Abstract
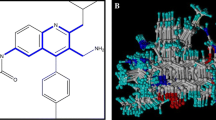
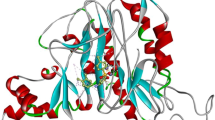
Methionine amino peptidases (MetAPs) are metalloproteases that remove co-translational N-terminal methionine from nascent polypeptide chains. Due to their essential role in protein synthesis, MetAPs are considered as potential targets for antibacterial drugs. In the present work, three-dimensional quantitative structure–activity relationship (3D-QSAR) studies were carried out on a series of pyridine-2-carboxylic acid thiazol-2-ylamide-based MetAP inhibitors using comparative molecular field analysis (CoMFA) and comparative molecular similarity indices analysis (CoMSIA) techniques. The models were developed using 30 training set molecules. The optimum CoMFA and CoMSIA models obtained for the training set were statistically significant with cross-validated correlation coefficients (q 2) of 0.799 and 0.704 and conventional correlation coefficients (r 2) of 0.989 and 0.954, respectively. These inhibitors were docked into MetAP active site. The CoMFA and CoMSIA field contour maps correlate well with the structural characteristics of the binding pocket of MetAP active site. Using the knowledge of structure–activity relationship and receptor–ligand interactions from 3D-QSAR model and the docked complexes, four new pyridine-2-carboxylic acid thiazol-2-ylamide analogs were designed. These analogs exhibit significantly better predicted activity than the reported molecules. The present work has implications for the development of novel antibiotics as potent MetAP inhibitors.





Similar content being viewed by others
References
Accelrys Software Inc., (2010) Discovery studio modeling environment, Release 2.1, San Diego: Accelrys Software Inc., http://accelrys.com/products/discovery-studio/. Accessed 26 Apr 2013
Angus DC, Linde-Zwirble WT, Lidicker J, Clermont G, Carcillo J, Pinsky MR (2001) Epidemiology of severe sepsis in the United States: analysis of incidence, outcome, and associated costs of care. Crit Care Med 29:1303–1310
Arfin SM, Kendall RL, Hall L, Weaver LH, Stewart AE, Matthews BW, Bradshaw RA (1995) Eukaryotic methionyl aminopeptidases: two classes of cobalt-dependent enzymes. Proc Natl Acad Sci USA 92:7714–7718
Bush BL, Nachbar RB Jr (1993) Sample-distance partial least squares: PLS optimized for many variables, with application to CoMFA. J Comput Aided Mol Des 7:587–619
Chang SY, McGary EC, Chang S (1989) Methionine aminopeptidase gene of Escherichia coli is essential for cell growth. J Bacteriol 171:4071–4072
Clark M, Cramer RD (1993) The probability of chance correlation using partial least squares (PLS). Quant Struct Act Relat 12:137–145
Copik AJ, Swierczek SI, Lowther WT, D’souza VM, Matthews BW, Holz RC (2003) Kinetic and spectroscopic characterization of the H178A methionyl aminopeptidase from Escherichia coli. Biochemistry 42:6283–6292
Cramer RD, Bunce JD, Patterson DE (1973) Crossvalidation, bootstrapping, and partial least squares compared with multiple regression in conventional QSAR studies. Quant Struct Act Relat 7:18–25
Cramer RD, Patterson DE, Bunce JD (1988) Comparative molecular field analysis (CoMFA). 1. Effect of shape on binding of steroids to carrier proteins. J Am Chem Soc 110:5959–5967
Folkers G, Merz A, Rognan D (1993) 3D-QSAR in drug design. ESCOM, The Netherlands
Frottin F, Martinez A, Peynot P, Mitra S, Holz RC, Giglione C, Meinnel T (2006) The proteomics of N-terminal methionine cleavage. Mol Cell Proteomics 5:2336–2349
Giglione C, Boularot A, Meinnel T (2004) Protein N-terminal methionine excision. Cell Mol Life Sci 61:1455–1474
Hoge CW, Gambel JM, Srijan A, Pitarangsi C, Echeverria P (1998) Trends in antibiotic resistance among diarrheal pathogens isolated in Thailand over 15 years. Clin Infect Dis 26:341–345
Jones G, Willett P, Glen RC (1995) Molecular recognition of receptor sites using a genetic algorithm with a description of desolvation. J Mol Biol 245:43–53
Jones G, Willett P, Glen RC, Leach AR, Taylor R (1997) Development and validation of a genetic algorithm for flexible docking. J Mol Biol 267:727–748
Kamath S, Buolamwini JK (2003) Receptor-guided alignment-based comparative 3D-QSAR studies of benzylidene malonitrile tyrphostins as EGFR and HER-2 kinase inhibitors. J Med Chem 46:4657–4668
Klebe G, Abraham U, Mietzner T (1994) Molecular similarity indices in a comparative analysis (CoMSIA) of drug molecules to correlate and predict their biological activity. J Med Chem 37:4130–4146
Kubinyi H (1993) QSAR: Hansch analysis and related approaches. VCH Verlagsgesellschaft mbH, Weinheim
Kunin CM (1993) Resistance to antimicrobial drugs—a worldwide calamity. Ann Intern Med 118:557–561
Li X, Chang YH (1995) Amino-terminal protein processing in Saccharomyces cerevisiae is an essential function that requires two distinct methionine aminopeptidases. Proc Natl Acad Sci USA 92:12357–12361
Li JY, Cui YM, Chen LL, Gu M, Li J, Nan FJ, Ye QZ (2004) Mutations at the S1 sites of methionine aminopeptidases from Escherichia coli and Homo sapiens reveal the residues critical for substrate specificity. J Biol Chem 279:21128–21134
Lowther WT, Matthews BW (2000) Structure and function of the methionine aminopeptidases. Biochim Biophys Acta 1477:157–167
Lowther WT, Matthews BW (2002) Metalloaminopeptidases: common functional themes in disparate structural surroundings. Chem Rev 102:4581–4608
Luo QL, Li JY, Liu ZY, Chen LL, Li J, Qian Z, Shen Q, Li Y, Lushington GH, Ye QZ, Nan FJ (2003) Discovery and structural modification of inhibitors of methionine aminopeptidases from Escherichia coli and Saccharomyces cerevisiae. J Med Chem 46:2631–2640
Martin GS, Mannino DM, Eaton S, Moss M (2003) The epidemiology of sepsis in the United States from 1979 through 2000. N Engl J Med 348:1546–1554
Meinnel T, Mechulam Y, Blanquet S (1993) Methionine as translation start signal: a review of the enzymes of the pathway in Escherichia coli. Biochimie 75:1061–1075
Miller CG, Kukral AM, Miller JL, Movva NR (1989) pepM is an essential gene in Salmonella typhimurium. J Bacteriol 171:5215–5217
Poreba M, Gajda A, Picha J, Jiracek J, Marschner A, Klein CD, Salvesen GS, Drag M (2012) S1 pocket fingerprints of human and bacterial methionine aminopeptidases determined using fluorogenic libraries of substrates and phosphorus based inhibitors. Biochimie 94:704–710
Rahal K, Wang F, Schindler J, Rowe B, Cookson B, Huovinen P, Marton A, Lalitha MK, Semina N, Kronvall G, Guzman M (1997) Reports on surveillance of antimicrobial resistance in individual countries. Clin Infect Dis Suppl 1:S169–S175
Sack RB, Rahman M, Yunus M, Khan EH (1997) Antimicrobial resistance in organisms causing diarrheal disease. Clin Infect Dis Suppl 1:S102–S105
Supuran CT, Scozzafava A, Clare BW (2002) Bacterial protease inhibitors. Med Res Rev 22:329–372
SYBYL 8.0 (2008) Tripos International, 1699 South Hanley Rd., St. Louis, Missouri, 63144, USA. http://www.tripos.com/. Accessed 13 Mar 2013
Varshavsky A (1997) The N-end rule pathway of protein degradation. Genes Cells 2:13–28
Vaughan MD, Sampson PB, Honek JF (2002) Methionine in and out of proteins: targets for drug design. Curr Med Chem 9:385–409
Viswanadhan VN, Ghose AK, Revankar GR, Robins RK (1989) Atomic physicochemical parameters for three dimensional structure directed quantitative structure-activity relationships. 4. Additional parameters for hydrophobic and dispersive interactions and their application for an automated superposition of certain naturally occurring nucleoside antibiotics. J Chem Inf Comput Sci 29:163–172
Wold S, Ruhe A, Wold H, Dunn WJ (1984) The collinearity problem in linear regression. The partial least squares approach to generalized inverses. SIAM J Sci Stat Comput 5:735–743
World Health Organization (1999) WHO report on infectious disease: removing obstables to healthy development. WHO: Geneva. http://www.who.int/infectious-disease-report/index-rpt99.html. Accessed 7 Aug 2013
Yang GF, Lu HT, Xiong Y, Zhan CG (2006) Understanding the structure-activity and structure-selectivity correlation of cyclic guanine derivatives as phosphodiesterase-5 inhibitors by molecular docking, CoMFA and CoMSIA analyses. Bioorg Med Chem 14:1462–1473
Ye QZ, Xie SX, Huang M, Huang WJ, Lu JP, Ma ZQ (2004) Metalloform-selective inhibitors of escherichia coli methionine aminopeptidase and X-ray structure of a Mn(II)-form enzyme complexed with an inhibitor. J Am Chem Soc 126:13940–13941
Acknowledgments
Research in VV’s laboratory is supported by RGYI Grant from Department of Biotechnology, Government of India and UPE-2 grant of University of Hyderabad. PAM is supported by the CSIR fellowship.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Meetei, P.A., Hauser, A.S., Raju, P.S. et al. Investigations and design of pyridine-2-carboxylic acid thiazol-2-ylamide analogs as methionine aminopeptidase inhibitors using 3D-QSAR and molecular docking. Med Chem Res 23, 3861–3875 (2014). https://doi.org/10.1007/s00044-014-0950-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-014-0950-z