Smyth GK. (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3:Article3
Garcia-Sastre A, Biron CA (2006) Type 1 interferons and the virus-host relationship: a lesson in detente. Science 312:879–882
Hermant P, Michiels T (2014) Interferon-lambda in the context of viral infections: production, response and therapeutic implications. J Innate Immun 6:563–574
Jewell NA, Cline T, Mertz SE, Smirnov SV, Flano E, Schindler C, Grieves JL, Durbin RK, Kotenko SV, Durbin JE (2010) Lambda interferon is the predominant interferon induced by influenza A virus infection in vivo. J Virol 84:11515–11522
Sadler AJ, Williams BR (2008) Interferon-inducible antiviral effectors. Nat Rev Immunol 8:559–568
Iwai A, Shiozaki T, Miyazaki T (2013) Relevance of signaling molecules for apoptosis induction on influenza A virus replication. Biochem Biophys Res Commun 441:531–537
Baccam P, Beauchemin C, Macken CA, Hayden FG, Perelson AS (2006) Kinetics of influenza A virus infection in humans. J Virol 80:7590–7599
Smith AM, Perelson AS (2011) Influenza A virus infection kinetics: quantitative data and models. Wiley Interdiscip Rev Syst Biol Med 3:429–445
Kawada J, Kimura H, Kamachi Y, Nishikawa K, Taniguchi M, Nagaoka K, Kurahashi H, Kojima S, Morishima T (2006) Analysis of gene-expression profiles by oligonucleotide microarray in children with influenza. J Gen Virol 87:1677–1683
Wang J, Nikrad MP, Travanty EA, Zhou B, Phang T, Gao B, Alford T, Ito Y, Nahreini P, Hartshorn K et al (2012) Innate immune response of human alveolar macrophages during influenza A infection. PLoS One 7:e29879
Shapira SD, Gat-Viks I, Shum BO, Dricot A, de Grace MM, Wu L, Gupta PB, Hao T, Silver SJ, Root DE et al (2009) A physical and regulatory map of host-influenza interactions reveals pathways in H1N1 infection. Cell 139:1255–1267
Zhang YH, Wang S, Yan HP, Zhao DT, Zhang X, Li A, Wang ZK, Wu H, Zhao Y (2012) Prognostic values of chemokines in different clinical outcomes of influenza A (H1N1)-infected patients. Zhonghua yi xue za zhi 92:2020–2022
Sivan G, Martin SE, Myers TG, Buehler E, Szymczyk KH, Ormanoglu P, Moss B (2013) Human genome-wide RNAi screen reveals a role for nuclear pore proteins in poxvirus morphogenesis. Proc Natl Acad Sci U S A 110:3519–3524
Mehraj V, Textoris J, Ben Amara A, Ghigo E, Raoult D, Capo C, Mege JL (2013) Monocyte responses in the context of Q fever: from a static polarized model to a kinetic model of activation. J Infect Dis 208:942–951
Pauwels M, van Weyenbergh J, Soumillion A, Proost P, De Ley M (1994) Induction by zinc of specific metallothionein isoforms in human monocytes. Eur J Biochem 220:105–110
Schoggins JW, Wilson SJ, Panis M, Murphy MY, Jones CT, Bieniasz P, Rice CM (2011) A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472:481–485
Balasubramaniam VR, Wai TH, Omar AR, Othman I, Hassan SS (2012) Cellular transcripts of chicken brain tissues in response to H5N1 and Newcastle disease virus infection. Virol J 9:53
de Saint-Vis B, Vincent J, Vandenabeele S, Vanbervliet B, Pin JJ, Ait-Yahia S, Patel S, Mattei MG, Banchereau J, Zurawski S et al (1998) A novel lysosome-associated membrane glycoprotein, DC-LAMP, induced upon DC maturation, is transiently expressed in MHC class II compartment. Immunity 9:325–336
Zhou Z, Xue Q, Wan Y, Yang Y, Wang J, Hung T (2011) Lysosome-associated membrane glycoprotein 3 is involved in influenza A virus replication in human lung epithelial (A549) cells. Virol J 8:384
Zak DE, Andersen-Nissen E, Peterson ER, Sato A, Hamilton MK, Borgerding J, Krishnamurty AT, Chang JT, Adams DJ, Hensley TR et al (2012) Merck Ad5/HIV induces broad innate immune activation that predicts CD8(+) T-cell responses but is attenuated by preexisting Ad5 immunity. Proc Natl Acad Sci U S A 109:E3503–E3512
Acknowledgments
This work was supported by the Wellcome Trust (074318 [J.C.K.]), the European Research Council under the European Union’s Seventh Framework Programme (FP7/2007-2013)/ERC Grant agreement no. 281824 (J.C.K.) and the NIHR Oxford Biomedical Research Centre. We thank the High-Throughput Genomics Group at the Wellcome Trust Centre for Human Genetics (funded by Wellcome Trust grant 090532/Z/09/Z and MRC Hub grant G0900747 91070) for the generation of gene expression data. We are grateful to Helen Lockstone and Natasha Sahgal for assistance with data analysis. We thank Adaikalavan Ramasamy for the reading of the manuscript and helpful suggestions.
Author information
Authors and Affiliations
Corresponding author
Additional information
Emma E. Davenport and Richard D. Antrobus contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1255 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Davenport, E.E., Antrobus, R.D., Lillie, P.J. et al. Transcriptomic profiling facilitates classification of response to influenza challenge. J Mol Med 93, 105–114 (2015). https://doi.org/10.1007/s00109-014-1212-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-014-1212-8