Abstract
Methionine aminopeptidase (MetAP2) is a metal-containing enzyme that removes initiator methionine from the N-terminus of a newly synthesized protein. Inhibition of the enzyme is crucial in diminishing cancer growth and metastasis. Fumagillin—a natural irreversible inhibitor of MetAP2—and its derivatives are used as potent MetAP2 inhibitors. However, because of their adverse effects, none of them has progressed to clinical studies. In search for potential reversible inhibitors, we built structure-based pharmacophore models using the crystal structure of MetAP2 complexed with fumagillin (PDB ID: 1BOA). The pharmacophore models were validated using Gunner–Henry scoring method. The best pharmacophore consisting of 1 H-bond donor, 1 H-bond acceptor, and 3 hydrophobic features was used to conduct pharmacophore-based virtual screening of ZINC15 database against MetAP2. The top 10 compounds with pharmacophore fit values > 3.00 were selected for further analysis. These compounds were subjected to absorption, distribution, metabolism, elimination, and toxicity (ADMET) prediction and found to have druglike properties. Furthermore, molecular docking calculations was performed on these hits using AutoDock4 to predict their binding mode and binding energy. Three diverse compounds: ZINC000014903160, ZINC000040174591, and ZINC000409110720 with respective binding energy/docking scores of − 9.22, − 9.21, and −817 kcal/mol, were submitted to 100 ns (MD) simulations using Nanoscale MD (NAMD) software. The compounds showed stable binding mode over time. Therefore, they may serve as a scaffold for further computational and experimental optimization toward the design of more potent and safer MetAP2 inhibitors.
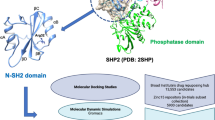
Graphic abstract






Similar content being viewed by others
References
Griffith EC et al (1998) Molecular recognition of angiogenesis inhibitors fumagillin and ovalicin by methionine aminopeptidase 2. Proc Natl Acad Sci 95(26):15183–15188. https://doi.org/10.1073/pnas.95.26.15183
O’Reilly MS, Brem H, Folkman J (1995) Treatment of murine hemangioendotheliomas with the angiogenesis inhibitor AGM-1470. J Pediatr Surg 30(2):325–330. https://doi.org/10.1016/0022-3468(95)90583-9
Rupnick MA et al (2002) Adipose tissue mass can be regulated through the vasculature. Proc Natl Acad Sci U S A 99(16):10730–10735. https://doi.org/10.1073/pnas.162349799
Takamiya Y et al (1994) AGM-1470 inhibits the growth of human glioblastoma cells in vitro and in vivo. Neurosurgery 34(5):869–875. https://doi.org/10.1227/00006123-199405000-00013
Yin SQ et al (2012) The development of MetAP-2 inhibitors in cancer treatment. Curr Med Chem 19(7):1021–1035. https://doi.org/10.2174/092986712799320709
Esa R et al (2020) The role of methionine Aminopeptidase 2 in Lymphangiogenesis. Int J Mol Sci. https://doi.org/10.3390/ijms21145148
McCandless SE et al (2017) Effects of MetAP2 inhibition on hyperphagia and body weight in Prader-Willi syndrome: a randomized, double-blind, placebo-controlled trial. Diabetes Obes Metab 19(12):1751–1761. https://doi.org/10.1111/dom.13021
Siddik MAB et al (2019) A MetAP2 inhibitor blocks adipogenesis, yet improves glucose uptake in cells. Adipocyte 8(1):240–253. https://doi.org/10.1080/21623945.2019.1636627
Proietto J et al (2018) Efficacy and safety of methionine aminopeptidase 2 inhibition in type 2 diabetes: a randomised, placebo-controlled clinical trial. Diabetologia 61(9):1918–1922. https://doi.org/10.1007/s00125-018-4677-0
Han Mİ et al (2019) Synthesis, molecular modeling, in vivo study, and anticancer activity of 1,2,4-triazole containing hydrazide–hydrazones derived from ( S)-naproxen. Archiv der Pharmazie. https://doi.org/10.1002/ardp.201800365
Yılmaz Ö et al (2020) Synthesis, anticancer activity on prostate cancer cell lines and molecular modeling studies of flurbiprofen-thioether derivatives as potential target of metap (type II). Med Chem 16(6):735–749. https://doi.org/10.2174/1573406415666190613162322
Cheruvallath Z et al (2016) Discovery of potent, reversible MetAP2 inhibitors via fragment based drug discovery and structure based drug design—part 1. Bioorg Med Chem Lett 26(12):2774–2778. https://doi.org/10.1016/j.bmcl.2016.04.073
McBride C et al (2016) Discovery of potent, reversible MetAP2 inhibitors via fragment based drug discovery and structure based drug design-Part 2. Bioorg Med Chem Lett 26(12):2779–2783. https://doi.org/10.1016/j.bmcl.2016.04.072
Heinrich T et al (2019) Identification of Methionine Aminopeptidase-2 (MetAP-2) Inhibitor M8891: a clinical compound for the treatment of cancer. J Med Chem 62(24):11119–11134. https://doi.org/10.1021/acs.jmedchem.9b01070
Weako J et al (2020) Identification of potential inhibitors of human methionine aminopeptidase (type II) for cancer therapy: structure-based virtual screening, ADMET prediction and molecular dynamics studies. Comput Biol Chem. https://doi.org/10.1016/j.compbiolchem.2020.107244
Heinrich T et al (2017) Novel reversible methionine aminopeptidase-2 (MetAP-2) inhibitors based on purine and related bicyclic templates. Bioorg Med Chem Lett 27(3):551–556
Liu S et al (1998) Structure of human methionine aminopeptidase-2 complexed with fumagillin. Science 282(5392):1324–1327. https://doi.org/10.1016/j.bmcl.2016.12.019
Guner O, Clement O, Kurogi Y (2004) Pharmacophore modeling and three dimensional database searching for drug design using catalyst: recent advances. Curr Med Chem 11(22):2991–3005. https://doi.org/10.2174/0929867043364036
Berman HM et al (2000) The protein data bank. Nucleic Acids Res 28(1):235–242
Çoruh I et al (2018) Synthesis, anticancer activity, and molecular modeling of etodolac-thioether derivatives as potent methionine aminopeptidase (type II) inhibitors. Archiv der Pharmazie. https://doi.org/10.1093/nar/28.1.235
Liu T et al (2010) Differential expression profiles of alternaria alternate genes in response to carbonyl sulfide fumigation. J Microbiol 48(4):480–485. https://doi.org/10.1007/s12275-010-9301-z
Kusaka M et al (1991) Potent anti-angiogenic action of AGM-1470: comparison to the fumagillin parent. Biochem Biophys Res Commun 174(3):1070–1076. https://doi.org/10.1016/0006-291x(91)91529-l
Arico-Muendel CC et al (2009) Carbamate analogues of fumagillin as potent, targeted inhibitors of methionine aminopeptidase-2. J Med Chem 52(24):8047–8056. https://doi.org/10.1021/jm901260k
Kass DJ et al (2012) Early treatment with fumagillin, an inhibitor of methionine aminopeptidase-2, prevents pulmonary hypertension in monocrotaline-injured rats. PLoS ONE 7(4):e35388. https://doi.org/10.1371/journal.pone.0035388
Ehlers T et al (2016) Methionine aminopeptidase type-2 inhibitors targeting angiogenesis. Curr Top Med Chem 16(13):1478–1488. https://doi.org/10.2174/1568026615666150915121204
Bernier SG et al (2004) A methionine aminopeptidase-2 inhibitor, PPI-2458, for the treatment of rheumatoid arthritis. Proc Natl Acad Sci U S A 101(29):10768–10773. https://doi.org/10.1073/pnas.0404105101
Sheppard GS et al (2004) 3-Amino-2-hydroxyamides and related compounds as inhibitors of methionine aminopeptidase-2. Bioorg Med Chem Lett 14(4):865–868
Kallander LS et al (2005) 4-Aryl-1,2,3-triazole: a novel template for a reversible methionine aminopeptidase 2 inhibitor, optimized to inhibit angiogenesis in vivo. J Med Chem 48(18):5644–5647. https://doi.org/10.1016/j.bmcl.2003.12.031
Wang GT et al (2007) Lead optimization of methionine aminopeptidase-2 (MetAP2) inhibitors containing sulfonamides of 5,6-disubstituted anthranilic acids. Bioorg Med Chem Lett 17(10):2817–2822. https://doi.org/10.1016/j.bmcl.2007.02.062
Kawai M et al (2006) Development of sulfonamide compounds as potent methionine aminopeptidase type II inhibitors with antiproliferative properties. Bioorg Med Chem Lett 16(13):3574–3577. https://doi.org/10.1016/j.bmcl.2006.03.085
Marino JP Jr et al (2007) Highly potent inhibitors of methionine aminopeptidase-2 based on a 1,2,4-triazole pharmacophore. J Med Chem 50(16):3777–3785. https://doi.org/10.1021/jm061182w
Morgen M et al (2016) Spiroepoxytriazoles are fumagillin-like irreversible inhibitors of MetAP2 with potent cellular activity. ACS Chem Biol 11(4):1001–1011. https://doi.org/10.1021/acschembio.5b00755
Mysinger MM et al (2012) Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J Med Chem 55(14):6582–6594. https://doi.org/10.1021/jm300687e
Kurogi Y, Guner O (2001) Pharmacophore modeling and three-dimensional database searching for drug design using catalyst. Curr Med Chem 8(9):1035–1055. https://doi.org/10.2174/0929867043364036
Sterling T, Irwin JJ (2015) ZINC 15 – ligand discovery for everyone. J Chem Inf Model 55(11):2324–2337. https://doi.org/10.1021/acs.jcim.5b00559
Lipinski CA (2004) Lead- and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1(4):337–341. https://doi.org/10.1016/j.ddtec.2004.11.007
Bhal SK et al (2007) The rule of five revisited: applying log D in place of log P in drug-likeness filters. Mol Pharm 4(4):556–560. https://doi.org/10.1021/mp0700209
Morris GM et al (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30(16):2785–2791. https://doi.org/10.1002/jcc.21256
Yang H et al (2019) admetSAR : web-service for prediction and optimization of chemical ADMET properties. Bioinformatics 35(6):1067–1069. https://doi.org/10.1093/bioinformatics/bty707
Lee J et al (2015) CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J Chem Theory Comput 12(1):405–413. https://doi.org/10.1021/acs.jctc.5b00935
Kim S et al (2017) CHARMM-GUI ligand reader and modeler for CHARMM force field generation of small molecules. J Comput Chem 38(21):1879–1886. https://doi.org/10.1002/jcc.24829
Phillips JC et al (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26(16):1781–1802. https://doi.org/10.1002/jcc.20289
Pettersen EF et al (2004) UCSF chimera—a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612. https://doi.org/10.1002/jcc.20084
Ahmed HEA, Zayed MF, Ihmaid S (2015) Molecular pharmacophore selectivity studies, virtual screening, and in silico ADMET analysis of GPCR antagonists. Med Chem Res 24(9):3537–3550. https://doi.org/10.1007/s00044-015-1389-6
Uba AI, Yelekçi K (2018) Pharmacophore-based virtual screening for identification of potential selective inhibitors of human histone deacetylase 6. Comput Biol Chem 77:318–330. https://doi.org/10.1016/j.compbiolchem.2018.10.016
Sakkiah S et al (2014) Dynamic and multi-pharmacophore modeling for designing polo-box domain inhibitors. PLoS ONE 9(7):e101405. https://doi.org/10.1371/journal.pone.0101405
van Breemen RB, Li Y (2005) Caco-2 cell permeability assays to measure drug absorption. Expert Opin Drug Metab Toxicol 1(2):175–185. https://doi.org/10.1517/17425255.1.2.175
Hewitt M et al (2009) In silico prediction of aqueous solubility: the solubility challenge. J Chem Inf Model 49(11):2572–2587. https://doi.org/10.1021/ci900286s
Schultes S et al (2010) Ligand efficiency as a guide in fragment hit selection and optimization. Drug Discov Today Technol 7(3):e157–e162. https://doi.org/10.1016/j.ddtec.2010.11.003
Uba AI et al (2019) Examining the stability of binding modes of the co-crystallized inhibitors of human HDAC8 by molecular dynamics simulation. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2019.1615989
Kleinjung J, Martínez L (2015) Automatic identification of mobile and rigid substructures in molecular dynamics simulations and fractional structural fluctuation analysis. Plos One. https://doi.org/10.1371/journal.pone.0119264
Acknowledgments
AIU and KY thank The Scientific and Technical Research Council of Turkey (TÜBITAK) for support.
Funding
This work received no funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors indicate no potential conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Albayati, S., Uba, A.I. & Yelekçi, K. Potential inhibitors of methionine aminopeptidase type II identified via structure-based pharmacophore modeling. Mol Divers 26, 1005–1016 (2022). https://doi.org/10.1007/s11030-021-10221-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-021-10221-7