Abstract
Sulfo-click is a chemoselective and biocompatible reaction between thioacids and sulfonyl azides that forms highly versatile N-acylsulfonamides, interesting bioisosteres of carboxylic acids. This reaction is useful for chemists and biologists and has many applications in medicinal chemistry, drug discovery, bioconjugation chemistry, and chemical biology. Sulfo-click amidations have been extensively used in kinetic target-guided synthesis (KTGS) to identify modulators of protein-protein interactions (PPIs). Different variants of KTGS screening, such as binary and multi-fragments, as well as one-pot deprotection/amidation strategies, have been successfully performed using sulfo-click chemistry. In this mini-review, we discuss the recent developments of sulfo-click amidation in KTGS and provide directions for future research.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The unique advantages of biocompatible click reactions that combine bio-orthogonality, efficiency, and chemo-selectivity have provided important developments in chemical biology and drug discovery. The sulfo-click amidation reaction [1,2,3,4] is an emergent biocompatible metal-free click chemistry reaction to yield N-acylsulfonamides, interesting bioisosteres of carboxylic acids. N-acylsulfonamides are a highly versatile class of compounds known for many decades and have recently regained new interest in medicinal chemistry and chemical biology.
The sulfo-click reaction with electron-deficient azides is believed to proceed stepwise by the nucleophilic addition of thioacids to give the intermediate I, followed by cyclization to the intermediate II, which undergoes a retro [3 + 2] cycloaddition leading to N-acylsulfonamide with elemental sulfur and dinitrogen as the sole byproducts (Scheme 1) [1, 2]. This versatile reaction has numerous applications in chemical biology and drug discovery. The sulfo-click amidation has been reported in the site-specific functionalization of peptides/proteins [5,6,7,8], fluorophores, and metal chelators [5]. It has also been used to detect thioacids in the bacterial proteome [9] and in the kinetic target-guided synthesis (KTGS) [10].
KTGS [10,11,12,13,14] is a promising tool for discovering biologically active compounds. KTGS brings the biological target to the forefront of the fragment-based lead discovery process. It is an unconventional discovery approach wherein the biological target is directly involved in assembling its own multidentate ligand. In KTGS, the biological target accelerates the reaction between complementary reactive fragments by bringing them in close proximity and proper orientation, thus allowing selectivity for some products over others (Fig. 1). KTGS has been successfully tested in various targets in vitro, from enzymes to RNA or protein-protein interaction sites to phosphate sites [10, 14].
Discussion
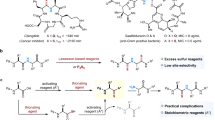
The chemical reaction employed by KTGS is key to its remarkable success. The sulfo-click amidation has been vital in developing KTGS for targeting protein-protein interactions (PPIs). Many biological processes rely on the interaction between different proteins, and therefore, PPIs represent a crucial and extensive category of potential targets for developing novel therapies [15, 16]. The first instance of KTGS being used to target PPIs was reported by Manetsch’s group [17], which identified PPI modulators through the sulfo-click amidation. A range of thioacids and sulfonyl azides were synthesized and employed in the KTGS strategy to identify potent inhibitors of B cell lymphoma-extra large (Bcl-xL). Bcl-xL is a crucial member of the Bcl-2 family that plays a critical role in regulating the intrinsic pathway of apoptosis. In this study, thioacids and sulfonyl azides were incubated as binary mixtures (18 different incubations, each incubation containing one thioacid and one sulfonyl azide) with and without Bcl-xL, and the formation of acylsulfonamide products was analyzed by LC-MS/MS. Only one of the 18 possible combinations displayed the desired Bcl-xL-templated effect (Scheme 2) [17]. The hit compound SZ4TA2 was then synthesized chemically and tested for its ability to inhibit the interaction between Bcl-xL and Bak using a fluorescence polarization (FP) competition assay. The hit compound SZ4TA2 displayed a remarkable ability to inhibit the interaction of Bcl-xL and Bak with an IC50 of 78.8 nM.
The study on the Bcl-xL-templated sulfo-click amidation was extended further using a more extensive fragment library to identify novel PPI inhibitors via the binary fragment KTGS strategy. Four of 81 possible combinations showed the templated effect, identifying three new hits, namely SZ7TA2, SZ9TA1, and SZ9TA5, and the previously identified hit SZ4TA2 (Scheme 3) [18]. All four hits were found to exhibit PPI modulatory activity when analyzed for their ability to modulate the PPI using a fluorescence-based competitive binding assay.
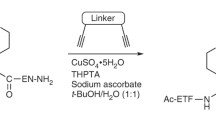
Although the sulfo-click amidation has been deemed reliable in various applications, it has not been widely used due to limitations regarding the preparation and handling of thioacids. Manetsch’s group successfully employed a one-pot deprotection/sulfo-click amidation approach in KTGS, overcoming this challenge. In this deprotection/sulfo-click amidation approach, 9-fluorenylmethyl (Fm) thioesters were quickly deprotected into thioacids using 5% piperidine in DMF and then reacted with sulfonylazides to produce N-acylsulfonamides (Scheme 4) [19]. The method has proven to be highly effective and has the potential to revolutionize the use of sulfo-click amidation in various applications.
While the binary KTGS approach using sulfo-click amidation has yielded promising results, the method requires LC-MS analysis of a large number of incubations, limiting the throughput of the KTGS screening platforms. As a result, the multi-fragment KTGS approach has been developed to improve the efficiency of KTGS in accessing a more extensive chemical space. The multi-fragment KTGS approach holds the potential to increase the screening platform’s throughput dramatically. The developed multi-fragment KTGS approach allowed the Manetsch laboratory to screen 1710 possible fragment combinations in 18 wells of a 96-well plate (190 fragment combinations in a single well; nine wells with the protein target and another nine wells without the protein target), utilizing an 83-member fragment library (Fig. 2) with and without myeloid cell leukemia-1 (Mcl-1) and analyzed by LC-MS/MS. Mcl-1, a protein belonging to the Bcl-2 family, is known for its anti-apoptotic properties. Its role in cancer cell survival and resistance to chemotherapy has made it a promising target for cancer therapy. Among the 51 total hits (Scheme 5), our findings identified 24 Mcl-1 inhibitors with single-digit micromolar IC50 values (Table 1) [20], as confirmed by fluorescence polarization (FP) studies. These results demonstrate that the multi-fragment sulfo-click KTGS approach is highly effective at developing hit compounds targeting Mcl-1. This finding is a breakthrough in KTGS screening as it reports the largest number of unique combinations per well ever recorded, opening up new possibilities for research in the fragment-based lead discovery field.
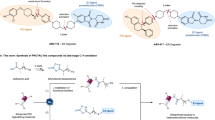
Next, the above-developed multi-fragment KTGS screening method was employed to identify inhibitors of pathogenic free-living amoebae (pFLA) glucokinase (Glck) enzymes. pFLA is notorious for inducing severe and potentially life-threatening infections of the central nervous system. Therefore, it is essential to investigate and explore novel strategies that can effectively prevent and treat these lethal infections. From an in-house library of 83 fragments (Fig. 2), 157 hit compounds were identified via the multi-fragment KTGS approach (Scheme 6). Among the total 157 hits, twelve inhibitors were identified against three pFLA Glck enzymes - Naegleria fowleri (Nf) Glck, Balamuthia mandrillaris (Bm) Glck, and Acanthamoeba castellanii (Ac) Glck (Table 2) [21]. The fragments were not selected based on predetermined binding potentials, indicating that KTGS, in combination with randomly designed fragments, can effectively screen chemical space and discover new compounds with medicinal chemistry potential. This demonstrates the usefulness of the multi-fragment KTGS screening in drug discovery research.
Conclusion
The sulfo-click amidation is a widely utilized biocompatible reaction that plays a crucial role in identifying protein-protein interaction modulators through KTGS. This reaction has proven highly effective in a binary fragment or multi-fragment screening format. A one-pot deprotection/amidation sulfo-click approach has been introduced and found applicable to KTGS addressing the thioacid’s handling and storage concerns. Notably, the use of the sulfo-click amidation method enabled the highest number of fragment combinations per well in any KTGS screening conducted thus far, thereby positioning the sulfo-click KTGS as a promising approach in fragment-based lead discovery.
Future prospective
Although there have been advancements in using sulfo-click amidation to identify PPI modulators via KTGS, there is still ample room for further development. Currently, only electron-deficient sulfonyl azides have been explored in binary and multi-fragment KTGS screening, which limits the available chemical space as a vast range of other electron-deficient azides, such as acyl/phosphoryl azides and electron-rich alkyl/aryl azides, remain unexplored. Additionally, optimizing the reaction conditions for performing it at 4 °C, the temperature at which most biological targets remain stable, could expand the usefulness of KTGS. Furthermore, the application of KTGS in cellular and in vivo using sulfo-click amidation has not yet been reported. We eagerly anticipate more advancements in sulfo-click amidation in KTGS in the near future.
Abbreviations
- KTGS:
-
Kinetic target-guided synthesis
- PPIs:
-
Protein-protein interactions
- RNA:
-
Ribonucleic acid
- Bcl-xL:
-
B cell lymphoma-extra large
- LC-MS/MS:
-
Liquid chromatography-tandem mass spectrometry
- FP:
-
Fluorescence polarization
- Fm:
-
9-Fluorenylmethyl
- DMF:
-
N, N-Dimethylformamide
- Mcl-1:
-
Myeloid cell leukemia-1
- pFLA:
-
Pathogenic free-living amoebae
- Glck:
-
Glucokinase
- Nf:
-
Naegleria fowleri
- Bm:
-
Balamuthia mandrillaris
- Ac:
-
Acanthamoeba castellanii
- Hs:
-
Homo sapiens
References
Shangguan N, Katukojvala S, Greenberg R, Williams LJ. The Reaction of Thio Acids with Azides: A New Mechanism and New Synthetic Applications. J Am Chem Soc. 2003;125:7754–5. https://doi.org/10.1021/ja0294919.
Kolakowski RV, Shangguan N, Sauers RR, Williams LJ. Mechanism of Thio Acid/Azide Amidation. J Am Chem Soc. 2006;128:5695–702. https://doi.org/10.1021/ja057533y.
Xie S, Fukumoto R, Ramström O, Yan M. Anilide Formation from Thioacids and Perfluoroaryl Azides. J Org Chem. 2015;80:4392–7. https://doi.org/10.1021/acs.joc.5b00240.
Clavé G, Dursun E, Vasseur JJ, Smietana M. An Entry of the Chemoselective Sulfo-Click Reaction into the Sphere of Nucleic Acids. Org Lett. 2020;22:1914–8. https://doi.org/10.1021/acs.orglett.0c00265.
Rijkers DTS, Merkx R, Yim CB, Brouwer AJ, Liskamp RMJ. ‘Sulfo-click’ for ligation as well as for site-specific conjugation with peptides, fluorophores, and metal chelators. J Pep Sci. 2010;16:1–5. https://doi.org/10.1002/psc.1197.
Yim CB, Dijkgraaf I, Merkx R, Versluis C, Eek A, Mulder GE, et al. Synthesis of DOTA-Conjugated Multimeric [Tyr3]Octreotide Peptides via a Combination of Cu(I)-Catalyzed “Click” Cycloaddition and Thio Acid/Sulfonyl Azide “Sulfo-Click” Amidation and Their in Vivo Evaluation. J Med Chem. 2010;53:3944–53. https://doi.org/10.1021/jm100246m.
Yim CB, Wildt B, Dijkgraaf I, Joosten L, Eek A, Versluis C, et al. Spacer Effects on in vivo Properties of DOTA-Conjugated Dimeric [Tyr3]Octreotate Peptides Synthesized by a “CuI-Click” and “Sulfo-Click” Ligation Method. ChemBioChem. 2011;12:750–60. https://doi.org/10.1002/cbic.201000639.
Zhang X, Li F, Lu XW, Liu CF. Protein C-Terminal Modification through Thioacid/Azide Amidation. Bioconjugate Chem. 2009;20:197–200. https://doi.org/10.1021/bc800488n.
Krishnamoorthy K, Begley TP. Reagent for the Detection of Protein Thiocarboxylates in the Bacterial Proteome: Lissamine Rhodamine B Sulfonyl Azide. J Am Chem Soc. 2010;132:11608–12. https://doi.org/10.1021/ja1034107.
Parvatkar PT, Wagner A, Manetsch R. Biocompatible reactions: advances in kinetic target-guided synthesis. Trends Chem. 2023;5:657–71. https://doi.org/10.1016/j.trechm.2023.06.002.
Hu X, Manetsch R. Kinetic target-guided synthesis. Chem Soc Rev. 2010;39:1316–24. https://doi.org/10.1039/B904092G.
Lossouarn A, Renard PY, Sabot C. Tailored Bioorthogonal and Bioconjugate Chemistry: A Source of Inspiration for Developing KineticTarget-Guided Synthesis Strategies. Bioconjugate Chem. 2021;32:63–72. https://doi.org/10.1021/acs.bioconjchem.0c00568.
Bosc D, Camberlein V, Gealageas R, Castillo-Aguilera O, Deprez B, Deprez-Poulain R. Kinetic Target-Guided Synthesis: Reaching the Age of Maturity. J Med Chem. 2020;63:3817–33. https://doi.org/10.1021/acs.jmedchem.9b01183.
Bosc D, Jakhlal J, Deprez B, Deprez-Poulain R. Kinetic target-guided synthesis in drug discovery and chemical biology: a comprehensive facts and figures survey. Future Med Chem. 2016;8:381–404. https://doi.org/10.4155/fmc-2015-0007.
Lu H, Zhou Q, He J, Jiang Z, Peng C, Tong R, et al. Recent advances in the development of protein–protein interactions modulators: mechanisms and clinical trials. Sig Transduct Target Ther. 2020;5:213 https://doi.org/10.1038/s41392-020-00315-3.
Parvatkar P, Kato N, Uesugi M, Sato S, Ohkanda J. Intracellular Generation of a Diterpene-Peptide Conjugate that Inhibits 14-3-3-Mediated Interactions. J Am Chem Soc. 2015;137:15624–7. https://doi.org/10.1021/jacs.5b09817.
Hu X, Sun J, Wang HG, Manetsch R. Bcl-XL-Templated Assembly of Its Own Protein-Protein Interaction Modulator from Fragments Decorated with Thio Acids and Sulfonyl Azides. J Am Chem Soc. 2008;130:13820–1. https://doi.org/10.1021/ja802683u.
Kulkarni SS, Hu X, Doi K, Wang HG, Manetsch R. Screening of Protein-Protein Interaction Modulators via Sulfo-Click Kinetic Target-Guided Synthesis. ACS Chem Biol. 2011;6:724–32. https://doi.org/10.1021/cb200085q.
Namelikonda NK, Manetsch R. Sulfo-click reaction via in situ generated thioacids and its application in kinetic target-guided synthesis. Chem Commun. 2012;48:1526–8. https://doi.org/10.1039/C1CC14724B.
Nacheva K, Kulkarni SS, Kassu M, Flanigan D, Monastyrskyi A, Iyamu ID, et al. Going beyond Binary: Rapid Identification of Protein−Protein Interaction Modulators Using a Multifragment Kinetic Target-Guided Synthesis Approach. J Med Chem. 2023;66:5196–207. https://doi.org/10.1021/acs.jmedchem.3c00108.
Kassu M, Parvatkar PT, Milanes J, Monaghan NP, Kim C, Dowgiallo M, et al. Shotgun Kinetic Target-Guided Synthesis Approach Enables the Discovery of Small-Molecule Inhibitors against Pathogenic Free-Living Amoeba Glucokinases. ACS Infect Dis. 2023;9:2190–201. https://doi.org/10.1021/acsinfecdis.3c00284.
Acknowledgements
We thank Northeastern University for providing all the necessary resources.
Funding
Open access funding provided by Northeastern University Library.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Parvatkar, P.T., Manetsch, R. The emergence of sulfo-click amidation in kinetic target-guided synthesis. Med Chem Res (2024). https://doi.org/10.1007/s00044-024-03236-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00044-024-03236-w