Abstract
Phylogenetic (i.e., leaf-labeled) trees play a fundamental role in evolutionary research. A typical problem is to reconstruct such trees from data like DNA alignments (whose columns are often referred to as characters), and a simple optimization criterion for such reconstructions is maximum parsimony. It is generally assumed that this criterion works well for data in which state changes are rare. In the present manuscript, we prove that each binary phylogenetic tree T with \(n\ge 20 k\) leaves is uniquely defined by the set \(A_k(T)\), which consists of all characters with parsimony score k on T. This can be considered as a promising first step toward showing that maximum parsimony as a tree reconstruction criterion is justified when the number of changes in the data is relatively small.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Mathematical phylogenetics is concerned with reconstructing the evolutionary (leaf-labeled) trees of species based on data. Typically, the data come in the form of an alignment (e.g., aligned DNA, RNA or proteins or aligned binary sequences like the absence or presence of certain morphological characteristics), whose columns are also often referred to as characters. While no tree reconstruction method can guarantee to recover the true tree for all data sets, it has long been known that in some special cases, a tree is uniquely defined by certain alignments. One such example is due to the classic theorem by Buneman [1]: If one takes a phylogenetic tree T and regards all its edges as bipartitions of the species set labeling the leaves, one can summarize the resulting binary characters in a set, which we refer to as alignment \(A_1(T)\). A re-formulated version of Buneman’s theorem states that \(A_1(T)\) then uniquely defines T [2]. Note that \(A_1(T)\) consists of all binary characters which induce precisely one change on T (namely on the edge which they correspond to).
A natural question arising from this scenario is if \(A_k(T)\) defines T for any value \(k\ge 1\), i.e., when k changes are allowed rather than just one. It has recently been shown that \(A_2(T)\) indeed defines T, but that unfortunately, for all \(k\ge 3\), \(A_k(T)\) does not define T whenever T has 2k leaves. In the present note, we will prove the positive result that \(A_k(T)\) always defines T whenever \(n \ge 20k\), where n denotes the number of leaves of T.
2 Preliminaries
2.1 Definitions and Basic Concepts
We start with some notation. Recall that a phylogenetic tree \(T=(V,E)\) on a species set \(X=\{1, \ldots , n\}\), is a connected acyclic graph with vertex set V and edge set E whose leaves are bijectively labeled by X. Such a tree T is also often referred to as phylogenetic X-tree. It is called binary if all its inner nodes have degree 3. Note that we consider two phylogenetic X-trees \(T=(V,E)\) and \(\widetilde{T}=(\widetilde{V},\widetilde{E})\) to be equal, denoted \(T=\widetilde{T}\), if there exists a map \(f:V\rightarrow \widetilde{V}\) such that \(e=\{u,v\} \in E \Longleftrightarrow \{f(u),f(v)\} \in \widetilde{E}\) and with the additional property that \(f(x)=x\) for all \(x \in X\). In other words, f is a graph isomorphism which preserves the leaf labeling.
Throughout this manuscript, when we refer to a tree T, we always mean a binary phylogenetic X-tree. However, for technical reasons, we sometimes also have to consider rooted trees: When an edge is removed from a binary (unrooted) tree, two subtrees remain, both of which have precisely one node of degree 2. This node is considered as the root of the respective subtree. The two subtrees adjacent to the root of a tree are called maximum pending subtrees of that tree, cf. Fig. 1.
In the present manuscript, whenever we refer to distances between vertices in a tree T, we mean the length of the unique path connecting these vertices in T. Moreover, we say that two leaves v and w form a cherry [v, w], if v and w are adjacent to the same inner node u of T.
Furthermore, recall that a bipartition \(\sigma \) of X into two non-empty disjoint subsets A and B is often called an X-split and is denoted by \(\sigma =A|B\). Recall that there is a natural relationship between X-splits and the edges of a phylogenetic X-tree T, because the removal of an edge e induces a bipartition \(\sigma _e\) of X. In the following, the set of all such induced X-splits of T will be denoted by \(\Sigma (T)\). Recall that for a binary phylogenetic X-tree T with \(|X|=n\) we have \(|\Sigma (T)|=2n-3\) [6, Prop. 2.1.3]. Moreover, following [3], the size of an X-split \(\sigma =A|B\) is defined as \(|\sigma |=\min \{|A|,|B|\}\). Given a set of X-splits, an element of this set with minimal size is called a minimal split.
Moreover, recall that a character f is a function from the taxon set X to a set \(\mathcal {C}\) of character states, i.e., \(f: X \rightarrow \mathcal {C}\). Note that a finite sequence of characters is also often referred to as alignment in biology. While in most biological cases, the order of the characters in an alignment plays an important role, for our purpose, it suffices to simply define an alignment as a multiset of characters. In this manuscript, we will only be concerned with binary characters (and thus also binary alignments), i.e., without loss of generality \(\mathcal {C}= \{a,b\}\).
There is a close relationship between X-splits and binary characters, because every X-split can be represented by a binary character by assigning the same state to taxa in the same subset. Throughout this manuscript, we assume for technical reasons and without loss of generality that \(f(1)=a\). If an X-split \(\sigma _e\) is induced by an edge e of a phylogenetic X-tree in the manner explained above, we also say that the corresponding binary character is induced by e.
Thus, a binary character \(f: X \rightarrow \{a,b\}\) assigns to each leaf of the tree a corresponding state. Now, an extension of such a character f on a tree T with vertex set V is a map \(g: V \rightarrow \{a,b\}\) such that \(g(x)=f(x)\) for all \(x \in X\). Moreover, we call \(ch(g) = \vert \{ \{u,v\} \in E, \, g(u) \ne g(v)\} \vert \) the changing number of g on T.
Another concept we need for the present manuscript is the so-called parsimony score l(f, T) of a character f on a tree T. Here, \(l(f,T) = \min \limits _{g} ch(g,T)\), where the minimum runs over all extensions g of f on T. The parsimony score of an alignment \(A=\{f_1,\ldots , f_m\}\) of characters is then defined as: \(l(A,T)=\sum \nolimits _{i=1}^m l(f_i,T)\).
Last, for a given tree T, we define \(A_k(T)\) to be the set consisting of all binary characters f with \(l(f,T)=k\). Following [2], we also refer to \(A_k(T)\) as the alignment induced by T and k.
2.2 Known Results
A basic result that we need throughout this manuscript is the following theorem, which counts the number of characters in \(A_k(T)\).
Theorem 1
[8] Let T be a binary phylogenetic X-tree with \(|X|=n\). Then, we have:
Another classic result that will play a fundamental role here is Menger’s theorem, which in the context of phylogenetics leads to the following proposition [6, Lemma 5.1.7 and Corollary 5.1.8]:
Proposition 1
[6, Adapted from Corollary 5.1.8] Let f be a binary character on X employing states from \(\mathcal {C}=\{a,b\}\), and let T be a binary phylogenetic X-tree. Then, l(f, T) is equal to the maximum number of edge-disjoint leaf-to-leaf paths of T, where each path connects one leaf in state a with one leaf in state b.
The next two statements are the classic theorem by Buneman and a direct consequence from it concerning \(A_1(k)\).
Theorem 2
(Buneman theorem [1, 6]) Let T and \(\widetilde{T}\) be two binary phylogenetic X-trees. Then, \(T=\widetilde{T}\) if and only if \(\Sigma (T)=\Sigma (\widetilde{T})\).
Corollary 1
Let T and \(\widetilde{T}\) be two binary phylogenetic X-trees. Then, \(T=\widetilde{T}\) if and only if \(A_1(T)=A_1(\widetilde{T})\).
The correctness of Corollary 1 follows directly from the 1:1 relationship between \(\Sigma (T)\) and \(A_1(T)\).
Last, we recall the following result from [2], which extends Corollary 1 to the case \(k=2\).
Proposition 2
(Adapted from Proposition 1 in [2]) Let T and \(\widetilde{T}\) be two binary phylogenetic X-trees. Then, \(T=\widetilde{T}\) if and only if \(A_2(T)=A_2(\widetilde{T})\).
3 Results
We are now in a position to state the main result of the present manuscript.
Theorem 3
Let \(k \in \mathbb {N}_{\ge 1}\) and let \(n \in \mathbb {N}\) such that \(n\ge 20k\). Let T and \(\widetilde{T}\) be two binary phylogenetic X-trees with \(|X|=n\). Then, \(T=\widetilde{T}\) if and only if \(A_k(T)=A_k(\widetilde{T})\).
Before we can prove Theorem 3, we need to state one more lemma.
Lemma 1
Let T be a binary phylogenetic X-tree with \(|X|=n\). Then, T has a set of \(\left\lfloor \frac{n}{2}\right\rfloor \) edge-disjoint leaf-to-leaf paths.
Proof
Note that if \(n<2\), then there is nothing to show. If \(n\ge 2\), we apply Theorem 1 to \(k=\left\lfloor \frac{n}{2}\right\rfloor \). For the case where n is even, this leads to:
Similarly, if n is odd, we have \(n\ge 3\) and \(k=\left\lfloor \frac{n}{2}\right\rfloor = \frac{n-1}{2}\) and therefore get
So in both cases, there is at least one character f on T with parsimony score \(k=\left\lfloor \frac{n}{2}\right\rfloor \). However, by Proposition 1, this immediately implies that there is also at least one choice of \(\left\lfloor \frac{n}{2}\right\rfloor \) edge-disjoint leaf-to-leaf paths in T. This completes the proof. \(\square \)
We are now finally in a position to prove Theorem 3, which is the main result of the present note.
Proof of Theorem 3
Note that the cases \(k=1\) and \(k=2\) are already proved by Corollary 1 and Proposition 2. Therefore, we may assume in the following that \(k\ge 3\).
Now, let \(T\ne \widetilde{T}\) be two phylogenetic X-trees with n leaves such that \(n \ge 20k\). Then \(\Sigma (T)\ne \Sigma (\widetilde{T})\) by Theorem 2, and as explained above we have \(|\Sigma (T)|=|\Sigma (\widetilde{T})|=2n-3\). Together, this implies that \(\Sigma (T)\setminus \Sigma (\widetilde{T}) \ne \emptyset \). Let \(\sigma =A|B \in \Sigma (T)\setminus \Sigma (\widetilde{T})\) be minimal, i.e., of minimal size. Without loss of generality, we assume \(|A|=|\sigma |\), i.e., \(|A|\le |B|\) and thus \(|B|\ge \frac{n}{2}\). Note that \(|A|\ge 2\) as \(\sigma \in \Sigma (T)\) but \(\sigma \not \in \Sigma (\widetilde{T})\) (otherwise, \(\sigma \) would be contained in both split sets as all X-trees contain edges leading to each of the leaves in X). Moreover, \(\sigma \) divides T into two subtrees \(T_A\) with leaf set A and \(T_B\) with leaf set B. In the following, we denote by \(T_{A_1}\) and \(T_{A_2}\) the two maximal pending subtrees of \(T_A\), which must exist as \(|A|\ge 2\), cf. Fig. 1. The taxon sets of \(T_{A_1}\) and \(T_{A_2}\) are denoted by \(A_1\) and \(A_2\), respectively. Note that \(|A_1|<|A|\) and \(|A_2|<|A|\), and also note that \(\Sigma (T)\) must contain the two X-splits \(\sigma _1=A_1|X\setminus A_1\) and \(\sigma _2=A_2|X\setminus A_2\) as \(T_{A_1}\) and \(T_{A_2}\) are subtrees of T.
(Taken from [2]) By removing an edge e from an unrooted phylogenetic tree T, it is decomposed into two rooted subtrees, \(T_A\) and \(T_B\). If, as in this figure, both of them consist of more than one node, then we can further decompose them into their two maximal pending subtrees, \(T_{A_1}\) and \(T_{A_2}\) or \(T_{B_1}\) and \(T_{B_2}\), respectively
Note that by the minimality of \(\sigma \), the X-splits \(\sigma _1\) and \(\sigma _2\) must be contained in \(\Sigma (T)\cap \Sigma (\widetilde{T})\). So we have \(\sigma _1\), \(\sigma _2\) \(\in \Sigma (\widetilde{T})\). By an analogous argument, \(\widetilde{T}\) must also contain all splits induced by edges of \(T_{A_1}\) and \(T_{A_2}\), respectively. So in fact, \(\widetilde{T}\) has \(T_{A_1}\) and \(T_{A_2}\) as subtrees, but as \(\sigma \not \in \Sigma (T)\cap \Sigma (\widetilde{T})\), they are not pending on the same inner node of \(\widetilde{T}\). In particular, this implies that \(\widetilde{T}\) can be represented as depicted in Fig. 2. Moreover, as \(\widetilde{T}\) does not contain split \(\sigma \), it must contain subtrees \(\widetilde{T}_{B_1},\ldots , \widetilde{T}_{B_m}\), where \(m\in \mathbb {N}_{\ge 2}\), along the path from the root \(\rho _1\) of \(T_{A_1}\) to the root \(\rho _2\) of \(T_{A_2}\). (Note that all these subtrees contain at least one leaf.) Without loss of generality, we assume that \(\widetilde{T}_{B_1}\) is pending on the same node as \(T_{A_1}\), and \(\widetilde{T}_{B_m}\) is pending on the same node as \(T_{A_2}\), cf. Fig. 2. In the following, we denote by \(B_i\) the set of leaves of subtree \(\widetilde{T}_{B_i}\) for all \(i=1,\ldots ,m\).
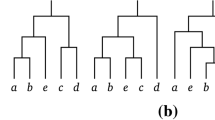
Tree \(\widetilde{T}\) as described in the proof of Theorem 3. Note that the subtree \(\mathring{T}\) might be empty, namely if \(m=2\). If \(m=3\), the proof considers \(\widetilde{T}_{B_2}\), but if \(m>3\), we can think of \(\mathring{T}\) as a rooted binary tree with root \(\mathring{\rho }\) and with \(\widetilde{b}\) leaves as described in the proof
Now we construct a binary character f such that \(l(f,\widetilde{T})=k>l(f,T)\) as follows:
-
We assign state a to all taxa in A.
-
Then, we choose two paths: one connecting the root \(\rho _1\) of \(T_{A_1}\) with the nearest leaf \(x_1\in \widetilde{T}_{B_1}\), and the other one connecting the root \(\rho _2\) of \(T_{A_2}\) with the nearest leaf \(x_2 \in \widetilde{T}_{B_m}\).
-
Then, we choose \(k-2\) additional edge-disjoint leaf-to-leaf paths in \(\widetilde{T}\), where all leaves considered are contained in \(B\setminus \{x_1,x_2\}\).
-
We then assign \(x_1\) and \(x_2\) state b. For all other \(k-2\) paths, we label one of its endpoints a and the other one b. All unlabeled leaves in B that are not contained in a path (if such leaves exist) also are assigned state b.
We subsequently prove \(l(f,\widetilde{T})=k>l(f,T)\), but before we do this, we first show that our construction is valid, i.e., that indeed \(k-2\) paths can be chosen as described above in the crucial third step.
Let \(p_1\) denote the path length of the path from \(\rho _1\) to \(x_1\) in \(\widetilde{T}\) and similarly, let \(p_2\) denote the path length of the path from \(\rho _2\) to \(x_2\) in \(\widetilde{T}\). Note that these two paths are edge-disjoint.
Now consider \(\widetilde{T}_{B_1}\) with its \(|B_1|\) leaves and root \(\widetilde{\rho }_1\): Before taking the path from \(\widetilde{\rho }_1\) to \(x_1\) (as a subpath of the path from \(\rho _1\) to \(x_1\)), by Lemma 1, there were \(\left\lfloor \frac{|B_1|}{2}\right\rfloor \) edge-disjoint leaf-to-leaf paths in \(\widetilde{T}_{B_1}\). Even without counting the number of such paths requiring edges of the already taken path, one can easily see that this path can at most reduce the number of edge-disjoint leaf-to-leaf paths in \(\widetilde{T}_{B_1}\) by \(p_1-2\) (as it contains \(p_1-2\) edges in \(\widetilde{T}_{B_1}\)). So in total, subtree \(\widetilde{T}_{B_1}\) allows for at least \(\left\lfloor \frac{|B_1|}{2}\right\rfloor - p_1+2\) edge-disjoint leaf-to-leaf paths. By the same argument, \(\widetilde{T}_{B_m}\) allows for at least \(\left\lfloor \frac{|B_m|}{2}\right\rfloor - p_2+2\) edge-disjoint leaf-to-leaf paths.
We now consider the leaves in \(B\setminus (B_1\cup B_m)\). Let \(\widetilde{b}\) denote the cardinality of this set. Note that \(\widetilde{b}\) can equal 0 (namely if \(m=2\)). On the other hand, if \(\widetilde{b}=1\) (i.e., if \(m=3\)), we can consider \(\widetilde{T}_{B_2}\), and otherwise, if \(\widetilde{b}\ge 2\) (i.e., \(m\ge 4\)) we can consider the rooted tree \(\mathring{T}\) containing subtrees \(\widetilde{T}_{B_2},\ldots , \widetilde{T}_{B_{m-1}}\) as depicted in the dashed box of Fig. 2. Again by Lemma 1, there are in all cases at least \(\left\lfloor \frac{\widetilde{b}}{2}\right\rfloor \) edge-disjoint leaf-to-leaf paths induced by \(B\setminus (B_1\cup B_m)\) in the tree under consideration (i.e., the empty tree, \(\widetilde{T}_{B_2}\) or \(\mathring{T}\), respectively). Note that each such path naturally corresponds to a path in \(\widetilde{T}\) which is edge-disjoint with all other paths chosen before.
So the total number \(\mathcal {P}\) of edge-disjoint leaf-to-leaf paths that are present in B which also do not intersect with the paths from \(\rho _1\) to \(x_1\) and \(\rho _2\) to \(x_2\), respectively, is bounded as follows:
Note that by the choice of \(x_1\) the height of \(\widetilde{T}_{B_1}\) is at least \(p_1-2\) and that in fact all leaves of \(\widetilde{T}_{B_1}\) have at least this distance to the root \(\widetilde{\rho }_1\) of \(\widetilde{T}_{B_1}\) (as the distance from \(x_1\) to \(\widetilde{\rho }_1\) and to \(\rho _1\) is minimal). So this leads to \(|B_1| \ge 2^{p_1-2}\) and thus \(p_1-2 \le \log _2|B_1|\). Analogously, we derive \(p_2-2 \le \log _2|B_m|\). Using this in the above inequality leads to:
where the last equation is due to \(|B_1|+|B_m|+\widetilde{b}=|B|\). Next, we use the fact that \(B_1\) and \(B_m\) are both proper subsets of B and thus we have \(\log _2|B_1| \le \log _2|B|\) as well as \(\log _2|B_m| \le \log _2|B|\). This leads to:
As stated above, we wish to select \(k-2\) edge-disjoint leaf-to-leaf paths in \(B\setminus \{x_1,x_2\}\), and we have shown that at least \(\frac{|B|}{2} -2 \log _2|B|-1.5\) such paths exist. So it remains to show that this number is at least \(k-2\) for \(k\ge 3\) and \(n\ge 20k\).
In this regard, we now set \(g(y):=\frac{1}{2}y-2 \log _2(y) -1.5\) and analyze this function. Using the first derivative \(g'(y)=\frac{1}{2}-\frac{2}{y \ln (2)}\), which equals 0 if and only if \(y= \frac{4}{\ln (2)} \approx 5.77\), as well as the second derivate \(g''(y)=\frac{2}{y^2 \ln (2)}>0 \ \ \forall y \ne 0\), it can be easily seen that g has a local minimum at \(y \approx 5.77\) and that for all values of y larger than this minimum, g is strictly monotonically increasing. Additionally, for \(y=30\) we have \(g(y)\approx 3.686>3= \frac{y}{10}\), so by the monotonicity of g and as \(g'(y)>\frac{1}{10}=\left( \frac{y}{10}\right) '\) for all \(y\ge 8\), we can conclude for all \(|B|\ge 30\):
Note that the third inequality is due to \(|B|\ge \frac{n}{2}\). Moreover, note that as \(k \ge 3\), we have \(n \ge 20k \ge 60\), which guarantees that \(|B|\ge 30\), so that the latter requirement is no restriction. Thus, it is indeed possible to choose \(k-2\) edge-disjoint leaf-to-leaf paths in B, additional to the two paths from \(\rho _1\) to \(x_1\) and \(\rho _2\) to \(x_2\), respectively.
Recall that we labeled all taxa in A with a, and \(x_1\) and \(x_2\) were labeled b. Moreover, we now have chosen \(k-2\) edge-disjoint leaf-to-leaf paths, and for each such path, we label one of its leaves a and the other one b. All other taxa not covered by such paths (if they exist) are labeled with b. We call the resulting character f.
Next, we show that \(l(f,\widetilde{T})=k\). Let us pick one leaf \(a_1\) from \(T_{A_1}\) and one leaf \(a_2\) from \(T_{A_2}\) and consider the paths from \(a_1\) to \(x_1\) and \(a_2\) to \(x_2\), respectively. Together with the \(k-2\) paths chosen from B, this leads to k edge-disjoint paths connecting leaves in state a with leaves in state b. So, by Proposition 1, we have \(l(f,\widetilde{T})\ge k\).
On the other hand, as all leaves in the subtrees \(T_{A_1}\) and \(T_{A_2}\) are in state a, f requires no substitutions in these subtrees. In fact, if we replace \(T_{A_1}\) and \(T_{A_2}\) by leaves \(a_1\) and \(a_2\) and call the resulting tree \(\widehat{T}\) and the resulting restriction of f on the remaining leaves \(\widehat{f}\), we have \(l(f,\widetilde{T})=l(\widehat{f},\widehat{T})\). However, as \(\widehat{f}\) has precisely k leaves in state a, the parsimony score of \(\widehat{f}\) on any tree can be at most k. (A change might be needed on all pending edges leading to the a-leaves, but more changes are definitely not required.) Therefore, we obtain \(l(\widehat{f},\widehat{T})\le k\) and thus also \(l(f,\widetilde{T})\le k\).
Altogether we conclude that \(l(f,\widetilde{T})= k\) and therefore \(f \in A_k(\widetilde{T})\).
Moreover, we now argue that \(l(f,T)<k\) and that therefore \(f \not \in A_k(T)\). In order to see this, we again replace the subtrees \(T_{A_1}\) and \(T_{A_2}\) by leaves \(a_1\) and \(a_2\), respectively, and we again obtain character \(\widehat{f}\), this time in combination with the restriction of T on the smaller leaf set, which we will call \(T'\). Now as above, we have \(l(f,T)=l(\widehat{f},T')\), and as there are only k leaves labeled a, we conclude as above that \(l(f,T)\le k\). However, in \(T'\), the leaves \(a_1\) and \(a_2\) form a cherry \([a_1,a_2]\), so no extension minimizing the changing number to give \(l(\widehat{f},T')\) can require a change for both of these leaves. (If node u incident to both \(a_1\) and \(a_2\) was in state b, there would be two changes on the cherry, but then it would be advantageous to change u to a, as in this case, while there might be an additional change on the edge corresponding to \(\sigma \), two changes, namely on the edges \(\{u,a_1\}\) and \(\{u,a_2\}\), could be saved.) Therefore, in total at most \(k-1\) changes are needed, so in fact, we have \(l(\widehat{f},T')<k\) and thus also \(l(f,T)<k\), which shows that \(f \not \in A_k(T)\).
In summary, we have found a character \(f \in A_k(\widetilde{T})\), for which we know that \(f \not \in A_k(T)\), which shows that \(A_k(\widetilde{T})\ne A_k(T)\). This completes the proof.
4 Discussion and Outlook
The main result of this manuscript, namely that all binary phylogenetic trees with \(n\ge 20k\) leaves are uniquely defined by their induced alignments \(A_k(T)\), can be seen as an extension of Corollary 1, which is a consequence of the classic Buneman theorem 2 [1], as well as Proposition 2. Based on these results as well as based on the computational considerations from [4], we conjecture that the factor of 20 in Theorem 3 can be reduced. However, note that it was already shown in [2] that a factor of 2 is not sufficient whenever \(k\ge 3\) and \(n=2k\). In any case, studying the gap between 2 and 20 is an interesting area for further investigations.
Moreover, note that in [2], the fact that \(A_2(T)\) defines T was only used as a first step (namely, a necessary pre-requisite) to show that T can also be recovered when using maximum parsimony as a criterion for tree reconstruction. In particular, it was shown there that for all \(n \ge 9\), T is the so-called unique maximum parsimony tree for \(A_2(T)\), i.e., \(T=T'\), where \(T'\) is the tree minimizing \(l(A_2(T),T')\). In this regard, we consider the present manuscript as an important step to answer the same question for \(A_k(T)\) whenever \(n \ge 20k\): Can T be uniquely recovered from \(A_k(T)\) when maximum parsimony is used for tree reconstruction? This question is mathematically intriguing, but also relevant for biologists, as maximum parsimony as a simple tree reconstruction criterion is often considered valid whenever the number of changes is relatively small (cf., e.g., [5, 7]). But of course whenever \(A_k(T)\) does not uniquely define T, no tree reconstruction method will be able to uniquely recover T from \(A_k(T)\), which highlights the importance of characterizing cases when \(A_k(T)\) indeed does characterize T. Theorem 3 of the present manuscript represents an essential step in this regard.
Data Availability
Data sharing is not applicable to this article as no new data were created or analyzed in this study.
References
Peter Buneman. The Recovery of Trees from Measures of Dissimilarity, pages 387–395. Edinburgh University Press, 1971.
Mareike Fischer. On the uniqueness of the maximum parsimony tree for data with up to two substitutions: An extension of the classic buneman theorem in phylogenetics. Molecular phylogenetics and evolution, 137:127–137, 2019.
Mareike Fischer and Volkmar Liebscher. On the balance of unrooted trees. J. Graph Algorithms Appl., 25:133–150, 2021.
Pablo A. Goloboff and Mark Wilkinson. On defining a unique phylogenetic tree with homoplastic characters. Molecular Phylogenetics and Evolution, 122:95 – 101, 2018.
J Lin and Masatoshi Nei. Relative efficiencies of the maximum-parsimony and distance-matrix methods of phylogeny construction for restriction data. Molecular biology and evolution, 8 3:356–65, 1991.
Charles Semple and Mike Steel. Phylogenetics (Oxford Lecture Series in Mathematics and Its Applications). Oxford University Press, 2003.
John Sourdis and Masatoshi Nei. Relative efficiencies of the maximum parsimony and distance-matrix methods in obtaining the correct phylogenetic tree. Molecular biology and evolution, 5 3:298–311, 1988.
Mike Steel. Phylogeny: Discrete and Random Processes in Evolution (CBMS-NSF Regional Conference Series). SIAM-Society for Industrial and Applied Mathematics, 2016. ISBN 978-1-611974-47-8.
Acknowledgements
I want to thank Mike Steel for helpful discussions on a previous version of this manuscript. Moreover, want to thank the German Academic Exchange Service DAAD for funding a conference trip to New Zealand in 2019, where I was first inspired to start working on this project. The rest of this project was completed as part of the joint research project DIG-IT!, which is kindly supported by the European Social Fund (ESF), reference: ESF/14-BM-A55-0017/19, and the Ministry of Education, Science and Culture of Mecklenburg-Vorpommern, Germany. Last but not least, I want to thank two anonymous reviewers, whose valuable suggestions helped to improve this manuscript.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author herewith certifies that she has no affiliations with or involvement in any organization or entity with any financial (such as honoraria; educational grants; participation in speakers’ bureaus; membership, employment, consultancies, stock ownership, or other equity interest; and expert testimony or patent-licensing arrangements) or non-financial (such as personal or professional relationships, affiliations, knowledge or beliefs) interest in the subject matter discussed in this manuscript.
Additional information
Communicated by Victor Chepoi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fischer, M. Defining Binary Phylogenetic Trees Using Parsimony. Ann. Comb. 27, 457–467 (2023). https://doi.org/10.1007/s00026-022-00627-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00026-022-00627-x