Abstract
Purpose
Most studies evaluating artificial intelligence (AI) models that detect abnormalities in neuroimaging are either tested on unrepresentative patient cohorts or are insufficiently well-validated, leading to poor generalisability to real-world tasks. The aim was to determine the diagnostic test accuracy and summarise the evidence supporting the use of AI models performing first-line, high-volume neuroimaging tasks.
Methods
Medline, Embase, Cochrane library and Web of Science were searched until September 2021 for studies that temporally or externally validated AI capable of detecting abnormalities in first-line computed tomography (CT) or magnetic resonance (MR) neuroimaging. A bivariate random effects model was used for meta-analysis where appropriate. This study was registered on PROSPERO as CRD42021269563.
Results
Out of 42,870 records screened, and 5734 potentially eligible full texts, only 16 studies were eligible for inclusion. Included studies were not compromised by unrepresentative datasets or inadequate validation methodology. Direct comparison with radiologists was available in 4/16 studies and 15/16 had a high risk of bias. Meta-analysis was only suitable for intracranial hemorrhage detection in CT imaging (10/16 studies), where AI systems had a pooled sensitivity and specificity 0.90 (95% confidence interval [CI] 0.85–0.94) and 0.90 (95% CI 0.83–0.95), respectively. Other AI studies using CT and MRI detected target conditions other than hemorrhage (2/16), or multiple target conditions (4/16). Only 3/16 studies implemented AI in clinical pathways, either for pre-read triage or as post-read discrepancy identifiers.
Conclusion
The paucity of eligible studies reflects that most abnormality detection AI studies were not adequately validated in representative clinical cohorts. The few studies describing how abnormality detection AI could impact patients and clinicians did not explore the full ramifications of clinical implementation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the developed world, first-line imaging is performed in almost all hospitals, and refers to imaging performed at onset, for example, a head computed tomography (CT) for an unconscious patient in the emergency department, or a head magnetic resonance imaging (MRI) for a patient with headache. First-line imaging is a high-volume task and a range of pathologies can be encountered. We distinguish this from second-line imaging where detailed biomarkers are extracted, based on prior clinical and first-line imaging information. Typically, second-line imaging is only performed in specialist hospitals where examples include large vessel occlusion imaging for stratifying stroke patients for thrombectomy treatment, or perfusion imaging for characterising brain tumours [1]. In comparison to first-line imaging, second-line imaging is a low-volume task.
Radiology workloads for first-line imaging have soared in the last decade due to changing demographics, increased screening for early diagnosis initiatives, and updated clinical pathway guidelines requiring imaging. In the years leading up to the coronavirus disease 2019 (COVID-19) pandemic, the number of brain MRI scans performed in, for example, the United Kingdom (UK) increased on average by 7.8% annually, and the demand for both CT and MRI reporting outpaced the growth in the radiology workforce [2, 3]. Reporting backlogs are problems of national importance in the UK, and analogous scenarios are seen in healthcare systems globally. Diagnostic delays cause poorer short and long-term clinical outcomes, with the late detection of illness inflating healthcare costs [4].
The automated detection of abnormalities in a scan using artificial intelligence (AI) has the potential to improve radiologist efficiency. The AI can be used to reorder radiology worklists by flagging abnormal scans, as a reader aid or even as a second reader to identify missed pathology. However, the considerable interest in introducing AI into clinical environments to improve productivity in the high volume first-line imaging tasks, may be clouded by two main challenges in most published studies. Firstly, many abnormality detection AI studies report the diagnostic accuracy using non-representative clinical datasets (e.g. intracranial hemorrhage alone versus healthy controls without any other pathology) [5], including commercially available AI solutions [6]. Indeed, few studies validate their findings on datasets that are representative of the scans seen in routine clinical practice which contain a wide variety of pathologies. A second concern is that many studies do not demonstrate the generalisability of AI models due to inadequate validation methodology [7, 8]. By validating abnormality detection AI on a hold-out subset from the same patient dataset, known as internal validation, it is unclear whether reported AI performance would translate to different patient populations scanned at different institutions. A recent systematic review analysing the enormous number of recent studies where AI was used for the detection of COVID-19 using chest imaging, found that all 62 included studies had no potential clinical use due to methodological biases such as the use of unrepresentative datasets and insufficient validation [9].
The aim of this systematic review was to determine the diagnostic performance and summarise the evidence supporting the use of those AI models carrying out first-line neuroimaging tasks. Critically, we ensured that our analyses were only focused on those studies that were not compromised by unrepresentative datasets or inadequate validation methodology. Therefore, we analysed those AI models that might conceivably be ready for use in the clinic. The primary objective was to determine the diagnostic accuracy of these AI models. A secondary objective was to determine the impact of AI on downstream clinical outcomes in those studies where this had been investigated.
Methods
This systematic review was conducted in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses of Diagnostic Test Accuracy (PRISMA-DTA) statement [10]. The review protocol is registered on the international prospective register of systematic reviews (PROSPERO), CRD42021269563.
Data Sources and Searches
The full strategy is listed in Supplementary Material 1. Searches were conducted on MEDLINE, EMBASE, the Cochrane Library and Web of Science for studies published until September 2021. Bibliographies from eligible studies and systematic reviews were searched for additional relevant studies. Conference abstracts and pre-prints were excluded. A full description of data extraction is provided in Supplementary Material 2.
Index Test, Reference Standard and Target Condition
The target condition of the systematic review was the abnormality detected, for example intracranial hemorrhage. The AI model detecting the target condition was the index test. The radiological review was designated as the reference standard.
Inclusion Criteria
We included studies where an AI model could predict if a given CT or MRI examination was abnormal. Only studies that validated AI models on test datasets that were separated from the training data temporally or geographically were included. Test datasets were required to have normal scans, scans with the target condition and scans with one or more non-target conditions, in order to be representative of clinical practice.
Exclusion Criteria
The motivation for the study was to review abnormality detection in first-line, clinical neuroimaging. Studies that only reported the accuracy of the AI model to make voxel-wise (e.g. segmentation studies) or slice-wise predictions but did not subsequently report at the examination level were excluded (unless examination level accuracy could be calculated from the published study data). Studies using second-line imaging exclusively (e.g. angiography, perfusion studies) were excluded. Psychiatric conditions were excluded if structural differences have only been shown in group-wise comparison studies (e.g. schizophrenia, autism spectrum disorder); all conditions that had a structural correlate often seen at the individual level were included (e.g. Alzheimer’s disease). Studies testing exclusively on pediatric populations were excluded. Studies not published in a peer-reviewed journal or without an English language translation were excluded [11].
Data Analysis
We used the QUality Assessment of Diagnostic Accuracy Studies‑2 (QUADAS-2) tool [12], tailored to the review question incorporating items from the Checklist for Artificial Intelligence in Medical Imaging (CLAIM) [13]; modified signalling questions are presented in Supplementary Material 3. The unit of analysis was the patient undergoing a CT or MRI examination. The primary outcome was diagnostic test accuracy. Secondary outcomes assessed whether the AI model had been implemented in clinical practice (i.e. in a pathway that could affect clinical outcomes rather than reporting diagnostic test accuracy on a retrospective test dataset in a dry laboratory setting), and if so, the associated performance metrics.
To determine the primary outcome measures, where published, the 2 × 2 contingency tables and the principal diagnostic accuracy measures of sensitivity (recall) and specificity were extracted for test datasets. The area under the receiver operating characteristic curve (ROC-AUC) values, and positive predictive values (PPV or precision) were also extracted where published. Where 2 × 2 contingency tables were not provided, the tables were populated based on the published study data; the calculations are outlined in Supplementary Material 4.
The PPV is important for abnormality detection (Supplementary Material 4) and is more informative than specificity for imbalanced datasets particularly when the prevalence of the target condition is small [14]. The calculation of PPV is, however, dependent on the prevalence of the target condition in a test dataset, where PPV increases with increasing prevalence assuming constant sensitivity and specificity; the calculation is outlined in Supplementary Material 4 [15].
To directly compare AI model performance, the PPV for each model must be adjusted for a uniform prevalence. There were sufficient studies in one subgroup (intracranial hemorrhage detection in CT scans) for the calculation of prevalence-adjusted PPV. Here, we chose a prevalence of 10% based on recent evidence of routine clinical practice in the UK [16]. The prevalence-adjusted PPV we subsequently quote can be interpreted as the PPV that would be expected for each model if the prevalence of ICH within the test dataset was 10%.
Meta-analysis
Meta-analysis was performed when four or more studies evaluated a given target condition within a specific modality [17]. Studies investigating the detection of intracranial hemorrhage on CT scans were the only subgroup of sufficient number and homogeneity to allow inclusion for meta-analysis.
A bivariate random effects model was used for meta-analysis, taking into account the within and between study variance, and the correlation between sensitivity and specificity across studies [18]. Sensitivities and specificities were presented for each study using forest plots, and pooled estimates for both measures were calculated. To investigate the impact of variables of interest contributing to heterogeneity, metaregression was performed with the variable of interest as a covariate for the bivariate model. Using the existing model parameters, the absolute differences in pooled sensitivity and specificity between subgroups of interest were computed.
Parameters of the model also allowed the estimation of the summary ROC (SROC) curve and the summary ROC-AUC (SROC-AUC). Using a resampling approach [19], the model estimates were also used to derive the pooled measures of balanced accuracy as well as the positive and negative likelihood ratios and the diagnostic odds ratio.
The meta-analysis was conducted by a statistician (M.G.) with 15 years of relevant experience. All the statistical analyses were performed in R (v 3.6.1, R Foundation for Statistical Computing, Vienna, Austria). The R package mada (v 0.5.10) was used for the bivariate model [20]. As some of the 2 × 2 contingency table input cell values derived from the individual studies contained zeros, we applied a continuity correction (0.5).
Results
Characteristics of Included Studies
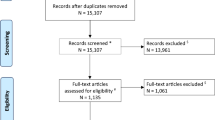
Database searches resulted in 42,870 unique results, of which 5734 potentially eligible full texts were assessed (Supplementary Fig. 1). Our criteria for clinically representative test datasets were not met in 1239 studies which were excluded. Additionally, we excluded 218 studies for using internal validation only. Only sixteen studies were of sufficient scientific rigour to be eligible for inclusion. The test datasets from the 16 studies comprised of 26,164 patients in total; however, the total number of patients in the training datasets could not be calculated as some commercial studies did not publish this data. Supervised convolutional neural networks (CNNs) to classify scans as normal or abnormal were used in 14/16 (88%) studies, with a variety of different model architectures and 5 studies (5/16, 31%) demonstrated the accuracy of commercially available AI models (from three AI vendors: Qure.ai (Mumbai, India); Aidoc (Tel Aviv, Israel); Avicenna.ai (La Ciotat, France) [21,22,23,24,25,26]). The largest subgroup of studies (11/16, 69%) employed CNNs to detect intracranial hemorrhage using CT [21,22,23,24,25, 27,28,29,30,31,32]. Other studies used CT and MRI to detect other single non-hemorrhage pathology (2/16, 13%) [21, 33], or multiple pathologies (4/16, 25%) [33,34,35,36]. The characteristics of each included study are summarised in Supplementary Material 5 with further AI model information shown in Table 1.
Assessment of Risk of Bias
The risk of bias evaluation for each study using the QUADAS‑2 tool is summarised in Fig. 1. A high risk of bias in at least one domain was shown in 15/16 (94%) of studies. The modified signalling questions used for assessing each study, and their explanations are in Supplementary Materials 3 and 6, respectively.
The following were the commonest sources of bias: eight studies (8/16, 50%) assessed AI model performance in laboratory conditions only (“analytical validation” [21, 25, 26, 28,29,30, 33, 35, 37]). In contrast, four studies (4/16, 25%) placed the AI model within the clinical pathway (“clinical validation”) [22,23,24, 32], which more closely resembles a “real world” environment and therefore the intended applicability. Seven studies (7/16, 44%) used temporal validation alone, and therefore had a high risk of bias for patient selection as there is limited assessment of generalisability [26, 30,31,32, 34,35,36], compared to 9/16 (56%) studies where AI models were externally validated on test data from other institutions [21,22,23,24,25, 27,28,29, 33]. Studies that used fewer than two radiologists to assess the images of a scan for their reference standard were considered at high risk of bias, as individual radiologists do not have perfect accuracy—five studies (5/16, 31%) were therefore considered to have high risk of bias as only the clinical report was reviewed [22, 27, 32, 33, 36]. One study had a high risk of bias as the reference standard was informed by the output of the AI model (the index test) [25]; this study was therefore excluded from the meta-analysis.
Analysis
The primary outcome for each study was diagnostic test accuracy, which is summarised in Table 2. Studies varied greatly in accuracy performance (sensitivity range: 0.70–1.00, specificity range: 0.51–1.00), but the highest accuracies were typically seen for intracranial hemorrhage detection using CT (Supplementary Fig. 2). Only two studies validated AI that used MRI; two AI models that detected any pathology using MRI had modest accuracies (sensitivity range: 0.78–1.00, specificity range: 0.65–0.80), compared to single pathology performance (Supplementary Fig. 3).
Diagnostic test accuracy of algorithms and comparators (single radiologists) in precision-recall space, for intracranial hemorrhage detection in CT imaging, when adjusted for an intracranial hemorrhage prevalence of 10%. The prevalence-adjusted positive predictive value (PPV) can be interpreted as the PPV you would expect for each model if the prevalence of ICH within the test dataset was 10%. An ideal classifier would be at the top-right corner at (1, 1). The size of each marker is proportional to the size of the test dataset. In studies where there was more than one operating point, we chose the operating point with the highest specificity. CQ500 = CQ500 external test dataset. Qure.ai, Aidoc and Avicenna.ai are commercial vendors for AI products
Diagnostic Test Accuracy of Radiologists Compared to AI
The performance of AI models against individual radiologists under laboratory conditions are available for four studies, summarised in Fig. 2. A full description of these studies can be found in Supplementary Material 7.
Clinical Implementation
All three (3/16, 19%) studies that also investigated clinical implementation performance, assessed the detection of intracranial hemorrhage using CT. Two (2/16, 13%) studies placed the AI model at the start of the clinical pathway before radiologist interpretation (pre-read triage) [22, 32], and 2/16 (13%) at the end after radiologist interpretation (post-read) [27, 32].
In the two studies where AI had been applied for pre-read triage, one showed a reduction in the time-to-report for non-urgent examinations which the AI flagged as abnormal, from a median time of 512 min to 19 min [32]. Radiologists in this study were unaware of the reprioritisation and were effectively blinded from the output of the AI. The other study was unblinded, as radiologists were made aware of examinations that the AI predicted to be abnormal [22]. This study demonstrated significant reductions in the mean time-to-report for flagged examinations for outpatients (674 to 70 min, p < 0.001), inpatients (390 to 352 min, p = 0.002), but not emergency cases (p = 0.37), or an undefined “other” class (p = 0.25) [22]. Importantly, neither study examined the extent and potential harms of delaying non-flagged studies, particularly AI false negatives which occurred in 26/347 (7.5%) [32] and 205/1760 (11.6%) [22] of outputs, respectively.
In two studies (2/16, 13%), AI had been applied as a second reader after radiologist interpretation and discrepancies between radiologists and AI were examined [27, 32]. AI was able to identify 4/347 (1.2%) and 2/5965 (0.03%) of intracranial hemorrhages that radiologists had missed (radiologist false negatives), respectively. If implemented, both studies estimated that the radiologist would be alerted that there was a discrepancy between them and the AI model in 10% (34/347) and 5% (313/5965) of cases, respectively, and 9 and 157 re-reviews would be required for 1 change in report, respectively [32]. In the second study, radiologist-positive and AI-negative discrepancies were also examined (59/5965, 1%) and found these were all AI false negatives—AI was unable to identify any radiologist overcalls (radiologist false positives). If AI were to be implemented to identify overcalls (radiologist false positives) as well as misses (radiologist false negatives), the radiologist would be alerted in 6% (372/5965) of cases and 186 re-reviews would be required for 1 change in a report [27].
Meta-analysis
Six (6/16, 38%) studies were unsuitable for meta-analysis. We excluded five studies (5/16, 31%) with heterogeneous modality or target conditions (one study detected hyperdense, intracranial hemorrhage only as might typically be seen in an acute setting, and did not consider isodense or hypodense intracranial hemorrhage as might typically be seen in a subacute or chronic setting) [26, 33,34,35,36]. One study (1/16, 6%) was excluded due to the fundamental methodological flaw of having a circular reference standard as described above [25]. The remaining subgroup of studies were those detecting intracranial hemorrhage using CT and applying CNNs, and consisted of 10 studies (10/16, 63%) [21,22,23,24, 27,28,29,30,31,32]; we included these studies for meta-analysis. Forest plots of sensitivity and specificity (Fig. 3) graphically showed a high level of heterogeneity. There was significant heterogeneity observed in both sensitivity and specificity; the χ2-test p-values were both < 0.001 and I2 statistics were 85.4% and 99.3%, respectively. The pooled sensitivity for intracranial hemorrhage detection in CT = 0.901 (95% confidence interval [CI] 0.853–0.935), and the pooled specificity = 0.903 (95% CI 0.826–0.948). The derived pooled measures of balanced accuracy = 0.931 (95% CI 0.889–0.957); positive likelihood ratio = 26.7 (95% CI 15.8–42.3); negative likelihood ratio = 0.106 (95% CI 0.0471–0.199); and diagnostic odds ratio = 280.0 (95% CI 128.0–533.0).
Heterogeneity was investigated using metaregression, which compared the pooled sensitivities and specificities of two subsets of the studies: three studies where different AI models were applied on the same test dataset (CQ500) [21, 28, 29], and three studies where the same AI model (Aidoc) was applied on different test datasets [22,23,24]. Using the Aidoc subset as a baseline, the CQ500 subset had higher pooled sensitivity (p = 0.008) and lower pooled specificity (p = 0.004), implying that AI model type and patient make-up in the test dataset contributed to the heterogeneity observed.
Individual study ROC point estimates resulted in a summary ROC (SROC) curve (Fig. 4), for which the summary ROC-AUC = 0.948.
Discussion
Summary
This study aimed to determine the diagnostic accuracy of AI systems used to identify abnormalities in first-line neuroimaging tasks. Any productivity gains in such tasks are important as they are high volume and performed in almost all hospitals. To ensure our analyses were only focused on those studies that were not compromised by unrepresentative datasets or inadequate validation methodology, we excluded many studies (1239) that did not validate the AI model using datasets from representative clinical cohorts and many (218) for validating without temporal or external validation. Only sixteen studies were of sufficient rigour to be eligible for inclusion; however, even for these studies the overall methodological quality remained low with a high risk of bias in 94% of studies. Furthermore, most included studies were retrospective, with only four studies validating their AI models in clinical environments prospectively in real time (i.e. clinical validation).
For CT imaging studies, a subgroup of 10 AI models used to detect intracranial hemorrhage using CNNs, had a pooled sensitivity and specificity of 0.90, with a summary ROC-AUC of 0.95. Metaregression suggested that differences in the AI model development and patient selection contributed to the significant heterogeneity observed in both pooled measures (sensitivity I2 = 85.4%, specificity I2 = 99.3%). Four CT imaging studies allowed direct comparison between AI models with radiologists under laboratory conditions—further discussion is provided in Supplementary Material 7.
For MRI, only two studies were included. Both studies validated AI models that detect all pathologies. Together with a third study that used CT, a limitation of the three AI models that detect all pathologies is that findings seen in healthy ageing such as small vessel disease and age-commensurate atrophy are considered abnormal—this is reflected in the high prevalence of what was assigned as pathological in their test datasets (64–81%) [33,34,35]. AI that overcalls all older patients as abnormal raises concerns for applicability in clinical practice.
There were only three clinical implementation studies where AI was placed within the clinical pathway, as pre-read triage and for post-read discrepancy identification [22, 27, 32]. No study demonstrated a downstream clinical or health economic benefit.
Strengths and Limitations
A strength of this study was that the search strategy was sensitive [38, 39]. This allowed the identification of a wide range of studies included in this review, many of which were missing in other systematic reviews for the general use of AI in neuroimaging [40,41,42,43]. We also included all AI methods, not just those limited to deep learning. Whilst broad inclusion is a study strength, it is also conceivable that summary performance accuracy might be diminished by the inclusion of older AI models; however, older AI models barely contributed to our results as 88% of all eligible studies, and 100% of studies in the meta-analysis subgroup used CNNs.
Another strength, unique to this study, was that the inclusion criteria were designed to only include studies where outcome metrics would have a reasonable chance of generalising to first-line neuroimaging in routine clinical practice. Therefore, the diagnostic test accuracies presented here are plausibly more generalisable than if less stringent inclusion criteria were used. Specifically, we first excluded studies that did not validate AI on temporally distinct or external test datasets. Second, we excluded studies that did not test on representative patient cohorts (which as a minimum standard required normal brains, the target condition and at least one non-target condition). As a result, we excluded studies that validated AI models on test datasets that contained the target condition and healthy controls only, which does not reflect the “real world”; we note that almost all ischemic stroke detection studies were therefore excluded [44].
A limitation is that meta-analysis was only suitable for one subgroup where there were sufficient homogeneous studies using the same imaging modality, target condition and AI model type. Another limitation is that no formal assessment of publication bias was undertaken; however, it is unlikely that our overall conclusions would change if studies with poorer AI model performances had been published.
Strategies for Implementing AI into Clinical Pathways
The standalone diagnostic accuracy of AI to detect abnormalities has been demonstrated to be high, particularly for intracranial hemorrhage in CT imaging. There was insufficient evidence, however, to suggest where such AI would be most useful in the clinical pathway. This included those being marketed commercially (Supplementary Material 8).
Both studies that investigated intracranial hemorrhage AI detectors for pre-read CT worklist triage found that the greatest reduction in reporting time was for outpatient examinations when compared to emergency or inpatient examinations [22, 32]. There was insufficient evidence, however, from these and any other studies regarding the downstream clinical benefit and cost-effectiveness of AI implementation. AI models with poor sensitivity in a pre-read triage setting would systemically increase the time to report AI false negative examinations as these would be put at the back of the reporting queue; it was unclear from both studies whether AI false negatives were significantly delayed and the extent of harm, if any, that was associated with this delay. For pre-read triage, it is also unknown whether knowing that AI puts flagged examinations to the front of the queue could have long-term consequences on radiologist performance. There is a similar question regarding AI intended as a second reader which may unintentionally affect the behaviour of radiologists; for example, the implementation of automated computer-assisted diagnosis (CAD) tools in mammography, to be used as a reader aid during radiologist interpretation, has previously been shown to reduce radiologist sensitivity [45] and overall accuracy [46].
One advantage of a post-read implementation is that radiologists are initially blinded to the AI decision. In a post-read setting, AI models could be used to flag discrepancies to determine potential radiologist “misses” or “overcalls” and allow a re-review. An AI model with poor specificity or PPV in this setting would create a high burden on radiologist time with a large number of false positive scans to re-review. In the two studies that investigated discrepancies, there appeared to be low additive diagnostic yield associated with a high rate of re-review. Therefore, further studies will be necessary to understand the cost-effectiveness of such post-read strategies.
Many AI models developed for high-sensitivity pre-read triage could plausibly be repurposed as high-PPV post-read discrepancy identifiers and vice versa simply by adjusting the operating threshold.; however, for any specific downstream clinical task it is necessary to further validate any predictive model following such recalibration.
Conclusion
We have analysed the evidence and presented the diagnostic performance of the current state-of-the-art AI detection models that can be applied to first-line neuroimaging. Such tasks are important as they are high volume and performed in almost all hospitals and offer considerable potential for the necessary productivity gains required in the twenty-first century. If the intended use of AI detection models is as a tool to improve radiologist efficiency rather than a replacement for radiologists, AI may be clinically useful even if the accuracy shown in our meta-analysis remains lower than that of radiologists for the task of intracranial hemorrhage detection; however, at present, there is insufficient evidence to recommend implementation of AI for abnormality detection, including hemorrhage detection, into any part of the clinical pathway. Importantly, the clinical and health economic benefits are currently unproven. For now, future research efforts should aim to minimise bias and demonstrate analytical validation through well-designed studies using clinically representative external test datasets which can unequivocally prove high performance accuracy and good generalisability. Following this, clinical trials will be required to confirm the performance findings in the “real world” and determine whether the clinical benefits of implementing AI in the clinical pathway outweigh the potential harm to patients. In addition to clinical validation, such trials could include health economic analyses to determine the costs incurred and benefits obtained within the wider healthcare system.
References
Booth TC, Luis A, Brazil L, Thompson G, Daniel RA, Shuaib H, et al. Glioblastoma post-operative imaging in neuro-oncology: current UK practice (GIN CUP study). Eur Radiol. 2021;31:2933–43.
Dixon S. Diagnostic imaging dataset annual statistical release 2020/21. 2021. https://www.england.nhs.uk/statistics/statistical-work-areas/diagnostic-imaging-dataset/diagnostic-imaging-dataset-2020-21-data/. Accessed 20 Mar 2023.
The Royal College of Radiologists London. Clinical radiology UK workforce census 2020 report. 2020. https://www.rcr.ac.uk/system/files/publication/field_publication_files/clinical-radiology-uk-workforce-census-2020-report.pdf. Accessed 20 Mar 2023.
World Health Organization. Cancer control: early detection. WHO Guide for effective programmes. 2007. http://apps.who.int/iris/bitstream/10665/43743/1/9241547338_eng. Accessed 15 Feb 2022.
Lee JY, Kim JS, Kim TY, Kim YS. Detection and classification of intracranial haemorrhage on CT images using a novel deep-learning algorithm. Sci Rep. 2020;10:1–7.
Rava RA, Seymour SE, LaQue ME, Peterson BA, Snyder KV, Mokin M, et al. Assessment of an artificial intelligence algorithm for detection of intracranial hemorrhage. World Neurosurg. 2021;150:e209–e17.
Titano JJ, Badgeley M, Schefflein J, Pain M, Su A, Cai M, et al. Automated deep-neural-network surveillance of cranial images for acute neurologic events. Nat Med. 2018;24:1337–41.
Hooper SM, Dunnmon JA, Lungren MP, Mastrodicasa D, Rubin DL, Ré C, et al. Impact of upstream medical image processing on downstream performanceof a head CT triage neural network. Radiol Artif Intell. 2021;3:200229.
Roberts M, Driggs D, Thorpe M, Gilbey J, Yeung M, Ursprung S, et al. Common pitfalls and recommendations for using machine learning to detect and prognosticate for COVID-19 using chest radiographs and CT scans. Nat Mach Intell. 2021;3:199–217.
McInnes MDF, Moher D, Thombs BD, McGrath TA, Bossuyt PM, Clifford T, et al. Preferred reporting items for a systematic review and meta-analysis of diagnostic test accuracy studies. JAMA. 2018;319:388–96.
Nussbaumer-Streit B, Klerings I, Dobrescu AI, Persad E, Stevens A, Garritty C, et al. Excluding non-English publications from evidence-syntheses did not change conclusions: a meta-epidemiological study. J Clin Epidemiol. 2020;118:42–54.
Whiting PF, Rutjes AWS, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, et al. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011;155:529–36.
Mongan J, Moy L, Charles E, Kahn J. Checklist for artificial intelligence in medical imaging (CLAIM): a guide for authors and reviewers. Radiol Artif Intell. 2020;2:e200029.
Abu-Akel A, Bousman C, Skafidas E, Pantelis C. Mind the prevalence rate: overestimating the clinical utility of psychiatric diagnostic classifiers. Psychol Med. 2018;48:1225–7.
Tenny S, Hoffman M. Prevalence. 2017. https://www.ncbi.nlm.nih.gov/books/NBK430685/. Accessed 20 Mar 2023.
Hocking KC, Wright CR, Alhun U, Hughes F, Balian VJ, Kabuli MAK, et al. Acute haemorrhage rate in 28,000 out-of-hours CT heads. Br J Radiol. 2022;94:20210580.
Ebrahimzadeh S, Islam N, Dawit H, Salameh J‑P, Kazi S, Fabiano N, et al. Thoracic imaging tests for the diagnosis of COVID-19. Cochrane Database Syst Rev. 2022. https://doi.org/10.1002/14651858.CD013639.pub5.
Reitsma JB, Glas AS, Rutjes AWS, Scholten RJPM, Bossuyt PM, Zwinderman AH. Bivariate analysis of sensitivity and specificity produces informative summary measures in diagnostic reviews. J Clin Epidemiol. 2005;58:982–90.
Zwinderman AH, Bossuyt PM. We should not pool diagnostic likelihood ratios in systematic reviews. Stat Med. 2008;27:687–97. https://doi.org/10.1002/sim.2992.
Doebler P. Mada: meta-analysis of diagnostic accuracy. 2015. http://www.cran.r-project.org/packages/mada. Accessed 20 Mar 2023.
Chilamkurthy S, Ghosh R, Tanamala S, Biviji M, Campeau NG, Venugopal VK, et al. Deep learning algorithms for detection of critical findings in head CT scans: a retrospective study. Lancet. 2018;392:2388–96.
Ginat D. Implementation of machine learning software on the radiology worklist decreases scan view delay for the detection of intracranial hemorrhage on CT. Brain Sci. 2021;11:832.
Ginat DT. Analysis of head CT scans flagged by deep learning software for acute intracranial hemorrhage. Neuroradiology. 2020;62:335–40.
Buls N, Watté N, Nieboer K, Ilsen B, de Mey J. Performance of an artificial intelligence tool with real-time clinical workflow integration—Detection of intracranial hemorrhage and pulmonary embolism. Phys Medica. 2021;83:154–60.
Voter AF, Meram E, Garrett JW, John-Paul JY. Diagnostic accuracy and failure mode analysis of a deep learning algorithm for the detection of intracranial hemorrhage. J Am Coll Radiol. 2021;18:1143–52.
McLouth J, Elstrott S, Chaibi Y, Quenet S, Chang PD, Chow DS, et al. Validation of a deep learning tool in the detection of intracranial hemorrhage and large vessel occlusion. Front Neurol. 2021;12:655.
Salehinejad H, Kitamura J, Ditkofsky N, Lin A, Bharatha A, Suthiphosuwan S, et al. A real-world demonstration of machine learning generalizability in the detection of intracranial hemorrhage on head computerized tomography. Sci Rep. 2021;11:1–11.
Monteiro M, Newcombe VFJ, Mathieu F, Adatia K, Kamnitsas K, Ferrante E, et al. Multiclass semantic segmentation and quantification of traumatic brain injury lesions on head CT using deep learning: an algorithm development and multicentre validation study. Lancet Digit Health. 2020;2:e314–e22.
Wang X, Shen T, Yang S, Lan J, Xu Y, Wang M, et al. A deep learning algorithm for automatic detection and classification of acute intracranial hemorrhages in head CT scans. Neuroimage Clin. 2021;32:102785.
Kuo W, Häne C, Mukherjee P, Malik J, Yuh EL. Expert-level detection of acute intracranial hemorrhage on head computed tomography using deep learning. Natl Acad Sci. 2019;116:22737–45.
Chang PD, Kuoy E, Grinband J, Weinberg BD, Thompson M, Homo R, et al. Hybrid 3D/2D convolutional neural network for hemorrhage evaluation on head CT. Am J Neuroradiol. 2018;39:1609–16.
Arbabshirani MR, Fornwalt BK, Mongelluzzo GJ, Suever JD, Geise BD, Patel AA, et al. Advanced machine learning in action: identification of intracranial hemorrhage on computed tomography scans of the head with clinical workflow integration. NPJ Digit Med. 2018;1:1–7.
Nael K, Gibson E, Yang C, Ceccaldi P, Yoo Y, Das J, et al. Automated detection of critical findings in multi-parametric brain MRI using a system of 3D neural networks. Sci Rep. 2021;11:1–10.
Finck T, Schinz D, Grundl L, Eisawy R, Yigitsoy M, Moosbauer J, et al. Automated pathology detection and patient triage in routinely acquired head computed tomography scans. Invest Radiol. 2021;56:571–8.
Gauriau R, Bizzo BC, Kitamura FC, Landi Junior O, Ferraciolli SF, Macruz FBC, et al. A deep learning—based model for detecting abnormalities on brain MR images for triaging: preliminary results from a multisite experience. Radiol Artif Intell. 2021;3:e200184.
Prevedello LM, Erdal BS, Ryu JL, Little KJ, Demirer M, Qian S, et al. Automated critical test findings identification and online notification system using artificial intelligence in imaging. Radiology. 2017;285:923–31.
FDA-NIH Biomarker Working Group. BEST (Biomarkers, EndpointS, and other Tools) resource. Silver Spring (MD), Bethesda (MD): Food and Drug Administration (US); National Institues for Health (US). 2016. https://www.ncbi.nlm.nih.gov/books/NBK326791/. Accessed 18 June 2022.
McKenzie J, Brennan S, Ryan R, Thomson H, Johnston R, Thomas J. Chapter 3: Defining the criteria for including studies and how they will be grouped for the synthesis. Cochrane Handb Syst Rev Interv version 63 (updated Febr 2022). 2022. https://training.cochrane.org/handbook/current/chapter-03. Accessed 31 July 2022.
Lefebvre C, Glanville J, Briscoe S, Featherstone R, Littlewood A, Marshall C, et al. Chapter 4: Searching for and selecting studies. Cochrane Handb Syst Rev Interv version 63 (updated Febr 2022). 2022. https://training.cochrane.org/handbook/current/chapter-04. Accessed 31 July 2022.
Yao AD, Cheng DL, Pan I, Kitamura F. Deep learning in neuroradiology: A systematic review of current algorithms and approaches for the new wave of imaging technology. Radiol Artif Intell. 2020; https://doi.org/10.1148/ryai.2020190026.
Aggarwal R, Sounderajah V, Martin G, Ting DSW, Karthikesalingam A, King D, et al. Diagnostic accuracy of deep learning in medical imaging: a systematic review and meta-analysis. npj Digit Med. 2021;4:1–23.
Liu X, Faes L, Kale AU, Wagner SK, Fu DJ, Bruynseels A, et al. A comparison of deep learning performance against health-care professionals in detecting diseases from medical imaging: a systematic review and meta-analysis. Lancet Digit Health. 2019;1:e271–e97.
Nagendran M, Chen Y, Lovejoy CA, Gordon AC, Komorowski M, Harvey H, et al. Artificial intelligence versus clinicians: systematic review of design, reporting standards, and claims of deep learning studies. BMJ. 2020; https://doi.org/10.1136/bmj.m689.
Murray NM, Unberath M, Hager GD, Hui FK. Artificial intelligence to diagnose ischemic stroke and identify large vessel occlusions: a systematic review. J Neurointerv Surg. 2020;12:156–64.
Lehman CD, Wellman RD, Buist DSM, Kerlikowske K, Tosteson ANA, Miglioretti DL. Diagnostic accuracy of digital screening mammography with and without computer-aided detection. JAMA Intern Med. 2015;175:1828–37.
Fenton JJ, Taplin SH, Carney PA, Abraham L, Sickles EA, D’Orsi C, et al. Influence of computer-aided detection on performance of screening mammography. N Engl J Med. 2007;356:1399–409. https://doi.org/10.1056/NEJMoa066099.
Acknowledgements
SA is supported by an Engineering and Physical Sciences Research Council (EPSRC) funded PhD studentship (EP/R513064/1). This research was also supported by the Wellcome/EPSRC Centre for Medical Engineering (WT 203148/Z/16/Z) (TB, MG, MM) which includes open access fees, The Royal College of Radiologists (TB) and King’s College Hospital Research and Innovation (TB).
Author information
Authors and Affiliations
Contributions
SA, DW, JC, MM and TCB conceived the study. SA, DW, and CS executed the search and extracted data. MG conceived and performed the meta-analysis and meta-regression. SA and TCB wrote the paper. All authors contributed to revisions of the manuscript and approved the final version. TCB is the study guarantor. The corresponding author attests that all listed authors meet authorship criteria and that no others meeting the criteria have been omitted.
Corresponding authors
Ethics declarations
Conflict of interest
S. Agarwal, D. Wood, M. Grzeda, C. Suresh, M. Din, J. Cole, M. Modat and T.C. Booth declare that they have no competing interests.
Additional information
Data Availability
Raw data are available on request from the corresponding author. The lead author and manuscript’s guarantor (TCB) affirm that the manuscript is an honest, accurate, and transparent account of the study being reported; that no important aspects of the study have been omitted; and that any discrepancies from the study as planned (and, if relevant, registered) have been explained.
Supplementary Information
62_2023_1291_MOESM1_ESM.docx
Supplementary Material: A systematic review of artificial intelligence for abnormality detection in high volume neuroimaging and sub-group meta-analysis for intracranial hemorrhage detection
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Agarwal, S., Wood, D., Grzeda, M. et al. Systematic Review of Artificial Intelligence for Abnormality Detection in High-volume Neuroimaging and Subgroup Meta-analysis for Intracranial Hemorrhage Detection. Clin Neuroradiol 33, 943–956 (2023). https://doi.org/10.1007/s00062-023-01291-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00062-023-01291-1