Abstract
Nuclear Cyclin D1 (Ccnd1) is a main regulator of cell cycle progression and cell proliferation. Interestingly, Ccnd1 moves to the cytoplasm at the onset of differentiation in neuronal precursors. However, cytoplasmic functions and targets of Ccnd1 in post-mitotic neurons are unknown. Here we identify the α4 subunit of gamma-aminobutyric acid (GABA) type A receptors (GABAARs) as an interactor and target of Ccnd1–Cdk4. Ccnd1 binds to an intracellular loop in α4 and, together with Cdk4, phosphorylates the α4 subunit at threonine 423 and serine 431. These modifications upregulate α4 surface levels, increasing the response of α4-containing GABAARs, measured in whole-cell patch-clamp recordings. In agreement with this role of Ccnd1–Cdk4 in neuronal signalling, inhibition of Cdk4 or expression of the non-phosphorylatable α4 decreases synaptic and extra-synaptic currents in the hippocampus of newborn rats. Moreover, according to α4 functions in synaptic pruning, CCND1 knockout mice display an altered pattern of dendritic spines that is rescued by the phosphomimetic α4. Overall, our findings molecularly link Ccnd1–Cdk4 to GABAARs activity in the central nervous system and highlight a novel role for this G1 cyclin in neuronal signalling.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cyclin D1 (Ccnd1) and its catalytic counterparts, cyclin-dependent kinase 4 (Cdk4) and Cdk6, are main regulators of cell proliferation by promoting the G1–S phase transition of the cell cycle (reviewed in [1]). Under favourable conditions, mitogenic signals induce the expression of Ccnd1, the formation of Ccnd1–Cdk4/6 complexes and their localization to the nucleus. There, they phosphorylate the Retinoblastoma repressor protein (pRB), which eventually leads to its inhibition and in turn releases the activity of the E2F transcription factor. This induces the expression of genes required to go through the next steps of the cell cycle. This pRB-dependent control of cell proliferation is considered the “canonical” function of Ccnd1 and it has been reported as very relevant in tumour proliferation. Accordingly, Ccnd1 is classified as an oncogene frequently amplified in many types of neoplasia, and different cancer treatments with Cdk4/6 inhibitors such as Palbociclib have been approved to block this function. However, Ccnd1 also operates through several other non-canonical pathways including some pRB- and Cdk-independent ones to regulate different cellular processes (reviewed in [2]). Thus, Ccnd1 promotes cell detachment, migration and invasion of normal and tumour cells independently of pRB status [3,4,5] by interacting with cytoplasmic and membranous targets, such as filamin A, PACSIN, Ral GTPases and paxillin [6,7,8,9]. For example, Ccnd1–Cdk4 promotes cell invasion and metastasis through the phosphorylation of paxillin and the activation of the Rac1 pathway in membrane ruffles [9]. Moreover, the expression of a membrane-associated form of Ccnd1 harbouring the farnesylation signal of K-Ras (Ccnd1-CAAX) maximises the ability of tumour cells to invade and metastasize [10]. Besides cell invasion, membrane-targeted Ccnd1 also influences cell signalling responses [11].
The first hint for a role of Ccnd1 in the nervous system came from the analysis of CCND1 knockout (KO) mice [12], which present certain behaviours (altered leg-clasping reflex) indicative of neurological abnormalities. Since then, several studies have substantiated the involvement of Ccnd1 in neuronal functioning. In neural stem cells, Ccnd1–Cdk4 has been involved in regulating the length of G1, which is thought to influence the decision of cells to proliferate or differentiate [13,14,15]. Also, under pathological conditions, nuclear Ccnd1 reactivation/upregulation in post-mitotic neurons has been observed to promote apoptosis (reviewed in [2]). Ccnd1 is located in the cytoplasm of differentiating post-mitotic cortical neurons and in neuroblastoma cell lines, although this has been proposed as a mechanism to prevent apoptosis [16] or to promote cell cycle withdrawal [17], respectively. Nevertheless, Ccnd1 could be exerting an active role in the cytoplasm. Interestingly, Ccnd1 is necessary for NGF-induced neuritogenesis in PC12 pheochromocytoma cell line [18]. Also cytoplasmic expression of Ccnd1–Cdk4 has been observed in the hippocampus during development [19] where it has been involved in neuronal plasticity [20]. But despite these works, the role and target/s of Ccnd1 in the nervous system and in particular its role in the cytoplasm of the neurons remains poorly understood.
γ-Aminobutyric acid (GABA) is the major inhibitor neurotransmitter in the central nervous system. Its actions are mediated through different types of GABA receptors: GABA A receptors (GABAARs), which are ligand-gated chloride ion channels, and GABA B receptors (metabotropic G protein-coupled). Deficits in GABAAR function are increasingly being involved in different pathologies, such as anxiety, cognitive deficits, depression, epilepsy, schizophrenia and substance abuse [21]. These receptors are also clinically relevant drug targets for general anaesthetic, anticonvulsant, anxiolytic, or sedative–hypnotic agents. GABAARs mediate both phasic and tonic inhibition: synaptic GABAARs are activated transiently by the GABA released from presynaptic vesicles, whereas extra-synaptic GABAARs are activated by low GABA concentrations in the extracellular space and thus mediate tonic inhibition. The tonic inhibitory currents control neuronal excitability and the strength of synaptic transmission [22].
GABAARs are hetero-pentamers that belong to the ligand-gated ion channel superfamily. A total of 19 subunits (as well as splice variants) have been cloned and grouped into eight subunit classes: α (1–6), β (1–3), γ (1–3), δ, ε, π, θ and ρ (1–3). GABAARs are usually formed by two α, two β, and one γ or one δ subunits. Each subunit contains a large amino-terminal extracellular domain and four transmembrane (TM) domains. Between TM3 and TM4 there is a large intracellular loop (ICL) where regulation by phosphorylation and protein interactions primarily occurs. The subunit composition dictates its subcellular location and function.
GABAARs containing the α4 subunit are expressed in the hippocampus, cortex and thalamus, and mediate extra-synaptic inhibition in thalamus and dentate gyrus [23]. At the onset of puberty, α4-β-δ GABAAR expression increases peri-synaptically at excitatory synapses in the hippocampus, contributing to anxiety, synaptic pruning and learning deficits in mice [24,25,26,27,28]. In fact, in the α4-KO mice, synaptic pruning is prevented, and synaptic plasticity and learning abilities are restored [23, 29, 30]. In the adult hippocampal neurogenesis, α4-containing GABAARs are involved in proliferation, migration and dendritic development [31]. Mutations in the GABAergic receptor subunit 4 gene GABRA4 [32,33,34] and reduced protein and mRNA levels of α4 [35, 36] are reported in patients of autism spectrum disorder. Furthermore, the α4-KO mice show a phenotype compatible with high-functioning autism [37].
Here we report that Ccnd1 interacts with the α4 subunit of GABAARs. Ccnd1–Cdk4 complex phosphorylates α4 in the intracellular loop between TM3 and TM4 and enhances its surface localization, affecting the GABAARs signalling and the pattern of dendritic spines. We propose a novel role for cytoplasmic Ccnd1–Cdk4/6 in regulating α4-containing GABAARs in the central nervous system.
Materials and methods
Cell culture
Mycoplasma-free HEK-293T cells were obtained from the American Type Culture Collection. Cells were maintained at 37 °C in a 5% CO2 incubator, and grown in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% Foetal Bovine Serum (FBS), 10 U/mL penicillin/streptomycin (P/S) and 4 mM glutamine. Mycoplasma detection tests were performed in the Cell culture service (SCT-CC). Transient transfection of vectors was performed with Lipofectamine 2000 (Invitrogen) according to manufacturer’s instructions.
Expression vectors
Human CCND1 was fused to three copies of the Flag or HA or one of Red monomer epitope under the CMV promoter in pcDNA3. Ccnd1–CAAX construct is a fusion of the 3′ end of the K-Ras ORF containing the CAAX motif (GGC TGT GTG AAA ATT AAA AAA TGC ATT ATA ATG TAA) to the 3′ end of the CCND1 ORF [10]. Mouse GABRA4 (IMAGE ID 6828002, Source BioScience) and Human GABRB3 (IMAGE ID 3871111, Source BioScience) were used to obtain N-terminal GST fusions in pGEX-KG (Clontech), N-terminal Flag fusions in pJEN1 (pcDNA3 derived, from Dr. Egea), N-terminal EGFP fusions in pEGFP-N1 (Clontech) or cloned into FCIV1 lentiviral vector (from Dr. Encinas). Standard PCR-mediated site-directed mutagenesis was used to obtain the non-phosphorylatable and phosphomimetic mutants of GABRA4 using the following primers: T423A and S431A: CGCCAAATCCATTCAGCAGGGC and CCGAAGCTAAGTGAGCCTTAGGCGCGGCTTCAGAAGACTCCTGGAC; S456A and S458A: CGGCCCCCGCTCCTCATGGCACATTGCGGC and CGGCAGATGAAAGACCTCTGGCTG; T500A: CGCCTCCTCCCCCTGCTCCAC and CCGCTGACACATTCCCAGCAGC; and T423E and S431E: GCGAACCAAATCCATTCAGCAGGGC and TAGCTAAGTGAGCCTTAGGCTCGGCTTCAGAAGACTC CTGGAC.
Immunoprecipitation assays
For co-immunoprecipitation experiments, HEK-293T cells were harvested 48 h after transfection. The cells were re-suspended in RIPA buffer (50 mM Tris pH7.4, 150 mM NaCl, 1% NP-40, 0.1% sodium deoxycholate, 0.1% SDS, 5 mM EDTA, protease and phosphatase inhibitors), rocked 1 h at 4 °C, and spun for 5 min at 600 g. The supernatants were incubated with protein G magnetic beads (Dynabeads, Invitrogen) for 30′ at 4 °C. After the preclearing, the samples were incubated with 2 μg of αFlag M2 antibody (Sigma) overnight and precipitated with protein G dynabeads. IP of anti-phospho-Threonine–Proline antibody (#9391, Cell Signalling) was performed similarly, but using protein A dynabeads. The beads were collected and washed three times with 1 mL of cold RIPA buffer, and bound proteins were separated by SDS-PAGE gels and visualised by western blots.
IP of endogenous α4 was carried out in hippocampus of adult female mice (≈ 60 days old) with a polyclonal antibody (12979-1-AP, Proteintech). Briefly, cleared cell extracts in RIPA buffer (with protease and phosphatase inhibitors) were immuno-precipitated with Protein A linked to magnetic beads (Invitrogen). An anti-Flag (#F7425, Sigma) was used as a mock control. Washes were performed with RIPA buffer without SDS.
Immunoblotting
For immunoblot, protein samples were resolved by SDS-PAGE, transferred to PVDF membranes (Millipore), and incubated with primary antibodies: anti-Ccnd1 (monoclonal DCS-6, BD biosciences, 1:500), anti-Gabra4 (polyclonal, AB5459, Merck, 1:1000, and polyclonal 12979-1-AP, Proteintech, 1:500), anti-Flag (polyclonal, 11508721, Invitrogen, 1:1000), anti-HA (rat monoclonal 3F10, Roche #11867431001, 1:5000), anti-TFR (monoclonal H68.4, Invitrogen #13-6800, 1:1,000) and anti-GST (goat polyclonal, Amersham #27-4577, 1:2000). Appropriate peroxidase-linked secondary antibodies (GE Healthcare UK Ltd) were detected using the chemiluminescent HRP substrate Immobilon Western (Millipore). Chemiluminescence was recorded with a ChemiDoc-MP imaging system (BioRad).
Production and purification of recombinant proteins
All of the GST fusions were expressed in E. coli BL21 (DE3) by adding 1 mM isopropyl-β-d-thiogalactopyranoside (IPTG) to LB broth cultures at a cell density of 0.3 A600 and subsequent incubation for 4 h at 30 °C. The proteins were purified using glutathione-Sepharose 4B beads as directed by the supplier (Amersham Biosciences) in 500 μl of lysis buffer containing 50 mM HEPES, pH 7.5, 150 mM NaCl, 1 mM EDTA, 0.5% Triton X-100, 10% glycerol, 1 mM DTT, and protease and phosphatase inhibitors. Concentration and purity of substrates were estimated by comparison to protein standards stained with Coomassie Brilliant Blue.
For in vitro transcription and translation, Flag-Ccnd1 was amplified by PCR (forward primer: CGCGCTAATACGACTCACTATAGGGAGACCCAAGCCCATGGGATCAC; reverse primer: TTTTTTTTTTTTTTTTTTTTTTTTTTTTTGGCTGATCAGCGAGCTCTAG), transcribed in vitro with the T7 RNA polymerase (New England Biolabs), and in vitro-translated with a Rabbit Reticulocyte Lysate system (Promega).
GST pull-down assay
For the GST pull-down assay, 400 ng of GST, GST-α4 or GST-Cter-α4 purified from E. coli immobilised on Glutathione-Sepharose 4B beads was incubated with Flag-Ccnd1 in binding buffer (20 mM HEPES–KOH, pH 7.5, 150 mM KCl, 5 mM MgCl2, 0.5 mM EDTA, 0.1% NP-40, 1 mM DTT, 1 mM PMSF, 10% glycerol, protease and phosphatase inhibitors) for 30 min at room temperature. After four washes with the same buffer, the samples were analysed by SDS-PAGE.
Kinase assay
Briefly, 0.2 mg substrate (either GST-α4, GST-Cter-α4 or GST-β3) was mixed with 1.5 μL of active Ccnd1–Cdk4 complex purified from baculovirus (Sigma C0620), 10 mM ATP, 7 mCi of γ-32P-ATP (PerkinElmer, 3000 Ci/mmol), and either DMSO or 2 μM of the Cdk4/6 inhibitor Palbociclib (Selleckchem, S1116) in 20 μL of kinase buffer (50 mM Tris–HCl pH 7.5, 10 mM MgCl2, 0.5 mM DTT, 1 mM EGTA and 2.5 mM b-glycerophosphate). This mixture (20 μL) was incubated for 30 min at 37 °C, then boiled in 2 × Laemmli buffer, and separated by electrophoresis. Phosphorylated proteins were visualised by autoradiography of the dried slab gels.
MS analysis of phosphopeptides
The C-terminus of α4 fused to GST was used in an in vitro kinase assay with Ccnd1–Cdk4 in the presence of ATP or in the absence of ATP as a control. Samples were subsequently subjected to SDS-PAGE. The gels were stained with Coomassie Brilliant Blue G-250 colloidal (EZBlue Gel Staining Reagent, Sigma). After washing with water, protein bands of interest were prepared and submitted to Fundació Institut d’Investigació Biomèdica de Bellvitge (IDIBELL) Proteomics Service (Barcelona) for analysis of the Chymotryptic peptide molecular masses by liquid chromatography–mass spectrometry. Gel slices were manually cut and proteomic service suggested for each band-assay to recover three slices (low 1, middle 2, and high 3 mobility) as phosphorylation could alter band mobility. Briefly, gel bands were washed with water, ammonium bicarbonate (50 mM) and 50% acetonitrile. Next, samples were reduced by incubation with DTT (10 mM) at 60 °C for 45 min and alkylated with iodoacetamide (50 mM) for 30 min, in the dark. Finally, proteins were digested with chymotrypsin (5 ng/μL) at 25 °C overnight (Trypsin gold, Promega). Digestion was stopped by addition of 5% formic acid and peptides extracted twice with 70% acetonitrile and 5% formic acid (10 min sonication). Peptide extracts were evaporated to dryness, re-suspended with 2% acetonitrile 0.1% formic acid and analysed by nano-HPLC–MS/MS.
Surface biotinylation
For surface biotinylation (all steps performed on ice), HEK-293T cells were washed twice with PBS and incubated with 1 mg/mL EZ-link sulpho-NHS-SS-biotin (Pierce), a non-permeable biotin, in PBS for 15 min. Following surface labelling, non-conjugated biotin was quenched by washing twice with TBS. Cells were lysed in RIPA buffer (50 mM Tris, pH7.4, 150 mM NaCl, 1% NP-40, 0.1% SDS, 5 mM EDTA, with phosphatase and protease inhibitors), and protein concentrations were determined using a DC protein assay (Bio-Rad). Equal amounts of protein (usually 1 mg in 500 μL) were incubated overnight with 20 μL NeutrAvidin-coated beads (Pierce) at 4 °C with constant rotation. On the next day, beads were washed three times with RIPA buffer and eluted samples were processed for immunoblotting.
Whole-cell currents measurements in tsA201 cells
Electrophysiological experiments were done with mycoplasma-free tsA201 cells (kindly provided by Prof. F. Ciruela, Universitat de Barcelona) and purchased from the American Type Culture Collection (ATCC, CRL-3216, RRID: CVCL_0063). Mycoplasma detection tests were performed with PlasmoTest (Invivogen, code: rep-pt1). Cells were transiently co-transfected with 1 μg total cDNA using polyethylenimine (PEI) transfection reagent (1 mg/mL) in a 3:1 ratio (PEI:DNA). In all transfections, the DNA ratio used was 1:1:2 (plasmid A: plasmid B: plasmid C), where A is α4, α4T423A/S431A or α4T423E/S431E, B is β3 and C is a control vector or Ccnd1–CAAX codifying plasmid. Cells were re-plated on poly-d-lysine-coated glass coverslips to allow optimal density with isolated cells. All experiments were performed 48 h after transfection.
For the measurements of GABA-evoked whole-cell currents in tsA201, cells were visualised with an inverted microscope (AxioVert A.1, Carl Zeiss) and maintained in extracellular flowing solution at a rate of 60 mL/h. The extracellular solution contained (in mM): 140 NaCl, 5 KCl, 10 HEPES, 11 glucose, 2.5 CaCl2, 1.2 MgCl2 (pH 7.4 with NaOH; osmolarity 305 mOsm/kg adjusted with sorbitol). Electrodes were fabricated from borosilicate glass (1.50 mm O.D., 1.16 I.D., Harvard Apparatus) pulled with a P-97 horizontal puller (Sutter Instruments) and polished with a forge (MF-830, Narishige) to a final resistance of 2–5 MΩ. The intracellular solution contained (in mM): 140 KCl, 2 MgCl2, 2 CaCl2, 10 HEPES, 1.1 EGTA, 2 Mg–ATP (pH 7.2 with KOH; osmolarity 295 mOsm/kg adjusted with sorbitol).
Macroscopic GABAA-mediated currents were recorded at room temperature (22–25 °C) in the whole-cell configuration from cells positive for Venus (α4 in FCIV1) and mCherry (Ccnd1–CAAX in pcDNA3 Red monomer) fluorescence at a holding membrane potential of − 60 mV. Currents were recorded with Axopatch 200B amplifier, filtered at 2 kHz and digitised at 5 kHz using Digidata 1440A interface with pClamp 10 software (Molecular Devices Corporation). Series resistance was typically 5–20 MΩ, and was monitored at the beginning and at the end of the experiment. Cells that showed a change in series resistance greater than 20% were rejected.
Rapid application of the agonist GABA (1–2 ms exchange) was applied by switching between a continuously flowing control solution (extracellular solution diluted by 4%) and a GABA-containing solution (1 μM GABA and 2.5 mg/mL of sucrose diluted in extracellular solution). Solution switching was achieved by piezoelectric translation of a theta-barrel application tool made from borosilicate glass (1.5 mm O.D.; Sutter Instruments) mounted on a piezoelectric translator (P-601.30; Physik Instrumente). Agonist was applied at 2 min interval and the magnitude of the peak current was measured (IGABA). Electrophysiological recordings were analysed using IGOR Pro (Wavemetrics Inc.) with NeuroMatic (Jason Rothman, UCL).
Preparation of organotypic hippocampal slice cultures
Organotypic hippocampal slice cultures were prepared as previously described [38]. Briefly, whole brain of newborn rats (postnatal days 5–7) was dissected and placed in ice-cold Ca2+-free dissection solution (10 mM D-glucose, 4 mM KCl, 26 mM NaHCO3, 8% sucrose, 5 mM MgCl2, 1:1000 Phenol Red) saturated with 5% CO2/95% O2, enabling the brain to chill for 1 min. Hippocampi were then isolated with the help of a magnifying glass, cut in slices (400 µm) in the same solution using a tissue chopper (Stoelting), and maintained at 35.5 °C in culture on permeable membranes in a medium containing 20% horse serum, 1 mM CaCl2, 2 mM MgSO4, 1 mg/L insulin, 0.0012% ascorbic acid, 30 mM HEPES, 13 mM d-glucose, and 5.2 mM NaHCO3. Culture medium was replaced with fresh one every 2–3 days. The slices were used at 4–8 days in vitro.
Whole-cell recordings of GABAA receptor-mediated currents in hippocampal slices
For some experiments, slices were treated for 2 h with either DMSO (control solution) or 2.5 μM Palbociclib (Cdk4/6 inhibitor) (Selleckchem), as indicated. For expression of recombinant α4, slices were transfected using the biolistic method (Helios gene gun system, Bio-Rad) and maintained in culture for 24 h. After that, slices were placed in a chamber and perfused with artificial cerebrospinal fluid [aCSF: 119 mM NaCl, 2.5 mM KCl, 1 mM NaH2PO4, 26 mM NaHCO3, 11 mM glucose, 1.2 mM MgCl2, 2.5 mM CaCl2; osmolarity adjusted to 290 mOsm; pH 7.5] supplemented with 100 μM AP5 (NMDA receptor antagonist), 10 μM CNQX (AMPA receptor antagonist), 1 μM strychnine (glycine receptor antagonist) as external solution, gassed with 5% CO2/95% O2 at 29 °C in the electrophysiology set-up. DMSO or Palbociclib was also included in the extracellular solution for the experiments with pre-treatment. Patch recording pipettes (4–6 MΩ) were pulled from thin-walled borosilicate capillary glass (World Precision Instruments [WPI], Sarasota, FL) on a P-97 electrode puller (Sutter Instrument, San Rafael, CA) and filled with high chloride internal solution (178 mM CsCl, 10 mM HEPES, 2.5 mM MgCl2, 4 mM Na2ATP, 0.4 mM Na3GTP, 10 mM sodium phosphocreatine, 0.6 mM EGTA; pH adjusted to 7.2, osmolarity adjusted to 290 mOsm). CA1 pyramidal neurons were recorded under whole-cell voltage clamp at − 60 mV using Multiclamp 700A/B amplifiers (Molecular Devices, San Jose, CA, USA). At least 10 min of stable holding currents was recorded. The tonic GABAA receptor-mediated current was measured as the outward shift in holding current following application of picrotoxin (100 μM). To measure miniature inhibitory postsynaptic currents (mIPSCs), 1 μM tetrodotoxin was added to the aCSF to block action potentials. Recordings were obtained with the gap-free mode of pClamp and mIPSCs were analysed with the event detection of pCLAMP software (Molecular Devices). Approximately 2 min of stable recordings before picrotoxin addition was used for the analysis of mIPSC amplitude and frequency.
Cortical neuron culture
For primary culture of cortical neurons, cortices were dissected from E15.5 CCND1 KO or WT mice in ice-cold HBSS-MHPS (Hank’s Balanced Salt Solution, 10 mM MgSO4, 10 mM HEPES pH 7.2 and 10 U/mL P/S). After dissection, cortices were incubated with papain solution (1 mg/mL papain in HBSS-MHPS) at 37 °C for 23 min. Enzymatic digestion was inhibited by washing the tissue 3 times with 10 mg/mL trypsin inhibitor (Fisher) in HBSS-MHPS. Additionally, one more wash was done with neuro-basal medium supplemented with 2% B27, 2 mM Glutamax and 1 mM sodium pyruvate (NB27, Fisher) at room temperature. The tissue was triturated by passing it through a flame-polished Pasteur pipette (< 20 times). After the mechanical disaggregation, the supernatant containing mostly single cells was centrifuged at 650 rpm for 4 min. The pellet was re-suspended in NB27 and the cells were counted in a Neubauer chamber. Cells were seeded at low density (25,000 cells/cm2) with NB27 medium in poly-d-lysine (0.5 mg/mL) and laminin (5 μg/mL)-coated plates, in 1:1 conditioned medium. One-third of the culture medium is replaced with fresh one every 3–4 days. Primary cortical neurons were transfected at 3 DIV using Lipofectamine 2000 (Invitrogen) following the manufacturer’s recommendations and the proportion 0.3 μg DNA:1.2 μL Lipofectamine2000. Cortical neurons were transfected with GFP alone or with Flag-tagged α4T423E/S431E together with GFP in a 4:1 proportion. The medium was changed 4 h after transfection and cells were fixed at 20 DIV.
Genotyping of CCND1 knockout mice
CCND1 knockout mice [12] were obtained from Charles River. Transgenic mice were maintained in a mixed background of C57BL/6 and SV129. Embryo’s tails were collected and warmed at 95 °C NaOH 50 mM for 45 min. After neutralisation with Tris 0.15 M pH 8.5, the samples were then centrifuged for 1 min at 10,000 rpm. Supernatant containing genomic DNA was used for CCND1 genotyping by PCR. The following primers were used: a common primer CTGTCGGCGCAGTAGCAGAGAGCTACAGAC, a CCND1 primer CGCACAGGTCTCCTCCGTCTTGAGCATGGC and a neomycin primer CTAGTGAGACGTGCTACTTCCATTTGTCACG. PCR was performed in a Thermal Cycler (T100 Bio-Rad) and PCR products were analysed in a 1.5% agarose gel followed by Ethidium Bromide staining. The expected band pattern is 249 bp for WT, and 394 bp for KO alleles.
Immunofluorescence and spine analysis
Cortical neurons transfected with GFP were fixed at 20 DIV using 4% PFA and 4% sucrose in PBS for 15 min at room temperature and then washed with PBS. Neurons were permeabilized for 5 min with 0.1% Triton X-100 in PBS and blocked with 3% BSA in PBS. Primary antibody α-GFP (A11120, ThermoFisher) was diluted 1:200 in 0.3% BSA in PBS. Proteins were detected by incubation with secondary antibody Alexa 488 rabbit anti-mouse (A11059, ThermoFisher). Images were acquired with an Olympus FV1000 confocal microscope using the following parameters: stacks of 10 slices were imaged every 0.37 µm, under a 60× objective. Laser power and PMT values were kept constant throughout images and conditions. Immunofluorescence quantification was performed using NeuronJ of ImageJ. Spine density and morphology analysis of WT and CCND1 KO cortical neurons was performed on 3D reconstructions of confocal z series acquired using a 2 × zoom. Spines were counted as mushroom-type if the spine head was wider than the spine neck. All analyses were done blind to the experimental condition.
Statistical analyses
Different statistical analyses were used in this work. In every analysis, the minimum level of statistical significance was a p value equal to 0.05. The significance level is represented in each graph as indicated. In order to evaluate possible differences between two experimental groups, a t test was performed. When there was a multiple comparison, one-way ANOVA with a post hoc test (Tukey) was performed. In whole-cell recordings in ts201A cells, comparisons between groups were done using the parametric Student’s t test. For tonic currents comparison, Wilcoxon test was performed and in mIPSC analysis. Mann–Whitney and Kolmogorov–Smirnov tests were done. Microsoft Excel and GraphPad Prism v5.0 (GraphPad) were used for statistical analysis and graphical representation.
Ethical considerations
All procedures with animals follow the protocols approved by the Institutional Committee of Care and Use of Animals (Comitè Institucional de Cura i Ús d’Animals), and experiments were approved by the Ethics Committee of the University of Lleida (CEEA 03-02/19). Animals were housed and maintained in the animal facility of the University of Lleida with 12 h:12 h light/dark cycle and food/water available ad libitum. For electrophysiological experiments, all biosafety procedures and animal care protocols were approved by the bioethics committee from the Consejo Superior de Investigaciones Cientificas and were performed according to the guidelines set out in the European Community Council Directives (2010/63/EU, 86/609/EEC). All efforts were made to minimise the number of animals and their suffering.
Results
The α4 subunit of GABAA receptors interacts with Ccnd1
To investigate the role of Ccnd1 in the nervous system, we performed a yeast two-hybrid screening using Ccnd1 as a bait and an adult mouse brain library as a prey [39]. We found that Ccnd1 interacts with the C-terminal half of the α4 GABAAR subunit. We obtained three independent clones spanning amino acids 354–552, 385–552 and 386–552 (Fig. 1A). This region of α4 contains the intracellular loop (ICL) between the TM3 and TM4 transmembrane domains, which is phosphorylated by protein kinase C (PKC, [40]). To confirm the interaction between the α4 subunit and Ccnd1, we carried out immunoprecipitation (IP) experiments with Flag-tagged Ccnd1 and the HA-tagged C-terminal region of α4 (354-552aa) expressed in HEK-293T cells. We observed specific co-IP of the C-terminal domain of α4 in the Ccnd1 IP (Fig. 1B). To test whether there is a direct interaction between Ccnd1 and the α4 subunit, we performed in vitro GST-pull-down assays. Fusions of GST-α4 or GST-C-terminal α4 (354-552aa) purified from E. coli were mixed with Flag-Ccnd1 produced by in vitro transcription and translation. Ccnd1 was co-precipitated when GST-α4 or GST-C-terminal α4 were pulled down, but not with GST alone (Fig. 1C). To analyse the endogenous interaction between Ccnd1 and the α4 GABAAR subunit, we immuno-precipitated α4 from the hippocampus of adult mice since co-expression of Ccnd1 and α4 mRNAs in the CA1, CA3 and dentate gyrus regions of the hippocampus is observed in the Allen mouse brain atlas (www.brain-map.org). As shown in Fig. 1D, the co-immunoprecipitation of endogenous Ccnd1 with the α4 subunit of GABAAR was weakly detected. Overall, our data indicate that there is a specific and direct interaction between Ccnd1 and the α4 subunit of GABAA receptors but this interaction may be transitory or represents a small percentage of the total proteins in vivo. In accordance with the weakness of the interaction, α4 could be a substrate of Ccnd1–Cdk4 complex.
Ccnd1 interacts with the α4 subunit of GABAA receptors. A Scheme representing the mouse α4 subunit of GABAARs (552aa). N-terminal (Nt) and C-terminal (Ct), intracellular and extracellular domains and the four transmembrane domains (TM1-4) are shown. In a yeast two-hybrid screening, Ccnd1 was found to interact with three independent clones of the C-terminal region of the α4 subunit (in yellow), spanning amino acids 354–552, 385–552, 386–552 (the starting amino acids are shown as stars). The three clones contained the intracellular loop (ICL) between TM3 and TM4. Putative phosphorylation sites for Cdk (SP or TP) in the ICL are shown in blue. B HEK-293T cells were transfected with the C-terminus of α4 subunit (354-552aa, tagged with HA: HA-Ct-α4) and Ccnd1 (tagged with Flag: Flag-Ccnd1). At 48 h after transfection, Flag-Ccnd1 was precipitated using the M2 Flag antibody and protein G dynabeads (Invitrogen). Input (IN) and immunoprecipitation (IP) samples were analysed by western blot using anti-HA and anti-Flag antibodies, to detect co-IP of the C-terminus of α4 with Ccnd1. C In vitro-translated Flag-Ccnd1 was incubated with recombinant GST or GST fusions of α4 full-length (GST-α4) or the C-terminus of α4 (GST-Ct-α4) purified from E. coli. Input (IN) and GST pull-down (PD) samples were analysed by western blot using anti-Flag antibody to detect Ccnd1 and anti-GST antibody to detect α4. Intact protein fusions are indicated. D Immunoprecipitation of endogenous α4 from adult mouse hippocampal extracts was performed using a rabbit polyclonal antibody against α4. As a control, a rabbit polyclonal antibody against the Flag epitope (FL) was used. Input (IN) and IP samples were analysed by western blot to detect Ccnd1 and α4 subunit. Ab antibody band
The α4 subunit of GABAARs is phosphorylated at T423 and S431 by Ccnd1–Cdk4 complex
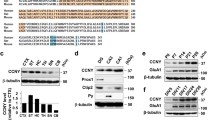
As Ccnd1 is the regulatory subunit of Cdk4/6 kinases, we analysed the sequence of α4 for putative phosphorylation sites by Cdks. We found five S/TP sites in the sequence of the intracellular loop (Fig. 1A). In an in vitro kinase assay, Ccnd1–Cdk4 complex phosphorylated recombinant GST-α4 and this phosphorylation was prevented by inhibiting the complex with the specific Cdk4/6 inhibitor Palbociclib (Fig. 2A). Besides, another subunit of the GABAARs, β3, which lacks S/TP sites, was not phosphorylated by the Ccnd1–Cdk4 complex (Fig. 2B). In order to know which residues of α4 were phosphorylated by Ccnd1–Cdk4, we mutated the five intracellular putative Cdk phosphorylation sites to non-phosphorylatable residues (alanine) generating three different mutant alleles: T423A and S431A (α4T423A/S431A); S456A and S458A (α4S456A/S458A); and T500A (α4T500A). Neither α4S456A/S458A nor α4T500A mutants showed a decrease in phosphorylation by Ccnd1–Cdk4 (Fig. 2C, D). However, a significant reduction of α4 phosphorylation by Ccnd1–Cdk4 was observed using the non-phosphorylatable mutant T423 and S431 (α4T423A/S431A: AA), either with the full-length or the C-terminus construct of α4 (Fig. 2C–F). Moreover, we analysed by mass spectrometry phosphopeptides from the C-terminus of α4 subunit phosphorylated by Ccnd1–Cdk4, and we found that both T423 and S431 were phosphorylated in vitro (Supplementary Table 1). Altogether, our results show that Ccnd1–Cdk4 complex is able to phosphorylate α4 subunit at T423 and S431.
Ccnd1–Cdk4 phosphorylates α4 at T423 and S431 residues. A In vitro radioactive kinase assay using purified recombinant GST-α4 fusion and Ccnd1–Cdk4 (Sigma). Cdk4/6 inhibitor Palbociclib was added at 1 µM concentration. Coomassie blue staining is shown as loading control (input). B Recombinant GST-α4 and GST-β3 fusions were tested for Ccnd1–Cdk4 kinase activity as in A. C Kinase assay was performed with Ccnd1–Cdk4 against GST-α4 wild type and the non-phosphorylatable mutants: T423A/S431A; S456A/S458A; and T500A. D Quantification of the phosphorylation in C showing the mean ± SEM. The amount is relative to a pRB control (n = 3, One-way ANOVA, post-Tukey analysis *p < 0.05). E Kinase assay was performed with Ccnd1–Cdk4 against the C-terminus of α4 wild type (GST-Ct-α4 wt) and two clones (#1 and #2) of the non-phosphorylatable mutant T423A/S431A (AA). F Quantification of the phosphorylation in E showing the mean ± SEM. The amount is relative to a pRB control (n = 3, One-way ANOVA, post-Tukey analysis ***p < 0.001). G Immunoprecipitation of Cdk substrates in HEK-293T cells transfected with α4 wild type (wt) or the non-phosphorylatable mutant T423A/S431A (AA), together with β3 and HA-Ccnd1-CAAX (D1) or HA-Ccnd1K112E-CAAX (D1KE). PhosphoTP (pTP) antibody was used for the immunoprecipitation (IP), and α4 was detected by immunoblot. Protein levels of α4, Ccnd1 (anti-HA antibody) and actin were also analysed in the whole-cell extract (WCE). H Quantification of the phosphorylation in G showing the mean ± SEM (n = 3 One-way ANOVA, post-Tukey analysis ***p < 0.001)
To investigate if the α4 subunit is also phosphorylated by Ccnd1–Cdk in vivo, we immuno-precipitated Cdk substrates with an anti-phospho-Threonine–Proline (pTP) antibody and the presence of phosphorylated α4 was assessed by western blot. We transiently transfected HEK-293T cells with α4, β3 subunits and Ccnd1 or Ccnd1K112E, a mutant of Ccnd1 that does not form an active complex with Cdk [41]. For the experiments where we co-expressed Ccnd1 or Ccnd1K112E, we used a variant that contains the CAAX motif of K-ras to maintain Ccnd1 attached to the cell membrane [10]. In the pTP IP, we observed an increase in the phosphorylated α4 subunit in the presence of Ccnd1-CAAX relative to the presence of Ccnd1K112E-CAAX (Fig. 2G, H). Moreover, when we used the non-phosphorylatable α4 mutant (α4T423A/S431A: AA), we did not observe higher levels of phosphorylated α4 in the presence of Ccnd1 (Fig. 2G, H). This data suggests that Ccnd1–Cdk4 can phosphorylate α4 in vivo at least at T423.
Ccnd1 increases the surface levels of the α4 subunit of GABAA receptors
Phosphorylation of several subunits of GABAA receptors can affect their abundance at the cell surface, where they are active [42]. To analyse whether phosphorylation by Ccnd1–Cdk4 may affect GABAAR localization, we studied the surface levels of the α4 subunit when co-expressed heterologous with Ccnd1 in HEK-293T cells. For this, we transiently expressed α4 and β3 in these cells and assessed the surface levels of α4 by biotin labelling of surface proteins (Fig. 3A). We used the transferrin receptor (TFRC) protein levels as a surface protein control and quantified the levels of α4 at the surface relative to the levels of α4 in the total cell extract (Fig. 3B). We observed a statistically significant increase in the surface levels of α4 (148 ± 20%, p = 0.04, Fig. 3A, B) only when Ccnd1–CAAX was also co-expressed. This increase was abolished in the presence of Ccnd1K112E–CAAX (98 ± 29%, p = 0.93), as well as when the non-phosphorylatable mutant α4T423A/S431A was used (107 ± 24%, p = 0.82), indicating that the observed effect depends on T423/S431 phosphorylation by Ccnd1–Cdk4. These results suggest that Ccnd1 increases the surface levels of the α4 subunit of GABAAR in a kinase-dependent manner.
Ccnd1 increases surface levels of α4 subunit. A HEK-293 T cells were transfected with β3 and α4-Flag or the non-phosphorylatable mutant (α4T423A/S431A-Flag: AA). Either HA-Ccnd1-CAAX (D1) or HA-Ccnd1K112E-CAAX (D1KE) or a control vector was co-transfected. A biotinylation assay was used to purify cell surface proteins. TFRC is used to normalise α4 levels. Representative blots with whole cell extracts and surface proteins are shown. Antibodies used for protein detection are: polyclonal rabbit anti-Flag to detect α4-Flag; TFRC, monoclonal mouse anti-TFRC. B Quantification of the experiment in A as a surface to WCE ratio. Bars show mean values ± S.E.M (n = 4 independent experiments). *p < 0.05 (one-way ANOVA and Tukey post-test). ns not significant
Ccnd1 and T423/S431 phosphorylation of α4 subunit diminishes GABAAR rundown response
Because Ccnd1–Cdk4 increases the surface localization of α4-containing GABAARs, we reasoned that it should affect GABA signalling. To study the electrophysiology of GABAARs, we carried out whole-cell patch-clamp recordings of HEK-293 tsA201 cells transiently expressing α4 and β3 subunits, since functional GABAA receptors can be formed in HEK-293T cells by co-expression of these subunits [40]. To measure the behaviour of the α4β3 GABAARs, 1 μM GABA was applied for 5 s every 2 min in these cells (Fig. 4). Receptor-mediated current amplitude decreased (rundown) over time (Fig. 4B, E, F): after 12 min of recording, GABA-mediated current was 60 ± 5% (n = 7) of the initial response. Co-expression of Ccnd1-CAAX with α4β3 receptors partially prevented their rundown in a significant manner (Fig. 4C, E, F). At 12 min after the start of the experiment, the GABA-mediated current amplitude, in the presence of Ccnd1-CAAX, was 80 ± 9% (n = 5) of the initial GABA-mediated response. To confirm that phosphorylation of T423/S431 in α4 is important for this effect, we co-transfected a phosphomimetic α4T423E/S431E (α4EE) mutant with β3 and observed that it completely obliterates the rundown of GABAARs (Fig. 4D–F). In addition, we determined that the expression of Ccnd1-CAAX together with the non-phosphorylatable allele of α4 (α4AA) did not reduced the rundown efficiency (Fig. 4E, F). Thus, our data suggest that Ccnd1 can reduce the rundown response of GABAARs through the phosphorylation of α4.
Ccnd1 and phosphorylation of α4 at T423/S431 decreases GABAA receptor rundown. A Schematic representation of the whole-cell patch-clamp arrangement for GABA currents recordings in HEK-tsA201 cells transfected with α4 and β3 subunits. Cells were rapidly switched from a control solution (in blue) to a 1 μM GABA-containing solution (in yellow). B Representative recording of whole-cell currents evoked by rapid application of 1 μM GABA (solid line above the trace) to HEK-tsA201 cells expressing wild-type α4β3 GABAARs. The GABA-evoked current at the beginning of the recording (time 0 min) and the response after 14 min (time 14 min) are shown. Dashed line denotes the zero current. Holding potential was held at − 60 mV. C Same as B, but from cells co-expressing wild-type α4β3 GABAARs plus Ccnd1-CAAX. Note that the rundown of the peak current at 14 min is smaller than in condition B. D Same as B, but from cells co-expressing phosphomimetic (T423E/S431E) GABAA receptor α4EEβ3. E Peak current values at different times of GABA stimulation. Grey circles, wild type receptor alone (α4β3); red circles, wild type receptor together with Ccnd1-CAAX (α4β3 + Ccnd1); grey triangles, non-phosphorylatable GABAAR alone (α4AAβ3); red triangles, non-phosphorylatable GABAAR together with Ccnd1-CAAX (α4AAβ3 + Ccnd1); blue squares, phosphomimetic GABAAR (α4EEβ3). F Bar graph showing the mean ± S.E.M. of the relative current at t = 12 min compared with current at time = 0 for cells expressing wild type α4β3 GABAARs with or without Ccnd1-CAAX (n = 7 and n = 5), α4AAβ3 GABAARs with or without Ccnd1-CAAX (both n = 6) or for α4EEβ3 GABAARs alone (n = 6). Open circles denote single experimental values. For E and F *p < 0.05, α4β3 vs. α4β3 + Ccnd1-CAAX; #p < 0.05 α4β3 vs. α4EEβ3; ns denotes not significant (p > 0.05). In whole-cell recordings in ts201A cells, comparisons between groups were done using the parametric student’s t test (Unpaired student’s t test)
Cdk4/6 inhibition decreases tonic currents and mIPSC amplitude in the hippocampus
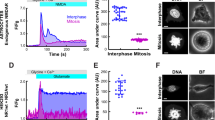
The α4 subunit is a component of extra-synaptic, high-affinity GABAARs, which mediate tonic inhibition in the thalamus and hippocampus [23]. To assess the importance of Ccnd–Cdk4/6 on GABAARs in a physiological context, we measured tonic currents in hippocampal slices of newborn rats. Organotypic hippocampal slice cultures were treated with DMSO (vehicle) or the Cdk4/6 specific inhibitor Palbociclib at 2.5 µM for 2 h (Fig. 5A). GABA inhibitory post-synaptic currents were evoked with single voltage pulses in Schaffer collaterals, and tonic currents were recorded from CA1 pyramidal neurons under whole-cell voltage-clamp configuration. The application of the antagonist picrotoxin (100 µM) causes a decrease in the operative extrasynaptic GABAA channels, which can be measured as an outward shift of the holding current [43] (Fig. 5B). This shift corresponds to the GABAAR tonic currents (Fig. 5C). Whilst the holding current in DMSO-treated cells was shifted by 21 ± 4 pA, the shift in Palbociclib-treated slices was only 9 ± 3 pA (Fig. 5B, C). Therefore, Ccnd–Cdk4/6 inhibition by acute Palbociclib application causes a significant decrease in the GABAAR tonic current.
Cdk4/6 inhibition by Palbociclib decreases GABA tonic currents and mIPSC amplitude and frequency in rat hippocampal slices. A Schematic representation of the experimental design for GABA currents recordings in CA1 neurons: hippocampal slices from newborn rats were cultured in vitro, treated for 2 h with DMSO or Palbociclib 2.5 µM, and tonic (extrasynaptic) or miniature inhibitory post-synaptic currents (mIPSCs, synaptic currents) were measured. B GABA inhibitory post-synaptic currents were evoked by stimulation of Schaffer collaterals and the tonic GABAA receptor-mediated current was measured as the outward shift in holding current following application of the GABAAR antagonist picrotoxin. CA1 pyramidal neurons were recorded under whole-cell voltage-clamp at − 60 mV. Effects of picrotoxin (100 µM) on holding currents recorded in DMSO- (grey rounds) or Palbociclib-treated (red squares) hippocampal slices are represented: the shift in holding currents is shown as double-headed arrows for DMSO (grey) and Palbociclib (red). Recordings are aligned at the time of picrotoxin addition (t = 0). Each point represents the mean value of holding current at a specific time for 10 cells. C Quantification of the shift in holding current (observed in B) for DMSO- and Palbociclib-treated cells (n = 10 for each condition), represented in a scatter dot plot showing mean values ± S.E.M (*p < 0.05, Wilcoxon–Mann–Whitney test). The shift in holding currents (corresponding to tonic currents) is reduced in Palbociclib-treated slices compared to vehicle. D To measure miniature inhibitory post-synaptic currents (mIPSCs), 1 µM tetrodotoxin was added to block action potentials. Representative traces of mIPSCs of DMSO- or Palbociclib-treated cells are shown. E Scatter dot plot showing mIPSC amplitude quantifications of DMSO- (n = 9) and Palbociclib-treated cells (n = 8). Bars show mean values ± S.E.M. *p < 0.05, Wilcoxon–Mann–Whitney test. Cumulative distribution of mIPSC amplitude (F) and frequency (G) from CA1 pyramidal neurons treated with either Palbociclib or control vehicle, as indicated. “n” represents number of miniature responses recorded from nine DMSO or eight Palbociclib-treated cells (***p < 0.001, Kolmogorov–Smirnov test)
Given that the α4 subunit can also form synaptic receptors when clustered together with βxγ2 subunits [44], we tested whether the acute Cdk4/6 inhibition may in turn affect GABAA receptor synaptic function. To this end, we measured miniature inhibitory post-synaptic currents (mIPSCs) in organotypic cultures treated with 2.5 µM Palbociclib for 2 h, in the presence of the action potential blocker tetrodotoxin (1 µM). We observed that the amplitude and frequency of the miniature responses were reduced after CcndCdk4/6 inhibitor treatment as compared to the control (DMSO) treatment (− 106 ± 20 pA, 0.9 ± 0.51 Hz for DMSO vs − 51 ± 5 pA, 0.48 ± 0.17 Hz for Palbociclib, Fig. 5D–G). Putting all these electrophysiological experiments together, our results suggest that Ccnd1–Cdk4/6 enhances GABAAR function in the hippocampus.
Phosphorylation status of α4 at T423/S431 modify tonic currents and mIPSC amplitude in the hippocampus
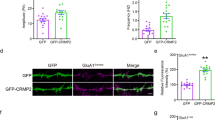
To further test whether these effects on neuronal GABAA receptor responses may be mediated by α4 phosphorylation, we expressed α4 phosphorylation mutants in hippocampal neurons from organotypic hippocampal slice cultures. GFP-tagged wild-type α4, non-phosphorylatable (T423A/S431A: AA) or phosphomimetic (T423E/S431E: EE) mutants were expressed for 24 h by biolistic (gene gun) transfection. Then, whole-cell voltage-clamp recordings were carried out from transfected neurons, guided from their GFP fluorescence. Recordings from un-transfected neurons were also performed, as control. Tonic currents were evaluated from the shift in holding current upon application of the GABAA receptor antagonist picrotoxin, as described above for Fig. 5B, C. As shown in Fig. 6A, B, expression of the non-phosphorylatable mutant (AA) significantly reduced the shift in holding current (tonic response), as compared to wt-transfected neurons. Conversely, expression of the phosphomimetic mutant (EE) significantly enhanced tonic responses, as compared to wt α4. Recordings from untransfected neurons produced similar tonic responses as those from neurons expressing wt α4. To note, absolute holding currents were not significantly different between transfected and un-transfected neurons after picrotoxin application, indicating that other ionic currents in the cell are not affected by the expression of recombinant α4 (Supplementary Fig. 2A).
α4 phosphorylation mutants alter GABA tonic currents and mIPSC amplitude and frequency in cultured hippocampal slices. A CA1 pyramidal neurons expressing wild-type α4 (WT), the non-phosphorylatable mutant T423A/S431A α4 (AA), the phosphomimetic mutant T423E/S431E α4 (EE) or untransfected neurons, as control, were recorded under whole-cell voltage-clamp configuration at − 60 mV from hippocampal slices. The shift in holding current upon addition of picrotoxin (t = 0) was monitored, as an indication of GABAA receptor-mediated tonic currents. ‘n’ represents the number of cells in each condition. B Quantification of the shift in holding current (tonic current, as observed in A) for untransfected neurons and for neurons expressing wild-type (WT), T423A/S431A (AA) or T423E/S431E (EE) α4. Values from individual cells are plotted with points, together with mean values ± S.E.M (‘p’ values according to Mann–Whitney test). Cumulative distribution of mIPSC amplitude (C) and frequency (D) from un-transfected neurons and from neurons expressing wild-type (WT), T423A/S431A (AA) or T423E/S431E (EE) α4. ‘n’ represents the number of miniature responses from all cells in each condition. ‘p’ values according to Kolmogorov–Smirnov test are shown for the comparison of wt α4-transfected cells with respect to T423A/S431A (AA), T423E/S431E (EE) or un-transfected cells. Average mIPSCs for each condition is shown as inset in (C). For all panels, wild-type α4 is represented with black symbols, T423A/S431A α4 with red symbols, T423E/S431E α4 with blue symbols and un-transfected neurons with grey symbols.
Finally, we also tested the effect of α4 phosphorylation in GABAA receptor synaptic currents by measuring the amplitude and frequency of miniature IPSCs (mIPSCs) in neurons expressing these α4 phosphorylation mutants. As shown in Fig. 6C, D, expression of the non-phosphorylatable mutant (AA) significantly reduced the amplitude and frequency of mIPSCs, as compared to wt-expressing neurons (left-shift in the cumulative distributions), whereas expression of the phosphomimetic mutant (EE) produced the opposite effect (right-shift in the mIPSC amplitude and frequency distributions). To note, mIPSCs were smaller (inset in Fig. 6C) and displayed slightly slower kinetics (Supplementary Fig. 2B) in neurons expressing recombinant α4, as compared to un-transfected neurons. These results are consistent with the reported smaller and slower currents of α4-containing GABAA receptors versus those containing the α1 subunit [45]. These observations suggest that recombinant α4 is assembling with endogenous subunits to form functional receptors in hippocampal neurons.
Overall, these electrophysiological experiments with hippocampal slices support the interpretation that α4 phosphorylation by Cdk4/6 increases GABAA receptor currents in neurons.
Dendritic spine pattern is altered in CCND1 KO neurons and rescued by the phosphomimetic T423E/S431E α4 subunit
Since Ccnd1 regulates α4-containing GABAA receptors, we wondered whether the absence of Ccnd1 could disrupt some α4-dependent functions in vivo. The α4 subunit of GABAARs has been involved in synaptic pruning in the hippocampus of adolescent mice decreasing the spine density and altering the spine pattern (for instance reducing mushroom spines) [27, 28]. Because Ccnd1 promotes the activity of α4-containing GABA receptors, we asked if the absence of Ccnd1 could disrupt the normal pattern of spine development. We analysed the number and morphology of dendritic spines in cortical neurons from WT and CCND1 KO mice. Cultured neurons were transfected with GFP at 3 DIV, and dendritic spines were analysed at 20 DIV, when spines are completely developed (Fig. 7A, B). The density of dendritic spines was not significantly changed by Ccnd1 expression (WT = 8.09 ± 0.20 vs CCND1 KO = 8.59 ± 0.24 spines/10 µm, non-significant, Fig. 7C). However, the density of more mature spines with the mushroom-shape was significantly increased in CCND1 KO neurons compared with neurons from WT animals (WT = 3.90 ± 0.15 vs KO = 4.56 ± 0.15 mushroom/10 µm, p < 0.05, Fig. 7D). Interestingly, the expression of the phosphomimetic α4-allele (α4EE) rescued the CCND1 KO phenotype reducing the density of mushroom spines (KO + α4EE = 3.72 ± 0.17 vs KO = 4.56 ± 0.15 mushroom/10 µm, p < 0.01, Fig. 7D). These results suggest that Ccnd1 could influence the dendritic spine pattern through α4 regulation.
The phosphomimetic α4 allele rescues the increase of mushroom dendritic spine density in CCND1 KO neurons. A Schematic representation of the experimental design for dendritic spine analysis of cortical neurons at 20 DIV. Cultured cortical neurons from WT and CCND1 KO mice were transfected at 3 DIV with GFP or GFP + α4EE and fixed at 20 DIV. Changes in spine density and morphology were assessed by analysing GFP-positive neurons with NeuronJ (see “Materials and methods”). B Representative confocal images of WT and CCND1 KO neurons (top) and higher magnification of dendrites with NeuronJ software (bottom) are shown. Scale bar: 10 µm. C Quantification of the number of total dendritic spines relative to 10 µm of neurite, from WT (n = 39), CCND1 KO (n = 42), WT + α4EE (n = 38) and CCND1 KO + α4EE (n = 37) neurons (4 mice/group). Bars show mean values ± S.E.M. ns non-significant, one-way ANOVA and Tukey post-test. D Quantification of the number of mushroom spines relative to 10 µm of neurite from the same samples as in C. Bars show mean values ± S.E.M. **p < 0.01 *p < 0.05, one-way ANOVA and Tukey post-test
Discussion
In neurons, Ccnd1–Cdk4 has been involved in promoting proliferation versus differentiation of stem cells during cortical development [13] and in the adult hippocampus [46]. However, the role of Ccnd1 in the regulation of neuronal signalling has never been investigated before. Interestingly, CCND1 KO mice show an abnormal limb-clasping reflex [12], being this alteration an early symptom of neurological defects associated with hippocampal neuron malfunctioning [47]. In this work, we have unveiled a novel role of Ccnd1 promoting the surface localization of α4-containing GABAA receptors (Fig. 8).
Ccnd1 interacts with the α4 subunit of GABAAR and Ccnd1–Cdk4 complex phosphorylates T423 and S431 in the intracellular loop of α4. This phosphorylation increases the surface levels of α4 subunit and the efficacy of GABAAR signalling in whole-cell patch-clamp recordings. Moreover, CcndCdk4/6 inhibition or the absence of phosphorylation of α4 decreases tonic currents and the amplitude of mIPSCs in the hippocampus of newborn rats. Accordingly, the density of mushroom dendritic spines is increased in cortical neurons from CCND1 KO mice, and this phenotype is rescued by the phosphomimetic allele of α4. Therefore, we propose that Ccnd1–Cdk4-mediated phosphorylation of α4 is an important physiological regulator of GABAAR-mediated inhibition.
Ccnd1 has been found in the cytoplasm of post-mitotic neurons [16]. Supporting this, the mRNA of Ccnd1 has also been found in the synaptic neuropil in the hippocampus [48], suggesting local translation of Ccnd1 in the dendrites. Also, Cdk4 has been localised in the cytoplasm in adolescent mouse hippocampal neurons [20]. However, as nuclear localization of Ccnd1 in post-mitotic neurons promotes apoptosis, its cytoplasmic localization has been suggested as a Ccnd1 sequestration mechanism to avoid cell death. Nevertheless, we have observed that membrane-associated Ccnd1 phosphorylates α4, increase the α4 surface localization and promotes GABAAR signalling. Then, our data strongly suggest that Ccnd1–Cdk4 plays an active role in the cytoplasm of post-mitotic neurons. Previously, Ccnd1–Cdk4 cytoplasmic activity has been associated with the regulation of cell motility and invasion through the interaction with membrane-associated proteins, such as Filamin A, Ral GTPases and Paxillin [6, 8, 9, 49]. However, a cytoplasmic role of Ccnd1 regulating neuronal signalling has never been described before.
In this work, we have proved that Ccnd1–Cdk4 phosphorylates α4 at T423 and S431 in the intracellular loop (ICL) in vitro. Moreover, in transiently transfected HEK-293T cells, we have detected Ccnd1-specific phosphorylation of α4 in vivo. Rundown of GABAA receptors involves phosphorylation–dephosphorylation processes [50,51,52]. The ICL between TM3 and TM4 of several subunits of GABAARs is regulated by phosphorylation, affecting the interaction of GABAA receptors with other proteins, the stability and the surface localization of the subunits, or the activity of the GABAA receptors [21]. For instance, PKC phosphorylates the α4 subunit in the intracellular loop (S443) increasing its stability, membrane insertion and the efficacy of GABAARs [40, 53]. Consistent with these previous data, we have determined by whole-cell patch-clamp recordings that the expression of Ccnd1 or the phosphomimetic allele of α4 (α4T423E/S431E) caused a decrease in the rundown of α4β3 containing GABAA receptors in HEK-293 cells. Moreover, Ccnd1 is not able to reduce the rundown of GABAA receptors containing the non-phosphorylatable α4 mutant (α4T423A/S431A). We propose that Ccnd1–Cdk4 enhances the response of α4-containing GABAA receptors by promoting the surface accumulation of α4 through its phosphorylation. On the one hand, Ccnd1 but not the inactive allele Ccnd1K112E is able to promote surface accumulation of α4. Moreover, Ccnd1 does not increase surface levels of the non-phosphorylatable mutant of α4 (α4T423A/S431A).
The adjustment of the surface levels of GABAAR subunits takes place through the regulation of export and internalisation processes. The ICL of β3 subunit is involved in internalisation of the receptors by clathrin. There is an atypical site for interaction with clathrin adaptor protein 2 (AP2) (KTHLRRRSSQLK), that contains the phosphorylation site for PKA, PKC and CamKII: it only binds AP2 when dephosphorylated [54]. Such a mechanism could also be happening for α4 subunit, that contains a similar motif (NMRKRTNAL) in the intracellular loop (aa 385–393). In fact, α4 has been described to interact directly with clathrin µ2 by immunoprecipitation [55]. Although T423 and S431 are not found inside the motif, phosphorylation of the ICL could be affecting the binding of α4 to clathrin. Another possibility is that the trafficking from the ER to the cytoplasmic membrane is affected. In fact, it is interesting to note that in Fig. 2G the D1KE samples had a higher level of α4 in the WCE which may coincide with a reduced transport of α4 to the surface and, a possible accumulation in the ER. Perhaps, phosphorylation at the ICL could affect the association of α4 with trafficking proteins. For instance, in the α4 and δ subunits, there are dibasic or multi-basic motifs in homologous positions that reduce export from the ER and increase transport back from the Golgi apparatus to the ER [56]. The mutation of dibasic motif to Ala in α4 enhances surface expression of the receptors. Thus, the increase of negative charges in the ICL could increase the α4 traffic to membranes.
Consistently with a role of Ccnd1–Cdk4/6 on GABAergic signalling, we have shown that inhibition of Cdk4/6 or the non-phosphorylatable mutant of α4 produces a decrease in tonic currents and the amplitude and frequency of mIPSCs in CA1 pyramidal neurons. Conversely, the phosphomimetic α4 allele increases both extra-synaptic and synaptic currents. Even though α4 is mostly extra-synaptic, it can also form synaptic GABAARs when assembled with γ2. Regarding synaptic currents, it could not be discarded that detection of synaptic alpha4-containing GABA receptors may have been influenced by spontaneous seizure-like events at the organotypic slice cultures. Concerning mIPSC responses, it is feasible that phosphorylation of α4 by CcndCdk4/6 complexes is still promoting its activity also when the α4 subunit is forming α4βxγ2 receptors at the synapse [57]. This would mean that α4 phosphorylation by CcndCdk4/6 has the same effect irrespective of whether the receptor localises synaptically or extra-synaptically. By contrast, α4-containing synaptic and extra-synaptic receptors show different modulation by PKA and PKC phosphorylation. PKA activation decreases synaptic α4 expression whilst increases extra-synaptic α4. Activation of PKC instead promotes α4 expression at the synapse and has no effect extra-synaptically [58].
Dendritic spines are the post-synaptic sites where most excitatory synapsis occurs. Their change in number and morphology underlies synaptic plasticity, important for learning and memory processes. GABA, through GABAARs activation promotes spine shrinkage and elimination [59]. Specifically, GABAARs containing α4βδ subunits have been involved in synaptic pruning at the CA1 and CA3 regions in the hippocampus [27, 28]: the expression of the α4 subunit of GABAARs increases in the hippocampus during adolescence [28] and the density of dendritic spines decreases post-pubertal, whilst in the GABRA4 KO mice this pruning does not occur. Therefore, the density of dendritic spines in the hippocampus of adult mice is increased in the GABRA4 KO mice [37]. The effect of α4 on synaptic pruning is mainly observed in mushroom- and stubby-shaped dendritic spines [27, 28]. According to a role of Ccnd1 in promoting α4-containing GABAARs activity, we observed an increase in the density of mushroom-type spines in CCND1 KO neurons, an effect rescued by the phosphomimetic allele of α4 at T423 and S431 (α4EE). Mushroom spines are considered to be one of the mature types of spines, involved in memory [60]. In this sense, the increase in α4βδ receptors at puberty impairs learning [25] and the GABRA4 KO mouse has increased spatial memory [37]. In the same direction, increased expression of Ccnd1 was found in CA1 region of aged learning-impaired rats [61] and the expression of Ccnd1 in the hippocampus of rats triggers deficits in spatial working memory [62]. Moreover, GABRA4 KO mice show an autistic-like behaviour, and autism has been associated with an increased number of spines [63]. The effect of Ccnd1 on GABA signalling and dendritic spines opens the possibility of exploring Ccnd1 functions in diseases related to the imbalance between inhibitory/excitatory signalling and dendritic spine abnormalities, such as autism spectrum disorders and epilepsy.
Data availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Sherr CJ, Roberts JM (2004) Living with or without cyclins and cyclin-dependent kinases. Genes Dev 18:2699–2711. https://doi.org/10.1101/gad.1256504
Hydbring P, Malumbres M, Sicinski P (2016) Non-canonical functions of cell cycle cyclins and cyclin-dependent kinases. Nat Publ Gr. https://doi.org/10.1038/nrm.2016.27
Neumeister P, Pixley FJ, Xiong Y et al (2003) Cyclin D1 governs adhesion and motility of macrophages. Mol Biol Cell. https://doi.org/10.1091/mbc.02-07-0102
Li Z, Wang C, Jiao X et al (2006) Cyclin D1 regulates cellular migration through the inhibition of thrombospondin 1 and ROCK signaling. Mol Cell Biol 26:4240–4256. https://doi.org/10.1128/mcb.02124-05
Cemeli T, Guasch-Vallés M, Nàger M et al (2019) Cytoplasmic cyclin D1 regulates glioblastoma dissemination. J Pathol 248:501–513. https://doi.org/10.1002/path.5277
Zhong Z, Yeow WS, Zou C et al (2010) Cyclin D1/cyclin-dependent kinase 4 interacts with filamin A and affects the migration and invasion potential of breast cancer cells. Cancer Res 70:2105–2114. https://doi.org/10.1158/0008-5472.CAN-08-1108
Meng H, Tian L, Zhou J et al (2011) Cell Cycle PACSIN 2 represses cellular migration through direct association with cyclin D1 but not its alternate splice form cyclin D1b. Cell Cycle 10:73–81. https://doi.org/10.4161/cc.10.1.14243
Fernández RMH, Ruiz-Miró M, Dolcet X et al (2011) Cyclin D1 interacts and collaborates with Ral GTPases enhancing cell detachment and motility. Oncogene 30:1936–1946. https://doi.org/10.1038/onc.2010.577
Fusté NP, Fernández-Hernández R, Cemeli T et al (2016) Cytoplasmic cyclin D1 regulates cell invasion and metastasis through the phosphorylation of paxillin. Nat Commun. https://doi.org/10.1038/ncomms11581
Fusté NP, Castelblanco E, Felip I et al (2016) Characterization of cytoplasmic cyclin D1 as a marker of invasiveness in cancer. Oncotarget 7:26979–26991
Chen K, Jiao X, Ashton A et al (2020) The membrane-associated form of cyclin D1 enhances cellular invasion. Oncogenesis. https://doi.org/10.1038/s41389-020-00266-y
Sicinski P, Donaher JL, Parker SB et al (1995) Cyclin D1 provides a link between development and oncogenesis in the retina and breast. Cell 82:621–630. https://doi.org/10.1016/0092-8674(95)90034-9
Lange C, Huttner WB, Calegari F (2009) Cdk4/CyclinD1 overexpression in neural stem cells shortens G1, delays neurogenesis, and promotes the generation and expansion of basal progenitors. Cell Stem Cell 5:320–331. https://doi.org/10.1016/j.stem.2009.05.026
Lim S, Kaldis P (2012) Loss of Cdk2 and Cdk4 induces a switch from proliferation to differentiation in neural stem cells. Stem Cells 30:1509–1520. https://doi.org/10.1002/stem.1114
Miyashita S, Owa T, Seto Y et al (2021) Cyclin D1 controls development of cerebellar granule cell progenitors through phosphorylation and stabilization of ATOH1. EMBO J 40:1–17. https://doi.org/10.15252/embj.2020105712
Sumrejkanchanakij P, Tamamori-Adachi M, Matsunaga Y et al (2003) Role of cyclin D1 cytoplasmic sequestration in the survival of postmitotic neurons. Oncogene 22:8723–8730. https://doi.org/10.1038/sj.onc.1206870
Sumrejkanchanakij P, Eto K, Ikeda MA (2006) Cytoplasmic sequestration of cyclin D1 associated with cell cycle withdrawal of neuroblastoma cells. Biochem Biophys Res Commun 340:302–308. https://doi.org/10.1016/j.bbrc.2005.11.181
Marampon F, Casimiro M, Fu M et al (2008) Nerve Growth factor regulation of cyclin D1 in PC12 cells through a p21RAS extracellular signal-regulated kinase pathway requires cooperative interactions between Sp1 and nuclear factor-kappaB. Mol Biol Cell 19:2566–2578. https://doi.org/10.1091/mbc.e06-12-1110
Schmetsdorf S, Gärtner U, Arendt T (2005) Expression of cell cycle-related proteins in developing and adult mouse hippocampus. Int J Dev Neurosci 23:101–112. https://doi.org/10.1016/j.ijdevneu.2004.07.019
Li C, Li X, Chen W et al (2007) The different roles of cyclinD1-CDK4 in STP and mGluR-LTD during the postnatal development in mice hippocampus area CA1. BMC Dev Biol 7:1–9. https://doi.org/10.1186/1471-213X-7-57
Nakamura Y, Darnieder LM, Deeb TZ, Moss SJ (2015) Regulation of GABAARs by phosphorylation. Adv Pharmacol 72:97–146. https://doi.org/10.1016/bs.apha.2014.11.008
Comenencia-Ortiz E, Moss SJ, Davies PA (2014) Phosphorylation of GABA A receptors influences receptor trafficking and neurosteroid actions. Psycopharmacology 231:3453–3465. https://doi.org/10.1007/s00213-014-3617-z
Chandra D, Jia F, Liang J et al (2006) GABAA receptor α4 subunits mediate extrasynaptic inhibition in thalamus and dentate gyrus and the action of gaboxadol. Proc Natl Acad Sci USA 103:15230–15235. https://doi.org/10.1073/pnas.0604304103
Shen H, Gong QH, Aoki C et al (2007) Reversal of neurosteroid effects at α4β2δ GABAA receptors triggers anxiety at puberty. Nat Neurosci 10:469–477. https://doi.org/10.1038/nn1868
Shen H, Sabaliauskas N, Sherpa A et al (2010) A critical role for α4βδ GABAA receptors in shaping learning deficits at puberty in mice. Science (80-) 327:1515–1518. https://doi.org/10.1126/science.1184245.A
Afroz S, Shen H, Smith SS (2017) α4βδ GABAA receptors reduce dendritic spine density in CA1 hippocampus and impair relearning ability of adolescent female mice: Effects of a GABA agonist and a stress steroid. Neuroscience 347:22–35. https://doi.org/10.1016/j.neuroscience.2017.01.051
Afroz S, Parato J, Shen H, Smith SS (2016) Synaptic pruning in the female hippocampus is triggered at puberty by extrasynaptic GABAA receptors on dendritic spines. Elife 5:1–23. https://doi.org/10.7554/eLife.15106
Parato J, Shen H, Smith SS (2019) α4βδ GABA A receptors trigger synaptic pruning and reduce dendritic length of female mouse CA3 hippocampal pyramidal cells at puberty. Neuroscience 398:23–36. https://doi.org/10.1016/j.neuroscience.2018.11.032
Shen H, Sabaliauskas N, Yang L et al (2017) Role of α4-containing GABAA receptors in limiting synaptic plasticity and spatial learning of female mice during the pubertal period. Brain Res 1654:116–122. https://doi.org/10.1016/j.brainres.2016.01.020
Moore MD, Cushman J, Chandra D et al (2010) Trace and contextual fear conditioning is enhanced in mice lacking the α4 subunit of the GABAA receptor. Neurobiol Learn Mem 93:383–387. https://doi.org/10.1016/j.nlm.2009.12.004
Duveau V, Laustela S, Barth L et al (2011) Spatiotemporal specificity of GABAA receptor-mediated regulation of adult hippocampal neurogenesis. Eur J Neurosci 34:362–373. https://doi.org/10.1111/j.1460-9568.2011.07782.x
Ma DQ, Whitehead PL, Menold MM et al (2005) Identification of significant association and gene-gene interaction of GABA receptor subunit genes in autism. Am J Hum Genet 77:377–388. https://doi.org/10.1086/433195
Collins AL, Ma D, Whitehead PL et al (2006) Investigation of autism and GABA receptor subunit genes in multiple ethnic groups. Neurogenetics 7:167–174. https://doi.org/10.1007/s10048-006-0045-1
Griswold AJ, Van Booven D, Cuccaro ML et al (2018) Identification of rare noncoding sequence variants in gamma-aminobutyric acid A receptor, alpha 4 subunit in autism spectrum disorder. Neurogenetics 19:17–26. https://doi.org/10.1007/s10048-017-0529-1
Fatemi SH, Reutiman TJ, Folsom TD et al (2010) mRNA and protein levels for GABAAα4, α5, β1 and GABABR1 receptors are altered in brains from subjects with autism. J Autism Dev Disord 40:612–624. https://doi.org/10.1007/s10803-009-0924-z.mRNA
Fatemi S, Reutiman T, Folsom TD, Thuras PD (2009) GABA A receptor downregulation in brains of subjects with autism. J Autism Dev Disord 39:223–230. https://doi.org/10.1007/s10803-008-0646-7
Fan C, Gao Y, Liang G et al (2020) Transcriptomics of Gabra4 knockout mice reveals common NMDAR pathways underlying autism, memory, and epilepsy. Mol Autism 11:1–17. https://doi.org/10.1186/s13229-020-0318-9
Gähwiler B, Campogna M, Debanne D et al (1997) Organotypic slice cultures: a technique has come of age. Trends Neurosci 20:471–477. https://doi.org/10.1016/S0166-2236(97)01122-3
Ruiz-Miró M, Colomina N, Fernández RMH et al (2011) Translokin (Cep57) interacts with cyclin D1 and prevents its nuclear accumulation in quiescent fibroblasts. Traffic 12:549–562. https://doi.org/10.1111/J.1600-0854.2011.01176.X
Abramian AM, Comenencia-Ortiz E, Vithlani M et al (2010) Protein kinase C phosphorylation regulates membrane insertion of GABA A receptor subtypes that mediate tonic inhibition. J Biol Chem 285:41795–41805. https://doi.org/10.1074/jbc.M110.149229
Baker GL, Landis MW, Hinds PW (2005) Multiple functions of D-type cyclins can antagonize pRb-mediated suppression of proliferation. Cell Cycle 4:329–337. https://doi.org/10.4161/cc.4.2.1485
Connelly WM, Errington AC, Di GG, Crunelli V (2013) Metabotropic regulation of extrasynaptic GABAA receptors. Front Neural Circuits 7:1–8. https://doi.org/10.3389/fncir.2013.00171
Bright DP, Smart TG (2013) Methods for recording and measuring tonic GABAA receptor-mediated inhibition. Front Neural Circuits 5:193. https://doi.org/10.3389/fncir.2013.00193
Tang X, Hernandez CC, Macdonald RL (2010) Modulation of spontaneous and GABA-evoked tonic α4β3δ and α4β3γ2L GABAA receptor currents by protein kinase A. J Neurophysiol 103:1007–1019. https://doi.org/10.1152/jn.00801.2009
Lagrange A, Botzolakis E, Macdonald R (2007) Enhanced macroscopic desensitization shapes the response of alpha4 subtype-containing GABAA receptors to synaptic and extrasynaptic GABA. J Physiol 1:655–676. https://doi.org/10.1113/jphysiol.2006.122135
Artegiani B, Lindemann D, Calegari F (2011) Overexpression of cdk4 and cyclinD1 triggers greater expansion of neural stem cells in the adult mouse brain. J Exp Med 208:937–948. https://doi.org/10.1084/jem.20102167
Yamada M, Takeshita T, Miura S et al (2001) Loss of hippocampal CA3 pyramidal neurons in mice lacking STAM1. Mol Cell Biol 21:3807–3819. https://doi.org/10.1128/MCB.21.11.3807-3819.2001
Cajigas IJ, Tushev G, Will TJ et al (2012) The local transcriptome in the synaptic neuropil revealed by deep sequencing and high-resolution imaging. Neuron 74:453–466. https://doi.org/10.1016/j.neuron.2012.02.036
Fernández-Hernández R, Rafel M, Fusté NP et al (2013) Cell cycle cyclin D1 localizes in the cytoplasm of keratinocytes during skin differentiation and regulates cell-matrix adhesion) Cyclin D1 localizes in the cytoplasm of keratinocytes during skin differentiation and regulates cell-matrix adhesion. Cell Cycle 12:2510–2517. https://doi.org/10.4161/cc.25590
Stelzer A, Kay A, Wong R (1988) GABAA-receptor function in hippocampal cells is maintained by phosphorylation factors. Science (80-) 241:339–341. https://doi.org/10.1126/science.2455347
Chen QX, Stelzer A, Kay AR, Wong RKS (1990) GABAA receptor function is regulated by phosphorylation in acutely dissociated guinea-pig hippocampal neurones. J Physiol 420:207–221
Gyenes M, Wang Q, Gibbs TT, Farb DH (1994) Phosphorylation factors control neurotransmitter and neuromodulator actions at the gamma-aminobutyric acid type A receptor. Mol Pharmacol 46:542–549
Abramian AM, Comenencia-Ortiz E, Modgil A et al (2014) Neurosteroids promote phosphorylation and membrane insertion of extrasynaptic GABAA receptors. Proc Natl Acad Sci USA 111:7132–7137. https://doi.org/10.1073/pnas.1403285111
Kittler JT, Chen G, Honing S et al (2005) Phospho-dependent binding of the clathrin AP2 adaptor complex to GABA A receptors regulates the efficacy of inhibitory synaptic transmission. Proc Natl Acad Sci USA 102:14871–14876. https://doi.org/10.1073/pnas.0506653102
Gonzalez C, Moss SJ, Olsen RW (2012) Ethanol promotes clathrin adaptor-mediated endocytosis via the intracellular domain of δ-containing GABAA receptors. J Neurosci 32:17874–17881. https://doi.org/10.1523/JNEUROSCI.2535-12.2012
Bracamontes JR, Li P, Akk G, Steinbach JH (2014) Mutations in the main cytoplasmic loop of the GABA A receptor a 4 and d subunits have opposite effects on surface expression. Mol Pharmacol 86:20–27. https://doi.org/10.1124/mol.114.092791
Wafford KA, Thompson SA, Thomas D et al (1996) Functional characterization of human γ-aminobutyric acidA receptors containing the a4 subunit. Mol Pharmacol 50(3):670–678
Bohnsack JP, Carlson SL, Morrow AL (2016) Differential regulation of synaptic and extrasynaptic α4 GABA(A) receptor populations by protein kinase A and protein kinase C in cultured cortical neurons. Neuropharmacology 105:124–132. https://doi.org/10.1016/j.neuropharm.2016.01.009
Hayama T, Noguchi J, Watanabe S et al (2013) GABA promotes the competitive selection of dendritic spines by controlling local Ca2+ signaling. Nat Neurosci 16:1409–1416. https://doi.org/10.1038/NN.3496
Bourne J, Harris KM (2007) Do thin spines learn to be mushroom spines that remember? Curr Opin Neurobiol 17:381–386. https://doi.org/10.1016/j.conb.2007.04.009
Burger C, López MC, Feller JA et al (2007) Changes in transcription within the CA1 field of the hippocampus are associated with age-related spatial learning impairments. Neurobiol Learn Mem 87:21–41. https://doi.org/10.1016/j.nlm.2006.05.003
Wu K, Li S, Bodhinathan K et al (2012) Enhanced expression of Pctk1, Tcf12 and Ccnd1 in hippocampus of rats: impact on cognitive function, synaptic plasticity and pathology. Neurobiol Learn Mem 97:69–80. https://doi.org/10.1016/j.nlm.2011.09.006
Penzes P, Cahill ME, Jones KA et al (2011) Dendritic spine pathology in neuropsychiatric disorders. Nat Neurosci 14:285–293. https://doi.org/10.1038/nn.2741
Acknowledgements
We are grateful to M. Encinas for plasmids. We thank Sònia Rius for technical assistance and members of the CYC lab for helpful considerations. The cell culture experiments were performed in the Cell Culture Scientific & Technical Service from Universitat de Lleida.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This work was funded by the Spanish Ministry of Innovation and Science MICINN (PID2019-104859GB-I00 to E.G. and PID2020-117651RB to J.A.E.) and by Generalitat de Catalunya (2017-SGR-569). M. Ventura Monserrat was supported by a predoctoral fellowship of University of Lleida.
Author information
Authors and Affiliations
Contributions
NP conceptualization, supervision, validation, investigation, methodology, writing-original draft, writing—review and editing; MVM validation, investigation, methodology; FF funding acquisition, data curation, writing—review and editing; JT funding acquisition, supervision, writing—review and editing; NC funding acquisition, methodology; FM-C investigation, methodology; JPP investigation, methodology; DS funding acquisition, investigation, validation, methodology; EL-M investigation, methodology; CG-V investigation, methodology; JAE supervision, validation, methodology, funding acquisition; JE funding acquisition, conceptualization, methodology; EG conceptualization, supervision, funding acquisition, investigation, writing—review and editing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
The animal procedures were performed according to the guidelines set out in the European Community Council Directives (2010/63/EU, 86/609/EEC) and in accordance with the Law 5/1995 and the Decree RD53/2013 (Catalan Government). The experiments were approved by the Ethics Committee of the University of Lleida (CEEA 03-02/19) and by the bioethics committee from the Consejo Superior de Investigaciones Cientificas.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pedraza, N., Monserrat, M.V., Ferrezuelo, F. et al. Cyclin D1—Cdk4 regulates neuronal activity through phosphorylation of GABAA receptors. Cell. Mol. Life Sci. 80, 280 (2023). https://doi.org/10.1007/s00018-023-04920-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-023-04920-7