Abstract
Identifying novel breast cancer biomarkers will improve patient stratification, enhance therapeutic outcomes, and help develop non-invasive diagnostics. We compared the proteomic profiles of whole-cell and exosomal samples of representative breast cancer cell subtypes to evaluate the potential of extracellular vesicles as non-invasive disease biomarkers in liquid biopsies. Overall, differentially-expressed proteins in whole-cell and exosome samples (which included markers for invasion, metastasis, angiogenesis, and drug resistance) effectively discriminated subtypes; furthermore, our results confirmed that the proteomic profile of exosomes reflects breast cancer cell-of-origin, which underscores their potential as disease biomarkers. Our study will contribute to identifying biomarkers that support breast cancer patient stratification and developing novel therapeutic strategies. We include an open, interactive web tool to explore the data as a molecular resource that can explain the role of these protein signatures in breast cancer classification.
Graphical Abstract

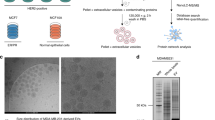
(A) We quantified proteomic profiles of breast cancer cells (BCCs) and breast cancer-derived exosomes (BCDEs) samples from four breast cancer cell lines representative of common breast cancer subtypes – Her2-positive (Her2, MDA-MB-453), Luminal A (LA, MCF7), Luminal B (LB, ZR-75), and triple-negative (TN, MDA-MB-231). (B) We independently performed four comparisons for BCCs and BCDEs, comparing each cell line against the remaining three. (C) We identified differentially-expressed proteins (DEPs) for BCCs and BCDEs, defined protein signatures, and functionally analyzed resulting networks. Through pathway inference analysis (PIA), we identified Kyoto Encyclopedia of genes and genomes (KEGG) subpaths and biological Gene Ontology (GO) terms that displayed differential activation. (D) We validated our proteomic signature using the Cancer Genome Atlas (TCGA) and the Cancer Proteome Atlas (TCPA) databases, verified that differentially-activated pathways in BCDEs caused a corresponding response in receptor cells, and confirmed that the BCDE proteomic signature reflects their cell-of-origin and identifies candidate disease biomarkers in liquid biopsies
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Despite ongoing advances, breast cancer remains a significant health problem. According to the World Health Organization, 2.3 million women worldwide were diagnosed with breast cancer in 2022, with 670,000 succumbing to the disease [1]. Breast cancer represents the leading cause of cancer-related deaths in women younger than 45, with incidence and mortality rates expected to increase significantly in the near future [1]. Unfortunately, breast cancer’s heterogeneity, aggressiveness, and biological complexity make developing safe and effective therapeutic approaches a challenging task [2].
Of the approximately ten different breast cancer subtypes currently described, the expression profile of the estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (Her2) characterize four clinically-relevant molecular subtypes - luminal A (LA; ER+, PR+, Her2-) (40%), luminal B (LB; ER+, PR+, Her2+) (15–20%), Her2-enriched (Her2; ER-, PR-, Her2+) (10–15%), and triple-negative (TN; ER-, PR-, Her2-) (15–20%) [3]. The development of the aggressive TN subtype usually involves metastasis to the brain, which associates with the failure of most therapeutic strategies [4, 5].
Current methods for cancer diagnostics include radiological imaging for detection and biopsy for confirmation, with the latter providing a means to establish the molecular subtype and prognosis; however, these invasive methodologies can provide false positives, and image-based diagnostic tools remain expensive and not widely available. Furthermore, conventional biopsies may not reflect the complete nature of primary/secondary solid tumors due to tumor heterogeneity and distinct genomic profiles [6]. Breast cancer subtype classifiers based on gene expression signatures have significantly improved patient stratification and supported clinical decision-making (7–8). Compared to traditional diagnostic methods, analysis of liquid biopsies of easily-accessible biological fluids (e.g., saliva, urine, and peripheral blood) represents a highly sensitive and minimally invasive diagnostic means to quickly provide results [9]. Liquid biopsies in breast cancer management have provided positive outcomes in early diagnosis, screening, and prognosis prediction [10,11,12]; however, few assays have been clinically validated and incorporated into clinical guidelines, such as those used to detect EGFR mutations in lung cancer, RAS mutations in colorectal carcinoma, or PIK3CA mutations in breast cancer. Additional approaches based on next-generation sequencing focus on analyzing alterations in cell-free tumor DNA isolated from plasma samples and have been gradually incorporated into the clinical setting [13,14,15].
The bodily fluids used for liquid biopsies contain extracellular vesicles such as exosomes and nanovesicles of endosomal origin, which play roles in short- and long-range cell-to-cell communication via the transfer of cargos that include lipids, proteins, and nucleic acid species. Exosomes play essential roles in physiological processes and have been implicated in controlling various disease-related pathways. They also represent a promising source of non-invasive biomarkers [16,17,18] as they are present at elevated levels in cancer patient blood samples [19,20,21], their protected cargo reflects their cell-of-origin (22–23), and tumor cell-derived exosomes possess pro-metastatic potential [24].
Herein, we present the proteomic and functional characterization of breast cancer cells (BCCs) and breast cancer cell-derived exosomes (BCDEs) representative of four clinically-recognized subtypes [25]. This analysis demonstrates intrinsic differences, correlations with clinical scenarios, and applicability as non-invasive biomarkers to improve breast cancer patient stratification and disease monitoring.
Results
Characterization of breast cancer cells and breast cancer cell-derived exosomes
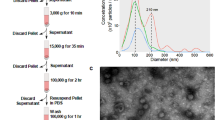
We analyzed BCCs and BCDEs from the MDA-MB-453 (Her2), MCF7 (LA), ZR-75 (LB), and MDA-MB-231 (TN) cell lines to identify potential diagnostic/prognostic biomarkers that may help to stratify patients and guide therapy choice (Fig. 1A describes the overall workflow). We isolated BCDEs by ultracentrifugation of BCC supernatants cultured in exosome-depleted media. We confirmed the receptor profile of BCCs by Western blotting and evaluated the presence of the CD63, CD9, and CD81 tetraspanins (TSPANs) (given their function in exosome-biogenesis and release) and other exosome-related proteins (Hsc70 and Alix) in BCCs and BCDE extracts. The absence of clathrin and calnexin from BCDE samples indicated the lack of contamination with extracellular vesicle populations derived from the plasma membrane or endoplasmic reticulum (Fig. 1B). We determined exosome size and concentration via nanoparticle tracking analysis (NTA) (Fig. 1C), finding that BCDEs possessed sizes of 110–125 nm and that BCCs exhibited specific exosome production/secretion profiles, with LA and LB BCCs giving rise to the most significant number of exosomes. We confirmed the significantly elevated level of BCDEs isolated from LA and LB BCCs found by NTA and the presence of the well-described cup-shaped BCDEs by transmission electron microscopy (TEM) (26–27) (Fig. 1D-E).
Differential protein expression analysis in BCCs and BCDEs
We preprocessed and normalized proteomic data from BCCs and BCDEs to ensure comparability; overall, we quantified 1668 (BCCs) and 389 (BCDEs) UniProtKB/SwissProt proteins. The exploratory data analysis by clustering and principal component analysis (PCA) on both data sets discounted abnormal sample behavior (Supplementary Figure S1). We compared BCCs and BCDEs from each subtype against the remaining three subtypes combined: TN vs. (Her2 + LA + LB), Her2 vs. (TN + LA + LB), LA vs. (TN + Her2 + LB) and LB vs. (TN + Her2 + LA), which will be referred to as TN, Her2, LA, and LB, respectively, for simplicity. Table S1 reports the differentially-expressed proteins (DEPs) in each BCC and BCDE comparison. Overall, BCCs displayed a more significant number of dysregulated proteins, although BCDEs exhibit more significant levels of alterations. We observed the most severe dysregulation (in terms of number) in the TN (265 DEPs) and Her2 comparisons (121 DEPs) in BCCs and the TN (109 DEPs) and Her2 (118 DEPs) comparisons in BCDEs.
We represented significant DEPs from each subtype comparison for BCCs and BCDEs and considered their direction of change (upregulation/downregulation) as shown by Venn diagrams (Fig. 1F). In general, we observed a very low level of intersections among subtypes, with the most common intersection being between LA and the remaining subtypes. Our findings also suggested that most DEPs displayed subtype-specificity in BCCs and BCDEs; the more significant number of specific DEPs makes this especially true for the TN subtype.
We also explored proteomic correlation levels between BCCs and BCDEs through the statistics of the 239 commonly measured proteins. This analysis demonstrated significant positive correlations between parental BCCs and BCDE samples and significant negative correlations between non-parental BCCs and BCDE samples, except for LA and LB (Fig. 1G; Tables S2 and 3). This result suggests that BCDE protein cargo reflects the proteomic profile of their cell of origin and provides evidence for exosomes as a potential source of non-invasive breast cancer biomarkers.
Characterization of BCCs and BCDEs and differential protein expression analysis. (A) Characterization workflow. (B) Western blot of BCC extracts confirming characteristic receptor expression and BCC/BCDE extracts for exosome marker expression (Clathrin/Calnexin as quality control for contamination with other extracellular vesicle types). (C) The concentration and distribution of BCDEs were analyzed with the NanoSight NS300 (vertical axis, x107 particles/ml) and by size (horizontal axis, in nanometers [nm]). (D) Quantitative analysis of BCDEs. (E) Representative TEM images of BCDEs. Scale bar = 200 nm. (F) Intersection of significantly dysregulated proteins by subtype in BCCs and BCDEs. Venn diagrams demonstrate the number of commonly significantly upregulated (“UP”) and downregulated (“DOWN”) proteins in intersections for subtype comparisons. (G) Statistical correlations between subtype comparisons in BCCs (columns) and BCDEs (rows). Correlations among parental BCCs and BCDEs are positive, while non-parental relations present near-zero or negative values. Significant correlations are depicted with a red star. n = 3 in each experiment; error bars represent mean ± SEM, *p < 0.01, **p < 0.001, ***p < 0.0001; one-way ANOVA with post-Bonferroni’s Multiple Comparison Test
Definition of a BCC protein signature with subtype discriminatory potential
We defined a 33-protein signature of BCCs by selecting each comparison’s top ten DEPs and removing duplicates (Table S2). Figure 2A depicts the expression level of said proteins, highlighting protein-specific profiles. Of note, 24 proteins presented a pattern of specific up- or down-regulation in one of the different subtypes (bold in Table S2). Proteins specifically overexpressed in (i) Her2 BCCs mainly associate with metabolism and energy production (e.g., KRT17 and CKB), (ii) LA and LB BCCs play roles in tumor growth and angiogenesis (e.g., GFRA1 and CTSD) and (iii) TN BCCs function in cytoskeletal remodeling, migration, epithelial-mesenchymal transition, and chemoresistance. Of note, specific signature proteins do not present subtype specificity; instead, they display a gradual change across subtypes (e.g., SLC25A22, PDLIM1, CAPN2, FASN, and REEP6), while others displayed specificity to both luminal subtypes (e.g., KRT8, KRT18, HSPB1, and EPHX1).
The 33-protein signature can stratify cancer subtypes, as seen in the sample classification after the unsupervised hierarchical clustering performed (Fig. 2B) and the inter-subtype separation in the PCA plot (Fig. 2C). Both approaches demonstrated the significant difference between TN and the remaining subtypes and the relative similarity between the LA and LB subtypes.
Using STRING, we explored the BCC signature protein-protein interaction (PPI) network and functional profile [28]. The resulting PPI network displayed significantly high levels of connections (p = 6.88e-15, Fig. 2D) and presented a clear structure with a prominent central cluster comprising two-thirds of the 33 proteins and multiple isolated groups of one to three proteins. Functional enrichment revealed twenty-one biological processes (BP), six molecular functions (MF), and thirty-three cellular component (CC) enriched GO terms. These included BP GO functions Cytoskeleton organization (GO: 0007010), Cornification (GO:0070268), Oxidation-reduction process (GO:0055114), Cellular modified amino acid metabolic process (GO:0006575), and CC GO function Invadopodium (GO:0071437), which we highlighted given the interesting pattern of upregulation/downregulation of their related proteins as this provides subtype-specific functional detail.
Her2-specific downregulated BCC proteins related to functions such as Cytoskeleton organization, Cornification, and Invadopodium and upregulated proteins related to Oxidation-reduction and Cellular modified amino acid metabolic processes. LA-specific downregulated BCC proteins related to Cytoskeleton organization, Cellular modified amino acid metabolic process, and Invadopodium, and upregulated proteins related to Cornification, while LB upregulated proteins related to Cornification and downregulated proteins related to Cytoskeleton organization and Invadopodium. Finally, TN-specific upregulated BCC proteins related to Cytoskeleton organization and Invadopodium, and downregulated proteins related to Cornification and Cellular modified amino acid metabolic process. Interestingly, keratins (KRTs) represent currently applied cancer diagnosis biomarkers: TPS (KRT18), TPACYK (KRT8/18), and CYFRA 21 − 1 (KRT19) [28,29,30,31]; here, they displayed common behavior, independent of the remaining proteins related to Cytoskeleton organization and opposite to KRT7 (Fig. 2E).
In summary, a 33-protein signature of BCCs supported the clear discrimination of breast cancer subtypes and, more importantly, provided subtype-specific functional detail that could explain the differential prognosis of each breast cancer subtype.
A 33-protein Signature of BCCs Discriminates Subtypes and Provides Subtype-specific Functional Detail. (A) Barplots reporting the expression levels of the 33 BCC signature proteins (Her2, LA, LB, and TN representing MDA-MB-453, MCF7, ZR-75, and MDA-MB-231). Boxes highlight upregulated (red) and downregulated (green) proteins for each BCC. (B) Heatmap of the expression levels of the 33 BCC signature proteins, with non-supervised clustering of the samples colored by the cell-of-origin. (C) The first and second components of a PCA of samples considering only the protein signature, colored by the cell-of-origin, with ellipse plots with a confidence level of 0.9. (D) PPI analysis of the BCC protein signature. Proteins colored by their association to GO functions Cytoskeleton organization (GO: 0007010), Cornification (GO:0070268), Oxidation-reduction process (GO:0055114), Cellular modified amino acid metabolic process (GO:0006575) and Invadopodium (GO:0071437). (E) Upregulated and downregulated proteins are depicted in red and blue for the specific Her2, LA, LB, and TN networks. For further information on the network, please visit https://version-11-5.string-db.org/cgi/network?networkId=bT6IRAwsY9iC
Definition of a BCDE protein signature with subtype discriminatory potential
We next defined a BCDE protein signature in the same manner by selecting the top ten DEPs in each subtype comparison and removing duplicates, which also comprised 33 proteins (Fig. 3A). Table S3 summarizes the upregulated and downregulated proteins in BCDE samples, highlighting their subtype specificity. In general, we observed a general downregulation of BCDE signature proteins in TN (25 proteins under the median) and Her2 (20 proteins under the median) and an upregulation in LA (22 proteins above the median) and LB (19 proteins above the median). Similar to the BBC signature, the BCDE signature could stratify cancer subtypes, as evidenced in sample classification after unsupervised hierarchical clustering (Fig. 3B) and inter-subtype separation in the PCA plot (Fig. 3C). Importantly, these findings suggest that the protein cargo of BCDEs partly reflects their cell of origin.
As before, we explored the BCDE signature PPI network and functional profile using STRING [28]. The resulting PPI network displayed significantly elevated levels of connections (p < 1.0e-16, Supplementary Figure S2) and possessed a similar structure to the BCC network, with a prominent central cluster comprising two-thirds of the total proteins and multiple isolated clusters of one to two proteins. Functional enrichment of signature proteins with STRING revealed 14 BCC-enriched GO terms in BCDEs; however, PPI analysis failed to reveal associations between protein clusters and biological functions in this case. Supplementary Figs. S2B-E report the different patterns of dysregulation in Her2, the less differentiated patterns in LA and LB, and the general downregulation of signature proteins in TN, respectively.
We next explored the efficacy of our protein signatures to stratify breast cancer subtypes through The Cancer Genome Atlas (TCGA) and The Cancer Proteome Atlas (TCPA). PCAs revealed that both BCC and BCDE protein signatures discriminated TN from the LA and LB subtypes in the TCGA dataset but had more difficulty separating Her2 and could not discriminate between LA and LB (Fig. 3D). Interestingly, the BCDE protein signature represented the most powerful means of separating subtypes, especially for the TN subtype, providing evidence for the potential of exosomes as a source of subtype biomarkers in liquid biopsies. In the TCPA dataset, BCC and BCDE signatures could discriminate the TN subtype from the remaining subtypes (Fig. 3D). While further studies will be needed to corroborate our findings, these preliminary results offer an interesting starting point when considering liquid biopsies as a tool for TN breast cancer diagnosis.
Validation of a 33-protein signature in BCDEs. (A) Barplots reporting the expression level of the 33 BCDE signature proteins (Her2, LA, LB, and TN representing MDA-MB-453, MCF7, ZR-75, and MDA-MB-231). Boxes highlight upregulated (red) and downregulated (green) proteins for each BCDE. (B) Heatmap of expression levels of the 33 BCDE signature proteins, with non-supervised clustering of the samples colored by the cell-of-origin. (C) The first and second components of a PCA of samples considering only the protein signature, colored by the cell-of-origin, with ellipse plots with a confidence level of 0.9. (D) PCA plots of samples in the TCGA (left) and TCPA (right) datasets for the BCC and BCDE signatures proteins. For both validation steps, Her2, LA (LumA), LB (LumB), and TN (Basal) represent MDA-MB-453, MCF7, ZR-75, and MDA-MB-231
Pathway analysis reveals functional differences in BCC subtypes
We next performed pathway inference analysis (PIA) with the Hipathia package [32,33,34,35] in BCCs to explore dysregulated pathways and subpathways for each subtype from the Kyoto Encyclopedia of Genes and Genomes (KEGG) signaling pathways (34–35). Hipathia divides KEGG pathways into subpathways, which are defined as all the nodes in a path ending in a particular final node (also called the effector protein). A subpathway exists for each final node within a KEGG signaling pathway; thus, Hipathia infers the activation level for each subpathway separately as an independent biological entity. After analyzing all available subpathways, we discovered 234 (Her2), 38 (LA), 26 (LB), and 146 (TN) altered subpathways in BCCs (Table S4).
A summary of the number of altered subpathways within each pathway provided potentially interesting differences between BCC subtypes. Figure 4A depicts the top altered pathways, which support the definition of two pattern types: (i) pathways altered mainly in one subtype (e.g., cAMP, HIF-1, and NF-kB signaling pathways in Her2; AGE-RAGE signaling pathway in diabetic complications in LB; or Chemokine signaling pathway in TN) and (ii) pathways altered at the same time in different subtypes (e.g., Toll-like receptor signaling pathway, Rap1 signaling pathway, or Proteoglycans in cancer).
The subpathways associated with the second group also presented with subtype-specific dysregulation patterns for BCCs. For instance, the Rap1 signaling pathway (Fig. 4B), which promotes tumor invasion and metastasis in breast cancer [36], includes seven dysregulated subpathways in at least one subtype (identified in Fig. 4C). The subroute ending in the THBS1 effector protein (the THBS1 subroute, in blue), which relates to defective angiogenesis in the pathway (Fig. 4C), displayed overactivation in Her2 and TN and underactivation in LA and LB. The ITGA2B subroute (lilac), which functions in cell adhesion and migration, displayed a similar pattern. Cell adhesion and migration also associated with the ACTB, PRKC1/PARD6A/PARD3, and ITGAL subpathways (pink, red, and orange, respectively), which displayed deactivation in Her2. Additionally, the ITGAL subroute displayed deactivation in LA and activation in TN. The MAPK1 subroute (yellow), related to proliferation and survival, displayed overactivation in Her2, while the RAC1 subroute (dark lilac), related to SMG crosstalk, displayed overactivation in TN. We observed similar differential patterns to the Rap1 signaling pathway in additional pathways, including the Proteoglycans in cancer, Pathways in cancer, Hepatitis B, and Toll-like receptor signaling pathways (Fig. 4D-G, with subpathways of Fig. 4D and E identified in Supplementary Figs. S3 and 4). Our results indicate that although the Rap1 signaling pathway becomes systematically altered in breast cancer, each subtype possesses specific alterations in the pathway (activating or deactivating specific subpathways) and causes different potential functional outcomes. This knowledge may support the development of tailored treatments that respond to each patient’s specific requirements.
Pathway analysis reveals altered functional outcomes according to BCC subtype. (A) Number of subpathways altered in the top altered KEGG pathways in each BCC (Her2, LA, LB, and TN representing MDA-MB-453, MCF7, ZR-75, and MDA-MB-231). Framed pathways are further represented in B and D-G. (B) The dotplot represents the altered subpathways’ statistical value within the Rap1 signaling KEGG pathway. (C) Representative example, detail of the Rap1 signaling KEGG pathway in each BCC. Specific parts of the altered subpathways are colored following the column Subpathway in (B). Common components are colored in light gray. For further, visit https://www.genome.jp/pathway/hsa04015. (D-G) Dot plots representing the statistical value of altered subpathways within the (D) Proteoglycans in Cancer, (E) Pathways in cancer, (F) Hepatitis, and (G) Toll-like receptor signaling pathway KEGG pathways in each BCC subtype
Functional impact of BCDE cargo in receptor cell pathways
We hypothesized that the functional characterization of BCDEs should be interpreted as the functional change that BCDEs induce in receptor cells; therefore, we analyzed altered pathways in BCDEs with Hipathia analogously to BCCs. We interpreted the PIA in BCDEs as the impact of BCDE protein cargo on signaling pathways in target cells, finding 49 (Her2), 47 (LA), 2 (LB), and 133 (TN) subpathways altered in BCDEs (Table S4). Summarizing the number of altered subpathways within each KEGG pathway, we found considerable differences among BCDEs (Fig. 5A), with specific significant alterations to Ovarian steroidogenesis, the Glucagon signaling pathway, and Adrenergic signaling in cardiomyocytes in TN BCDEs, Prostate cancer in Her2 BCDEs, and Retrograde endocannabinoid signaling in LA BCDEs. We also discovered KEGG pathways commonly altered in all BCDEs, such as Adherens Junction (Fig. 5B), whose altered subpathways are characterized in Fig. 5C. We also found the ErbB signaling pathway, Proteoglycans in cancer, and Pathways in cancer KEGG pathways altered in Her2 and TN BCDEs (Fig. 5D and F).
Interestingly, the Her2 and TN subtypes present with overexpressed pathways whose effector proteins relate to migration and invasion processes. For instance, the Adherens junction subpathway with SNAI2 effector protein (plays an essential role in promoting invasion and metastasis) and the subpathway from Proteoglycans in cancer ending in HOXD10 (related to cell migration and invasion) become upregulated in Her2 (Supplementary Fig. S5). Furthermore, the activity of specific proteins (PXN, PTK2, PRKCA, NUDT16L1, HSPB2, and ACTB) through the Proteoglycans in cancer pathway (related to migration and invasion) became increased in TN. The LA and LB subtypes present with the overactivation of subpathways concerning the effector protein CTNND1 (orange and red in Fig. 5B), which functions in angiogenesis and metastasis repression.
Pathway analysis reveals altered functional outcomes according to BCDE subtypes. (A) The number of subpathways in the top ten altered KEGG pathways in each BCDE (Her2, LA, LB, and TN representing MDA-MB-453, MCF7, ZR-75, and MDA-MB-231). Framed pathways are further represented in B and D-E. (B) The dotplot represents the altered subpathways’ statistical value within the Adherens junction KEGG pathway. (C) Representative example, detail of KEGG pathway Adherens junction. Specific parts of the altered subpathways are colored following the column Subroute in B. All nodes in subpathway CTNND1** (red) are included in subroute LEF1 CTNNB1 (orange) except for pl20ctn, representing node CTNND1**. For further details, visit https://www.genome.jp/pathway/hsa04520. (D-F) Dot plots representing the statistical value of the altered subpathways within the (D) Proteoglycans in Cancer, (E) ErbB signaling pathway, and (F) Pathways in Cancer KEGG pathways in each analyzed BCDE
BCDEs differentially influence breast cancer progression as a reflection of the cell of origin
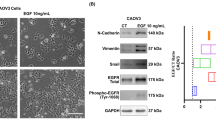
We next conducted two experiments to corroborate our hypothesis that the BCDE protein cargo functionally impacts target cells. As seen in Fig. 5, the functional results suggested that TN and Her2 BCDEs impact migration and invasion, while LA and LB BCDEs mainly impact the vascular endothelium and angiogenesis events during cancer progression; thus, we performed tube formation and scratch wound healing assays in the presence of BCDEs from specific subtypes to evaluate their potential to alter pathways according to their cargo.
Phase-contrast microscopy images from tube formation assays using human umbilical vein endothelial cells (HUVECs) revealed that LA and LB BCDEs contributed to more complete and complex nets with a more significant number of tube-like structures than TN and Her2 BCDEs or PBS (as vehicle control treatment) (Fig. 6A). The quantification of tubes, loops, and branching points (as representative measures of angiogenesis) demonstrated that LB BCDEs significantly increased the ability of HUVECs to form these structures compared to PBS (Fig. 6B-D). LA BCDEs significantly increased loop numbers compared to PBS (Fig. 6C) while LB BCDEs significantly increased loop numbers compared to TN BCDEs (Fig. 6C) and tube-like structure formation and branching points compared to TN and Her2 BCDEs (Fig. 6B and D).
In the scratch wound healing assay, we exposed fibroblasts to BCDEs and monitored scratch closure over 24 h as a proxy for migration and invasion. In this case, TN (but not Her2, LA, or LB) BCDEs significantly increased fibroblast migration compared to PBS and LA BCDEs (Fig. 6E).
In summary and in agreement with the previous pathways subroute analysis (Fig. 5), the tube formation assay results suggest that LA and LB BCDEs have a more significant impact on angiogenesis than TN and Her2 BCDEs, with LB BCDEs having the most significant impact. The scratch wound healing assay results suggest that TN BCDEs have a more pronounced invasive and metastatic potential. Together, these results support the proposed interpretation of the functional results of the PIA analysis as the different functional impact of BCDE subtypes in recipient cells according to their cell of origin.
Differential contribution of BCDEs to migration and angiogenesis according to subtype. (A) Bright-field images of tube formation assays. HUVECs were seeded (4.0 × 104) on 48-well plates coated with Matrigel and incubated with PBS or BCDEs (5 µg). (B-D) Quantitative analysis representative of (B) tubes, (C) loops, and (D) branching points formed by HUVECs cultured on Matrigel-coated plates after a 12 h incubation with BCDEs. (E) Quantitative analysis of wound healing assays using fibroblasts (expressed as % fibroblast migration). Fibroblasts were treated with 5 µg BCDEs and monitored for 24 h; fibroblasts were treated with PBS as a control. Bars represent mean ± SEM, **p < 0.01, ***p < 0.001; one-way ANOVA with the post, “Bonferroni’s Multiple Comparison Test of three different experiments, n = 3 wells in each experiment
ExoCellBC web tool
We have made all data and results available within the ExoCellBC web tool (https://bioinfo.cipf.es/metafun-ExoCellBC), which is freely accessible to any user and allows the confirmation of the results described in this manuscript and the exploration of other results of interest. This easy-to-use resource is divided into different sections: [1] summary of analysis results in each phase. Then, for each of the datasets (BCC and BCDE), the detailed results of the [2] exploratory analysis, [3] differential expression, and [4] pathway analysis, as well as [5] the protein-protein interaction analysis results. The user can interact with the web tool through graphics and tables and search for specific information for a protein or pathway.
Discussion
While advances such as molecular characterization and transcriptional profiling have prompted a degree of improvement [37,38,39,40,41], breast cancer patient stratification remains a challenge. Proteomics has provided an additional step towards fully describing tumor heterogeneity and the associated underlying mechanisms (by allowing the detection of protein markers); meanwhile, liquid biopsies [42] have seen an increase in use as a non-invasive means of diagnosing, monitoring, and treating cancer. Here, we thoroughly explored the proteomic profile of BCCs and BCDEs representative of the most common breast cancer subtypes.
We present a 33-protein signature of deregulated BCC proteins (potential biomarkers) to potentially guide therapeutic management, representing breast cancer subtype heterogeneity (Fig. 2). Specifically, deregulated proteins in Her2 (KRT17, SQOR, CKB, CKMT1B/CKMT1A, and ALDH4A1) highlight high metabolic rates and energy demands and their ability to grow and spread more rapidly than other subtypes (43–44). Çubuk et al. previously described the importance of crosstalk between metabolism and signaling pathways in BC [45]. Specifically, deregulated proteins in luminal subtypes (GFRAI, CA2, and UGDH in LA) (IARS, SLCA3, and CTSD in LB) associated with angiogenesis, suggesting the use of antiangiogenic therapies [46,47,48]. LA and LB shared similar profiles, with commonly deregulated proteins, including the KRTs; indeed, CK7/8, 18, and 19 expression indicated a differentiated glandular phenotype and a better prognosis than Her2 or TN (49–50). Finally, specifically upregulated proteins in TN (e.g., TUBB6, MSN, GSTP1, ANXA1, VIM, and ITGB1) participate in cytoskeletal remodeling, migration, epithelial-mesenchymal transition, metastasis, and chemoresistance, which reflects this subtype’s invasive and metastatic nature (Fig. 2E).
The PPI analysis of BCC signature proteins revealed an elevated level of interconnection, with the functional analysis encountering 21 enriched functions (Fig. 4). Proteins related to Cellular modified amino acid metabolic processes (which could influence proliferation, survival, and drug resistance) [43,44,45, 51] became upregulated in Her2 and downregulated in LA and TN. Proteins related to Cornification became downregulated in Her2 and TN and upregulated in LA and LB. Of note, KRT7 - which displayed a different behavior from KRT8, KRT18, and KRT19 - has been identified as a prognostic breast cancer biomarker, with elevated gene expression associated with poorer outcomes, especially in basal and HER2 + subtypes. Elevated KRT7 levels have also been linked to a poorer 12-month response to treatment, particularly for patients undergoing platinum-based chemotherapy. Of note, KRT18 and 19 have been recognized as robust diagnostic and prognostic biomarkers for various human cancers, including breast cancer, where overexpression correlates with poorer patient prognosis and the loss of estrogen/progesterone receptors. Proteins related to Cytoskeleton organization but not to Cornification became downregulated in Her2, LA, and LB and upregulated in TN, revealing the highly invasive mesenchymal-like breast cancer phenotype of the TN subtype (36, 52–53). Proteins related to Oxidation-reduction process displayed a heterogeneous pattern, with a more significant upregulation observed in Her2 in accordance with their dysregulated metabolism [44].
Exosomes represent a potent intercellular communication mechanism that mediates tumor-host crosstalk by modifying the activity of signaling pathways in recipient cells and inducing gene transcription alterations [54]. Recent research highlighted the role of exosomes as biomarkers in liquid biopsies [55,56,57,58]. We reported a 33-protein signature in BCDEs, which stratified breast cancer subtypes and recapitulated the cell-of-origin phenotype, indicating the potential of BCDEs as diagnostic/prognostic biomarkers in liquid biopsies [59] (Fig. 3).
Among the 33 BCDE proteins, we noted the specific upregulation of EDIL3 - previously observed in invasive breast cancer cell extracellular vesicles [60] - in TN BCDEs. Her2 BCDEs specifically upregulate SUSD2, ERBB2, CKB, and CLCA2, which function in drug resistance, energy transduction, and non-invasive phenotypes (61–62). Proteins specifically enriched in LA BCDEs reflected the tumorigenic potential and drug resistance mechanisms of these less invasive breast cancer subtypes. The immunohistochemical detection of mucins (e.g., MUC5AC and MUC5B) correlates with several clinicopathologic characteristics in breast cancer patients [63]; moreover, MUC5B expression represents an early event in mucinous breast cancer development (64–65). Therefore, Mucin proteins in exosomes may represent a prognostic/diagnostic biomarker in LA breast cancer.
The presence of the CTSD extracellular membrane protein in LB BCDEs may support the design of targeted therapeutic strategies for LB metastasis to enhance the selectivity/efficiency of current treatments. CTSD, a member of the cathepsin family, plays roles in various physiological/pathological processes, including cancer progression and metastasis. The cathepsins resident in the extracellular space (including CTSD) can cleave extracellular matrix proteins, cell receptors, or cytokines, thereby influencing the tumor microenvironment and promoting metastasis. In the context of breast cancer, changes in cathepsin expression during metastatic progression have been observed, indicating the potential role of CTSD in modulating the tumor microenvironment during metastasis [48]. The potential of CTSD as a therapeutic target is further supported by research into lysosomal functions, given the involvement of CTSD in lysosomal degradation pathways. This opens the possibility of developing targeted therapeutic strategies that leverage the role of CTSD in the extracellular space and disease processes. In summary, the presence of CTSD in LB BCDEs presents an opportunity to explore targeted therapeutic interventions that take advantage of its extracellular functions, potentially impacting cancer metastasis.
In parallel, we detected the presence of SLC7A5/SLC3A2 in LB BCDEs, whose co-expression has previously been reported in a subgroup of ER+/Her2- breast cancer patients who fail to benefit from endocrine therapy [66]. The BCDE signature also includes BASP1, which displayed upregulation in LA and TN and has been reported to interact with ERα and enhance the anti-cancer effects of tamoxifen breast cancer treatment, enhancing patient survival [67]; however, BASP1 expression also associates with lung cancer brain metastasis (68–69). Identifying these proteins in exosomes highlights their potential as a source of biomarkers in liquid biopsies [69].
We validated the BCC and BCDE signatures on TCGA (transcriptomics) and TCPA (proteomics) datasets, finding promising separations between subtypes. Interestingly, the BCDE signature discriminated TN from remaining subtypes, thereby representing an interesting starting point when considering BCDEs as TN biomarkers in liquid biopsies. While our models were based on purified BCCs and BCDEs, TCGA and TCPA data are obtained from tissues comprising many cell types, which could limit the discriminatory power of the reported signatures. Even given these limitations, the validation of the proteomic signatures with TCGA/TCPA datasets provides evidence for the robust nature of our findings (Fig. 5).
Additional functional results in BCC by PIA revealed the specific dysregulation of different parts of hallmark-related signaling pathways for each BCC subtype (Rap1 and Toll-like signaling pathways, Proteoglycans in cancer, and Pathways in cancer). This information may guide drug development and disease management for each subtype. For instance, the AGE-RAGE signaling pathway, which modulates cellular redox balance and is relevant for tumor blood endothelial cell survival in the tumor microenvironment (70–71), displayed significant alterations in LA and LB BCCs. An analysis of the top ten altered subpathways in each subtype highlighted their main features: metabolism and energy (Her2), actin cytoskeletal regulation, VEGF signaling (LA and LB), and invasiveness (TN) [49]. While LA and LB BCCs display a degree of invasiveness, their proteomic and functional profiles highlight their less aggressive nature. The PIA analysis proved that while BCC subtypes alter common signaling pathways, the proteins involved and subpathways affected may differ. Identifying these vital differential players may be crucial in improving patient disease management.
Concerning the pathway functional analysis performed on BCDEs, the application of functional analysis methods to omics data has helped to understand the biological implications of the genes/proteins altered in a study; however, this approach assumes that the genes/proteins are accompanied by the necessary cell machinery allowing them to perform their function. In the case of our BCDE dataset, we cannot assume that proteins within BCDEs have the same functions as they would have in a cell, nor that they interact following the same pathways as in a cell. Thus, the interpretation of a pathway and functional characterization in BCDEs cannot be performed in a manner homologous to BCCs, and BCDEs cannot be considered the performers of the altered functions. Since BCDEs transport proteins to receptor cells, and different protein cargos will affect target cells differently, we hypothesized that the functional characterization of BCDEs should be interpreted as the functional change BCDEs would induce in receptor cells.
The PIA performed in BCDEs revealed that TN BCDEs demonstrated significant differences from the remaining subtypes, suffering from alterations in pathways and subpathways related to Adherens junction, Pathways in cancer, ErbB signaling pathways, and Proteoglycans in cancer. These findings highlight the role of TN BCDEs in proliferation, invasion, metastasis, and drug resistance. Pathways specifically upregulated in LA and LB BCDEs instead related to angiogenic processes. In vitro assays (wound healing and tube formation) supported the differential role of BCDEs in specific oncogenic processes (migration and angiogenesis) depending on BCC phenotypes. LA and LB BCDEs had a more substantial influence on angiogenesis than TN and Her2 BCDEs, which correlated well with the proteomic results. Overall, our findings provide evidence that the specific cargo and cell origin influence the role of BCDEs in cancer progression.
To the best of our knowledge, we present the first proteomic and functional characterization of BCC and BCDE samples from cell lines representative of clinical breast cancer subtypes, revealing potential subtype-specific biomarkers and disease mechanisms. Our study identified protein signatures in BCCs and BCDEs that stratified subtypes and indicated that the protein cargo of exosomes functionally impacts receptor proteins in a manner that depends on the cell of origin. Of note, this preliminary study on purified cells does not consider heterogeneity in in vivo tumor cells and tissues. Furthermore, we focused on specific breast cancer subtypes that may not fully represent the entire range of molecular subtypes, especially for the highly heterogeneous TN subtype. Further studies in patient-derived samples will be necessary to corroborate our findings and define a TN proteomic signature with diagnostic capacity.
In conclusion, our study revealed protein signatures in BCCs and BCDEs that define subtypes, with significant positive correlations observed between BCCs and BCDEs. The BCDE signature offers a promising starting point for further analysis of the role of exosomes as biomarkers of the TN subtype in liquid biopsies; moreover, we provide evidence that BCDEs from different subtypes impact receptor cells differentially, triggering changes that replicate the functional profile of their cells of origin. These findings agree with related studies, reinforcing that BCDEs recapitulate the phenotype of parental cells and validate their role as potential biomarkers in liquid biopsies. The information and web tool may help guide further studies to improve multigene panels, personalized therapeutic approaches, and monitor cancer progression.
Data availability
The mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifiers PXD024304 (proteomic and pathway profiling of four BCCs) and PXD024300 (proteomic and pathway profiling of four BCDEs). Results data are available in the ExoCellBC web tool (https://bioinfo.cipf.es/metafun-ExoCellBC).
References
a) https://www.who.int/news-room/fact-sheets/detail/breast-cancer; b) Anastasiadi Z, Lianos GD, Ignatiadou E, Harissis HV, Mitsis M (2017) Breast cancer in young women: an overview. Updat Surg. 69(3):313-7
Januškevičienė I, Petrikaitė V (2019) Heterogeneity of breast cancer: the importance of interaction between different tumor cell populations. Life Sci 239:117009
Perou CM, Sørlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA et al (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752
Abouharb S, Ensor J, Loghin ME, Katz R, Moulder SL, Esteva FJ et al (2014) Leptomeningeal disease and breast cancer: the importance of tumor subtype. Breast Cancer Res Treat 146(3):477–486
Bai X, Ni J, Beretov J, Graham P, Li Y (2021) Triple-negative breast cancer therapeutic resistance: where is the Achilles’ heel? Cancer Lett 497:100–111
Nikanjam M, Kato S, Kurzrock R (2022) Liquid biopsy: current technology and clinical applications. J Hematol OncolJ Hematol Oncol 15(1):131
Sarhangi N, Hajjari S, Heydari SF, Ganjizadeh M, Rouhollah F, Hasanzad M (2022) Breast cancer in the era of precision medicine. Mol Biol Rep 49(10):10023–10037
Falato C, Schettini F, Pascual T, Brasó-Maristany F, Prat A (2023) Clinical implications of the intrinsic molecular subtypes in hormone receptor-positive and HER2-negative metastatic breast cancer. Cancer Treat Rev 112:102496
De Rubis G, Rajeev Krishnan S, Bebawy M (2019) Liquid biopsies in Cancer diagnosis, monitoring, and prognosis. Trends Pharmacol Sci 40(3):172–186
Alimirzaie S, Bagherzadeh M, Akbari MR (2019) Liquid biopsy in breast cancer: a comprehensive review. Clin Genet 95(6):643–660
Mazzitelli C, Santini D, Corradini AG, Zamagni C, Trerè D, Montanaro L et al (2023) Liquid Biopsy in the management of breast Cancer patients: where are we now and where are we going. Diagn Basel Switz 13(7):1241
de Freitas AJA, Causin RL, Varuzza MB, Calfa S, Hidalgo Filho CMT, Komoto TT et al (2022) Liquid Biopsy as a Tool for the diagnosis, treatment, and monitoring of breast Cancer. Int J Mol Sci 23(17):9952
Verner EL, Jackson JB, Severson E, Valkenburg KC, Greer AE, Riley DR et al (2023) Validation of the Labcorp Plasma Focus Test to Facilitate Precision Oncology through cell-free DNA genomic profiling of solid tumors. J Mol Diagn JMD 25(7):477–489
Pancrazzi A, Bloise F, Moncada A, Perticucci R, Vecchietti S, Pompili F et al BL-MOL-AR project, preliminary results about Liquid Biopsy: Molecular Approach Experience and Research Activity in Oncological Settings. Glob Med Genet. 10(3):172–187
Maity AP, Gangireddy M, Degen KC, Al-Saleem FH, Bramson J, Ciocca V et al Impact of Simultaneous Circulating Tumor DNA and Tissue Genotyping in the Workup of Stage IV Lung Adenocarcinoma on Quality of Care in an Academic Community Medical Center. JCO Oncol Pract;19(8):620-5
Mitchell MI, Ma J, Carter CL, Loudig O (2022) Circulating Exosome Cargoes Contain functionally diverse Cancer biomarkers: from Biogenesis and function to purification and potential translational utility. Cancers 14(14):3350
Pinzani P, D’Argenio V, Del Re M, Pellegrini C, Cucchiara F, Salvianti F et al (2021) Updates on liquid biopsy: current trends and future perspectives for clinical application in solid tumors. Clin Chem Lab Med 59(7):1181–1200
Srivastava A, Moxley K, Ruskin R, Dhanasekaran DN, Zhao YD, Ramesh R (2018) A non-invasive liquid biopsy screening of urine-derived exosomes for miRNAs as biomarkers in Endometrial Cancer patients. AAPS J 20(5):82
Andreola G, Rivoltini L, Castelli C, Huber V, Perego P, Deho P et al (2002) Induction of lymphocyte apoptosis by tumor cell secretion of FasL-bearing microvesicles. J Exp Med 195(10):1303–1316
Huber V, Fais S, Iero M, Lugini L, Canese P, Squarcina P et al (2005) Human colorectal cancer cells induce T-cell death through release of proapoptotic microvesicles: role in immune escape. Gastroenterology 128(7):1796–1804
Logozzi M, De Milito A, Lugini L, Borghi M, Calabrò L, Spada M et al (2009) High levels of exosomes expressing CD63 and caveolin-1 in plasma of melanoma patients. PLoS ONE 4(4):e5219
Ha DH, Kim HK, Lee J, Kwon HH, Park GH, Yang SH et al (2020) Mesenchymal Stem/Stromal cell-derived exosomes for Immunomodulatory Therapeutics and skin regeneration. Cells 9(5):1157
Cano A, Ettcheto M, Bernuz M, Puerta R, Esteban de Antonio E, Sánchez-López E et al (2023) Extracellular vesicles, the emerging mirrors of brain physiopathology. Int J Biol Sci 19(3):721–743
Hoshino A, Costa-Silva B, Shen TL, Rodrigues G, Hashimoto A, Tesic Mark M et al (2015) Tumour exosome integrins determine organotropic metastasis. Nature 527(7578):329–335
Masood S (2016) Breast cancer subtypes: morphologic and biologic characterization. Womens Health Lond Engl 12(1):103–119
Jung MK, Mun JY (2018) Sample Preparation and Imaging of exosomes by Transmission Electron Microscopy. J Vis Exp. (131):56482
Andreu Z, Masiá E, Charbonnier D, Vicent MJ (2023) A Rapid, Convergent Approach to the identification of Exosome inhibitors in breast Cancer models. Nanotheranostics 7(1):1–21
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J et al (2019) STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47(D1):D607–D613
Bodenmüller H, Donié F, Kaufmann M, Banauch D (1994) The tumor markers TPA, TPS, TPACYK and CYFRA 21 – 1 react differently with the keratins 8, 18 and 19. Int J Biol Markers 9(2):70–74
Lane EB, Alexander CM (1990) Use of keratin antibodies in tumor diagnosis. Semin Cancer Biol 1(3):165–179
van Dalen A (1996) Significance of cytokeratin markers TPA, TPA (cyk), TPS and CYFRA 21.1 in metastatic disease. Anti-cancer Res 16(4B):2345–2349
Hidalgo MR, Cubuk C, Amadoz A, Salavert F, Carbonell-Caballero J, Dopazo J (2017) High throughput estimation of functional cell activities reveals disease mechanisms and predicts relevant clinical outcomes. Oncotarget 2017; 8(3), 5160–5178
Amadoz A, Hidalgo MR, Çubuk C, Carbonell J, Dopazo J (2018) A comparison of mechanistic signaling pathway activity analysis methods. Brief Bioinform 20:1655–1668
Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45(D1):D353–D361
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30
Zhang YL, Wang RC, Cheng K, Ring BZ, Su L (2017) Roles of Rap1 signaling in tumor cell migration and invasion. Cancer Biol Med 14(1):90–99
Chia SK, Bramwell VH, Tu D, Shepherd LE, Jiang S, Vickery T et al (2012) A 50-gene intrinsic subtype classifier for prognosis and prediction of benefit from adjuvant tamoxifen. Clin Cancer Res off J Am Assoc Cancer Res 18(16):4465–4472
Pu M, Messer K, Davies SR, Vickery TL, Pittman E, Parker BA et al (2020) Research-based PAM50 signature and long-term breast cancer survival. Breast Cancer Res Treat 179(1):197–206
Weigelt B, Geyer FC, Reis-Filho JS (2010) Histological types of breast cancer: how special are they? Mol Oncol 4(3):192–208
Senkus E, Kyriakides S, Ohno S, Penault-Llorca F, Poortmans P, Rutgers E et al (2015) Primary breast cancer: ESMO Clinical Practice guidelines for diagnosis, treatment and follow-up. Ann Oncol off J Eur Soc Med Oncol 26(Suppl 5):v8–30
Coates AS, Winer EP, Goldhirsch A, Gelber RD, Gnant M, Piccart-Gebhart M et al (2015) Tailoring therapies–improving the management of early breast cancer: St Gallen International Expert Consensus on the primary therapy of early breast Cancer 2015. Ann Oncol off J Eur Soc Med Oncol 26(8):1533–1546
Caputo V, Ciardiello F, Corte CMD, Martini G, Troiani T, Napolitano S (2023) Diagnostic value of liquid biopsy in the era of precision medicine: 10 years of clinical evidence in cancer. Explor Target Anti-Tumor Ther 4(1):102–138
Holloway RW, Marignani PA (2021) Targeting mTOR and glycolysis in HER2-Positive breast Cancer. Cancers 13(12):2922
Ligorio F, Pellegrini I, Castagnoli L, Vingiani A, Lobefaro R, Zattarin E et al (2021) Targeting lipid metabolism is an emerging strategy to enhance the efficacy of anti-HER2 therapies in HER2-positive breast cancer. Cancer Lett 511:77–87
Cankut, Çubuk, Carlos, Loucera (2023) María Peña-Chilet, and Joaquin Dopazo. Crosstalk between Metabolite Production and Signaling Activity in Breast Cancer. Int J Mol Sci. 24(8): 7450
Chen M, Ba H, Lu C, Dai J, Sun J (2018) Glial cell line-derived neurotrophic factor (GDNF) promotes angiogenesis through the demethylation of the fibromodulin (FMOD) promoter in Glioblastoma. Med Sci Monit 24:6137–6143
Yu Yming, Xu L, Li H, ran, Zhang T qi, Qian G, Li Lfeng et al (2021) IARS2 regulates proliferation, migration, and angiogenesis of human umbilical vein endothelial cells. Rev Assoc Médica Bras. 67(4):555 – 60
Pranjol MZI, Gutowski NJ, Hannemann M, Whatmore JL (2018) Cathepsin D non-proteolytically induces proliferation and migration in human omental microvascular endothelial cells via activation of the ERK1/2 and PI3K/AKT pathways. Biochim Biophys Acta BBA -. Mol Cell Res 1865(1):25–33
Abd El-Rehim DM, Pinder SE, Paish CE, Bell J, Blamey R, Robertson JF et al (2004) Expression of luminal and basal cytokeratins in human breast carcinoma. J Pathol 203(2):661–671
Dai X, Cheng H, Bai Z, Li J (2017) Breast Cancer cell line classification and its relevance with breast tumor subtyping. J Cancer 8(16):3131–3141
Bian X, Liu R, Meng Y, Xing D, Xu D, Lu Z (2021) Lipid metabolism and cancer. J Exp Med 218(1):e20201606
Itatani Y, Kawada K, Inamoto S, Yamamoto T, Ogawa R, Taketo M et al (2016) The role of chemokines in promoting Colorectal Cancer Invasion/Metastasis. Int J Mol Sci 17(5):643
Theocharis AD, Skandalis SS, Neill T, Multhaupt HAB, Hubo M, Frey H et al (2015) Insights into the key roles of proteoglycans in breast cancer biology and translational medicine. Biochim Biophys Acta BBA - Rev Cancer 1855(2):276–300
Liu J, Ren L, Li S, Li W, Zheng X, Yang Y et al (2021) The biology, function, and applications of exosomes in cancer. Acta Pharm Sin B 11(9):2783–2797
Zhou L, Wang W, Wang F, Yang S, Hu J, Lu B et al (2021) Plasma-derived exosomal miR-15a-5p as a promising diagnostic biomarker for early detection of endometrial carcinoma. Mol Cancer 20(1):57
Castillo J, Bernard V, San Lucas FA, Allenson K, Capello M, Kim DU et al (2018) Surfaceome profiling enables isolation of cancer-specific exosomal cargo in liquid biopsies from pancreatic cancer patients. Ann Oncol off J Eur Soc Med Oncol 29(1):223–229
Zheng M, Hou L, Ma Y, Zhou L, Wang F, Cheng B et al (2019) Exosomal let-7d-3p and miR-30d-5p as diagnostic biomarkers for non-invasive screening of cervical cancer and its precursors. Mol Cancer 18(1):76
DeCastro J, Littig J, Chou PP, Mack-Onyeike J, Srinivasan A, Conboy MJ et al (2021) The microfluidic toolbox for analyzing exosome biomarkers of aging. Mol Basel Switz 26(3):535
Lobb RJ, Hastie ML, Norris EL, Van Amerongen R, Gorman JJ, Möller A (2017) Oncogenic transformation of lung cells results in distinct exosome protein profile similar to the cell of origin. Proteomics 17(23–24):1600432
Lee JE, Moon PG, Cho YE, Kim YB, Kim IS, Park H et al (2016) Identification of EDIL3 on extracellular vesicles involved in breast cancer cell invasion. J Proteom 131:17–28
Zhang P, Lin Y, Liu Y (2021) CLCA2 suppresses the proliferation, migration and invasion of cervical cancer. Exp Ther Med 22(1):776
Purrington KS, Knight J, Dyson G, Ali-Fehmi R, Schwartz AG, Boerner JL et al (2020) CLCA2 expression is associated with survival among African American women with triple negative breast cancer. Sang QXA, editor. PLOS ONE. 15(4):e0231712
Xu Y, Miao C, Jin C, Qiu C, Li Y, Sun X et al (2018) SUSD2 promotes cancer metastasis and confers cisplatin resistance in high grade serous ovarian cancer. Exp Cell Res 363(2):160–170
Mukhopadhyay P, Chakraborty S, Ponnusamy MP, Lakshmanan I, Jain M, Batra SK (2011) Mucins in the pathogenesis of breast cancer: implications in diagnosis, prognosis and therapy. Biochim Biophys Acta 1815(2):224–240
Kim D, Jung WH, Koo JS (2012) Expression of MUC1, MUC2, MUC5AC and MUC5B in mucinous lesions of the breast. Pathobiology 79(3):144–153
Alfarsi LH, El-Ansari R, Craze ML, Masisi BK, Mohammed OJ, Ellis IO et al (2020) Co-expression Effect of SLC7A5/SLC3A2 to predict response to endocrine therapy in oestrogen-receptor-positive breast Cancer. Int J Mol Sci 21(4):1407
Marsh LA, Carrera S, Shandilya J, Heesom KJ, Davidson AD, Medler KF et al (2017) BASP1 interacts with oestrogen receptor α and modifies the tamoxifen response. Cell Death Dis 8(5):e2771
Lin CC, Huang YK, Cho CF, Lin YS, Lo CC, Kuo TT et al (2020) Targeting positive feedback between BASP1 and EGFR as a therapeutic strategy for lung cancer progression. Theranostics 10(24):10925–10939
Hoshino A, Kim HS, Bojmar L, Gyan KE, Cioffi M, Hernandez J et al (2020) Extracellular vesicle and particle biomarkers define multiple human cancers. Cell 182(4):1044–1061e18
Kennon AM, Stewart JA (2021) RAGE differentially altered in vitro responses in vascular smooth muscle cells and adventitial fibroblasts in Diabetes-Induced Vascular calcification. Front Physiol 12:676727
Waghela BN, Vaidya FU, Ranjan K, Chhipa AS, Tiwari BS, Pathak C (2021) AGE-RAGE synergy influences programmed cell death signaling to promote cancer. Mol Cell Biochem 476(2):585–598
Acknowledgements
The authors thank Dr. Stuart P. Atkinson for his collaboration in manuscript preparation and English revision.
Funding
This study was supported by the European Research Council (grant ERC-CoG-2014-648831 “MyNano”) and the H2020 EU-OpenScreen-Drive project (grant agreement No. 823893). Our screening platform is part of the European Infrastructure, ERIC EU-OpenScreen, and the Excellence Network, TeNTaCLES (Translational NeTwork for the CLinical application of Extracellular Vesicle-Tentacles-RED2018-102411-T). This research was also supported by the Spanish Ministry of Science and Innovation (PID2019-108806RB-I00; PID2020-117211GB-I00; PID2021-124430OA-I00 funded by MICIU/AEI/https://doi.org/10.13039/501100011033 and by FEDER, UE; and partially funded by the Institute of Health Carlos III (project IMPaCT-Data, IMP/00019), co-funded by ERDF and CIBERONC (project n. CB22/12/00048). The proteomic analysis was performed in the proteomics facility of SCSIE University of Valencia, which belongs to ProteoRed, PRB2-ISCIII, supported by grant PT13/0001. Agreement 2019 between IVO and the Association “Mou-te per elles.”
Author information
Authors and Affiliations
Contributions
Z.A., M.R.H., F.G.-G., and M.J.V. initiated, designed, and supervised the study; Z.A. and E.M. performed most experiments; Z.A., E.M., and M.J.V. contributed to experimental work and data analysis; M.R.H., S.R.-G., P.M.-M., and F.G.-G. contributed to bioinformatic analysis; Z.A., M.R.H., E.M., J.A.L.-G., F.G.-G., and M.J.V. prepared the manuscript. All authors discussed the results and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not Applicable.
Consent for publication
This publication has full consent from all authors.
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Andreu, Z., Hidalgo, M.R., Masiá, E. et al. Comparative profiling of whole-cell and exosome samples reveals protein signatures that stratify breast cancer subtypes. Cell. Mol. Life Sci. 81, 363 (2024). https://doi.org/10.1007/s00018-024-05403-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-024-05403-z