Abstract
Polyploidisation—and, additionally, agmatoploidy (concerted fission of chromosomes) in some plant groups—have significantly contributed to the diversification of alpine plant species. Both processes have driven the diversification of Luzula sect. Luzula, leading to a number of different karyotypes, rendering it one of the most intricate plant groups in the Alps. For the Eastern Alps eight species with six karyotypes were reported, but their distribution is insufficiently known. A herbarium revision of 1044 specimens revealed that L. alpina, L. campestris, L. exspectata, L. multiflora and L. sudetica are widespread across the Eastern Alps; L. exspectata is thus new for the Northern Alps and new for Germany, France and possibly Croatia. Luzula divulgata is distributed in the easternmost Alps and adjacent areas, with only a few indications for the western Eastern Alps. Luzula divulgatiformis is new for the Alps where it was recorded in the Southern Alps and southerly adjacent areas. Luzula campestris, L. divulgata and L. divulgatiformis are distributed from lowlands to the montane belt, L. alpina, L. exspectata and L. sudetica are high-elevation species and L. multiflora occurs from lowlands to the alpine belt. Additionally, we estimated genome size (GS) and karyotypes for 20 populations and revealed four karyotypes corresponding to three ploidy levels. The GS of diploid L. exspectata and L. sudetica was 0.83 pg DNA, tetraploid L. alpina had a double (1.63 pg) and hexaploid L. multiflora a triple (2.48 pg) GS. Our study underlines the importance of large-scale herbarium revisions of intricate taxa, combined with cytological methods, even in well-studied mountain areas such as the Alps and poses new hypotheses regarding the evolution of this polyploid–agmatoploid group.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With more than 4000 vascular plant species, the Alps are one of the hotspots of European biodiversity, harbouring many endemics; in many respects they are the best-studied mountain range of the world (Merxmüller 1952, 1953, 1954; Pawłowski 1970; Tribsch and Schönswetter 2003; Tribsch 2004; Schönswetter et al. 2005; Aeschimann et al. 2004, 2011a, b; Taberlet et al. 2016; Kadereit 2017). The Eastern Alps, harbouring 3720 vascular plant taxa (Aeschimann et al. 2011b) consist of the siliceous Central Alps flanked to the south and north by the peripheral Southern and Northern Limestone Alps (Möbus 1997). This tripartite structure of different bedrocks highly affects the distribution ranges of alpine plants, as substrate is one of the key factors influencing plant distributions (Alvarez et al. 2009). There are several calciphilous species restricted to either the calcareous Southern or Northern Alps and some taxa occur in both mountain ranges (Merxmüller 1952, 1953, 1954). Especially the Southern Alps are rich in endemics (Pawłowski 1970; Tribsch 2004; Kadereit 2017) as the southern margin of the Eastern Alps acted as a major refugium for calcicolous plants during the Last Glacial Maximum (Tribsch and Schönswetter 2003), when the Eastern Alps were almost entirely covered with a continuous ice sheet (van Husen 1987).
One of the plant groups in which species distribution ranges in the Eastern Alps have likely been influenced both by the bedrock (Aeschimann et al. 2004) and the Pleistocene glaciations is Luzula sect. Luzula (Juncaceae). It comprises 59 species and numerous infraspecific taxa, which are very similar in habit and difficult to distinguish morphologically (Záveská Drábková 2013). The main differences are in minor quantitative traits, which are very variable (Kirschner 1993, 2002; Bačič et al. 2007b, 2016). For a reliable identification of taxa, genome size (GS) data coupled with karyological investigations are ideal, but time consuming to obtain; furthermore, the latter rely on freshly collected seeds. This approach is thus not suitable for large-scale studies, but can be substituted by detailed revision of herbarium specimens employing measurements of micromorphological characters.
Diversification of Luzula has been strongly influenced by polyploidisation, which was an important driving force in the evolution of Angiosperms (Soltis et al. 2009; Husband et al. 2013; Weiss-Schneeweiss et al. 2013) and is also one of the major factors affecting high mountain biodiversity in the Alps (Burnier et al. 2009; Guggisberg et al. 2009; Casazza et al. 2012; Pachschwöll et al. 2015). Contrary to true polyploidy, where GS at least initially increases linearly with the increased ploidy level, in the case of agmatoploidy (Malheiros and Gardé 1950; Löve et al. 1957; Luceña and Guerra 1996) the increase in chromosome number is not correlated with an increase of GS; instead, the nuclear DNA amount is retained (Mello-Sampayo 1961; Bačič et al. 2007a; Guerra 2016). Agmatoploidy is based on fission of holocentric (holokinetic) chromosomes; in contrast to the “normal type” of monocentric chromosomes, where spindle microtubules attach to a single kinetochore during mitosis or meiosis, in holocentric chromosomes they attach along the whole length through diffuse kinetochores (Bureš et al. 2013). In addition, holocentric sister chromatids are interconnected along their whole length before anaphase disjunction and their fragmentation is thus not necessarily deleterious as the fragmented parts are successfully segregated in daughter nuclei (Nordenskiöld 1951; Haizel et al. 2005; Nagaki et al. 2005; Bozek et al. 2012; Bureš et al. 2013; Guerra 2016). The term agmatoploidy was in the past applied also for the opposite process, that is the concerted fusion of holocentric chromosomes, generating a karyotype with only half of the original chromosome number, but Luceña and Guerra (1996) introduced the term symploidy for this phenomenon. In addition to complete agmatoploidy and symploidy also partial agmatoploidy and symploidy have been observed, with only a subset of chromosomes in the nucleus fragmented or fussed resulting in a continuous series of chromosome numbers (Kirschner 1992a; Guerra 2016; Bureš et al. 2013).
Holocentric chromosomes alongside agmatoploidy and/or symploidy have been observed in at least nine animal and at least five angiosperm lineages (Melters et al. 2012; Bureš et al. 2013), namely Cyperaceae/Juncaceae/Thurniaceae (Nordenskiöld 1951; Löve et al. 1957; Hipp et al. 2009; Bozek et al. 2012), Chionographis (Melanthiaceae; Tanaka and Tanaka 1977), Cuscuta (Convolvulaceae; Pazy and Plitmann 1995), Droseraceae (Melters et al. 2012; Kolodin et al. 2018) and Myristica (Myristicaceae; Flach 1966), but might be even more common (Zedek and Bureš 2018). In Luzula true agmatoploidy and symploidy were observed, whereas in other taxa only partial agmato—and/or symploidy were evidenced (Guerra 2016).
Whereas in the case of true polyploidy the monoploid GS can decrease significantly after polyploidisation, a phenomenon termed genome downsizing (Verma and Rees 1974; Leitch and Bennett 2004; Renny-Byfield et al. 2013), little is known about the changes in GS in the case of agmatoploidy (Guerra 2016). Agmatoploids should have DNA contents similar to their original, non-fragmented complements (Guerra 2016), which was confirmed by early DNA content estimations based on arbitrary values obtained from photometric studies in Luzula species (Halkka 1964). However, later studies revealed quantitative changes in the DNA content of members of agmatoploid series (e.g., Barlow and Nevin 1976). Such changes can, similarly as in organisms with monocentric chromosomes, be connected to proliferation or removal of retrotransposons, which can act very rapidly over short evolutionary periods (Zedek et al. 2010; Bureš et al. 2013). Nishikawa et al. (1984) and Roalson et al. (2007) observed a decreasing GS in correlation with increasing chromosome number in holocentric Carex and related genera, which could be explained by the loss of minute chromosome fragments during chromosomal fission or by an unrecognised mechanism of DNA diminution (Bureš et al. 2013). On the contrary, Schönswetter et al. (2007) suggested processes such as heterochromatin amplification to be responsible for a GS increase of only 6% in Juncus biglumis L. after doubling of 60 chromosomes, but were not able to confirm that the increase in chromosome number was indeed due to agmatoploidy.
The basic chromosome number in Luzula is assumed to be x = 6, which can be altered due to polyploidy, agmatoploidy and symploidy, leading to a somatic chromosome number ranging from 6 to 66 (Bozek et al. 2012). As a result of agmatoploidy, chromosomes of three different sizes have been reported. The largest, full-size chromosomes are designated as AL-type, intermediate, half-size chromosomes as BL-type, and the smallest, quarter-size chromosomes as CL-type (Nordenskiöld 1951; Bačič et al. 2007a; Bozek et al. 2012). Alongside agmatoploidy, true polyploidy also played an important role in the diversification of Luzula sect. Luzula (Malheiros and Gardé 1950; Nordenskiöld 1951, 1956; Kirschner 1992a, b; Bačič et al. 2007a; Bozek et al. 2012). The most common karyotypes in the section are thus attributable to either true polyploidy (2n = 12 AL, diploid; 2n = 24 AL, tetraploid; 2n = 36 AL, hexaploid), complete agmatoploidy (2n = 24 BL, 2n = 48 CL) or mixed karyotypes, i.e., partial agmatoploidy (2n = 12 AL + 24 BL) (Nordenskiöld 1951, 1956; Kirschner 1992a, 2002; Bačič et al. 2007a; Bozek et al. 2012; Záveská Drábková 2013; Bačič et al. 2016). In a phylogenetic study of Luzula based on plastid trnL-F and internal transcribed spacer (ITS) sequences (Bozek et al. 2012), relationships among the Alpine species and most other members of L. sect. Luzula remained unresolved, rendering conclusions regarding the direction of chromosome evolution difficult.
The distribution of species of L. sect. Luzula is insufficiently known due to frequent misidentifications (Bačič et al. 2007b) and the recent description of two new species, rendering many previous records unreliable (Bačič et al. 2007a). Before the two new species were discovered based on their peculiar karyotype, plants belonging to L. exspectata were mostly treated as L. alpina, L. multiflora and L. sudetica (particularly in Slovenia) or remained unidentified as L. multiflora agg., whereas those belonging to L. divulgatiformis were identified as L. campestris, L. multiflora and L. campestris agg. (Bačič et al. 2007a, b; Bačič, personal observations). For the Alps six species from L. sect. Luzula have been reported by Aeschimann et al. (2004). Later, L. exspectata Bačič & Jogan has been described from the Southern Alps, and L. divulgatiformis Bačič & Jogan from southerly adjacent areas (Bačič et al. 2007a, b). Both were discovered also in the Balkan Peninsula (Bačič et al. 2016), indicating that they might be more widespread, suggesting the occurrence of eight species in the Alps (Table 1).
Luzula campestris (L.) DC. s. str. and L. pallescens Sw. are diploid (2n = 12 AL), the former is distributed in Europe and North Africa (Kirschner 2002) and widespread in the Alps (Aeschimann et al. 2004; Fischer et al. 2008). The latter is widely distributed in northern Eurasia (Kirschner 2002), whereas in the Alps only a few localities in Austria have been reported (Fischer et al. 2008). True polyploids reported for the Alps are tetraploid (2n = 24 AL) and hexaploid (2n = 36 AL) populations of L. multiflora (Ehrh.) Lej. s. str. and tetraploid (2n = 24 AL) L. divulgata Kirschner. Luzula multiflora is a widespread and variable species, distributed in North America and Eurasia and introduced to many other areas (Kirschner 2002); it is also common in the Alps (Aeschimann et al. 2004). On the other hand, L. divulgata is an European species distributed from eastern Sweden and northern Poland over Central Europe to Bulgaria and Bosnia and Herzegovina (Kirschner 2002); in the Alps, this species has only been reported for the easternmost Northern Alps in Austria (Aeschimann et al. 2004; Fischer et al. 2008).
In addition, three true agmatoploid species have been reported or are expected in the Alps. Luzula exspectata and L. divulgatiformis have half-sized chromosomes (2n = 24 BL). The former is known from the Southern Alps (Bačič et al. 2007a, b) and the Balkan Peninsula (Bačič et al. 2016), whereas the latter has up to now only been recorded for the Balkan Peninsula (Bačič et al. 2007a, b, 2016). The only species with quarter-sized chromosomes (2n = 48 CL) reported for the Alps, where it is widespread (Aeschimann et al. 2004), is L. sudetica (Willd.) Schult. It is distributed from Iceland and Scandinavia in the north to the Pyrenees, northern Italy and the Balkan Peninsula in the south (Kirschner 2002). The only species in the Alps with a peculiar mixed karyotype (2n = 12 AL + 24 BL) is L. alpina Hoppe, which probably originated after hybridisation of two species and polyploidisation and is thus considered an allopolyploid partial agmatoploid (Kirschner 1992a). It is a common species in the Alps (Aeschimann et al. 2004), which has a disjunct partial distribution area in the eastern Pyrenees (Kirschner 2002). Whereas L. campestris, L. divulgata, L. divulgatiformis, L. multiflora and L. pallescens are mostly distributed in lowland, colline and montane zones, L. alpina, L. exspectata, L. multiflora and L. sudetica are typical species of the subalpine and alpine vegetation belts (Aeschimann et al. 2004; Bačič et al. 2007a, b).
Taking into account the morphological similarity of the taxa and frequent misidentifications, our aim here is to contribute to the knowledge of the distribution of the mostly mountain-dwelling taxa from Luzula sect. Luzula in the Eastern Alps. This is the first large-scale chorological study since the description of L. divulgatiformis and L. exspectata. Our study is based on the revision of 1044 herbarium specimens from eight herbaria, covering mostly the Eastern Alps, but providing some data also for the Western Alps (e.g., for the Aosta valley). Our specific aims are to test (1) whether L. divulgatiformis and L. exspectata have crossed the main chain of the Central Alps and can also be found in the Northern Alps and (2) whether there are differences in the distribution of species with respect to substrate (siliceous vs. calcareous) and elevation. In addition, we (3) provide additional GS and karyological data for the Alpine populations and (4) test whether changes in GS (i.e., genome increase or downsizing) occurred after polyploidisation and agmatoploidisation in Luzula sect. Luzula, taking into account all previously published GS data obtained with the same methodology.
Materials and methods
Revision of herbarium material and distribution maps
We revised 1044 specimens from herbaria (abbreviations follow Thiers 2016) in Austria (IB, WU, private herbarium collection W. Gutermann), Italy (AO, ROV, private herbarium collections C. Argenti from Belluno, M. Bovio from Aosta valley, F. Martini from Lombardia) and Slovenia (LJU; see Online Resource 1 for details); some of these specimens were collected by us between 2007 and 2012. Even if several herbaria were examined, most of them contain specimens from specific geographic regions of the Alps, notable exceptions being WU and the private herbarium collection W. Gutermann. Consequently, the distributional data presented here are geographically biased and some regions remain poorly sampled, but we are confident that the available data allow for general conclusions about the distribution of the investigated species in the Eastern Alps.
Plants were identified by T. Bačič using identification keys and other relevant literature (e.g., Kirschner 1993, 1994, 2002; Bačič et al. 2007a, b, 2016; Pignatti 2017); the most important characters are described below and can also be found in the identification key (see Results). The identification of the majority of the specimens could not be verified karyologically (for the others see below); rather, we estimated the ploidy level of the plants by measuring stomatal lengths as suggested by Jarolímová and Kirschner (1995) and utilised by Bačič et al. (2007a, b, 2016). The maximal length of the guard cells of 20–25 stomata was measured in the middle of the second cauline leaf from the top (abaxial epidermis of rehydrated herbarium material). Plants with stomatal lengths < 38 μm were considered diploid and those with stomatal lengths > 40 μm to be polyploid, whereas those in-between were classified as uninformative.
Luzula campestris and L. divulgata have globular seeds with large caruncles, long anthers with short filaments, long styles and stigmas. Luzula campestris has underground stolons and an almost smooth margin of cauline leaves, whereas L. divulgata is caespitose without stolons, and has densely papillose–serrate cauline leaf margins. All other studied species differ from the two in having ovoid, oblong–ovoid or ellipsoidal seeds with shorter caruncles and anthers. In general habit, L. divulgatiformis resembles L. divulgata, but in size of flower characters it approaches lowland forms of L. multiflora with its styles, stigmas and anther/filament length ratio somewhat longer than in L. multiflora. The stomatal length clearly distinguishes diploid L. divulgatiformis [(24) 30–37 (38) μm] from polyploid L. multiflora [(37) 39–48 (54) μm]. Luzula pallescens, which is frequently confused with pale forms of L. multiflora, is well characterised by having very small seeds and cylindrical clusters of many small flowers with pale tepals. An important, though less practical characteristic of L. pallescens is that its peduncles are densely papillose (at least in the upper half, viewed best at 150× magnification).
Of the investigated high mountain species, L. sudetica is morphologically the most distinct, recognisable by small seeds with very short caruncles and relatively small flower parts with dark tepals and capsules. Luzula alpina is characterised by dark brown to blackish flowers with long acuminate tepals, relatively large flower parts and seeds, often congested inflorescences and conspicuously broad basal leaves (Kirschner 1993). Mountain forms of Luzula multiflora can be very similar to L. alpina, dark-flowered, with similar seed, stamen and style/stigma characters and sometimes even with congested inflorescences, but their basal leaves are not conspicuously broad. As they are of the same ploidy, stomatal lengths and genome size measurements are not useful to distinguish them. Luzula exspectata is morphologically similar to L. alpina and the mountain form of L. multiflora, it strongly overlaps with these two species in floral and seed characters, and is rather variable in the width of the basal leaves and inflorescence habit, which is usually partly congested. However, as L. exspectata is diploid, it can be differentiated from polyploid L. alpina and L. multiflora by having shorter stomata and smaller genome size.
Species distribution maps of all revised specimens were produced in ArcGIS 10.3 (ESRI, Redlands, California, USA). As most of the specimens were lacking coordinates, we determined them approximately, using Google Earth, Austrian Map online (http://www.austrianmap.at) and Geopedia (www.geopedia.si). Most of the coordinates in Online Resource 1 are thus imprecise, but serve well for displaying general distribution patterns. In addition, we obtained data on elevational distribution of the taxa from herbarium labels; in cases where an elevation range was given on the herbarium label, both the minimum and maximum values were considered. The number of elevation data was N = 99 for L. alpina, N = 296 for L. campestris, N = 35 for L. divulgata, N = 32 for L. divulgatiformis, N = 153 for L. exspectata, N = 208 for L. multiflora and N = 114 for L. sudetica. We presented differences in elevational distribution among species in boxplots and statistically tested them using one-way analyses of variance (ANOVA) and Tukey’s post hoc test. Difference in elevational range between low-elevation (L. campestris, L. divulgata, L. divulgatiformis) and high-elevation species (L. alpina, L. exspectata, L. sudetica) was tested by Student t test; L. multiflora was not included in this analysis as its distribution ranges from lowlands to the alpine belt.
Chromosome number determination and genome size measurement
For the karyological analyses we collected seeds from 20 populations of four species during field excursions between 2005 and 2012 in the Austrian, Italian and Slovenian Alps (see Online Resource 1 for voucher information). Seeds were germinated in Petri dishes until the rootlet was at least 3 mm long. Root tips of young seedlings were fixed, stained and squashed to prepare microscopic slides, as previously described in Bačič et al. (2007a, 2016). Briefly, root tips were fixed in 4% phosphate-buffered neutral formaldehyde for 90 min at room temperature and post-fixed in several changes of 3: 1 methanol: acetic acid for at least 24 h at 4 °C. Fixed root tips were hydrolysed in 5 N HCl for 90 min at 20 °C and stained with Feulgen reagent overnight at 4 °C. Stained root tips were washed in several changes of SO2–water, and squash preparations of dissected root meristems were prepared in 45% acetic acid. Nuclei were frozen on the slide by dry-ice freezing, dehydrated in 96% ethanol for max 5 min and air dried in the dark. The amount of nuclear DNA (C-value) was measured densitometrically by interphase-peak DNA-image cytometry (Vilhar et al. 2001), using an Axioscope MOT light microscope (Carl Zeiss, Germany) with a 63 × oil immersion objective, CCD camera (Sony DXC-950P), frame grabber (Matrox Meteor) and computer with KS 400 ver. 3.0 image analysis software package (Carl Zeiss, Germany). The image analysis system was densitometrically calibrated prior to each set of measurements, and integrated optical density (IOD) was measured for approximately 200 interphase nuclei per slide. Pisum sativum cv. ‘Kleine Rheinländerin’ with 2C-value of 8.84 pg DNA (Greilhuber and Ebert 1994) was used as calibration standard to convert arbitrary units of IOD to pg DNA. Root tips of P. sativum were processed simultaneously with the Luzula specimens during all experimental procedures, i.e., germination, fixation, staining and measurement. The amount of DNA in the unreplicated (G1 phase) nuclei was expressed as 2C-value; for simplicity, we apply the term genome size (GS) to these values hereafter. The same slides were used to count chromosomes. Statistical analyses were performed with Microsoft Excel and Prism 3.02 (GraphPad) software.
Statistical analyses of genome size changes following polyploidisation and agmatoploidisation
To study if genome downsizing or increase occurred after agmatoploidisation or true polyploidisation in Luzula sect. Luzula, all available GS data obtained by image densitometry (i.e., the data published here supplemented with those from Bačič et al. 2007a, b, 2016) were recalculated to the monoploid GS (1x). In total, 103 monoploid GS data were analysed, including 72 diploids (L. campestris, L. divulgatiformis, L. exspectata, L. sudetica, L. taurica with karyotypes 12 AL—16 samples, 24 BL—47 samples and 48 CL—9 samples), 17 tetraploids (L. alpina, L. divulgata, L. multiflora with karyotypes 24 AL—9 samples and 12 AL + 24 BL—8 samples) and 14 hexaploids (L. multiflora with karyotype 36 AL). The GS was compared among the three ploidy levels and among different karyotypes within the same ploidy level using t test or one-way analysis of variance (ANOVA) and Tukey’s post test. The range of the GS within each of the recovered ploidy levels (‘intra-ploidy variation’) was calculated as the ratio of the highest and the lowest DNA content.
Results
Distribution of species in the Eastern Alps based on the revision of herbarium specimens
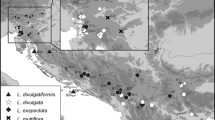
The revision of 1044 specimens confirmed the presence of seven species (Fig. 1, Online Resource 1) in the Eastern Alps—L. alpina (91 specimens), L. campestris (307 specimens), L. divulgata (27 specimens), L. divulgatiformis (31 specimens), L. exspectata (158 specimens), L. multiflora (250 specimens) and L. sudetica (111 specimens). Twenty-eight specimens could not be identified reliably and are therefore shown with white dots in Fig. 1 and marked with “cf.” in Online Resource 1. In addition, 69 specimens could not be determined to the species level, mostly due to poorly developed or collected herbarium material and high morphological variability of the group, and were thus labelled as L. campestris agg. Whereas L. alpina, L. campestris, L. exspectata, L. multiflora and L. sudetica are widespread across the Eastern Alps, L. divulgata is mostly limited to the easternmost Northern Alps and adjacent areas, with only a few indications for the western Eastern Alps, and L. divulgatiformis occurs only in the Southern Alps and southerly adjacent areas (Fig. 1).
Based on the results of herbarium revision and previously published data (mostly following Bačič et al. 2007b and Kirschner 1993, 1994, 2002), we present an updated identification key for the Alpine taxa of Luzula sect. Luzula in Online Resource 2.
Elevational distribution and substrate preferences
The box plots displaying the elevational distribution of the taxa (Fig. 2) reveal that L. alpina, L. exspectata and L. sudetica are mountain species mostly absent from lowlands, and reaching elevations above 2600 m a.s.l. Luzula multiflora can occur at higher elevations, but most of the localities were below 1500 m a.s.l. Luzula campestris, L. divulgata and L. divulgatiformis are lowland and montane species; only L. campestris was recorded also above 2000 m a.s.l. The combined elevational data for the three lowland species ranged from 20 to 2100 m a.s.l. and significantly differed (p < 0.0001, t test) from the three montane species which ranged from 550 to 2620 m a.s.l. Significant differences in elevational distributions were confirmed also by ANOVA for all the species (p < 0.0001), except for the comparisons of L. divulgatiformis vs. L. divulgata and L. campestris, and L. alpina vs. L. sudetica (Tukey’s post test p > 0.05).
Elevational distribution of species of Luzula sect. Luzula based on the revision of 1044 herbarium specimens, with geographical focus on the Eastern Alps (N = 99 for L. alpina, N = 296 for L. campestris, N = 35 for L. divulgata, N = 32 for L. divulgatiformis, N = 153 for L. exspectata, N = 208 for L. multiflora, N = 114 for L. sudetica)
From the data on the herbarium labels we can conclude that L. alpina and L. sudetica prefer siliceous substrate, but can also be found on limestone on locally acidic soils, whereas L. exspectata predominantly grows on limestone, but can occasionally be found on siliceous bedrock. Of the lower elevation species, L. divulgatiformis prefers calcareous and L. divulgata siliceous substrate, whereas L. campestris and L. multiflora are not linked to either of them.
Chromosome number and genome size variation
Cytogenetic analyses of 20 populations revealed four karyotypes corresponding to three ploidy levels (Fig. 3). The GS values for all analysed populations are presented in Online Resource 1 and, along with the previously published data for all Alpine species (Kirschner 2002; Bačič et al. 2007a, 2016), in Table 1. Most populations belonged to L. alpina (N = 7) with 2n = 12 AL + 24 BL and L. sudetica (N = 6) with 2n = 48 CL, followed by L. exspectata (N = 4) with 2n = 24 BL and hexaploid L. multiflora with 2n = 36 AL (N = 3). The estimated GS in the investigated samples ranged from 0.69 to 2.54 pg DNA. The GS of diploid L. exspectata and L. sudetica was 0.83 ± 0.01 and 0.83 ± 0.04 pg, respectively, tetraploid L. alpina had approximately double (1.63 ± 0.03 pg) and hexaploid L. multiflora triple GS (2.48 ± 0.03 pg) of the diploids. The coefficient of variation for GS measured in different populations of the same species was below 5% in all species except in L. sudetica (CV = 11.49%).
Genome size (2C-value) of investigated taxa of Luzula sect. Luzula. Each dot represents a 2C-value measured in 100–200 interphase nuclei of a single Luzula root tip and calibrated to Pisum sativum. Lines represent mean 2C-values for each taxon. The number of measured root tips (N) for each species is given in parentheses. Luzula exspectata (N = 16); L. sudetica (N = 28); L. alpina (N = 43); L. multiflora (N = 19)
When comparing the GS across all populations belonging to Luzula sect. Luzula analysed until now using densitometry (i.e., this study, Bačič et al. 2007a, 2016; Fig. 4), the mean 2C-values were 0.90 ± 0.01 pg DNA in diploids (N = 72, CV = 11.21%), 1.80 ± 0.05 pg in tetraploids (N = 17, CV = 10.58%) and 2.64 ± 0.04 pg in hexaploids (N = 14, CV = 5.98%). Differences among mean 2C-values of agmatoploids within the same ploidy level were non-significant (p = 0.2449 for diploids and p = 0.5156 for tetraploids), but the values differed significantly between any two ploidy levels (p < 0.0001, ANOVA and Tukey’s post-test). The intra-ploidy variation in GS was the highest in diploids (1.71-fold), intermediate in tetraploids (1.39-fold) and the lowest in hexaploids (1.21-fold). However, when comparing the monoploid (1x) GS across all three ploidy levels (Fig. 4 insert), no significant differences were apparent (p = 0.8136): 1x-values were 0.45 ± 0.01 pg DNA for diploids, 0.45 ± 0.01 pg for tetraploids and 0.44 ± 0.01 pg for hexaploids. Moreover, when comparing the GS among different karyotypes of the same ploidy level, no significant differences were observed. In diploids, the mean 2C-value of the 12 AL karyotype was 0.88 ± 0.02 pg DNA, for 24 BL karyotype 0.91 ± 0.02 pg and for 48 CL karyotype 0.84 ± 0.04 pg (p = 0.2449). The same was true for tetraploids with 24 AL karyotype (1.80 ± 0.05 pg) and 12 AL + 24 BL karyotype (1.74 ± 0.07 pg; p = 0.4847).
Genome size (2C-value) variation across three ploidy levels (2x, 4x and 6x) in Luzula sect. Luzula (data from this study, Bačič et al. 2007a, 2016). The insert shows the monoploid (1x) genome size, calculated from 2C-values of all three ploidy levels (N = 177 for diploids, N = 44 for tetraploids, N = 41 for hexaploids). Each dot represents a 2C-value measured in 100–200 interphase nuclei of a single Luzula root tip and calibrated to Pisum sativum
Discussion
Distribution patterns and variability of morphological characters
Our revision of 1044 herbarium specimens confirmed the presence of seven species belonging to Luzula sect. Luzula in the Eastern Alps and significantly enlarged the previously known distribution areas of L. divulgata (likely new for Italy), L. divulgatiformis (new for Italy) and L. exspectata (new for the Northern Alps). We acknowledge that the distributional data presented here are geographically biased but clearly show the large-scale distribution patterns of the investigated species in the Eastern Alps.
The only species reported for the Alps (Aeschimann et al. 2004; Fischer et al. 2008) but not confirmed by our revision is L. pallescens. In the Eastern Alps, this species has only been reported from Austria (Fischer et al. 2008), namely from several localities in Lower Austria (Niederösterreich) and Upper Styria (Obersteiermark; Melzer 1996), one locality in Carinthia (Kärnten) and one in North Tyrol (Nordtirol; Melzer 1982). In Austria it is a rare and critically endangered species (Fischer et al. 2008), therefore it is not surprising that it was not confirmed by our herbarium revision. Whereas the records from the eastern margins of the Alps are likely reliable as they are westerly adjacent to species’ continuous distribution area, the two isolated records from Carinthia and Tyrol should be checked in the field in the future.
A similarly scattered distribution in the Eastern Alps as for L. pallescens was reported for L. divulgata, which is also mostly restricted to the easternmost Alps and adjacent areas in Austria (Aeschimann et al. 2004; Fischer et al. 2008), but known from a single locality (Hohenweiler) in the western Eastern Alps in Vorarlberg [westernmost Austria; based on herbarium specimen revised by Kirschner; see Polatschek (2001)]. Our revision confirms that the species is most common in pre-Alpine lowland areas of eastern Austria (Fig. 1), but three specimens indicate its presence in the western Eastern Alps in Italy (Veneto: Belluno, Lombardia: Brescia) and Austria (Nordtirol: Ötztal). The species is rare in the Eastern Alps, but more widespread than believed before.
Similarly as L. divulgata and L. pallescens, L. divulgatiformis is relatively thermophilous; it is a common submediterranean species growing in open wood- and scrublands, forest margins and clearings as well as nutrient-poor karstic meadows and pastures below 1000 m a.s.l. (Fig. 2). The herbarium revision revealed its presence in similar habitats in the Southern Alps and southerly adjacent areas in Italy (Lago di Garda region in Lombardia and Trentino, and surroundings of Padova in Veneto), where it is probably more common.
All other species confirmed by the herbarium revision are distributed throughout the Eastern Alps (Fig. 1), which, with exception of L. exspectata, is in accordance with previously published data (e.g., Kirschner 2002; Aeschimann et al. 2004; Fischer et al. 2008). Of all mountain species L. sudetica is morphologically and ecologically most distinct, growing in moist meadows and bogs in the montane and alpine zones (Kirschner 2002). In the Eastern Alps it is most common in the siliceous Central Alps, but also in the Southern Alps (Fig. 1).
Luzula campestris and L. multiflora are predominantly lowland taxa (Fig. 2), the former growing in dry to moist meadows, and the latter preferring moist habitats, at least in the lowlands. Both, but especially L. multiflora, can be found also at higher elevations above 2000 m a.s.l. (Fig. 2). Whereas L. campestris is morphologically distinct and we were thus able to reliably identify all herbarium specimens, the identification of L. multiflora is more difficult, especially for tetraploid plants from higher elevations, which are difficult to distinguish from L. alpina and L. exspectata.
Luzula alpina is a common Alpine species (Kirschner et al. 1988), thriving from the montane to the alpine zone (Fig. 2). As it is acidophilic and grows mostly on siliceous substrate, it is most common in the Central Alps; there are, however, also many localities in the Southern Alps, rendering this species’ distribution in the Alps similar to that of L. sudetica (Fig. 1). Our revision of the herbarium material confirmed previous observations (Kirschner 1993) that although L. alpina typically has congested inflorescences, plants with pedunculate inflorescence clusters are not rare. In addition, the basal leaves of L. alpina are typically conspicuously broad as reported previously (e.g., Kirschner 1993; Fischer et al. 2008), but can also be much narrower, making its separation from L. multiflora difficult. Since both species are tetraploid, stomata-length and genome size do also not discriminate between them, but are crucial characters to distinguish morphologically and ecologically similar diploid L. exspectata.
Luzula exspectata is the most common (sub)alpine species of L. sect. Luzula on calcareous substrate in the Southern Alps as indicated by Bačič et al. (2007a, b, 2016), but also has scattered localities in the Central and the Northern Alps (Fig. 1) and we here report its first record for Germany (Bayern: Allgäuer Alpen). It mostly grows in oligotrophic, humus-rich grasslands on calcareous substrate, but can also be found on siliceous substrate. It is likely that this species is a common representative of L. sect. Luzula also at higher elevations of the Northern Alps, but additional studies are needed to confirm this. It also remains uncertain how common this species is in the Western Alps, for which we report a first record for France (Provence-Alpes-Côte d’Azur: Haut-Queyras). In addition, it is very likely that this species occurs also in Croatia (Istra: Učka), even if the specimen from there could not be identified unambiguously (Fig. 1).
High morphological variability, as shown in Online Resource 2 and 3, and intermediate stomatal lengths are the reasons, why seven of 91 revised specimens of L. alpina, 13 of 158 revised specimens of L. exspectata and three of 250 revised specimens of L. multiflora could not be unambiguously identified (“cf.” in Fig. 1 and Online Resource 1).
Elevational distribution and substrate preferences
Elevation data derived from herbarium labels (Fig. 2) mostly correspond to the ranges given in Flora Alpina (Aeschimann et al. 2004) for the studied taxa. In addition to the zones indicated there, our data confirm the presence of L. alpina also in the upper montane zone and L. sudetica is likely more common and not only sporadically occurring in the alpine zone. Whereas among the predominately low-elevation species diploid (L. campestris), polyploid (L. divulgata, L. multiflora) and agmatoploid (L. divulgatiformis) species can be found, only polyploids or agmatoploids can be found at higher elevation, even if L. campestris reaches up to 2000 m.
Similarly, substrate preferences correspond largely to those given in Flora Alpina. The only exception is L. alpina, for which equal occurrence frequency was indicated for calcareous, siliceous and mixed substrates by Aeschimann et al. (2004). Our data, however, indicate that this species prefers siliceous substrate, but can occasionally be found on limestone on locally acidic soils and thus the same substrate preferences should be given as for L. sudetica. This discrepancy is likely due to confusion of L. alpina with L. exspectata in the past, as the latter was described after the publication of Flora Alpina (Aeschimann et al. 2004). There were no clear patterns in substrate specificity in the Alps in relation to the karyotype of the species: the three predominately silicicolous species are polyploids or agmatoploids, calcicolous L. divulgatiformis and L. exspectata are agmatoploids, and species without substrate specificity are di- and polyploid.
Chromosome number and genome size variation
Until present, GS has been estimated for 23 Luzula species (ca. 20% of all species; Bennett and Leitch 2012), of which 11 belong to Luzula sect. Luzula (Záveská Drábková 2013). The present study attributed additional estimates for L. alpina, L. exspectata, L. multiflora and L. sudetica, all in the range of previously published results obtained with DNA-image cytometry (Bačič et al. 2007a, 2016) and flow cytometry (Bozek et al. 2012).
According to the prediction of the agmatoploid origin, the GS should be approximately the same in 12 AL, 24 BL and 48 CL cytotypes, and the GS of the partial agmatoploid (2n = 12 AL + 24 BL) should be equal to the one in tetraploids with the 24 AL karyotype (Barlow and Nevin 1976; Guerra 2016). The statistical tests performed for all available GS data showed no significant differences among 12 AL, 24 BL and 48 CL karyotypes and between 24 AL and 12 AL + 24 BL karyotypes. No significant changes in GS thus took place after the fission of chromosomes, which is in line with some previous suggestions (Halkka 1964; Kuta et al. 2004; Bačič et al. 2007a; Bozek et al. 2012), but has never been tested statistically on a larger sample before. On the contrary, some other studies (e.g., Barlow and Nevin 1976; Roalson et al. 2007; Bureš et al. 2013) showed changes in GS after fission of chromosomes, but these studies were mostly performed on other plant groups and with a smaller sample. This discrepancy clearly suggests that additional studies, both in Luzula as well as other plant groups with holocentric chromosomes are needed to shed light on genome evolution after agmatoploidisation.
In addition, also the monoploid GS of true diploid, tetraploid and hexaploid species is not significantly different, as revealed by the comparison of GS of 72 diploid, 17 tetraploid and 14 hexaploid populations (this study; Bačič et al. 2007a, 2016; Fig. 3). Genome downsizing, which often occurs after polyploidisation (Verma and Rees 1974; Leitch and Bennett 2004; Renny-Byfield et al. 2013), did thus not occur in Luzula sect. Luzula.
Hypotheses about the origin of the Alpine species of Luzula sect. Luzula
There are no phylogenetic studies providing sufficient resolution in the phylogenetic trees to enable conclusions regarding the origin and evolution of the Alpine species. It is therefore only possible to provide hypotheses about possible phylogenetic relationships based on cytogenetic and morphological traits. Only three European species are true diploids (12 AL) and thus putative parental species of all other taxa, namely L. campestris, L. pallescens and L. taurica (Kirschner 1992a). Luzula campestris is morphologically very divergent from other Alpine taxa with exception of L. divulgata (24 BL), for which Kirschner (1992a) suggested an autopolyploid origin from L. taurica (V. I. Krecz) Novikov, a species distributed between the Balkan Peninsula and the Caucasus (Kirschner 2002). Similarly, L. divulgatiformis and L. expectata (both 24 BL) might have originated from L. taurica via chromosome fission. Kirschner (1992a) suggested that the alpine tetraploid L. multiflora subsp. multiflora (24 AL) is of autopolyploid origin derived from L. pallescens (12 AL), whereas the lowland hexaploid populations (36 AL) are probably of allopolyploid origin, which was also suggested for other tetraploid taxa of L. multiflora s. lato (Kirschner 1995, 1996). Likewise, based on the seed characters, Kirschner (1992a) suggested that L. pallescens (12 AL) could have been an ancestor of L. sudetica (48 CL), but the link with 24 BL between them is missing (extinct or not yet sampled). Alternatively, based on the karyotype, L. exspectata (24 BL) might have been ancestor of L. sudetica, but the seed morphology does not support this hypothesis. Luzula alpina (12 AL + 24 BL) is of putative allopolyploid origin (Kirschner 1992a), with L. exspectata (24 BL) being one of the putative parents (Bačič et al. 2007a, b), whereas the other could be L. taurica (12 AL), which has similar seed characters, but divergent flower characters. Given unclear origin and relationships among the taxa of L. sect. Luzula, phylogenetic studies with genetic markers providing sufficient resolution are needed in the future. They will also enable a reconstruction of genome and karyotype evolution within the section.
Conclusions
Our study underlines the importance of both polyploidy and agmatoploidy for the evolution of Alpine species from L. sect. Luzula, as all high-elevation taxa are either agmatoploid (L. exspectata and L. sudetica), polyploid (L. multiflora) or both processes have been involved in their origin (L. alpina). It thus seems plausible that holocentric chromosomes confer selective advantage compared to monocentric chromosomes (Kolodin et al. 2018). Zedek and Bureš (2018) suggested that agmatoploidy can be evolutionary advantageous in extreme environments, e.g., in times of exposure to factors that cause chromosomal fragmentation. One of such factors is UV radiation, which increases with elevation (Zedek and Bureš 2018) and agmatoploid plants such as Luzula might, therefore, be in advantage in high mountain habitats. In addition, increased recombination rates associated to increased chromosome numbers due to fission of holocentric chromosomes may allow for more rapid rates of adaptation to novel environments (Escudero et al. 2012), which might have been beneficial also during the spread of Luzula species in the Alps after the retreat of the Pleistocene glaciers in the Holocene. No changes in GS happened after the fission of chromosomes in L. sect. Luzula, but actual processes accompanying agmatoploidy and subsequent evolution of genome remain poorly understood. In addition, the distribution of L. divulgata and L. pallescens centred in the easternmost Northern Alps and that of L. divulgatiformis centred in the Southern Alps emphasizes the importance of these areas as glacial refugia (Tribsch 2004). Last but not least, our study emphasizes the importance of taxonomic studies and large-scale herbarium revisions of intricate plant groups even in the Alps, one of the best-studied areas of the world.
References
Aeschimann D, Lauber K, Moser DM, Theurillat J-P (2004) Flora Alpina. Zanichelli, Berlin
Aeschimann D, Rasolofo N, Theurillat J-P (2011a) Analyse de la Flore des Alpes. 1: Historique et Biodiversité. Candollea 66:27–55. https://doi.org/10.15553/c2011v661a2
Aeschimann D, Rasolofo N, Theurillat J-P (2011b) Analyse de la Flore des Alpes. 2: Biodiversité et Chorologie. Candollea 66:225–253. https://doi.org/10.15553/c2011v662a1
Alvarez N, Thiel-Egenter C, Tribsch A, Holderegger R, Manel S, Schönswetter P, Taberlet P, Brodbeck S, Gaudeul M, Gielly L, Küpfer P, Mansion G, Negrini R, Paun O, Pellecchia M, Rioux D, Schüpfer F, van Loo M, Winkler M, Gugerli F (2009) History or ecology? Substrate type as a major driver of spatial genetic structure in Alpine plants. Ecol Lett 12:632–640. https://doi.org/10.1111/j.1461-0248.2009.01312.x
Bačič T, Jogan N, Dolenc Koce J (2007a) Luzula sect. Luzula in the south-eastern Alps: karyology and genome size. Taxon 56:129–136. https://doi.org/10.1007/s12224-016-9235-2
Bačič T, Dolenc Koce J, Jogan N (2007b) Luzula sect. Luzula (Juncaceae) in the South-Eastern Alps: morphology, determination and geographic distribution. Bot Helv 117:75–88. https://doi.org/10.1007/s00035-007-0783-1
Bačič T, Frajman B, Dolenc Koce J (2016) Diversification of Luzula sect. Luzula (Juncaceae) on the Balkan Peninsula: a cytogenetic approach. Folia Geobot 51:51–63. https://doi.org/10.1007/s12224-016-9235-2
Barlow PW, Nevin D (1976) Quantitative karyology of some species of Luzula. Plant Syst Evol 125:77–86. https://doi.org/10.1007/BF00986773
Bennett MD, Leitch IJ (2012) Plant DNA C-values database (release 6.0, Dec. 2012). http://data.kew.org/cvalues/(download. Accessed 18 Oct 2014
Bozek M, Leitch AR, Leitch IJ, Záveská Drábková L, Kuta E (2012) Chromosome and genome size variation in Luzula (Juncaceae), a genus with holocentric chromosomes. Bot J Linn Soc 170:529–541. https://doi.org/10.1111/j.1095-8339.2012.01314.x
Bureš P, Zedek F, Marková M (2013) Holocentric chromosomes. In: Wendel J, Greilhuber J, Doležel J, Leitch IJ (eds) Plant genome diversity. Vol. 2. Physical structure of plant genomes, vol 2. Springer, Heidelberg, pp 187–208. https://doi.org/10.1007/978-3-7091-1160-4_12
Burnier J, Buerki S, Arrigo N, Küpfer P, Alvarez N (2009) Genetic structure and evolution of alpine polyploid complexes: Ranunculus kuepferi (Ranunculaceae) as a case study. Mol Ecol 18:3730–3744. https://doi.org/10.1111/j.1365-294X.2009.04281.x
Casazza G, Granato L, Minuto L, Conti E (2012) Polyploid evolution and Pleistocene glacial cycles: a case study from the alpine primrose Primula marginata (Primulaceae). BMC Evol Biol 12:56. https://doi.org/10.1186/1471-2148-12-56
Escudero M, Hipp AL, Hansen TF, Voje KL, Luceño M (2012) Selection and inertia in the evolution of holocentric chromosomes in sedges (Carex, Cyperaceae). New Phytol 195:237–247. https://doi.org/10.1111/j.1469-8137.2012.04137.x
Fischer MA, Oswald K, Adler W (2008) Exkursionsflora für Österreich, Liechtenstein und Südtirol. 3. Auflage.: Biologiezentrum der Oberösterreichischen Landesmuseen. Linz
Flach M (1966) Diffuse centromeres in a dicotyledonous plant. Nature 209:1369–1370. https://doi.org/10.1038/2091369b0
Greilhuber J, Ebert I (1994) Genome size variation in Pisum sativum. Genome 37:646–655. https://doi.org/10.1139/g94-092
Guerra M (2016) Agmatoploidy and symploidy: a critical review. Genet Mol Biol 39:492–496. https://doi.org/10.1590/1678-4685-gmb-2016-0103
Guggisberg A, Mansion G, Conti E (2009) Disentangling reticulate evolution in an arctic-alpine polyploid complex. Syst Biol 58:55–73. https://doi.org/10.1093/sysbio/syp010
Haizel T, Lim YK, Leitch AR, Moore G (2005) Molecular analysis of holocentric centromeres of Luzula species. Cytogenet Genome Res 109:134–143. https://doi.org/10.1159/000082392
Halkka O (1964) A photometric study of the Luzula problem. Hereditas 52:81–88
Hipp AL, Rothrock PE, Roalson EH (2009) The evolution and chromosome arrangements in Carex (Cyperaceae). Bot Rev 75:96–109
Husband BC, Baldwin SJ, Suda J (2013) The incidence of polyploidy in natural plant populations: major patterns and evolutionary processes. In: Leitch IJ, Greilhuber J, Doležel J, Wendel JF (eds) Plant Genome Diversity, vol 2. Springer, New York, pp 255–276
Jarolímová V, Kirschner J (1995) Tetraploids in Luzula multiflora (Juncaceae) in Ireland: karyology and meiotic behaviour. Folia Geobot Phytotx 30:389–396. https://doi.org/10.1007/BF02803970
Kadereit JW (2017) The role of in situ species diversification for the evolution of high vascular plant species diversity in the European Alps: a review and interpretation of phylogenetic studies of the endemic flora of the Alps. Perspect Plant Ecol Evol Syst 26:28–38. https://doi.org/10.1016/j.ppees.2017.03.002
Kirschner J (1992a) Karyological differentiation of Luzula sect. Luzula in Europe. Thaiszia 2:11–39
Kirschner J (1992b) Luzula sect. Luzula (Juncaceae) in Spain. Plant Syst Evol 200:1–11. https://doi.org/10.1007/BF00984744
Kirschner J (1993) Taxonomic survey of Luzula sect. Luzula (Juncaceae) in Europe. Folia Geobot Phytotax 28:141–182. https://doi.org/10.1007/BF02854853
Kirschner J (1994) Luzula. In: Fischer M (ed) Exkursionsflora von Österreich. Ulmer Verlag, Wien, pp 1080–1083
Kirschner J (1995) Allozyme analysis of Luzula sect. Luzula (Juncaceae) in Ireland: evidence of the origin of tetraploids. Folia Geobot Phytotax 30:283–290. https://doi.org/10.1007/BF02803711
Kirschner J (1996) Luzula multiflora subsp. hibernica, a new tetraploid of Luzula sect. Luzula (Juncaceae) from Ireland. Watsonia 21:89–97
Kirschner J (2002) Luzula. In: Kirschner J (ed) Juncaceae 1: Rostkovia to Luzula, Species Plantarum: Flora of the World Part 6. National Library of Australia, Canberra, pp 18–188
Kirschner J, Engelskjøn T, Knaben GS (1988) Luzula alpina Hoppe, a neglected Alpine species. Preslia 60:97–108
Kolodin P, Cempírková H, Bureš P, Horová L, Veleba A, Francová J, Adamec L, Zedek F (2018) Holocentric chromosomes may be an apomorphy of Droseraceae. Plant Syst Evol 304:1289–1296. https://doi.org/10.1007/s00606-018-1546-8
Kuta E, Bohanec B, Dubas E, Vižintin L, Przywara L (2004) Chromosome and nuclear DNA study on Luzula: a genus with holokinetic chromosomes. Genome 47:1–11. https://doi.org/10.1139/g03-121
Leitch IJ, Bennett MD (2004) Genome downsizing in polyploidy plants. Biol J Linn Soc 82:651–663. https://doi.org/10.1111/j.1095-8312.2004.00349.x
Löve A, Löve D, Raymond M (1957) Cytotaxonomy of Carex section Capillares. Can J Bot 35:715–761
Luceño M, Guerra M (1996) Numerical variations in species exhibiting holocentric chromosomes: a nomenclatural proposal. Caryologia 49:301–309
Malheiros N, Gardé A (1950) Fragmentation as a possible evolutionary process in the genus Luzula DC. Genet Iber 2:257–262
Mello-Sampayo T (1961) Differential polyteny and karyotype evolution in ‘Luzula’, a critical interpretation of morphological and cytophotometric data. Genet Iber 13:1–22
Melters DP, Paliulis LV, Korf IF, Chan SWL (2012) Holocentric chromosomes: convergent evolution, meiotic adaptations, and genomic analysis. Chromosome Res 20:579–593
Melzer H (1982) Neues zur Gefäßpflanzenflora Kärntens. Carinthia II 172(92):241–252
Melzer H (1996) Neues zur Flora von Steiermark, XXXIV. Mitt Naturwiss Vereins Steiermark 125:121–136
Merxmüller H (1952) Untersuchungen zur Sippengliederung und Arealbildung in den Alpen. Teil 1. Jahrb Ver Schutze Alpenpflanzen -tiere 17:96–133
Merxmüller H (1953) Untersuchungen zur Sippengliederung und Arealbildung in den Alpen. Teil 2. Jahrb Ver Schutze Alpenpflanzen -tiere 18:135–158
Merxmüller H (1954) Untersuchungen zur Sippengliederung und Arealbildung in den Alpen. Teil 3. Jahrb Ver Schutze Alpenpflanzen -tiere 19:97–139
Möbus G (1997) Geologie der Alpen. Sven von Loga, Köln
Nagaki K, Kashihara K, Murata M (2005) Visualization of diffuse centromeres with centromere-specific histone H3 in the holocentric plant Luzula nivea. Plant Cell 17:1886–1893. https://doi.org/10.1105/tpc.105.032961
Nishikawa K, Furuta Y, Ishitobi K (1984) Chromosomal evolution in genus Carex as viewed from nuclear DNA content, with special reference to its aneuploidy. Jpn J Genet 59:465–472
Nordenskiöld H (1951) Cyto-taxonomical studies in the genus Luzula. I. Somatic chromosomes and chromosome numbers. Hereditas 37:325–355. https://doi.org/10.1111/j.1601-5223.1951.tb02898.x
Nordenskiöld H (1956) Cyto-taxonomical studies in the genus Luzula. II. Hybridisation experiments in the Luzula campestris-multiflora complex. Hereditas 42:7–73. https://doi.org/10.1111/j.1601-5223.1956.tb03011.x
Pachschwöll C, Escobar García P, Winkler M, Schneeweiss GM, Schönswetter P (2015) Polyploidisation and geographic differentiation drive diversification in a European high mountain plant group (Doronicum clusii aggregate, Asteraceae). PLoS ONE 10:e0118197. https://doi.org/10.1371/journal.pone.0118197
Pawłowski B (1970) Remarques sur l’endémisme dans la flore des Alpes et des Carpates. Vegetatio 21:181–243
Pazy B, Plitmann U (1995) Chromosome divergence in the genus Cuscuta and its systematic implications. Caryologia 48:173–180
Pignatti S (2017) 2. Luzula Lam. et DC. In: Pignatti S (ed) Flora d’Italia vol 1, 2nd edn. Edagricole-New Business Media, Bologna, pp 407–417
Polatschek A (2001) Flora von Nordtirol, Osttirol und Vorarlberg 4. Wolfgang Meighörner, Innsbruck
Renny-Byfield S, Kovarik A, Kelly LJ, Macas J, Novak P, Chase MW, Nichols RA, Pancholi MR, Grandbastien MA, Leitch AR (2013) Diploidization and genome size change in allopolyploids is associated with differential dynamics of low- and high-copy sequences. Plant J 74:829–839. https://doi.org/10.1111/tpj.12168
Roalson E, McCubbin AG, Whitkus R (2007) Chromosome evolution in Cyperales. In: Columbus JT, Friar EA, Porter JM, Prince LM, Simpson MG (eds) Monocots: comparative biology and evolution of Poales. Allen, Claremont, pp 62–71
Schönswetter P, Stehlik I, Holderegger R, Tribsch A (2005) Molecular evidence for glacial refugia of mountain plants in the European Alps. Mol Ecol 14:3547–3555. https://doi.org/10.1111/j.1365-294X.2005.02683.x
Schönswetter P, Suda J, Popp M, Weiss-Schneeweiss H, Brochmann C (2007) Circumpolar phylogeography of Juncus biglumis (Juncaceae) inferred from AFLP fingerprints, cpDNA sequences, nuclear DNA content and chromosome numbers. Mol Phylogenet Evol 42:92–103. https://doi.org/10.1016/j.ympev.2006.06.016
Soltis DE, Albert VA, Leebens-Mack J, Bell CD, Paterson AH, Zheng C, Sankoff D, Depamphilis CW, Wall PK, Soltis PS (2009) Polyploidy and angiosperm diversification. Am J Bot 96:336–348. https://doi.org/10.3732/ajb.0800079
Taberlet P, Zimmermann NE, Englisch T, Tribsch A, Holderegger R, Alvarez N, Niklfeld H, Coldea G, Mirek Z, Moilanen A, Ahlmer W, Marsan PA, Bona E, Bovio M, Choler P, Cieślak E, Colli L, Cristea V, Dalmas JP, Frajman B, Garraud L, Gaudeul M, Gielly L, Gutermann W, Jogan N, Kagalo AA, Korbecka G, Küpfer P, Lequette B, Letz DR, Manel S, Mansion G, Marhold K, Martini F, Negrini R, Niño F, Paun O, Pellecchia M, Perico G, Piękoś-Mirkowa H, Prosser F, Puşcaş M, Ronikier M, Scheuerer M, Schneeweiss GM, Schönswetter P, Schratt-Ehrendorfer L, Schüpfer F, Selvaggi A, Steinmann K, Thiel-Egenter C, van Loo M, Winkler M, Wohlgemuth T, Wraber T, Gugerli F, IntraBioDiv Consortium (2016) Genetic diversity in widespread species is not congruent with species richness in alpine plant communities. Ecol Lett 15:1439–1448
Tanaka T, Tanaka T (1977) Chromosome studies in Chionographis (Liliaceae): 1. On the holokinetic nature of chromosomes in Chionographis japonica Maxim. Cytologia 42:753–763
Thiers B (2016) Index Herbariorum: a global directory of public herbaria and associated staff. New York Botanical Garden’s Virtual Herbarium. http://sweetgum.nybg.org/science/ih/
Tribsch A (2004) Areas of endemism of vascular plants in the eastern Alps in relation to Pleistocene glaciation. J Biogeogr 31:747–760. https://doi.org/10.1111/j.1365-2699.2004.01065.x
Tribsch A, Schönswetter P (2003) Patterns of endemism and comparative phylogeography confirm palaeoenvironmental evidence for Pleistocene refugia in the eastern Alps. Taxon 52:477–497
Van Husen D (1987) Die Ostalpen in den Eiszeiten. Geologische Bundesanstalt, Wien
Verma SC, Rees H (1974) Nuclear DNA and evolution of allotetraploid Brassicaceae. Heredity 33:61–68. https://doi.org/10.1038/hdy.1974.64
Vilhar B, Greilhuber J, Dolenc Koce J, Temsch EM, Dermastia M (2001) Plant genome size measurement with DNA image cytometry. Ann Bot 87:719–728. https://doi.org/10.1006/anbo.2001.1394
Weiss-Schneeweiss H, Emadzade K, Jang T-S, Schneeweiss GM (2013) Evolutionary consequences, constraints and potential of polyploidy in plants. Cytogenet Genome Res 140:137–150. https://doi.org/10.1159/000351727
Záveská Drábková L (2013) A survey of karyological phenomena in the Juncaceae with emphasis on chromosome number variation and evolution. Bot Rev 79:401–446. https://doi.org/10.1007/s12229-013-9127-6
Zedek F, Bureš P (2018) Holocentric chromosomes: from tolerance to fragmentation to colonization of the land. Ann Bot 121:9–16. https://doi.org/10.1093/aob/mcx118
Zedek F, Šmerda J, Šmarda P, Bureš P (2010) Correlated evolution of LTR retrotransposons and genome size in the genus Eleocharis. BMC Plant Biol 10:e265
Acknowledgements
Open access funding provided by University of Innsbruck and Medical University of Innsbruck. We thank W. Gutermann, C. Argenti, M. Bovio, F. Prosser and F. Martini for herbarium material, P. Schönswetter, Š. Novak, M. Turjak, D. Kutnjak, I. Dakskobler and S. Strgulc Krajšek for collecting Luzula plants and seeds, and A. Čurman for genome size measurements. M. Magauer and M. Nolf helped with production of some figures. H. Niklfeld provided distribution data for Austria and P. Schönswetter contributed valuable comments on an earlier version of the manuscript. We thank two reviewers and the editor J. Stöcklin for their constructive feedback.
Funding
This study was financially supported by the Slovenian Research Agency (research programme Plant biology P1-0212) and a bilateral project “Scientific and Technological Cooperation Austria-Slovenia” (ICM, OeAD-GmbH, SI09/2018 to BF; Slovenian Research Agency; BI-AT/18-19-029 to JDK).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The authors declare that no ethical approval was needed for this research.
Informed consent
No humans or animals have been a subject of this research.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Bačič, T., Dolenc Koce, J. & Frajman, B. Diversification and distribution patterns of Luzula sect. Luzula (Juncaceae) in the Eastern Alps: a cytogenetic approach combined with extensive herbarium revisions. Alp Botany 129, 149–161 (2019). https://doi.org/10.1007/s00035-019-00219-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00035-019-00219-1