Abstract
To inform the management of wild fish populations, it is equally important to understand both the ecological connectivity of habitat patches, apparent at annual and seasonal scales, and the genetic connectivity, emerging at evolutionary scales across generations. Ecological connectivity indicates the potential for rapid recolonization upon local depletion, while genetic connectivity informs about the conservation needs related to the evolution of subpopulations and ecotypes in metapopulations. We combined acoustic biotelemetry and pooled-genome sequencing to study a northern pike (Esox lucius) population as a model of a freshwater piscivore that inhabits a network of shallow brackish lagoons in the southern Baltic Sea. We found limited ecological connectivity among genetically similar subpopulations of pike, suggesting a metapopulation structure characterized by discrete local subpopulations with infrequent migrations between them. Connectivity of different lagoons increased during spawning, suggesting directed spawning migrations to either freshwater rivers or low salinity patches in connected lake-like bays. Spawning site fidelity to either brackish or freshwater spawning sites was observed, further contributing to the reproductive isolation of certain subpopulations. The genetic population structure aligned with salinity gradients and geographical distance and was significant between pairs of rivers draining into the lagoon network, but it was unrelated to ecological connectivity. The results collectively suggest that local subpopulations may not rapidly replenish upon local depletion and that even weak connectivity among subpopulations was sufficient to maintain genetic homogeneity across lagoons with similar salinity levels. Effective management and conservation of species forming metapopulations, such as the coastal northern pike studied here, necessitate localized approaches that adapt fishing mortality to local abundance and promote access to specific habitats, especially rivers, during spawning to conserve the entire genetic biodiversity and foster resilience of the metapopulation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Connectivity is a critical determinant of population dynamics, genetic differentiation, and biodiversity conservation because it affects key processes such as migration and dispersal, population growth, gene flow, local adaptation, and ultimately population resilience (Schindler et al. 2010; Luque et al. 2012; Kool et al. 2013). Although high connectivity can also lead to negative demographic consequences in some circumstances, such as through the rapid spread of disease (e.g., Borg et al. 2017) or the facilitation of natural predation affecting source-sink dynamics (e.g., Olin et al. 2024), in general, it has been shown to mitigate local and regional ecological perturbations and overexploitation, e.g., by increasing overall population resilience and allowing negatively affected areas to be repopulated in cases of localized extinctions (Hilborn et al. 2003; Gido et al. 2015).
In large aquatic systems, subpopulations of fish (defined as behaviorally and genetically differentiated groups forming their own reproductive units) are connected either via the passive dispersal of eggs and larvae or via the active movement of juveniles and adults (Brown et al. 2016). While passive dispersal is often studied using hydrodynamic models predicting particle movement (Palmas et al. 2017), for monitoring the active movement of juveniles or adults, telemetry offers a suitable toolbox to determine exchange processes, provided that the spatial scale of study is tractable (Matley et al. 2022). The ultimate outcome of both passive and active dispersal can also be inferred from genetic techniques that assess the differentiation or relatedness among subpopulations and reveal the heritable consequences of dispersal (Riginos et al. 2014).
Population genetic approaches primarily give an insight into genetic population structure resulting from demographic effects, genetic drift, and selection. For example, the number of dispersers between discrete local subpopulations should directly influence the extent to which gene flow affects population structure over generational timescales (Lowe and Allendorf 2010; Cayuela et al. 2018). Although patterns of genetic connectivity emerging from population genetic analyses are fundamental for delimiting the stocks and identifying evolutionarily significant management units (Palsbøll et al. 2007; Hawkins et al. 2016), such techniques are not always well aligned to capture ecological connectivity among habitats at year-to-year or seasonal scales (Lowe and Allendorf 2010; Hawkins et al. 2016). In contrast to genetic connectivity, which provides information on the degree to which gene flow affects evolutionary processes over generational scales (Hedgecock et al. 2007; Lowe and Allendorf 2010), ecological connectivity is of central importance for shorter-term ecological and fishery dynamics, such as population growth and vital rates influenced by dispersal as well as local abundance (Nichols et al. 2000; Runge et al. 2006). Decrease or interruption in ecological connectivity may not have an immediate effect on population genetic structure (Marandel et al. 2018), yet it is highly relevant to local management decisions because it affects, for instance, the risk of localized overfishing, which may be overlooked when solely long-term evolutionary outcomes are considered (Hawkins et al. 2016). That is because even very small levels of exchange may contribute enough gene flow so that the subpopulations in different habitats remain genetically homogeneous (e.g., Cowen et al 2007; Hawkins et al. 2016; Cayuela et al. 2018). For example, in a metapopulation, defined as an assemblage of discrete local groups with limited dispersal between them (Hanski and Simberloff 1997), ecological connectivity may be sufficient to maintain genetic panmixia, but low enough to render subpopulations largely demographically independent, which may inhibit the rapid recovery of subpopulations when a local mortality event occurs (Hawkins et al. 2016; Olin et al. 2024). Safeguarding biocomplexity in such weakly connected metapopulations, which incorporates the diversity of spawning strategies and other behavioral adjustments in animals living in complex ecological systems, has been shown to be critical for achieving long-term stability and high productivity (Hilborn et al. 2003; Schindler et al. 2010).
Combining methods that track ecological (on year-to-year or seasonal time scales) and genetic connectivity (on generational evolutionary time scales) can play an important role in fully characterizing the degree of population connectivity and in tailoring appropriate management and conservation recommendations to different objectives (Lowe and Allendorf 2010; Travis et al. 2012; Hawkins et al. 2016). Over time, telemetry researchers and evolutionary geneticists have independently developed increasingly fine-tuned methods (see, e.g., Matley et al. (2022); Nathan et al. (2022) for telemetry; and Benestan (2020); Hohenlohe et al. (2021) for genetics), however, both communities remain largely siloed (but see Müller et al. (2023) for a review of the growing body of work that integrates both data types). Our work combines these two tool sets by integrating and linking behavioral and genetic data to analyze population structure and understand the complex role of dispersal and connectivity on ecological and evolutionary time scales in a coastal population of the freshwater piscivore northern pike (Esox lucius, hereafter pike) in the southern Baltic Sea, which has colonized a network of spatially vast brackish lagoons into which a set of rivers drain. Based on an improved understanding of the behavior and genetic population structure, we derive implications for management and conservation of pike in brackish lagoons in the southern Baltic Sea.
The Baltic Sea is one of the world’s largest brackish water bodies, characterized by a strong salinity gradient from sea salinity (30 practical salinity units, PSU) in its western part, connected to the North Sea, to almost fresh water (2 PSU) in the northeast (Schubert et al. 2017). In addition, at local levels, salinity gradients are pronounced, especially in nearshore coastal areas, such as, for example, a lagoon network around the island of Rügen, Germany, in the southern Baltic Sea, where salinities in different lagoons can range from almost freshwater oligohaline (< 5 PSU) to mesohaline conditions (< 12 PSU) (Arlinghaus et al. 2023b). These regional and local ecological gradients in salinity have shaped a unique species assemblage, comprising marine, estuarine, and freshwater species (Wennerström et al. 2013). Pike, a large-sized predator typical of freshwaters in the northern hemisphere (Craig 2008), is widely distributed throughout nearshore coastal waters of the Baltic Sea, where salinity does not exceed 18 PSU (Dahl 1961), taking advantage of an abundant foraging environment that provides access to energy-rich marine prey like Atlantic herring (Clupea harengus) (Winkler 1987).
For freshwater fish, successful survival at these salinity levels requires either evolutionary physiological adaptations allowing them to complete their life cycle in brackish water (Jacobsen et al. 2017) or development of behavioral traits, such as anadromy, that allows them to forage in brackish areas while continuing to spawn in the adjacent freshwater habitats (Engstedt et al. 2010; Ferguson et al. 2019; Aguirre et al. 2022). Migration during the spawning time can contribute to the reproductive isolation of subpopulations, for example, by affecting the timing of spawning among different groups (isolation by time, e.g., Brannon et al. 2004) or their spawning site preferences (isolation by location, e.g., Neville et al. 2006; Sunde et al. 2022). The evolution of migration tendencies also contributes to isolation-by-environment, e.g., through physiological adaptation to spawn in different salinity levels in pike (Sunde et al. 2022). All these spatiotemporally varying processes might lead to intraspecific differentiation along the ecological gradients, such as salinity, or various habitat patches, where in some species and particular situations, a continuum of genetically differentiated ecotypes and/or life-history strategies will be expressed and co-exist (Clemens and Schreck 2021; Stronen et al. 2022).
Earlier studies have shown that Baltic pike have developed three distinct reproductive strategies to successfully spawn in varying salinity levels. Part of the population has undergone local adaptation and can carry out their complete life cycle, including reproduction, in brackish conditions up to 10 PSU (Jørgensen etal 2010; Sunde et al. 2022). Other individuals undertake anadromous spawning migrations from brackish feeding grounds to freshwater rivers and wetlands (Engstedt et al. 2010; Sunde et al. 2019; Roser et al. 2023), while some fully reside in freshwater rivers throughout the year, making only occasional forays into brackish areas (Birnie-Gauvin et al. 2019). In addition, natal homing and spawning-site fidelity, mechanisms that contribute to reproductive isolation, are common in pike (Miller et al. 2001; Bosworth and Farrell 2006), and both have been reported in Baltic pike (Diaz-Suarez et al. 2022; Engstedt et al. 2014; Nordahl et al. 2019).
The presence of genetically differentiated ecotypes with different reproductive strategies may have a strong influence on both genetic and ecological connectivity within the pike population. As there is no potential for dispersal in pike during the adhesive egg and larval stages (Bry 1996), pike dispersal is based solely on the movements of juveniles and adults. However, pike is classically described as a sedentary, phytophilic ambush predator that has a rather small home range outside spawning time (Diana et al. 1977; Kobler et al. 2008; Craig 2008), although some studies in freshwater lakes showed that some individuals can be quite mobile and utilize all available habitats (Haugen et al. 2006). In the Baltic Sea, mark-recapture (Karås and Lehtonen 1993) and acoustic telemetry (Jacobsen et al. 2017; Flink et al. 2023; Dhellemmes et al. 2023b) studies showed relatively stationary behavior and rather small home ranges in coastal pike. However, during the spawning period, which usually takes place from March to May depending on latitude, Baltic pike exhibit increased mobility as they seek to reach the spawning grounds either in freshwater rivers (Tibblin et al. 2016) or in brackish lagoons (Flink et al. 2023). This suggests that subpopulations mix in various combinations throughout the year: brackish water residents and anadromous fish intermingle in foraging habitats but separate during spawning as anadromous fish move to freshwater habitats where they, in turn, share space with resident freshwater pike. Thus, on the one hand, the general sedentary lifestyle of pike suggests that ecological connectivity between parts of the metapopulation may be low, potentially fostering adaptive divergence on small geographic scales of a few kilometers due to limited exchange between groups residing in different coastal sites (Nordahl et al. 2019). However, on the other hand, the occasional bursts of movements during spawning may connect sites that are otherwise disconnected and potentially contribute to a gene flow among various subpopulations in different connected patches (e.g., lagoons of similar salinity levels; Möller et al. 2021).
Genetic research on pike across the Baltic Sea showed population structuring shaped by pattern of isolation by distance, where geographically close subpopulations are more similar than geographically distant ones (Laikre et al. 2005; Wennerström et al. 2017). Such patterns can be explained by the phytophilic, macrophyte-bound pike having low movement activity and limited dispersal outside vegetated nearshore areas (Sunde et al. 2022). There is also genetic evidence for differences between sympatric anadromous and brackish pike ecotypes, likely in response to physiological salinity adaptations and/or natal homing and site fidelity to different rivers (Nordahl et al. 2019; Sunde et al. 2022), which was also supported by otolith microchemical analyses (Engstedt et al. 2010; Möller et al. 2019). Similarly, studies in the coastal lagoons of the southern Baltic Sea, our study area, showed the influence of salinity on genetic structure, with pike in certain oligohaline lagoons differing from pike in nearby mesohaline lagoons, which, in the absence of physical barriers between these areas, suggests that physiological reasons, i.e., adaptation to salinity differences among lagoons, may be limiting gene flow between them (Möller et al. 2021; Roser et al. 2023). Furthermore, Roser et al. (2023) demonstrated the occurrence of freshwater spawning activity in our study system and showed that putative anadromous subpopulations appear to be genetically intermediate between mesohaline brackish and freshwater or oligohaline brackish stocks.
Substantial declines in pike abundances have been documented in many Baltic coastal areas in recent decades (van Gemert et al. 2022; Bergström et al. 2022; Olsson et al. 2023), calling for well-informed management actions. To contribute to the understanding and conservation of these declining stocks, our study focused on addressing the following key questions:
-
1.
Does the sedentary lifestyle of lagoon pike cause low ecological connectivity between parts of the population, increasing the potential for local overfishing?
-
2.
Do spawning migrations enhance both ecological and genetic connectivity, or do they rather promote reproductive isolation and population differentiation through fostering behaviorally differentiated ecotypes such as brackish residents, freshwater residents, and anadromous fish?
-
3.
Do patterns of space use and reproductive behavior of pike align with the overall genetic structure of the population, or do environmental factors such as salinity gradients and geographic distances have a strong influence on current genetic differentiation patterns?
We hypothesized that the pike population in the study area around the island of Rügen, Germany (see Arlinghaus et al. 2023b for a full review) is (H1) composed of several subpopulations with relatively stationary space use and low ecological connectivity among them, (H2) shows spawning site fidelity and behaviorally differentiated ecotypes, and (H3) its genetic structure is driven both by limited ecological connectivity and by environmental factors such as salinity gradients and geographic distances.
Methods
Study area
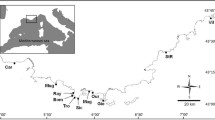
The study area comprises the network of interconnected coastal lagoons (locally known as Bodden) bordered by the islands of Fischland-Darß-Zingst, Hiddensee, Rügen and Usedom in the southern Baltic Sea (54.41N, 13.37E; area covered by our sampling approximately 1200 km2, total area approximately 2000 km2; Fig. 1). As a result of geographical characteristics of the region (e.g., varying patterns of land barriers between lagoons and open Baltic Sea and freshwater discharge from rivers), these lagoons exhibit significant hydrochemical variability, including salinity, water temperature, Secchi depth, and nutrient concentrations (supplementary materials, Table S1) (Arlinghaus et al. 2023b). The most pronounced is the salinity gradient, with higher salinity levels in the northwest mesohaline areas (e.g., in Vitter, Schaproder, and Kubitzer Bodden) and lower salinity levels in the southeast oligohaline lagoons (Peenestrom and Stettiner Haff). Our study site also comprises several freshwater rivers, the most important of which are the Barthe (west) and Peene (east), as well as several small rivers such as the Sehrowbach and Duwenbeek (Fig. 1).
Map of the study area displaying the positions of the acoustic telemetry receivers and fish tagging sites. Full names of the areas (freshwater in blue font): Barthe Barthe river, BAT Barther Bodden, BRG Breeger Bodden, BRT Breetzer Bodden, Duwenbeek Duwenbeek river, Gate GB-BS Gate between Greifswalder Bodden and open Baltic, Gate GB-P Gate between Greifswalder Bodden and Peenestrom, Gate G-KB Gate between Grabow and Kubitzer Bodden, Gate KB-S Gate between Kubitzer Bodden and Strelasund, Gate KSB-BS Gate between Schaproder/Kubitzer Bodden and open Baltic Sea, Gate P-SH Gate between Peenestrom and Stettiner Haff, Gate S-GB Gate between Strelasund and Greifswalder Bodden, Gate VB-BS Gate between Vitter Bodden and open Baltic Sea, Gate VB-WB Gate between Vitter Bodden and Wieker Bodden, GB Greifswalder Bodden (South), GJB Großer Jasmunder Bodden, Grabow Grabow, KB Kubitzer Bodden, KJB Kleiner Jasmunder Bodden, Landowbach Landowbach river, P Peenestrom, Peene Peene river, RB Rügischer Bodden, Ryck Ryck river, S Strelasund, SAB Saaler Bodden, SB Schaproder Bodden, Sehrowbach Sehrowbach river, VB Vitter Bodden, WB Wieker Bodden
Acoustic telemetry
To study ecological connectivity via pike migration and dispersal across the study area, a total of 389 adult pike (mean total length = 76.9 ± SD 12.4 cm; female, 226; male, 162; unknown, 1) were tagged in February–March 2020 (N = 301), November–December 2020 (N = 17), March–April 2021 (N = 63), and February 2022 (N = 8) (for tagging locations, see Fig. 1). The sampling methods included rod and reel fishing, fyke nets, gillnets, and electrofishing. Pike were fitted with Lotek acoustic transmitters (N = 120, MM-R-16 50 HP, approximately 6-year battery life, dry weight = 35 g, in-water weight = 18.9 g; N = 245, MM-R-16 33 HP, approximately 3.5-year battery life, dry weight = 26.7 g, in-water weight = 13.6 g, random pulse rate: 60–180 s, frequency = 69 kHz, Lotek Wireless Inc., ON, Canada; 24 transmitters were redeployed; the probability of detection by receivers was independent of tag type, see supplementary materials). Tagging locations covered all large lagoons, specifically Grabow, Schaproder and Kubitzer Bodden, Grosser Jasmunder Bodden, Strelasund, Greifwalder Bodden, and Peenestrom (Fig. 1). Some fish were tagged just before the spawning time in obstruction-free rivers (Barthe, Peene, Duwenbeek, and Sehrowbach; Fig. 1) assuming these represented migratory ecotypes moving back after spawning to brackish feeding grounds and potentially returning to freshwater bodies during the next spawning (i.e., anadromous ecotype) (Roser et al. 2023).
Pike movements were monitored for 3 years (March 2020–March 2023), using an array of 140 acoustic receivers (Vemco VR2Tx, frequency: 69kHz, MAP-113, Innovasea Systems Inc., Massachusetts, U.S.A.) deployed at 146 different locations across the study area. The receiver array covered the areas important to pike fisheries (Arlinghaus et al. 2023a), with higher receiver density in the western and northern lagoons and lower density in the Greifswalder Bodden and Peenestrom (Fig. 1). The receivers had mostly non-overlapping detection ranges, as the array was designed to monitor the broad movement ranges and connectivity between the areas of interest. In narrow links between the lagoons, a “gate” format with denser receiver deployment allowed for more focused monitoring of pike movements between the adjacent lagoons (Fig. 1). The receiver data were downloaded, processed, and filtered once a year in winter (using R package ATfiltR (Dhellemmes et al. 2023a), see supplementary materials for details). The detection data were aggregated at the daily level so that it provided records of each fish’s daily presence or absence on the array and the list of receivers where they were detected on a given day. Additionally, the dataset was completed with tagging, recapture, and opportunistic active tracking (using a manual VR100 receiver, Innovasea Systems Inc., Massachusetts, USA) locations and dates, which were attributed to specific geographical sections as described below.
We divided the study area into 29 sections, encompassing freshwater rivers (e.g., Peene), individual lagoons (e.g., Grabow, Kubitzer Bodden), and gates (e.g., Gate between Grabow and Kubitzer Bodden), and clustered the receivers in these sections into corresponding groups (Fig. 1). The choice of sections was based on environmental differences, particularly in salinity (supplementary materials, Table S1), and on geographical designations commonly used by locals for better stakeholder relevance and facilitation of their interpretation of the results. We treated gates as separate sections due to their higher receiver density, allowing for the minimization of bias in relevant metrics and analyses (more details below).
Movement networks: movement and ecological connectivity
To evaluate movement-based ecological connectivity, we constructed networks as unipartite undirected networks in which nodes represented the distinct geographic sections (i.e., with sets of receivers grouped according to their section, Fig. 1), and edges reflected subsequent detections between the sections. Only apparent movements, i.e., consecutive detections of an individual on two different receivers, were considered to create the network. Hence, the resulting maps with spatially explicit networks showed aggregated pike movement within and between the defined sections represented by nodes. To test whether the observed patterns of movement differed from random, the observed individual networks were compared with those generated from null models produced following the method by Lea et al. (2016) (see supplementary materials for details).
To quantify the networks, three metrics were used: (1) node strength reflected the number of movements within each node and provided a measure of occupancy (Barrat et al. 2004; Lea et al. 2016); (2) edge weight showed local connectivity and the strength of connections between node pairs, calculated as the total number of internodal transits (Barrat et al. 2004; Jacoby et al. 2012); and (3) edge density described the overall connectivity within the network, representing a proportion of edges present in a network out of the total edges possible (Jacoby et al. 2012). Networks were created and analyzed using R packages igraph (Csárdi and Nepusz 2006) and circlize (Gu et al. 2014).
To ensure comparability across different sections with varying receiver coverage and unequal number of tagged pike (Fig. 1), we adjusted node and edge metrics. Node strength was weighted by receiver density and mean distances between receivers (in water, i.e., without crossing land areas, calculated using actel (Flávio and Baktoft 2021)), and by the number of fish tagged in each section. Edge weight was adjusted by distance (in water) between connected nodes to give higher value to longer transits. The network was computed monthly for the entire population, and the adjustments were applied to monthly metrics, accounting for new receiver deployments, additional pike tagging, and reported fish mortalities. Lastly, 3-year average values were calculated annually and seasonally (winter, December–February; spring, March–May; etc.).
Pike movement between brackish and freshwater habitats
We conducted an analysis of pike movement between different habitats within our study area to investigate potential differences in their habitat preferences and habitat exchange behaviors, particularly between brackish and freshwater environments. This was done to investigate whether pike tagged in these different habitats show preferences for staying in them or moving to other habitats. Variations in dominant preferences per habitat type would shed light on the presence of behaviorally differentiated ecotypes in the studied pike population, in particular the presence of an anadromous ecotype. To do this, we categorized the receiver stations, as well as tagging, recapture, and active tracking locations into the habitat types: Brackish for lagoon areas, Estuary for the lagoonal areas within 1 km of a river mouth, Freshwater downstream for river sections up to 1 km from a river mouth, and Freshwater upstream for river sections beyond 1 km from a river mouth. For each pike, the location of the first capture/tagging was taken as the starting point and all transitions within and between different habitats were tracked. If no data were available for an individual after a previous observation, it was categorized as moving to the No Data category. We then summarized all documented transits to represent the movements of the entire population between habitats throughout the study period. This was visualized in R using circular plots from circlize (Gu et al. 2014).
Spawning site fidelity
To assess pike spawning site fidelity, we examined whether they were observed in the same areas during the spawning seasons (March–May) in different years (2020–2022) by checking for repeated logging at the same receiver/s. Further, we examined whether pike were tagged, recaptured, or detected by active tracking in the same area, which also indicates a return to the same spawning grounds, but more punctually. Recaptures were reported by fishers and anglers, as documented in the database set up for reports.
Population genetics and its link to ecological connectivity, geographical distance, and salinity gradients
We took advantage of already available whole-genome (pool-sequencing) data published in Roser et al. (2023), who genotyped the entire genome of animals from 11 study locations in the study area (N = 45–50 per location, in total 535 individuals; see details in supplementary materials, Fig. S2). This genomic investigation focused on pike sampled from lagoons and selected freshwater rivers, aiming to fully characterize the genetic diversity within the study region and discern patterns of population differentiation. To assess genetic differentiation among the study locations, FST was employed, a metric quantifying genetic variance among populations (Holsinger and Weir 2009). The resulting pairwise FST values indicated a distinct separation between mesohaline brackish-water sites in Bodden (e.g., Greifswalder Bodden, Barther Bodden, Schaproder/Kubitzer Bodden) and larger freshwater rivers (Barthe and Peene), as well as oligohaline lagoons (e.g., Peenestrom). Within this differentiation, putative anadromous populations in smaller rivers (e.g., Sehrowbach) exhibited a more intermediate genetic position, suggesting divergence of anadromous pike from brackish water pike as well as from populations from different freshwater sites (see Roser et al. (2023) for details; supplementary materials, Fig. S2).
The locations with both genetic and telemetry data available included mesohaline brackish-water (Barther Bodden, Kubitzer/Schaproder Bodden, Großer Jasmunder Bodden, Greifswalder Bodden), possibly resident freshwater (Barthe and Peene river), oligohaline brackish (Peenestrom), and a putative anadromous population (Sehrowbach). Another telemetry-based network analysis was conducted using only these eight areas as nodes to align the network to the genetic sampling locations (supplementary materials, Fig. S3), with edge weight used as an ecological connectivity measure.
To determine the extent to which ecological connectivity and environmental factors such as geographic distance and salinity difference correlate with the estimated levels of genetic differentiation among the key lagoons and rivers, we used partial Mantel tests using the package vegan (Oksanen et al. 2022), which allow for the control of one variable. We ran three partial Mantel tests: (1) between pairwise linear FST (i.e., \({F}_{ST}/(1-{F}_{ST})\); Rousset 1997) and pairwise ecological connectivity while controlling for geographic distance in water; (2) between pairwise linear FST and pairwise average salinity difference while controlling for geographic distance in water; and (3) between pairwise linear FST and geographic distance in water while controlling for pairwise average salinity differences.
Further, we fit a generalized linear model (GLM) using the package glmmTMB (Brooks et al. 2023) to compare the relative effect of each of the following variables: ecological connectivity (continuous: edge weight), salinity (continuous: PSU), presence of freshwater habitat in a pair (categorical: none, one, or both), and geographic distance (continuous: km). Linear FST was used as the response variable, and a beta distribution was used as FST values ranged between 0 and 1 (Nurbaev and Balanovskaia 1998). We included the identity of one location in the pair as a random effect in the model to account for the potential impact of our telemetry design on the results (e.g., different receiver coverage per area). We identified the explanatory variables that improved model fit by comparing the Akaike information criterion (AIC) of the full model with the AIC of stepwise simplified models using the MASS package with a two-unit difference of AIC indicating significantly different fit between the models (Venables and Ripley 2002; Burnham and Anderson 2004). The fit of the obtained most parsimonious model was assessed using the uniformity, dispersion, and outliers tests in the DHARMa package (Hartig and Lohse 2022). Fit was considered appropriate if no tests were significant.
For the Mantel tests and GLM, geographic distances and salinity differences among pairs of sites were calculated as follows: distances were calculated as the mean distance in water between all pairs of receivers located in different areas, and salinity differences were calculated on the basis of estimates presented in Arlinghaus et al. (2023b) (Table S1). All the continuous fixed effects were mean-centered, and their standard deviation was set to 1 to allow for a direct comparison of effect strength in the models.
All data handling and analysis were performed in R (R Core Team 2023), except for the random network calculations, which were done in Python 3.8.10 via the Anaconda 3 distribution (see supplementary materials for details).
Results
Descriptive information
Out of 389 tagged pike, 343 (88%) were detected on 138 receiver stations (out of the original 146 locations, 13 receivers were lost and 5 moved in 2021, Fig. 1). A total of 47 individuals (12%) were never detected, with the largest proportion tagged in the Peene river (40%) and Sehrowbach river (33%), and 54 individuals (14%) were detected by only one receiver during the entire study period, and it is not clear whether they resided in the respective areas, were not moving because they died, or had equipment malfunction (they were included in the analysis nevertheless). Overall, a total of 8,041,130 detections were recorded between March 2020 and March 2023. After removing duplicates, the dataset comprised 4,318,623 detections. Out of these, 399,337 (9%) indicated movements, regarded as consecutive logs of an individual on distinct receivers, while all other records were at unchanging locations, pointing to the predominantly sedentary behavior and site fidelity of pike. The testing of the individual pikes’ networks against those created with random walks showed that pike movements were nonrandom, as the edge density of the data and random networks were significantly different (Wilcoxon one-sample signed rank test, p-value < 0.05).
Movement and ecological connectivity
The movement network structure showed that pike movement linked most of the lagoons and freshwater rivers in the study area as far as geography allowed, but all these links, as well as the overall level of ecological connectivity, were weak (Fig. 2A, B). Most of the movements were local, occurring within selected (mostly original tagging) lagoons and freshwater rivers, where 75% of the transitions were recorded (Fig. 2A, B). When exchange levels were higher, they mainly happened between lagoons and nearby gates, still reflecting rather local movements [e.g., Kubitzer Bodden (KB) and Gate between Kubitzer Bodden and Strelasund (Gate KB-S), Fig. 2B.]
Ecological connectivity. (A) Map displaying the network structure of the pike movements, where node size corresponds to the total number of detections within each area and edge color represents the frequency of internodal movements. (B) Circular plot displaying proportions of movements within the areas and between the pairs of areas (the graph displays apparent movements, i.e., consecutive detections on two different receivers). (C) Monthly dynamics of overall ecological connectivity within the study system, represented by edge density network metric. See full names of the areas in Fig. 1
Strong seasonal differences in pike movement and connectivity were observed, with peak connectivity between the sections of the study area in spring, corresponding to the well-known spawning time of pike (Fig. 2C; Fig. S4). Connectivity was notably lower during other seasons, reaching its minimum in winter, indicating very low movement between the sections during this time (Fig. 2C; Fig. S4).
In spring, despite the movement network displaying high connectivity, with most regions linked by pike movement (Appendix, Fig. S4, B), some sections, such as lagoons Greifswalder Bodden (GB) and Peenestrom (Gate GB-P and P), showed no exchange movements despite being geographically close. Similarly, and only a small increase in movement was recorded between Rügischer Bodden (RB, northern Greifswalder Bodden) and Peenestrom (Gate GB-P and P) (Fig. S4, B). These two lagoons thus remained weakly connected to each other throughout the year despite their geographic proximity (Fig. S4).
Pike movement between brackish and freshwater habitats
The analysis of the movement between different habitats revealed distinct behavioral patterns among groups of pike, pointing at their differentiating ecotypes and spawning strategies. Pike tagged in brackish water tended to remain in brackish habitats. Most of their movement occurred either within brackish waters (53%) or between brackish waters and estuaries (40% of all movements to and from brackish habitats, Fig. 3). Correspondingly, movements to and from estuaries were mostly connected to brackish habitats (53%) or happened entirely within estuaries (27% out of all movements to and from estuaries, Fig. 3).
Pike movements between different habitat types: brackish—lagoon areas, estuary—lagoonal areas within 1 km of a river mouth, freshwater downstream—river sections up to 1 km from a river mouth, freshwater upstream—river sections beyond 1 km from a river mouth. The graph shows observed pike transits from (colored bars), to, and within the habitats, summarized to represent the entire population’s movements. If data on an individual were unavailable after the previous observation, it was categorized as moving to No data.
Pike tagged in freshwater rivers showed overall more diversity in habitat use. Individuals originating from upstream sections of larger rivers Barthe and Peene primarily remained in freshwater habitats (40% of movements remained upstream, and 22% happened between upstream and downstream freshwater habitats). Some of these fish ventured to estuaries (24% out of all movements to and from upstream freshwater habitats), but very few entered brackish lagoons (2%, Fig. 3).
In contrast, individuals tagged or observed in downstream sections of rivers and specifically in the small rivers Sehrowbach and Duwenbeek were frequently found in estuaries (47% out of all movements to and from downstream freshwater habitats) and brackish lagoons (12%, Fig. 3). This suggests that these pike have a greater tolerance for salinity variations and may represent an ecotype that uses both freshwater and brackish water habitats.
Spawning site fidelity
A total of 369 individuals were observed (tagged, recaptured, recorded by receivers or by active tracking) in at least one spawning season (March–May). Among these, almost half (N = 151, 43%) were only seen in one spawning season (32 of them were known to have died due to fishing harvest or naturally). The remaining half (N = 208, 56%) had records available for two (N = 143) or all three (N = 65) spawning seasons (March–May 2020–2022), which allowed for analyses of spawning site fidelity.
Pike showed strong spawning site fidelity: out of the 208 individuals observed over multiple spawning seasons, the vast majority (97%, N = 201) were found in the same section of the study area (lagoon, freshwater river, or gate) in at least two spawning seasons in 3 years of observations. Among them, only a third (29%, N = 59) were documented to have left the section outside of spawning, indicating a return migration to the spawning site, and 85% (N = 178) were detected at the same receiver in at least two spawning seasons, meaning that they used the exact geographical locations within the respective section. Among those, 80% (N = 142) did not visit exactly the same receivers outside of the spawning season.
Population genetics and its link to ecological connectivity, geographical distance, and salinity gradients
The partial Mantel tests revealed no significant effect of (movement-based) ecological connectivity on pairwise linear FST (r = −0.032; p = 0.56; 9999 permutations, Fig. 4A), regarded as a measure of genetic connectivity. Furthermore, there were significant effects of salinity differences among sites (r = 0.681; p = 0.004; 9999 permutations, Fig. 4C), indicating that populations differed more in their genotypes when they were from sites with stronger salinity differences. Both tests were controlled for geographic distance in water. The partial Mantel tests between the linear FST and geographic distance while controlling for salinity differences also showed a significant correlation (r = 0.749; p = 0.008; 9999 permutations), indicating that larger geographic distance was associated with greater degrees of genetic population differentiation (isolation by distance, Fig. 4B).
Effects of ecological connectivity, geographic distance, salinity difference, and the presence of the freshwater in a pair on genetic distance (pairwise linear FST). Shaded areas and error bars indicate 95% confidence intervals. All continuous explanatory variables are z-scaled to facilitate comparison of slopes. Green titles: results of the partial Mantel tests; in purple: GLM model fit for the predictors included into the most parsimonious model.
The most parsimonious generalized linear model also included geographic distance and salinity difference as well as presence of a freshwater site in a pair, all revealing significant associations with genetic distance (pairwise linear FST) except for when only one of the areas in a pair was of freshwater category (Table 1, Fig. 4). This suggests that genetic differentiation is increased among pairs of rivers. Salinity difference had the strongest relative effect on genetic distance, followed by geographic distance, and the differentiation in pairs that contained two freshwater sites. By contrast, including ecological connectivity did not improve the model fit (supplementary materials, Table S2), suggesting no influence of movement-based ecological connectivity on genetic connectivity in the lagoon pike population.
Discussion
A solid understanding of population structure and the complex role of dispersal and connectivity on evolutionary and ecological time scales greatly benefits from integrating behavioral and genetic data (Cowen et al. 2007; Lowe and Allendorf 2010; Hawkins et al. 2016; Marandel et al. 2018; Müller et al. 2023). In this study, we combined whole-genome sequencing from individual pike pooled at capture site levels (lagoons and rivers draining into the lagoon network) and associated measures of genetic connectivity with behavioral observations over 3 study years using acoustic telemetry to investigate ecological and genetic connectivity in the northern pike population inhabiting brackish lagoons surrounding the German islands of Fischland-Darß, Hiddensee, Rügen, and Usedom in the southern Baltic Sea. We found support for our first hypothesis (H1) that the study population was composed of several subpopulations with relatively stationary space use and low movement-based ecological connectivity among them, resembling a metapopulation structure. There were also indications supporting our second hypothesis (H2), suggesting potential spawning site fidelity and the presence of behaviorally differentiated ecotypes. In relation to our third hypothesis (H3), we found that the genetic structure of the studied pike populations was affected by salinity differences among study sites, by geographic distances, and by differentiation among river pairs, suggesting that evolutionary adaptations to local salinities, restricted movement in space, and potential natal homing to individual rivers for anadromous subpopulations mixing with freshwater residents structured the gene flow in the study area.
The limited movement-based ecological connectivity observed within the study area was in line with earlier studies that have described pike as a rather sedentary species (Diana et al. 1977; Cook and Bergersen 1988; Kobler et al. 2008). Previous studies in our study area reported rather small core home ranges for the lagoon pike (1.5 km2 in the lagoon system of more than 2000 km2) (Dhellemmes et al. 2023b), with maximal distances among two farthest recorded positions for an individual being on average 11.7 km, with most individuals having low maximal dispersal and only very few being explorers (Dhellemmes et al. 2023c). Marc-recapture studies on pike in the Baltic showed even smaller dispersal range, with recapture distances of 10 km being exceptional (Karås and Lehtonen 1993). Studies of anadromous pike populations in the Baltic reported similar dispersal distances, both using telemetry (Flink et al. 2023) and mark-recapture (Tibblin et al. 2023). Our network analyses provided further support for within-lagoon behavioral lifestyles, which renders the stock vulnerable to local pressures from harvesting or natural predation, similar to findings in Sweden (Olin et al. 2024). Nevertheless, most lagoons and freshwater tributaries in the study area were still connected through occasional adult pike movements as far as geography allowed (Fig. 2), pointing at a metapopulation-like demographic structure, consisting of an assemblage of discrete local groups with limited dispersal between them (Hanski and Simberloff 1997).
Most of the overall movement-based connectivity was gained in spring during the spawning season (March–May) when almost all the areas became connected by pike movement with traffic between some increasing sharply (Fig. A2). These findings align with earlier research indicating that pike activity peaks during spawning due to migration to spawning grounds (Cook and Bergersen 1988; Skov et al. 2018), a behavior also observed in brackish water pike populations (Tibblin et al. 2016; Flink et al. 2023; Dhellemmes et al. 2023b). The observed increased mobility level is likely due to migrations of anadromous pike to freshwater (Tibblin et al. 2015; Roser et al. 2023), but is also due to movements by brackish-water-adapted individuals to their specific spawning sites (Jacobsen et al. 2017). The pike in the Baltic Sea tend to preferentially aggregate in brackish sheltered bays for spawning (Flink et al. 2023), and if reaching these entails moving from more offshore feeding sites then the fish will engage in a spawning “run”.
Previous studies indicated that pike spawning migrations are primarily motivated by their fidelity to specific spawning sites (Miller et al. 2001; Craig 2008), which was also documented in the Baltic Sea (Larsson et al. 2015; Nordahl et al. 2019; Diaz-Suarez et al. 2022). Our findings align with that as most fish (98%) observed in at least two spawning seasons were documented to occupy the same area during spawning. However, telemetry tracking cannot definitively confirm that pike spawn where they are located during telemetry, as they may also miss spawning. In fact, it is possible that the inferred spawning site coincides with their year-round habitat, supported by the fact that 71% of pike did not leave their spawning area for the rest of the year. However, previous translocation experiments in the Rügen lagoons reported strong evidence for spawning site fidelity and revealed that brackish pike translocated to freshwater rivers returned to brackish sites and freshwater fish translocated to brackish water similarly returned to their rivers, suggesting adaptation to specific spawning habitats that vary in salinity (Dhellemmes et al. 2023d). Laboratory experiments further support this, showing that brackish-adapted pike struggle to reproduce successfully in freshwater, while the opposite is true for freshwater-adapted pike (Arlinghaus et al. 2023a, b). Additionally, the genetic differences in pike sampled from different rivers during spawning point toward natal site fidelity (Nordahl et al. 2019; Roser et al. 2023; Sunde et al. 2022), although experiments exposing larval pike to different river odors to confirm imprinting remain to be done. These findings underscore the importance of physiological adaptation to specific salinities in explaining the movements and spawning site fidelity observed in our tracking study.
Distinct behavioral patterns were evident among groups of pike observed in different habitats, particularly in terms of their movements between fresh and brackish waters. These variations hint at the presence of diverse pike ecotypes, as shown in previous work across the Baltic (Larsson et al. 2015), and shed light on their spawning strategies. First, the majority of pike tagged in brackish water lagoons remained in the lagoons throughout the year, including spawning season (Fig. 3), implying that they also spawned there and thus had undergone evolutionary adaptation to brackish spawning and recruitment, similar to reports from a telemetry study in a comparable Danish lagoon (Jacobsen et al. 2017). Pike from freshwater rivers showed more diversity in habitat use. Individuals originating from upstream sections of the larger rivers predominantly stayed in freshwater habitats and rarely ventured into brackish lagoons (Fig. 3). Comparable observations were reported in a study from Denmark by Birnie-Gauvin et al. (2019), who found a freshwater pike stock in a coastal river of the Baltic Sea that only occasionally visited the estuary but were otherwise freshwater residents. By contrast, individuals tagged or observed downstream, particularly in small rivers (e.g., Sehrowbach), were seen both in estuaries and brackish lagoons (Fig. 3), suggesting their greater tolerance to salinity variations and pointing to potentially anadromous subpopulations. Anadromy is well documented in pike in many Baltic Sea studies (e.g., Engstedt et al. 2010; Tibblin et al. 2016) including in our study area (Möller et al. 2019; Roser et al. 2023), although here this pike ecotype is rare today (< 6% among brackish samples) (Möller et al. 2019; Arlinghaus et al. 2023a). This is most likely due to extensive blocking of freshwater access since the 1970s (Roser et al. 2023), which might have advanced the selection pressures to adapt to spawning in brackish water.
Our association tests among ecological and genetic connectivity revealed that although movement-based ecological connectivity was generally limited, even such low exchange levels support sufficient gene flow to homogenize subpopulations between directly adjacent lagoons of similar mesohaline salinity (see also Möller et al. 2021). Accordingly, our measures of current movement-based connectivity had no significant effect on genetic divergence. The presence of sufficient gene flow among various mesohaline brackish lagoons is exemplified by little genetic differentiation among all mesohaline lagoons (Greifswalder Bodden, Großer Jasmunder Bodden, Barther Bodden, Kubitzer, and Schaproder Bodden; Roser et al. (2023)), which was also found in an earlier microsatellite-based study in Rügen pike by Möller et al. (2021). Moreover, evaluation of restricted ecological connectivity via telemetry suggests a finer demographic population structure than genetics data would imply, indicating that many local subgroups in different lagoons are genetically related, but spatially ecologically disaggregated due to infrequent exchange. Such low levels of ecological connectivity leave the local populations susceptible to local overfishing and other stressors, eroding their overall resilience (Kool et al. 2013; Gido et al. 2015; Olin et al. 2024), a key finding that an isolated genetic study would not have captured.
Geographic distance emerged as a significant driver of genetic differentiation, consistent with localized patterns of pike movement (Diana et al. 1977; Craig 2008), so that populations separated by larger distances displayed greater differentiation as they became increasingly reproductively isolated. This is in agreement with previous research in our study area (Möller et al. 2021) and in the Baltic Sea in general (Laikre et al. 2005; Wennerström et al. 2017; Nordahl et al. 2019; Sunde et al. 2022), which all showed that genetic structure is associated with geographic distance along the Baltic coast at both large (e.g., from Denmark to Finland) as well as smaller scales (e.g., within a lagoon system like our study site).
Our association models also showed a significant influence of salinity levels and the presence of freshwater in each pair of sites on genetic differentiation among these sites. This emphasizes that, in addition to geography, genetic differentiation is also driven by local adaptation to salinity, possibly assisted by natal homing to selected rivers (see also Nordahl et al. 2019), consistent with the observed habitat utilization strategies in the population, which revealed brackish residents as well as more migratory ecotypes. The whole-genome sequencing at the site level also showed a surprisingly high genetic differentiation among a mesohaline and an oligohaline lagoon, namely the Greifswalder Bodden and Peenestrom (Roser et al. 2023). This lagoon pair also exhibited the lowest ecological connectivity of all site pairs, although we might have underestimated the levels of movement here due to low receiver density in the area and considering the number of pike tagged (Fig. 1). Nonetheless, our results align with findings by Möller et al. (2021), who also reported that pike in most adjacent lagoons were genetically similar, except for fish from mesohaline Greifswalder Bodden and oligohaline Peenestrom lagoons, which are geographically adjacent but have very different salinities. This suggests that divergence may be rather driven by physiological dispersal barriers related to reproductive salinity tolerance (Möller et al. 2021; Sunde et al. 2022). Our results support these earlier findings by revealing how the behavior of pike contributes to reproductive isolation and population differentiation, both in terms of geography and site fidelity to selected rivers, as well as in terms of salinity adaptation (Nordahl et al. 2019; Sunde et al. 2022).
Implications for management and conservation
Evolutionary and ecological scales complement each other and are often interlinked in eco-evolutionary processes (Travis et al. 2012). Understanding both ecological and genetic connectivity is pivotal for effectively managing lagoon pike populations and fish populations in general (Hawkins et al. 2016). Both perspectives offer complementary views on population structure on different timescales, sometimes leading to divergent conclusions on suitable management and conservation strategies. In the case of the lagoon pike, whereas high genetic connectivity implied limited differentiation between the pike subpopulations in different mesohaline lagoons and thus a possibility to manage all lagoon pike as a single large stock, limited movement exchange between individual lagoons results in low ecological connectivity, hindering the recolonization potential and increasing the vulnerability of local subpopulations to local overfishing or other adverse events (e.g., large natural predation by fish-eating birds or seals, Olin et al. 2024). Hence, the effective management and conservation of the metapopulation of pike would require managing units on a more localized spatial scale in ecological time (e.g., year-to-year or seasonal), so that the population can withstand local stress factors and sustain harvest for both recreational and commercial fisheries that co-exploit the stock (Gido et al. 2015; Hawkins et al. 2016; Olin et al. 2024). At the same time, it is critical to maintain and foster ecotypic and genetic biocomplexity at the entire metapopulation level and across different rivers, which is recognized as a critical factor for building resilience and maintaining fisheries productivity (Schindler et al. 2010). To that end, a subpopulation-tailored approach is needed to support different pike ecotypes that are genetically and phenotypically differentiated. Anadromous subpopulations can be directly supported only at the stage when they migrate to freshwater habitats to spawn, e.g., by restoring connectivity between the freshwater and brackish realms through the removal of barriers associated with wetland management and agriculture to allow anadromous subpopulations to reach their historical spawning grounds (Roser et al. 2023). Another key action is control of excessive captures via passive gear (e.g., gill nets) in migration routes prior to spawning as they might preferentially target mobile pike during their spawning migration. Such measures, and especially the restoration of wetlands, were shown to be effective in enhancing the abundance and size structure of adult pike in Baltic coastal habitats (Larsson et al. 2015; Tibblin et al. 2023) and will also help to maintain a genetic ecotype that is presently rare in the Bodden lagoons. For supporting the brackish-adapted subpopulations, it is equally important to maintain connectivity to allow for the dispersal from foraging sites to low salinity spawning bays (Flink et al. 2023). Further research may focus on precisely identifying the lagoon spawning locations that can be seasonally protected to enhance the stock. Until such studies become available, it is safe to assume that enclosed vegetated bays, providing shelter from wave action and allowing for freshwater inflow from rivers and ditches to reduce local salinity, are important spawning grounds (Eklöf et al. 2023; Flink et al. 2023) whose protection would favor brackish-water-adapted pike in the Rügen lagoons.
Data availability statement
The telemetry data are available in the European Tracking Network repository (Dhellemmes and Arlinghaus 2021). All genetic sequence reads were archived at the European Nucleotide Archive under Accession nos ERR10795327 to ERR10795337 (study accession nr PRJEB59012) (http://www.ebi.ac.uk/ena/).
References
Aguirre WE, Reid K, Rivera J et al. (2022) Freshwater colonization, adaptation, and genomic divergence in threespine stickleback. Integr Comp Biol 62:388–405. https://doi.org/10.1093/icb/icac071
Arlinghaus R, Braun M, Dhellemmes F et al. (2023a) BODDENHECHT - Ökologie Nutzung und Schutz von Hechten in den Küstengewässern Mecklenburg-Vorpommerns. Berichte des IGB, Band 33. Leibniz-Institut für Gewässerökologie und Binnenfischerei, Berlin
Arlinghaus R, Rittweg T, Dhellemmes F et al. (2023b) A synthesis of a coastal northern pike (Esox lucius) fishery and its social-ecological environment in the southern Baltic Sea: Implications for the management of mixed commercial-recreational fisheries. Fish Res 263:106663. https://doi.org/10.1016/j.fishres.2023.106663
Barrat A, Barthélemy M, Pastor-Satorras R, Vespignani A (2004) The architecture of complex weighted networks. Proc Natl Acad Sci 101:3747–3752. https://doi.org/10.1073/pnas.0400087101
Benestan L (2020) Population genomics applied to fishery management and conservation. In: Oleksiak MF, Rajora OP (eds) Population genomics: marine organisms. Springer International Publishing, Cham, pp 399–421
Bergström U, Larsson S, Erlandsson M et al. (2022) Long-term decline in northern pike (Esox lucius L.) populations in the Baltic Sea revealed by recreational angling data. Fish Res 251:106307. https://doi.org/10.1016/j.fishres.2022.106307
Birnie-Gauvin K, Birch Højrup L, Kragh T et al. (2019) Getting cosy in freshwater: assumed to be brackish pike are not so brackish after all. Ecol Freshw Fish 28:376–384. https://doi.org/10.1111/eff.12460
Borg NJ, Mitchell MS, Lukacs PM et al. (2017) Behavioral connectivity among bighorn sheep suggests potential for disease spread. J Wildl Manag 81:38–45. https://doi.org/10.1002/jwmg.21169
Bosworth A, Farrell JM (2006) Genetic divergence among northern pike from spawning locations in the upper St. Lawrence river. N Am J Fish Manag 26:676–684. https://doi.org/10.1577/M05-060.1
Brannon EL, Powell MS, Quinn TP, Talbot A (2004) Population structure of Columbia river basin chinook salmon and steelhead trout. Rev Fish Sci 12:99–232. https://doi.org/10.1080/10641260490280313
Brooks M, Bolker B, Kristensen K, et al. (2023) glmmTMB: generalized linear mixed models using template model builder
Brown CJ, Harborne AR, Paris CB, Mumby PJ (2016) Uniting paradigms of connectivity in marine ecology. Ecology 97:2447–2457. https://doi.org/10.1002/ecy.1463
Bry C (1996) Role of vegetation in the life cycle of pike. In: Craig JF (ed) Pike. Springer, Netherlands, pp 45–67
Burnham KP, Anderson DR (2004) Multimodel inference: understanding AIC and BIC in model selection. Sociol Methods Res 33:261–304. https://doi.org/10.1177/0049124104268644
Cayuela H, Rougemont Q, Prunier JG et al (2018) Demographic and genetic approaches to study dispersal in wild animal populations: a methodological review. Mol Ecol 27:3976–4010. https://doi.org/10.1111/mec.14848
Clemens BJ, Schreck CB (2021) An assessment of terminology for intraspecific diversity in fishes, with a focus on “ecotypes” and “life histories.” Ecol Evol 11:10772–10793. https://doi.org/10.1002/ece3.7884
Cook MF, Bergersen EP (1988) Movements, habitat selection, and activity periods of northern pike in Eleven Mile Reservoir, Colorado. Trans Am Fish Soc 117:495–502. https://doi.org/10.1577/1548-8659(1988)117%3c0495:mhsaap%3e2.3.co;2
Cowen R, Gawarkiewicz G, Pineda J et al. (2007) Population connectivity in marine systems: an overview. Oceanog 20:14–21. https://doi.org/10.5670/oceanog.2007.26
Craig JF (2008) A short review of pike ecology. Hydrobiologia 601:5–16. https://doi.org/10.1007/s10750-007-9262-3
Crane DP, Miller LM, Diana JS et al. (2015) Muskellunge and northern pike ecology and management: important issues and research needs. Fisheries 40:258–267. https://doi.org/10.1080/03632415.2015.1038382
Csárdi G, Nepusz T (2006) The igraph software package for complex network research
Dahl J (1961) Alder og vækst hos danske og svenske brak- vandsgedder. Ferskvandfiskeribladet 59:34–38
Dhellemmes F, Aspillaga E, Monk CT (2023a) ATfiltR: a solution for managing and filtering detections from passive acoustic telemetry data. MethodsX 10:102222. https://doi.org/10.1016/j.mex.2023.102222
Dhellemmes F, Aspillaga E, Rittweg T et al. (2023b) Body size scaling of space use in coastal pike (Esox lucius) in brackish lagoons of the southern Baltic Sea. Fish Res 260:106560. https://doi.org/10.1016/j.fishres.2022.106560
Dhellemmes F, Arlinghaus R (2021) Boddenhecht telemetry dataset. https://marineinfo.org/id/dataset/7859
Dhellemmes F, Lukyanova O, Radinger J, et al. (2023c) Bewegungsökologie des Boddenhechts: Standhecht oder Wanderhecht und die Rolle der Umwelt. Berichte IGB 33:237–265
Dhellemmes F, Lukyanova O, Roser P, Arlinghaus R (2023d) Anadromie und Homing von Boddenhechten. Berichte IGB 33:227–296
Diana JS, Mackay WC, Ehrman M (1977) Movements and habitat preference of northern pike (Esox lucius) in Lac Ste. Anne Alberta. Trans Am Fish Soc 106:560–565. https://doi.org/10.1577/1548-8659(1977)106%3c560:MAHPON%3e2.0.CO;2
Diaz-Suarez A, Noreikiene K, Kisand V et al. (2022) Temporally stable small-scale genetic structure of northern pike (Esox lucius) in the coastal Baltic Sea. Fish Res 254:106402. https://doi.org/10.1016/j.fishres.2022.106402
Eklöf JS, Sundblad G, Erlandsson M et al. (2020) A spatial regime shift from predator to prey dominance in a large coastal ecosystem. Commun Biol 3:1–9. https://doi.org/10.1038/s42003-020-01180-0
Eklöf JS, Hansen JP, Eriksson BK et al. (2023) Effects of seasonal spawning closures on pike (Esox lucius L.) and perch (Perca fluviatilis L.) catches and coastal food webs in the western Baltic Sea. Fish Res 263:106674. https://doi.org/10.1016/j.fishres.2023.106674
Engstedt O, Stenroth P, Larsson P et al. (2010) Assessment of natal origin of pike (Esox lucius) in the Baltic Sea using Sr: Ca in otoliths. Environ Biol Fishes 89:547–555. https://doi.org/10.1007/s10641-010-9686-x
Engstedt O, Engkvist R, Larsson P (2014) Elemental fingerprinting in otoliths reveals natal homing of anadromous Baltic Sea pike (Esox lucius L.). Ecol Freshw Fish 23:313–321. https://doi.org/10.1111/eff.12082
Ferguson A, Reed TE, Cross TF et al. (2019) Anadromy, potamodromy and residency in brown trout Salmo trutta: the role of genes and the environment. J Fish Biol 95:692–718. https://doi.org/10.1111/jfb.14005
Flávio H, Baktoft H (2021) actel: standardised analysis of acoustic telemetry data from animals moving through receiver arrays. Methods Ecol Evol 12:196–203. https://doi.org/10.1111/2041-210X.13503
Flink H, Tibblin P, Hall M et al. (2023) Variation among bays in spatiotemporal aggregation of Baltic Sea pike highlights management complexity. Fish Res 259:106579. https://doi.org/10.1016/j.fishres.2022.106579
Freeland J, Kirk H, Petersen S (2011) Molecular ecology: Joanna R. Freeland and Heather Kirk; Stephen Petersen, 2nd ed., 1st impression. Wiley-Blackwell, Oxford; Hoboken, NJ
Gido KB, Whitney JE, Perkin JS, Turner TF (2015) Fragmentation, connectivity and fish species persistence in freshwater ecosystems. In: Closs GP, Olden JD, Krkosek M (eds) Conservation of freshwater fishes. Cambridge University Press, Cambridge, pp 292–323
Gu Z, Gu L, Eils R et al. (2014) circlize implements and enhances circular visualization in R. Bioinformatics 30:2811–2812. https://doi.org/10.1093/bioinformatics/btu393
Hansen JP, Sundblad G, Bergström U et al. (2019) Recreational boating degrades vegetation important for fish recruitment. Ambio 48:539–551. https://doi.org/10.1007/s13280-018-1088-x
Hanski I, Simberloff D (1997) The metapopulation approach, its history, conceptual domain and application to conservation. In: Hanski I, Gilpin ME (eds) Metapopulation biology: ecology, genetics, and evolution. Academic Press, San Diego, pp 5–26
Hansson S, Bergström U, Bonsdorff E et al. (2018) Competition for the fish–fish extraction from the Baltic Sea by humans, aquatic mammals, and birds. ICES J Mar Sci 75:999–1008. https://doi.org/10.1093/icesjms/fsx207
Hartig F, Lohse L (2022) DHARMa: residual diagnostics for hierarchical (multi-level/mixed) regression models
Haugen TO, Winfield IJ, Vøllestad LA et al. (2006) The ideal free pike: 50 years of fitness-maximizing dispersal in Windermere. Proc R Soc B Biol Sci 273:2917–2924. https://doi.org/10.1098/rspb.2006.3659
Hawkins SJ, Bohn K, Sims DW et al. (2016) Fisheries stocks from an ecological perspective: disentangling ecological connectivity from genetic interchange. Fish Res 179:333–341. https://doi.org/10.1016/j.fishres.2016.01.015
Hedgecock D, Barber P, Edmands S (2007) Genetic approaches to measuring connectivity. Oceanog 20:70–79. https://doi.org/10.5670/oceanog.2007.30
Hilborn R, Quinn TP, Schindler DE, Rogers DE (2003) Biocomplexity and fisheries sustainability. Proc Natl Acad Sci 100:6564–6568. https://doi.org/10.1073/pnas.1037274100
Hohenlohe PA, Funk WC, Rajora OP (2021) Population genomics for wildlife conservation and management. Mol Ecol 30:62–82. https://doi.org/10.1111/mec.15720
Holsinger KE, Weir BS (2009) Genetics in geographically structured populations: defining, estimating and interpreting FST. Nat Rev Genet 10:639–650. https://doi.org/10.1038/nrg2611
Jacobsen L, Bekkevold D, Berg S et al. (2017) Pike (Esox lucius L.) on the edge: consistent individual movement patterns in transitional waters of the western Baltic. Hydrobiologia 784:143–154. https://doi.org/10.1007/s10750-016-2863-y
Jacoby DMP, Brooks EJ, Croft DP, Sims DW (2012) Developing a deeper understanding of animal movements and spatial dynamics through novel application of network analyses. Methods Ecol Evol 3:574–583. https://doi.org/10.1111/j.2041-210X.2012.00187.x
Jørgensen AT, Hansen BW, Vismann B et al. (2010) High salinity tolerance in eggs and fry of a brackish Esox lucius population. Fish Manag Eco 17:554–560. https://doi.org/10.1111/j.1365-2400.2010.00755.x
Karås P, Lehtonen H (1993) Patterns of movement and migration of pike (Esox lucius L.) in the Baltic sea. Nord J Freshw Res 68:72–79
Kobler A, Klefoth T, Wolter C et al. (2008) Contrasting pike (Esox lucius L.) movement and habitat choice between summer and winter in a small lake. Hydrobiologia 601:17–27. https://doi.org/10.1007/s10750-007-9263-2
Kool JT, Moilanen A, Treml EA (2013) Population connectivity: recent advances and new perspectives. Landsc Ecol 28:165–185. https://doi.org/10.1007/s10980-012-9819-z
Laikre L, Miller LM, Palmé A et al. (2005) Spatial genetic structure of northern pike (Esox lucius) in the Baltic Sea. Mol Ecol 14:1955–1964. https://doi.org/10.1111/j.1365-294X.2005.02570.x
Larsson P, Tibblin P, Koch-Schmidt P et al. (2015) Ecology, evolution, and management strategies of northern pike populations in the Baltic Sea. Ambio 44:451–461. https://doi.org/10.1007/s13280-015-0664-6
Lea JSE, Humphries NE, von Brandis RG et al. (2016) Acoustic telemetry and network analysis reveal the space use of multiple reef predators and enhance marine protected area design. Proc R Soc B Biol Sci. https://doi.org/10.1098/rspb.2016.0717
Lehtonen H, Leskinen E, Selén R, Reinikainen M (2009) Potential reasons for the changes in the abundance of pike, Esox lucius, in the western Gulf of Finland, 1939–2007. Fish Manag Ecol 16:484–491. https://doi.org/10.1111/j.1365-2400.2009.00701.x
Lowe WH, Allendorf FW (2010) What can genetics tell us about population connectivity? Mol Ecol 19:3038–3051. https://doi.org/10.1111/j.1365-294X.2010.04688.x
Luque S, Saura S, Fortin M-J (2012) Landscape connectivity analysis for conservation: insights from combining new methods with ecological and genetic data. Landsc Ecol 27:153–157. https://doi.org/10.1007/s10980-011-9700-5
Marandel F, Lorance P, Andrello M et al. (2018) Insights from genetic and demographic connectivity for the management of rays and skates. Can J Fish Aquat Sci 75:1291–1302. https://doi.org/10.1139/cjfas-2017-0291
Matley JK, Klinard NV, Martins APB et al. (2022) Global trends in aquatic animal tracking with acoustic telemetry. Trends Ecol Evol 37:79–94. https://doi.org/10.1016/j.tree.2021.09.001
Miller LM, Kallemeyn L, Senanan W (2001) Spawning-site and natal-site fidelity by northern pike in a large lake: mark-recapture and genetic evidence. Trans Am Fish Soc 130:307–316. https://doi.org/10.1577/1548-8659(2001)130%3c0307:SSANSF%3e2.0.CO;2
Möller S, Winkler HM, Klügel A, Richter S (2019) Using otolith microchemical analysis to investigate the importance of brackish bays for pike (Esox lucius Linnaeus, 1758) reproduction in the southern Baltic Sea. Ecol Freshw Fish 28:602–610. https://doi.org/10.1111/eff.12478
Möller S, Winkler HM, Richter S, Bastrop R (2021) Genetic population structure of pike (Esox lucius Linnaeus, 1758) in the brackish lagoons of the southern Baltic Sea. Ecol Freshw Fish 30:140–149. https://doi.org/10.1111/eff.12571
Müller MF, Banks SC, Crewe TL, Campbell HA (2023) The rise of animal biotelemetry and genetics research data integration. Ecol Evol 13:e9885. https://doi.org/10.1002/ece3.9885
Nathan R, Monk CT, Arlinghaus R et al. (2022) Big-data approaches lead to an increased understanding of the ecology of animal movement. Science 375:1780. https://doi.org/10.1126/science.abg1780
Neville HM, Isaak DJ, Dunham JB et al. (2006) Fine-scale natal homing and localized movement as shaped by sex and spawning habitat in Chinook salmon: insights from spatial autocorrelation analysis of individual genotypes. Mol Ecol 15:4589–4602. https://doi.org/10.1111/j.1365-294X.2006.03082.x
Nichols JD, Hines JE, Lebreton J-D, Pradel R (2000) Estimation of contributions to population growth: a reverse-time capture-recapture approach. Ecology 81:3362. https://doi.org/10.2307/177500
Nilsson J, Flink H, Tibblin P (2019) Predator–prey role reversal may impair the recovery of declining pike populations. J Anim Ecol 88:927–939. https://doi.org/10.1111/1365-2656.12981
Nordahl O, Koch-Schmidt P, Sunde J et al. (2019) Genetic differentiation between and within ecotypes of pike (Esox lucius) in the Baltic Sea. Aquat Conserv Mar Freshw Ecosyst 29:1923–1935. https://doi.org/10.1002/aqc.3196
Nurbaev SD, Balanovskaia EV (1998) Interpopulation diversity of the gene pool: beta distribution of Wright’s F(ST) statistics. Genetika 34(7):1004–1008
Oksanen J, Simpson GL, Blanchet FG, et al. (2022) vegan: community ecology package
Olin AB, Bergström U, Bodin Ö et al. (2024) Predation and spatial connectivity interact to shape ecosystem resilience to an ongoing regime shift. Nat Commun 15:1304. https://doi.org/10.1038/s41467-024-45713-1
Olsson J (2019) Past and current trends of coastal predatory fish in the Baltic sea with a focus on perch, pike, and pikeperch. Fishes 4:1–26. https://doi.org/10.3390/fishes4010007
Olsson J, Andersson ML, Bergström U et al. (2023) A pan-Baltic assessment of temporal trends in coastal pike populations. Fish Res 260:106594. https://doi.org/10.1016/j.fishres.2022.106594
Palmas F, Olita A, Addis P et al. (2017) Modelling giant red shrimp larval dispersal in the Sardinian seas: density and connectivity scenarios. Fish Oceanogr 26:364–378. https://doi.org/10.1111/fog.12199
Palsbøll PJ, Bérubé M, Allendorf FW (2007) Identification of management units using population genetic data. Trends Ecol Evol 22:11–16. https://doi.org/10.1016/j.tree.2006.09.003
R Core Team (2023) R: A language and environment for statistical computing
Riginos C, Buckley YM, Blomberg SP, Treml EA (2014) Dispersal capacity predicts both population genetic structure and species richness in reef fishes. Am Nat 184:52–64. https://doi.org/10.1086/676505
Roser P, Dhellemmes F, Rittweg T et al. (2023) Synthesizing historic and current evidence for anadromy in a northern pike (Esox lucius L.) meta-population inhabiting brackish lagoons of the southern Baltic Sea, with implications for management. Fish Res 263:106670. https://doi.org/10.1016/j.fishres.2023.106670
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228. https://doi.org/10.1093/genetics/145.4.1219
Runge JP, Runge MC, Nichols JD et al. (2006) The role of local populations within a landscape context: defining and classifying sources and sinks. Am Nat 167:925–938. https://doi.org/10.1086/503531
Schindler DE, Hilborn R, Chasco B et al. (2010) Population diversity and the portfolio effect in an exploited species. Nature 465:609–612. https://doi.org/10.1038/nature09060
Schubert H, Telesh I, Schubert H, Telesh I (2017) Estuaries and coastal lagoons. In: Snoeijs-Leijonmalm P, Schubert H, Radziejewska T (eds) Biological oceanography of the Baltic sea. Springer, Dordrecht, pp 483–509
Skov C, Lucas MC, Jacobsen L (2018) Spatial ecology. In: Skov C, Nilsson PA (eds) Biology and ecology of pike. Taylor and Francis, Boca Raton FL, USA. pp 83–120
Stronen AV, Norman AJ, Vander Wal E, Paquet PC (2022) The relevance of genetic structure in ecotype designation and conservation management. Evol Appl 15:185–202. https://doi.org/10.1111/eva.13339
Sunde J, Larsson P, Forsman A (2019) Adaptations of early development to local spawning temperature in anadromous populations of pike (Esox lucius). BMC Evol Biol. https://doi.org/10.1186/s12862-019-1475-3
Sunde J, Yıldırım Y, Tibblin P et al. (2022) Drivers of neutral and adaptive differentiation in pike (Esox lucius) populations from contrasting environments. Mol Ecol 31:1093–1110. https://doi.org/10.1111/mec.16315
Tibblin P, Forsman A, Koch-Schmidt P et al. (2015) Evolutionary divergence of adult body size and juvenile growth in sympatric subpopulations of a top predator in aquatic ecosystems. Am Nat 186:98–110. https://doi.org/10.1086/681597
Tibblin P, Forsman A, Borger T, Larsson P (2016) Causes and consequences of repeatability, flexibility and individual fine-tuning of migratory timing in pike. J Anim Ecol 85:136–145. https://doi.org/10.1111/1365-2656.12439
Tibblin P, Bergström K, Flink H et al. (2023) Higher abundance of adult pike in Baltic Sea coastal areas adjacent to restored wetlands compared to reference bays. Hydrobiologia. https://doi.org/10.1007/s10750-023-05216-4
Travis JMJ, Mustin K, Bartoń KA et al. (2012) Modelling dispersal: an eco-evolutionary framework incorporating emigration, movement, settlement behaviour and the multiple costs involved. Methods Ecol Evol 3:628–641. https://doi.org/10.1111/j.2041-210X.2012.00193.x
van Gemert R, Koemle D, Winkler H et al. (2022) Data-poor stock assessment of fish stocks co-exploited by commercial and recreational fisheries: applications to pike (Esox lucius) in the western Baltic Sea. Fish Manage Ecol 29:16-28
Venables WN, Ripley BD (2002) Modern applied statistics with S, 4th edn. Springer, New York
Wennerström L, Laikre L, Ryman N et al. (2013) Genetic biodiversity in the Baltic Sea: species-specific patterns challenge management. Biodivers Conserv 22:3045–3065. https://doi.org/10.1007/s10531-013-0570-9
Wennerström L, Olsson J, Ryman N, Laikre L (2017) Temporally stable, weak genetic structuring in brackish water northern pike (Esox lucius) in the Baltic Sea indicates a contrasting divergence pattern relative to freshwater populations. Can J Fish Aquat Sci 74:562–571. https://doi.org/10.1139/cjfas-2016-0039
Winkler HM (1987) Einige Bemerkungen zur Ernährung des Hechts (Esox lucius L.) in den Küstengewässern der DDR. Wissenschaftliche Zeitschrift der Wilhelm-Pieck-Universität Rostock, Naturwissenschaftliche Reihe 36:53–56
Acknowledgements
First, we thank the European Union and State of Mecklenburg-Vorpommern, Ministry of Agriculture and Environment, for funding our project via the European Maritime and Fisheries Fund (grant/award numbers MV-I.18-LM-004 and B730117000069; BODDENHECHT). We thank the project leaders (Mr. Blume) and administrators (Mr. Bachmann) at the Ministry as well as our collaborators at the Landesforschungsanstalt für Landwirtschaft, Institut für Fischerei (C. Kühn) for immense support. This project would not have been possible without the involvement of the Institut für Fisch und Umwelt (FIUM) in Rostock, in particular P. Möller who organized the receiver downloads. We also thank the Wasserstraßen- und Schifffahrtsamt (WSA) Stralsund for approving the geographical location of our receivers in the study area. We thank the Angler Association of Mecklenburg-Vorpommern for authorizing sampling via electrofishing in the freshwater rivers. We thank the Landesamt für Landwirtschaft, Lebensmittelsicherheit und Fischerei Mecklenburg-Vorpommern (LALLF) for approving our sampling methods, including the animal care protocol for tagging (Aktenzeichen 7221.3-1-052/19). We thank the Staatliches Amt für Landwirtschaft und Umwelt Vorpommern (StALU) for granting us access to Nature Conservation Areas, the Nationalparkamt Vorpommern for temporal access to national parks and Biosphärenreservatsamt Südost-Rügen for access to the biosphere reserves. Extensive field research was conducted to obtain our data, and we thank H. Hansen, C. Monk, and J. Droll for their contributions to the early phases of the project, D. Niessner for coordinating the fisher and angler recapture reports, P. Roser for curating the pike capture database, and the Innovasea field team for support in the installation and maintenance of the telemetry system. We thank Sören Möller and Helmut Winkler for providing genetics samples from Stettiner Haff, and Malte Dittmann for organizing and performing the laboratory work for the genetic data analysis for this study at the University of Oldenburg. Finally, we thank the reviewers for the excellent feedback that helped improve the paper.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
O.L. was responsible for conceptualization, methodology, investigation, formal analysis, visualization, writing—original draft, and writing—review and editing; F.D. for conceptualization, data curation, methodology, investigation, formal analysis, software, supervision, and writing—review and editing; S.D. for formal analysis, investigation, writing—review and editing; A.N. for formal analysis, investigation, and writing—review and editing; and R.A. for conceptualization, methodology, funding acquisition, supervision, project administration, validation, and writing—review and editing.
Corresponding author
Ethics declarations
Competing interests statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethics statement
The research was completed following German legislation for animal experimentation, approved by Landesamt für Landwirtschaft, Lebensmittelsicherheit und Fischerei Mecklenburg-Vorpommern—Veterinärdienste und Landwirtschaft—under grant no. 7221.3-1-052/19.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lukyanova, O., Dhellemmes, F., Dennenmoser, S. et al. Combining biotelemetry and genetics provides complementary insights relevant to the management and conservation of a freshwater predator (Esox lucius) living in brackish lagoons. Aquat Sci 86, 77 (2024). https://doi.org/10.1007/s00027-024-01090-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00027-024-01090-x