Abstract
Gastric cancers are highly heterogeneous malignant tumors. To reveal the relationship between differentiation status of cancer cells and tumor immune microenvironments in gastric cancer, single-cell RNA-sequencing was performed on normal mucosa tissue, differentiated gastric cancer (DGC) tissue, poorly differentiated gastric cancer (PDGC) tissue and neuroendocrine carcinoma (NEC) tissue sampled from surgically resected gastric cancer specimens. We identified the signature genes for both DGC and PDGC, and found that signature genes of PDGC strongly enriched in the epithelial–mesenchymal transition (EMT) program. Furthermore, we found that DGC tends to be immune-rich type whereas PDGC tends to be immune-poor type defined according to the density of tumor-infiltrating CD8+ T cells. Additionally, interferon alpha and gamma responding genes were specifically expressed in the immune-rich malignant cells compared with immune-poor malignant cells. Through analyzing the mixed adenoneuroendocrine carcinoma, we identified intermediate state malignant cells during the trans-differentiation process from DGC to NEC, which showed double-negative expressions of both DGC marker genes and NEC marker genes. Interferon-related pathways were gradually downregulated along the DGC to NEC trans-differentiation path, which was accompanied by reduced CD8+ cytotoxic T-cell infiltration. In summary, molecular features of both malignant cells and immune microenvironment cells of DGC, PDGC and NEC were systematically revealed, which may partially explain the strong tumor heterogeneities of gastric cancer. Especially along the DGC to NEC trans-differentiation path, immune-evasion was gradually enhanced with the decreasing activities of interferon pathway responses in malignant cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Worldwide, gastric cancer ranks fifth in incidence and fourth in mortality, especially in Eastern Asia and Eastern Europe, where the incidence is significantly higher than that in other areas [1, 2]. Gastric cancer is a malignant entity of various pathological types, including well/moderately differentiated adenocarcinoma, poorly differentiated adenocarcinoma, mucous adenocarcinoma and signet ring cell carcinoma [3, 4]. In addition to the above “inter-tumor” heterogeneity, significant “intra-tumor” heterogeneity exists in gastric cancer [5, 6]. Clinically, it is common for different tumor components to coexist in the same gastric cancer, including differentiated adenocarcinoma, poorly differentiated adenocarcinoma or signet ring carcinoma [3, 7]. Some specific tumor components, such as hepatoid adenocarcinoma, neuroendocrine carcinoma and adenocarcinoma with enterocyte differentiation, make this heterogeneity more drastic and result in an even worse prognosis [8,9,10,11,12].
With the development of high-throughput sequencing technology, the understanding of the biological characteristics of gastric cancer has gradually deepened [13,14,15,16]. The Cancer Genome Atlas (TCGA) and Asian Cancer Research Group (ACRG) have defined specific molecular subtypes of gastric cancer, which to some extent has promoted progress in the clinical treatment of gastric cancer [17, 18]. Recent advances in single-cell sequencing techniques have led to further insights into the carcinogenesis and progression of gastric cancer [19,20,21,22,23,24], particularly the tumor microenvironment [25,26,27,28,29]. However, there is still a lack of research on the mechanisms of how tumor epithelial cells in different subtypes of gastric cancer affect the tumor immune microenvironment differently, and few studies have emphasized on the heterogeneity of gastric cancer. Here, utilizing a single-cell transcriptomic sequencing approach, we compared the differences between differentiated gastric cancer and other types of gastric cancer components, thus further elucidating the heterogeneity of gastric cancer and unveiling the relationship between gastric cancer cells and immune infiltration.
Materials and methods
Patients and samples
Normal mucosa was sampled at least 5 cm away from the tumor border. Samples were obtained immediately after surgical resection. Each sample was divided into 3 parts: one for single-cell RNA sequencing, one for pathological examination and one for cryopreservation. The histopathological information of the corresponding tissues was evaluated by two independent pathologists. Tumor staging was classified according to the 2017 TNM classification of the American Joint Committee on Cancer (AJCC).
Single-cell cDNA amplification and library construction
We performed single-cell separation, DNA amplification and library construction following the manufacturer’s guidelines for chromium single-cell sequencing technology from 10X Genomics (USA). The sequencing libraries were constructed using the Chromium Single Cell 3ʹ Library and Gel Bead Kit V3. Finally, the prepared libraries were quality checked and sequenced on an Illumina NovaSeq 6000 platform.
Laser capture microdissection
The tumor tissues were formalin-fixed and paraffin-embedded (FFPE) and then cut into 5-μm-thick sections, fixed with 95% ethanol and subjected to hematoxylin and eosin staining for conventional histological examination by three independent pathologists to determine the location of malignant cells. Then, experimental tissues (8 μm thick) that were placed onto membrane slides (MMI, Germany) for laser capture microdissection were fixed with 95% ethanol and stained with eosin. Samples on the slides were isolated by laser capture microdissection using the CellCut Plus system (MMI, Germany). The dissected tissues were lysed, and genomic DNA was extracted using the DNA Extraction Kit for FFPE samples (Amoy Diagnostics, China) according to the manufacturer’s protocol.
Whole-exome sequencing (WES)
Extracted genomic DNA (approximately 200 ng) was fragmented by a Covaris system (Thermo Fisher Scientific, USA) into fragments of 150–200 bp. We used the DNA Clean & Concentrator Kit (Tianmo Biotech, China) for purification according to the manufacturer’s protocol. Next, libraries for WES capture were used with SureSelectXT Human All Exon V6 (Agilent Technologies, USA) following the provided instructions. The products were sequenced on an Illumina NovaSeq 6000 platform.
Immune-histochemical staining
Tissues were cut into 5-μm-thick sections and were then deparaffinized and treated with 3% H2O2-CH3OH for 15 min to block endogenous peroxidase. The slices were submerged in pH 6.0 or 9.0 buffer for antigen retrieval and then incubated at 37 °C for 2 h with primary antibodies, including anti-CLDN3 (Abcam, ab214487, 1:500), anti-FABP1 (Abcam, ab171739, 1:4000), anti-PIGR (Abcam, ab275020, 1:500), anti-PHGR1 (SinoBiological, 204852-T08, 1:100), anti-MUC1 (Abcam, ab70475, 1:100), anti-CD8 (ZSGB-Bio, ZA-0508), anti-MUC6 (ZSGB-Bio, ZM-0396), anti-MUC5AC (ZSGB-Bio, ZA-0664), anti-Syn (ZSGB-Bio, ZM-0246) and anti-CgA (ZSGB-Bio, ZM-0076). After incubation with horseradish peroxidase (HRP)‐conjugated IgG (ZSGB‐Bio, China) at room temperature for 30 min, the cells were stained with a 3,3N-diaminobenzidine tetrahydrochloride (DAB) detection kit (ZSGB‐Bio, China). The expression and localization of proteins were detected under light microscopy.
Processing of single-cell RNA-seq data
We processed our data following the manufacturer’s instructions of 10X Cell Ranger (version 3.1.0) with default arguments used to process the raw data, and we used the human GRCh38 genome as the reference genome. We retained cells with more than 500 detected genes, more than 1000 transcripts and less than 50% mitochondrial genes because of the high percentages of mitochondrial genes in parts of normal gastric cells. The unique molecular identifiers (UMIs) of each cell were normalized and transformed through log-transformation. The log-normalized values were used in the downstream analyses.
Clustering and differentially expressed gene analysis
The R package Seurat (version 3.2.2) was used to perform the downstream analyses, and Harmony (version 1.0) was used to correct batch effects of different experimental batches [30, 31]. Individual cells were clustered through a graph-based clustering approach of Seurat. Differentially expressed genes (DEGs) of different cell types were identified through the FindAllMarkers function of Seurat using the Wilcoxon test and the fold change with a value of 2. The detailed parameters were as follows: test.use = 'wilcox', min.pct = 0.25, and logfc.threshold = log(1.5). Metascape (http://metascape.org/) was used to perform gene enrichment analysis [32].
CNVs inferred by single-cell RNA-seq data
The R package CopyKAT (version 1.0.4) was used to infer CNVs based on the gene expression of all the epithelial cells in our single-cell RNA data [33]. The parameters ngene.chr = 5, win.size = 25, and KS.cut = 0.05 were used, and epithelial cells from the adjacent normal tissues were assigned as normal epithelial cells during the analysis. According to CNV outputs inferred by gene expression, the mean of squares of deviation as the measurement represents the CNV levels of single cells.
Classification of malignant cells and non-malignant epithelial cells
With the identified clusters of epithelial cells, cell clusters of epithelial cells with high CNV levels were manually annotated as malignant cells, and other clusters of epithelial cells with low CNV levels were annotated as non-malignant cells. The classification strategy is as following. First, we confirmed the CNV patterns of cancer cells by WES data from the corresponding laser capture microdissected tissues, which guaranteed a high cancer cell purity by combining pathology information (Fig. S2A). Second, we combined both clustering and CNV information to distinguish malignant cells from non-malignant epithelial cells. Non-malignant epithelial cells clustered together with normal epithelial cells from normal mucosa. And malignant cells were separated from non-malignant epithelial cells in tumor samples and epithelial cells from normal mucosa clearly (Fig. 1F and Fig. S2B). Third, non-malignant epithelial cells from different tumor samples clustered together with normal epithelial cells and mixed very well, while malignant cells were separated among tumor samples from different patients (Fig. S1G). The above information confirms the validity of our strategy to better distinguish malignant cells from non-malignant epithelial cells in tumor samples.
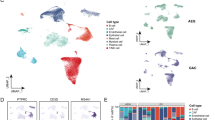
Representative single-cell transcriptome landscape of gastric cancer. A UMAP plot showing different cell types (n = 46,883), with pathological information including differentiated gastric cancer (DGC), poorly differentiated gastric cancer (PDGC) and neuroendocrine carcinoma (NEC). B The expression of corresponding markers for different cell types. C UMAP plot for one normal mucosa tissue (n = 2232). D Heatmap showing chromosomal landscape of large-scale copy number variations (CNV) inferred from single cell RNA data. E Violin plot showing CNV deviation of non-malignant and malignant epithelial cells from each case. F UMAP plot showing different clusters of epithelial cells. G Heatmap showing differentially expressed genes (DEGs) between malignant and non-malignant epithelial cells. Part gene names are displayed and the full gene list is in Supplementary Table S4. H Gene regulatory networks (GRNs) identified in malignant and non-malignant epithelial cells. Part transcription factor names are displayed. I Gene enrichment analysis showing the signaling pathways relatively up- or downregulated in malignant compared with non-malignant epithelial cells. J Heatmap showing the expressing scores of hallmark gene sets in malignant and non-malignant epithelial cells. K Violin plot showing the expression levels of interferon alpha and gamma response genes in different epithelial clusters
CNVs inferred by WES data
For the WES data, low-quality and adapter-contaminated reads were removed and trimmed by Trimmomatic (version 0.39), and then clean reads were aligned to the human GRCh38 genome using BWA (version 0.7.17). Control-freec (version 11.5) was used to estimate CNVs, outputting the copy numbers of 10 M segments.
Identification of the tumor differentiation gene set
DEGs of differentiated gastric cancer (DGC) compared with poorly differentiated gastric cancer (PDGC) were identified with the FindMarker function built in the Seurat package. Genes highly correlated with the DEGs of DGC were selected as candidate genes, with a Pearson correlation greater than 0.5. Among these candidate genes, we used the lasso regression and random forest methods to remove the uncertain genes and identify the tumor differentiation gene set. The glmnet function of the package glmnet (version 2.0–16) was applied to perform lasso regression, with the parameters alpha = 1 and family = 'binomial'. The randomForest function of the randomForest package (version 4.6–15) was used to perform random forest, with the parameters ntree = 1500 and mtry = 10. With prior knowledge about tumor differentiation, genes with low contributions assessed by lasso regression and random forest and with little association with tumor differentiation were removed from the candidate gene set, including TM4SF20, HEPH, LINC01133, PPP1R1B, AKR1C3 and REEP6. Finally, we retained genes whose expressing level (the log-normalized value) is higher than the ln value of 0.5. The retained genes were identified as the tumor differentiation gene set and was shown in Supplementary Table S6.
Identification of gene expression patterns in gastric cancer cells
To identify diverse gene expression programs of malignant epithelial cells, we first used the AUCell package (version 1.6.1) to score malignant cells in each sample for highly variable genes and carried out feature extraction using nonnegative matrix factorization (NMF) with the default “scd” decomposition algorithm and 10 factors in the factorization per tumor sample with the function implemented in the NNLM package (version 0.4.2). Then, the Pearson correlation coefficients of factors from different tumor samples were clustered and manually divided into several programs. These programs were annotated with specific functional genes and pathways.
Gene regulatory network (GRN) analysis
The R package SCENIC (version 1.1.3) was used to assess the transcription factor strength [34]. We used the GENIE3 method to detect correlations based on the dataset of motifs located 20 kb around TSSs from the cisTarget database.
Gene set signature and signaling pathway enrichment analysis
To assess the expression levels of different signatures at the single-cell resolution, the AddModuleScore function of Seurat was used to score gene sets of single cells. We also used the Gene Set Variation Analysis (GSVA) enrichment scores to describe enrichment scores of the hallmark gene sets at the single-cell levels through the R package GSVA (version 1.32.0). The “ssgsea” method of the GSVA package was used [35].
Quantifying metabolic activity at single-cell resolution
The R package scMetabolism (version 0.2.1) was used to quantify different types of metabolic activities at the single-cell level [36]. We used the parameters of method = 'VISION' and metabolism.type = 'KEGG' to perform the analysis.
Pseudotime analysis
The R package Monocle2 (version 2.12.0) with the DDR-Tree method and default parameters was used to perform the single-cell trajectory analysis [37]. We used the DEGs or marker genes of individual clusters as ordering gene sets. The cell trajectories were inferred with the default parameters after dimension reduction and cell ordering.
Cell communication analysis
The toolkit NATMI was used to perform cell communication analysis [38]. The ExtractEdges function calculates the expression and specificity of ligand–receptor interactions, and the DiffEdges function identifies changes in ligand–receptor interactions between two conditions. The built-in “lrc2p” database was used to predict interactions, the weight of edges was calculated by the mean method, and the detection threshold value was set to 0.2.
Survival analysis
Survival analysis of gastric cancer samples from the TCGA dataset based on the expression status of identified genes was carried out by the survival package (version 0.4.8) and the survminer package (2.44–1). The assumption of the Cox proportional hazards model was tested using cox with 0.1 as the cut-off value, and the Cox proportional hazards model was fit using patient groups divided by the median gene expression level.
Results
Representative single-cell transcriptome landscape of gastric cancer obtained by a pathology-informed sampling strategy
We applied a pathology-informed sampling strategy and droplet-based scRNA-seq (10X Genomics) on 14 tissues sampled from surgically resected gastric cancer specimens, including 1 normal mucosa tissue, 6 differentiated gastric cancer (DGC) tissues, 6 poorly differentiated gastric cancer (PDGC) tissues and 1 neuroendocrine carcinoma (NEC) tissue, all of which were confirmed by side-by-side histopathological examination (Supplementary Table S1). The neuroendocrine carcinoma tissue and a corresponding adenocarcinoma tissue were from the same tumor pathologically diagnosed as mixed adenoneuroendocrine carcinoma (MANEC).
After stringent quality control, 46,883 cells were ultimately retained for further analyses, which detected a median of 1723 genes and 5480 unique molecular identifiers (UMIs) per cell (Fig. S1A). After dimension reduction and clustering, 8 clusters of cells were identified, and the captured cells were clearly divided into epithelial cells (EPCAM and KRT8), fibroblasts (THY1 and DCN), endothelial cells (PECAM1 and CDH5), myeloid cells (C1QA and CD163), T/NK cells (CD3D and CD3E), B cells (CD79A and MS4A1), plasma cells (MZB1) and mast cells (KIT and TPSAB1) according to the expression of well-known marker genes (Supplementary Table S2 and Fig. 1A, B). The percentages of these cell types differed in tumor samples, and epithelial cells and T/NK cells comprised the majority (Fig. S1B–E).
We captured one normal mucosa tissue in our data, and epithelial cells of this sample were divided into progenitor cells, surface cells, chief cells, parietal cells and endocrine cells according to well-known markers (Fig. 1C and Fig. S1F). Then, we estimated CNV levels of epithelial cells from tumor tissues, with high-CNV and low-CNV epithelial cells existed in every tumor sample (Fig. 1D, E). Cells with high CNVs were confirmed by WES data from the corresponding laser capture microdissected tissue (Fig. S2A). To classify malignant cells from non-malignant epithelial cells in our data, we combined clustering and CNV information to distinguish malignant cells from non-malignant cells (Fig. S1G, H), resulting in 11 malignant cell clusters and 7 non-malignant cell clusters (Supplementary Table S3, Fig. 1F and Fig. S1I, J). Non-malignant epithelial cells in tumor samples from different patients clustered together with the epithelial cells from normal mucosa, while malignant cells were clustered by samples and separated from non-malignant epithelial cell clusters (Fig. S2B, C). By comparing malignant and non-malignant epithelial cells, we found that malignant cells lost digestive and defense functions but enhanced protein synthesis activities and metabolism (Supplementary Table S4 and Fig. 1G, I). We also assessed the gene regulatory networks (GRNs) of malignant and non-malignant cells and clustered them based on GRNs that classified epithelial cells according to different patients (Fig. S2D). GRNs, such as those for YY1, RAD21, RARG, TP63 and CERBPB, were identified in malignant cells, while several GRNs, including those for SP1, IRF7, IRF9, FOXA1 and KLF2, were identified in non-malignant epithelial cells (Fig. 1H).
We assessed different signatures of malignant clusters and found that endocrine cells represented some pancreatic beta cell-like signatures, which displayed secretion-related features (Fig. 1J and Fig. S2G). Moreover, the T1 and T6 clusters represented interferon alpha and gamma response signatures, which implies that the T1 and T6 clusters showed unique immunological related features (Fig. 1J, K and Fig. S2E, F). We also evaluated the metabolism-related activities in epithelial clusters and found that glycerolipid and glycerophospholipid metabolism activities were higher in T6 than in other malignant clusters (Fig. S2H, I).
Differentiation signature of cancer cells identified by comparing DGC with PDGC
We compared the transcriptomes of DGCs and PDGCs with a clear pathological diagnosis to clarify their molecular differences (Fig. 2A, B). As for DEGs of malignant cells from PDGC and DGC, the expressions of CLDN3, FABP1, S100A10 and PHGR1 were significantly higher in DGCs, while epithelial–mesenchymal transition (EMT)-related genes such as VIM, LAMB3, LAMC2, COL6A1 and COL17A1 were specifically expressed in PDGCs¸ indicating that the EMT signature was enriched in PDGCs (Supplementary Table S5, Fig. 2C, D and Fig. S3A). Several EMT-related terms such as focal adhesion, response to wounding, positive regulation of cell migration and cell–cell adhesion were enriched in PDGC (Fig. 2E). Compared with non-malignant epithelial cells, there were more upregulated and downregulated DEGs in PDGCs than in DGCs (Fig. S3B, S3D). In addition, HIF1A was mainly expressed in PDGCs, indicating potentially stronger hypoxia in PDGCs (Fig. S3E).
Comparison between differentiated gastric cancer (DGC) and poorly differentiated gastric cancer (PDGC). A The percentage of malignant cells in DGC, PDGC and neuroendocrine carcinoma (NEC). B UMAP plot showing distribution of epithelial cells from DGC, PDGC and NEC. C Heatmap showing differentially expressed genes between malignant epithelial cells from DGC and PDGC. Part gene names are displayed and the full gene list is in Supplementary Table S5. D Violin plot showing the expression levels of EMT-associated genes in DGC and PDGC. E Gene enrichment analysis showing the functions relatively upregulated in PDGC. F Non-negative matrix factorization (NMF) showing transcriptomic programs featured by DGC and PDGC. G Violin plot showing the expression levels of genes involved in differentiation score from DGC and PDGC (****—p value < 0.0001 calculated by wilcox test). H UMAP plot showing the levels of differentiation score. I The correlation between the tumor differentiation score and tumor score. J Kaplan–Meier survival (days) curve showing the survival of intestinal-type gastric cancer cases in the TCGA dataset with high and low differentiation score
To assess the gene expression programs of DGCs and PDGCs, we used the nonnegative matrix factorization (NMF) method to analyze malignant cells of different cancer types. We found that the EMT, epithelial-related and metabolism/energy programs were enriched in PDGCs which was confirmed by gene enrichment analysis, whereas absorption/transportation, cell cycle, endocytosis, interleukin-related signaling, metabolism/energy, protein maturation, protein processing and response to external stimulus programs were enriched in DGCs (Fig. 2F).
Based on upregulated genes highly correlated with malignant cells of DGCs, we used the lasso regression and random forest methods to trim low-contribution genes. Finally, we identified a gene set associated with the differentiation status of gastric cancer, including AGR3, CLDN3, CLDN4, FABP1, LGALS4, PHGR1, MYH14 and S100A14 (Supplementary Table S6 and Fig. S3F, G). The identified tumor differentiation gene set had significantly higher expression in DGCs than in PDGCs (Fig. 2G and Fig. S3H). This gene signature clearly defined DGC, especially in the T1 and T6 clusters in our dataset (Fig. 2H). Additionally, we proposed a score based on the upregulated markers of all malignant cells as the tumor score. The tumor score had a negative correlation with our identified tumor differentiation gene set, which indicated that a higher differentiation level of cancer cells represented a less malignant signature of the tumor (Fig. 2I). Moreover, in the TCGA database, the higher expression of our differentiation signature gene set was clearly associated with better overall survival in intestinal-type gastric cancer (Fig. 2J).
Differentiation signatures partially overlapped with immune infiltration-related genes
Next, to study the relationship between the differentiation signature of the cancer cells and characteristics of the corresponding immune microenvironments, we divided the patients into the immune-poor type and immune-rich type according to the density of tumor-infiltrating CD8+ T cells [39]. We found that the proportion of malignant cells was higher in the immune-poor type, which indicated the high density of malignant cells in immune-poor tumors, as indicated by H&E staining (Fig. 3A, B). We identified markers such as S100A2, CAV1, CAV2 and ANXA1 and GO terms including focal adhesion, integrin-related pathways and response to wounding in the immune-poor type, while genes such as LAGLS4, CLDN3 and PIGR and GO terms including granulocyte migration, neutrophil degradation and endocytosis were identified in the immune-rich type (Supplementary Table S7, Fig. 3C and Fig. S4A). Moreover, gastric cancer samples of MSI-H and MSI-L in the TCGA dataset were used to confirm DEGs between immune-poor and immune-rich types, and as expected, markers of malignant cells in the immune-poor type tumor tissues had a higher expression in MSI-L samples, while markers of malignant cells in the immune-rich type tumor tissues had a higher expression in MSI-H samples in the TCGA dataset (Fig. 3D). The percentage of malignant cells from immune-rich and immune-poor types differed in malignant clusters, and malignant cells in the immune-rich type tumor tissues were enriched in the T1, T6 and T7 clusters (Fig. S4C). Moreover, we analyzed the pseudotime trajectory of malignant cells according to the immune-related gene sets, and the path of malignant cells from immune-rich types to immune-poor types was identified, indicating a continuous transition between these two types (Fig. S4B).
Transcriptomic profiling of immune infiltration-related genes. A The percentage of malignant cells in immune-rich and immune-poor type samples. B UMAP plot showing distribution of epithelial cells from immune-rich and immune-poor type samples. C Heatmap showing differentially expressed genes between immune-rich and immune-poor type gastric cancer. Part gene names are displayed and the full gene list is in Supplementary Table S7. D Violin plot showing expression levels of markers for immune-rich malignant cells in microsatellite instability-high (MSI-H) and microsatellite instability-low (MSI-L) samples from the TCGA dataset (ns—P value > = 0.05, *—P value < 0.05, **—P value < 0.01, ***—P value < 0.001, ****—P value < 0.0001 calculated by t test). E Violin plot showing expression levels of hallmark signatures and GO gene sets of immune response in immune-rich and immune-poor type gastric cancer. F Violin plot showing expression levels of genes in response to interferon gamma and antigen presentation-related genes in immune-rich and immune-poor type gastric cancer. G Violin plot showing expression levels of exhausted ligand-related and costimulatory ligand-related genes in immune-rich and immune-poor type gastric cancer. H Hallmark gene signatures highly correlated with exhausted ligand and costimulatory ligand gene sets. I Violin plot showing expression levels of interferon alpha/gamma response genes, exhausted ligand and costimulatory ligand-related response genes in MSI-H gastric cancer samples in MSI-H and MSI-L samples from the TCGA dataset (ns—P value > = 0.05, *—P value < 0.05, **—P value < 0.01, ***—P value < 0.001, ****—P value < 0.0001 calculated by t test). J Violin plot showing differentiation score in immune-rich and immune-poor type gastric cancer. K Representative images of hematoxylin–eosin staining, and immune-histochemical staining of CD8 and other differentially expressed proteins in immune-rich and immune-poor type samples. Scale bar, 300 μm
We utilized several hallmark signatures and GO gene sets of immune responses to score tumor cells from immune-rich and immune-poor types. The cancer cells in immune-rich type tumors tended to have higher expression of interferon alpha and interferon gamma response genes (Fig. 3E and Fig. S4D). Genes in response to interferon gamma, including CCL20, CCL25 and CD74, and antigen presentation-related genes, such as HLA-DPA1, HLA-DRA and HLA-DRB, were significantly upregulated in the cancer cells of immune-rich type tumors (Fig. 3F). Moreover, to identify the responses of tumor cells to infiltrative T cells, we studied the exhausted and costimulatory receptors and found that the cancer cells of immune-rich type tumors showed higher expression of both the corresponding exhausted ligand-related and costimulatory ligand-related genes compared with the immune-poor type (Fig. 3G). Meanwhile, the hallmark gene signatures highly correlated with exhausted ligand and costimulatory ligand gene sets were enriched in immune-related activities, such as interferon gamma response, interferon alpha response, inflammatory responses and complement activities (Fig. 3H). We also screened these features in the TCGA dataset and found higher expression of interferon alpha/gamma response genes, exhausted ligand and costimulatory ligand-related response genes in MSI-H gastric cancer samples, which were characterized by heavy immune infiltration (Fig. 3I).
As seen from the heatmap, the DEGs between the immune-poor type and the immune-rich type in general overlapped with those between PDGC and DGC, and the malignant cells in the immune-rich type had a higher differentiation score than those in the immune-poor type (Fig. 3J, K). We also identified genes that were highly correlated with interferon gamma and the interferon alpha pathway, and several genes were also differentially expressed in the malignant cells between the immune-rich and immune-poor types (Fig. S4E). For example, PHGR1 and CLDN3 showed significantly higher expression in the malignant cells of the immune-rich type and was mainly expressed in the T1 and T6 clusters (Fig. S4F). Meanwhile, their expression was associated with better prognosis of intestinal-type gastric cancer in the TCGA database (Fig. S4G). Finally, we also identified metabolic activities highly correlated with interferon alpha/gamma signatures and found that lipid-related signatures, such as retinol metabolism and steroid hormone biosynthesis, were positively correlated with the immune-rich type (Fig. S4H, I), while pyruvate-related signatures, such as purine metabolism, pyruvate metabolism and pyrimidine metabolism, were negatively correlated with the immune-rich type [40].
Tumor infiltrating T-cell characteristics and their interactions with malignant cells
To delineate NK/T-cell clusters and the association between T cells and malignant cells in detail, we analyzed all the T cells and divided them into 17 clusters based on the corresponding markers, including 3 NK or γ/δ T-cell clusters, 4 CD8+ T-cell clusters, 6 CD4+ T-cell clusters and 4 other T-cell clusters of specific signatures (Fig. 4A, B). These clusters were distributed relatively evenly in different tumor samples (Fig. 4C). Of these T-cell clusters, STMN1+ T cells represent the cell cycle signature and contain both CD8+ and CD4+ T cells (Fig. S5A). For these four CD8+ T cell clusters (GZMB+ CD8+ T cells, GZMH+ CD8+ T cells, GZMK+ CD8+ T cells and TNFSF9 CD8+ T cells), GZMB+ CD8+ T cells and GZMH+ CD8+ T cells have a stronger cytotoxic ability than others, while GZMB+ CD8+ T cells have a specific exhausted signature compared with other CD8+ T cells (Fig. 4D). We screened out exhaustion-related genes such as PDCD1, CTLA4 and HAVCR, which are mainly expressed in GZMB+ CD8+ T cells (Fig. S5B).
Tumor infiltrating T cell characteristics and interactions with malignant cells. A UMAP plot showing different clusters of T cells. B The expression of corresponding markers for different T cell clusters. C The percentage of different types of T cells in each sample. D Costimulatory, cytotoxic, exhausted and resident T cell signature score in different types of CD8+ T cells. E UMAP plot showing distribution of T cells across PDGCs and DGCs, immune-rich and immune-poor types. F Heatmap showing differentially expressed genes between GZMB+ CD8+ T cells from immune-rich and immune-poor type gastric cancer. Part gene names are displayed. G Interactions between epithelial cells and CD8+ T cells inferred from cell communication analysis using the toolkit NATMI
T cells from both DGCs and the immune-rich type account for the majority of T cells we captured (Fig. 4E). Moreover, all of the four main functional CD8+ T cell types were significantly enriched in the immune-rich type, which was consistent with the pathological findings (Fig. S5C, D). We found that GZMB+ CD8+ T cells from the immune-rich type had a higher expression of CXCL13 (Fig. 4F and Fig. S5E), which was also reported to be associated with good responses to immunotherapies in triple-negative breast cancer [41]. Also, we found that energy-producing functions such as oxidative phosphorylation, glycolysis/gluconeogenesis and the citrate cycle had a positive correlation with cytotoxic and exhausted signatures (Fig. S5F, G), indicating that the cytotoxic and exhausted functions of CD8+ T cells probably consume more energy.
Then we conducted an in-depth analysis of cell–cell communications. For different clusters of malignant cells and CD8+ T cells, we found that exhaustion-related ligands were mainly enriched in the T1, T4 and T6 malignant clusters, and exhaustion-related receptors were mainly enriched in GZMB+ CD8+ T cells, especially interactions such as LAGLS3-LAG3, CEACAM1-HAVCR2 and NECTIN4-TIGIT, which indicated that these pairs of malignant cells and CD8+ T cells were involved in T-cell exhaustion (Fig. 4G). Regarding the communications between immune types and different types of CD8+ T cells, we also identified exhaustion-related interactions between malignant cells and GZMB+ CD8+ T cells, while the corresponding ligands were distributed in both immune-poor and immune-rich malignant cells, especially interactions such as LAGLS3-LAG3 and HLA-B-KLRD1 (Fig. 4G). Moreover, we identified interactions between malignant cells and CD4+ T cells. Interactions such as NECTIN4-TIGIT and MYL9-CD69 were distributed in almost all the interaction pairs of different types of malignant cells and CD4+ T cells, but they were more specific in CXCL13+ CD4+ T cells (Fig. S5H).
Transition of differentiation status in mixed adenoneuroendocrine carcinoma
Of these 12 patients we sequenced, 2 had MANEC, which was defined as a type of gastric cancer with the morphological and immunophenotypic characteristics of both classic adenocarcinoma and neuroendocrine carcinoma. Combining morphological and immune-histochemical analysis, we identified three prevalent components in these 2 MANEC tumors: differentiated adenocarcinoma, intermediate state cancer cells and neuroendocrine carcinoma. The intermediate state cancer cells referred to tumor components that were morphologically intermediate between differentiated adenocarcinoma and neuroendocrine carcinoma but did not express the well-known markers of either differentiated adenocarcinoma (MUC5AC and MUC6) or neuroendocrine carcinoma (Syn and CgA), although it can be regarded as differentiated adenocarcinoma in pathological examination (Fig. 5A).
Transition of differentiation status in mixed adenoneuroendocrine carcinoma. A Representative images of hematoxylin–eosin staining, and immune-histochemical staining of MUC6, MUC5AC, Syn and CgA in adenocarcinoma, intermediate state and neuroendocrine carcinoma (NEC) components from the same mixed adenoneuroendocrine carcinoma. Scale bar, 300 μm. B Pseudotime trajectory of all the epithelial cells from non-malignant epithelial cells, malignant cells from differentiated gastric adenocarcinoma (DGC) to NEC. C Heatmap showing genes that changed gradually along the trans-differentiation path of DGC to NEC. Part gene names are displayed. D Gene regulatory networks identified in malignant cells of NEC and DGC. Part TF names are displayed. E Heatmap showing the expressing scores of hallmark gene sets along the trans-differentiation path of DGC to NEC. F PCA plot showing the expression of CHGA and CTAG2 in non-malignant epithelial cells, and epithelial cells from DGC and NEC. G Kaplan–Meier survival curve showing the survival (days) of gastric cancer cases in the TCGA dataset with high and low expression levels of APOA1 and CGA
Since both patients had differentiated gastric adenocarcinoma with neuroendocrine carcinoma components, we combined single-cell transcriptome data from all differentiated gastric cancer, intermediate state and neuroendocrine carcinoma components to explore the origin of neuroendocrine components. DEGs and differentially expressed TFs between DGCs and NECs were analyzed in our dataset, and endocrine-related signatures and genes specific to NECs, such as CGA, CHGA, and CHGB, as well as NEC specific TFs, such as HMGN3, HOXD11 and FEV, were identified (Fig. S6D). The PCA reduction of all the epithelial cells showed a gradual change from differentiated gastric adenocarcinoma to an intermediate state and then to neuroendocrine carcinoma (Fig. S6A). Based on PCA components, we created the pseudotime path of all the epithelial cells from non-malignant epithelial cells, DGCs to NECs, with three poles of epithelial cells and different signatures identified (Fig. 5B). We identified genes that changed gradually along the trans-differentiation path. KRT20, PHGR1, PIGR and CLDN3 were gradually downregulated, while PDCD5, CPNE1, PDCD2, GLO1, APOA1 and CGA were upregulated (Fig. 5C and Fig. S6B), indicating that the differentiation score gradually decreased as the expression of the neuroendocrine markers increased. Differentiation-related TFs such as KLF2, KLF3 and FOXA3 were downregulated, and endocrine-related TFs such as HOXA7, FEV and LMX1A were upregulated along the trans-differentiation path (Fig. 5D). Especially, potential targets of immunotherapy including MUC1, CEACAM5 and CLDN18 were gradually downregulated along the trans-differentiation path. On the contrary, CTAG2, a cancer-testis antigen, was gradually upregulated toward NEC status (Fig. 5F). Moreover, the expression of APOA1 and CGA was associated with poorer prognosis in gastric cancer from the TCGA dataset, and CGA is a typical marker of NECs (Fig. 5G). We noticed that malignant cells of the intermediate state expressed neither the well-known DGC signature genes (MUC1, KRT20, PHGR1) nor the NEC signature genes (SCG3, CGA, CHGB), which was consistent with the aforementioned morphological and immune-histochemical staining findings. Using the hallmark gene set, we can see the changing signatures along the trans-differentiation path, and signatures such as unfolded protein response, fatty acid metabolism, glycolysis and MYC targets were specifically enriched in the intermediate state (Fig. 5E). Several genes including LGALS1, DDIT4 and EIF4G1 were also enriched in the intermediate state (Fig. S6C).
The negative correlation of interferon-related pathways was also found along the DGC to NEC trans-differentiation path (Fig. 5E and Fig. S6E, F). The decrease of immune responding signatures in malignant cells was accompanied by reduced CD8+ T cell infiltration in IHC staining (Fig. S6B). Analysis of specific genes in interferon-related pathways also confirmed a similar correlation, such as the expression of CASP1, CD74, B2M, IFITM3 and IFI27, which have negative correlations with NEC signatures (Fig. S6G).
Transcriptomic characteristics of myeloid cells in gastric cancer
Macrophages account for the main part of myeloid cells (9 identified clusters, with different marker genes such as FCN1, C1QA, CCL18, HSPA18, SPP1, CXCL10, INHBA, CDKN1C and MT1G), and other cell clusters were also identified, such as CD1C+ DCs, CCR7+ DCs, LTB+ DCs, FCGR3B+ neutrophils and cycling myeloid cells, according to the corresponding marker genes. Of them, FCN1+ macrophages and C1QA+ macrophages account for the largest proportion of myeloid cells (Fig. 6A, C). For macrophages, several gene sets, such as the M1 polarization gene set, M2 polarization gene set, pro-inflammatory gene set and anti-inflammatory gene set, were used to score macrophages, and all clusters of macrophages tended to exhibit M2 polarization signatures (Fig. 6B). We identified consistent positive correlation of the above signatures with metabolic activities, such as fatty acid biosynthesis and nicotinate-related metabolism, as well as diverse correlations with energy metabolism, such as the TCA cycle and oxidative phosphorylation (Fig. 6D).
Transcriptomic characteristics of myeloid cells in gastric cancer. A UMAP plot showing different clusters of myeloid cells in gastric cancer. B The expressing scores of M1 polarization gene set, M2 polarization gene set, pro-inflammatory gene set and anti-inflammatory gene set in myeloid cells. C The expression of corresponding markers for different myeloid cell clusters. D Correlation of metabolic activity-related gene sets with M1 polarization gene set, M2 polarization gene set, pro-inflammatory gene set and anti-inflammatory gene set in myeloid cells. E The percentage of different types of myeloid cells in immune-rich and immune-poor type gastric cancer, and differentiated gastric cancer (DGC), poorly differentiated gastric cancer (PDGC) and neuroendocrine carcinoma (NEC). F UMAP plot showing the distribution of myeloid cells across NEC, PDGCs and DGCs, immune-rich and immune-poor types. G Interactions between immune-poor/rich type epithelial cells and macrophages inferred from cell communication analysis using the toolkit NATMI
Moreover, we screened the distribution of myeloid cells in diverse classifications, such as DGCs and PDGCs, and immune-rich and immune-poor types. We observed that the proportion of INHBA+ macrophages increased in the immune-poor type compared with the immune-rich type and increased in NECs compared with DGCs, which was in accordance with the aforementioned findings and indicated that INHBA+ macrophages might influence gastric cancers by repressing the immune responses (Fig. 6E, F). Finally, we identified ligand–receptor interaction pairs of malignant cells and macrophages. For the immune-rich and immune-poor types of malignant cells, we identified the specific interaction TNFSF10-TNFRSF11B in the pairs of CDKN1C+ macrophages-malignant cells and CXCL10+ macrophages-malignant cells, which corresponds to the inhibitory function of TNFRSF11B in immune responses (Fig. 6G).
Discussion
Our pathology-informed sampling strategy provided exact pathological information on the tumor tissue we sequenced to minimize the influence of the high intra-tumoral heterogeneities, resulting in good discriminations between DGC and PDGC as well as between the immune-poor type and immune-rich type. Additionally, the CNV information inferred from RNA data could also be validated by WES data from corresponding tissue by laser capture microdissection [42]. Taking advantage of this and the single-cell sequencing technology [43], we could clearly distinguish malignant cells from non-malignant epithelial cells by combining clustering and CNV information and achieve a more accurate clustering of tumor cells. By comparing DGC and PDGC, we identified the EMT signature enriched in PDGC, which is consistent with the malignant type associated with the poor differentiation state. More interestingly, our differentiation signatures overlapped with immune infiltration-related genes screened by comparing the immune-rich type with the immune-poor type.
The most interesting finding of our study is the transition of differentiation status in mixed adenoneuroendocrine carcinoma. A pseudotime trajectory of all the malignant cells from DGCs to NECs was constructed, and genes that changed gradually along the trans-differentiation path were identified, indicating a convergence to a small-cell neuroendocrine state reported by Nikolas et al. [44]. MUC1, CEACAM5 and CLDN18 that are potential targets of immunotherapy were gradually downregulated, indicating the increasing evasion of immune attack during this neuroendocrine differentiation process. The malignant cells of intermediate state showed double-negative signatures for DGC and NEC marker genes, representing a possible “stem/progenitor cell” signature that might be associated with gastric cancer progression. Similar to studies in prostate cancer and non-small-cell lung cancer, many transcription factors were found to be differentially expressed in this trans-differentiation path [45, 46]. The new finding is the negative correlation of interferon-related pathways found along the DGC to NEC trans-differentiation path, which was also accompanied by reduced CD8+ T lymph cell infiltration in IHC staining, supporting neuroendocrine differentiation as a mechanism for immune evasion in gastric cancer, which needs further functional investigations in the future.
Availability of data and material
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive in National Genomics Data Center (National Genomics Data Center Members and Partners, 2020), Beijing Institute of Genomics (China National Center for Bioinformation), Chinese Academy of Sciences, under accession number HRA002108 that are publicly accessible at http://bigd.big.ac.cn/gsa-human.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J (2016) Cancer statistics in China, 2015. CA Cancer J Clin. https://doi.org/10.3322/caac.21338
Sentani K, Imai T, Kobayashi G, Hayashi T, Sasaki N, Oue N, Yasui W (2021) Histological diversity and molecular characteristics in gastric cancer: relation of cancer stem cell-related molecules and receptor tyrosine kinase molecules to mixed histological type and more histological patterns. Gastric Cancer 24:368–381
Tanaka H, Yoshii M, Imai T, Tamura T, Toyokawa T, Muguruma K, Hirakawa K, Ohira M (2021) Clinical significance of coexisting histological diffuse type in stage II/III gastric cancer. Mol Clin Oncol 15:234
Alsina M, Gullo I, Carneiro F (2017) Intratumoral heterogeneity in gastric cancer: a new challenge to face. Ann Oncol 28:912–913
Smyth EC, Nilsson M, Grabsch HI, van Grieken NC, Lordick F (2020) Gastric cancer. Lancet 396:635–648
Okagawa Y, Sumiyoshi T, Kondo H, Tomita Y, Uozumi T, Iida R, Sakano H, Tokuchi K, Jin T, Yoshida M, Sakurada A, Fujii R, Minagawa T, Morita K, Yane K, Ihara H, Hirayama M, Oyamada Y, Okushiba S (2021) Comparison of clinicopathological features and long-term prognosis between mixed predominantly differentiated-type and pure differentiated-type early gastric cancer. BMC Cancer 21:235
Lin J, Zhao Y, Zhou Y, Tian Y, He Q, Lin J, Hao H, Zou B, Jiang L, Zhao G, Lin W, Xu Y, Li Z, Xue F, Li S, Fu W, Li Y, Xu Z, Li Y, Chen J, Zhou X, Zhu Z, Cai L, Li E, Li H, Zheng C, Li P, Huang C, Xie J (2021) Comparison of survival and patterns of recurrence in gastric neuroendocrine carcinoma, mixed adenoneuroendocrine carcinoma, and adenocarcinoma. JAMA Netw Open 4:e2114180
Lin JX, Wang ZK, Hong QQ, Zhang P, Zhang ZZ, He L, Wang Q, Shang L, Wang LJ, Sun YF, Li ZX, Liu JJ, Ding FH, Lin ED, Fu YA, Lin SM, Xie JW, Li P, Zheng CH, Huang CM (2021) Assessment of clinicopathological characteristics and development of an individualized prognostic model for patients with hepatoid adenocarcinoma of the stomach. JAMA Netw Open 4:e2128217
Koh J, Nam SK, Kwak Y, Kim G, Kim KK, Lee BC, Ahn SH, Park DJ, Kim HH, Park KU, Kim WH, Lee HS (2021) Comprehensive genetic features of gastric mixed adenoneuroendocrine carcinomas and pure neuroendocrine carcinomas. J Pathol 253:94–105
Ishida S, Akita M, Fujikura K, Komatsu M, Sawada R, Matsumoto H, Saegusa J, Itoh T, Kakeji Y, Zen Y (2021) Neuroendocrine carcinoma and mixed neuroendocrine-non-neuroendocrine neoplasm of the stomach: a clinicopathological and exome sequencing study. Hum Pathol 110:1–10
Murakami T, Yao T, Mitomi H, Morimoto T, Ueyama H, Matsumoto K, Saito T, Osada T, Nagahara A, Watanabe S (2016) Clinicopathologic and immunohistochemical characteristics of gastric adenocarcinoma with enteroblastic differentiation: a study of 29 cases. Gastric Cancer 19:498–507
von Loga K, Woolston A, Punta M, Barber LJ, Griffiths B, Semiannikova M, Spain G, Challoner B, Fenwick K, Simon R, Marx A, Sauter G, Lise S, Matthews N, Gerlinger M (2020) Extreme intratumor heterogeneity and driver evolution in mismatch repair deficient gastro-oesophageal cancer. Nat Commun 11:139
Togasaki K, Sugimoto S, Ohta Y, Nanki K, Matano M, Takahashi S, Fujii M, Kanai T, Sato T (2021) Wnt signaling shapes the histologic variation in diffuse gastric cancer. Gastroenterology 160:823–830
Röcken C, Amallraja A, Halske C, Opasic L, Traulsen A, Behrens HM, Krüger S, Liu A, Haag J, Egberts JH, Rosenstiel P, Meißner T (2021) Multiscale heterogeneity in gastric adenocarcinoma evolution is an obstacle to precision medicine. Genome Med 13:177
Sundar R, Liu DH, Hutchins GG, Slaney HL, Silva AN, Oosting J, Hayden JD, Hewitt LC, Ng CC, Mangalvedhekar A, Ng SB, Tan IB, Tan P, Grabsch HI (2021) Spatial profiling of gastric cancer patient-matched primary and locoregional metastases reveals principles of tumor dissemination. Gut 70:1823–1832
Cristescu R, Lee J, Nebozhyn M, Kim KM, Ting JC, Wong SS, Liu J, Yue YG, Wang J, Yu K, Ye XS, Do IG, Liu S, Gong L, Fu J, Jin JG, Choi MG, Sohn TS, Lee JH, Bae JM, Kim ST, Park SH, Sohn I, Jung SH, Tan P, Chen R, Hardwick J, Kang WK, Ayers M, Hongyue D, Reinhard C, Loboda A, Kim S, Aggarwal A (2015) Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat Med 21:449–456
Cancer Genome Atlas Research Network (2014) Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513:202–209
Zhang P, Yang M, Zhang Y, Xiao S, Lai X, Tan A, Du S, Li S (2019) Dissecting the single-cell transcriptome network underlying gastric premalignant lesions and early gastric cancer. Cell Rep 27:1934–1947
Bockerstett KA, Lewis SA, Noto CN, Ford EL, Saenz JB, Jackson NM, Ahn TH, Mills JC, DiPaolo RJ (2020) Single-cell transcriptional analyses identify lineage-specific epithelial responses to inflammation and metaplastic development in the gastric corpus. Gastroenterology. https://doi.org/10.1053/j.gastro.2020.08.027
Wang R, Dang M, Harada K, Han G, Wang F, Pool PM, Zhao M, Tatlonghari G, Zhang S, Hao D, Lu Y, Zhao S, Badgwell BD, Blum MM, Shanbhag N, Estrella JS, Roy-Chowdhuri S, Abdelhakeem A, Wang Y, Peng G, Hanash S, Calin GA, Song X, Chu Y, Zhang J, Li M, Chen K, Lazar AJ, Futreal A, Song S, Ajani JA, Wang L (2021) Single-cell dissection of intratumoral heterogeneity and lineage diversity inmetastatic gastric adenocarcinoma. Nat Med 27:141–151
Kim J, Park C, Kim KH, Kim EH, Kim H, Woo JK, Seong JK, Nam KT, Lee YC, Cho SY (2022) Single-cell analysis of gastric pre-cancerous and cancer lesions reveals cell lineage diversity and intratumoral heterogeneity. NPJ Precis Oncol 6:9
Kumar V, Ramnarayanan K, Sundar R, Padmanabhan N, Srivastava S, Koiwa M, Yasuda T, Koh V, Huang KK, Tay ST, Ho S, Tan A, Ishimoto T, Kim G, Shabbir A, Chen Q, Zhang B, Xu S, Lam KP, Lum H, Teh M, Yong WP, So J, Tan P (2022) Single-cell atlas of lineage states, tumor microenvironment, and subtype-specific expression programs in gastric cancer. Cancer Discov 12:670–691
Zhang M, Hu S, Min M, Ni Y, Lu Z, Sun X, Wu J, Liu B, Ying X, Liu Y (2021) Dissecting transcriptional heterogeneity in primary gastric adenocarcinoma by single cell RNA sequencing. Gut 70:464–475
Jeong HY, Ham IH, Lee SH, Ryu D, Son SY, Han SU, Kim TM, Hur H (2021) Spatially distinct reprogramming of the tumor microenvironment based on tumor invasion in diffuse-type gastric cancers. Clin Cancer Res 27:6529–6542
Kwon M, An M, Klempner SJ, Lee H, Kim KM, Sa JK, Cho HJ, Hong JY, Lee T, Min YW, Kim TJ, Min BH, Park WY, Kang WK, Kim KT, Kim ST, Lee J (2021) Determinants of response and intrinsic resistance to PD-1 blockade in microsatellite instability-high gastric cancer. Cancer Discov 11:2168–2185
Li X, Sun Z, Peng G, Xiao Y, Guo J, Wu B, Li X, Zhou W, Li J, Li Z, Bai C, Zhao L, Han Q, Zhao RC, Wang X (2022) Single-cell RNA sequencing reveals a pro-invasive cancer-associated fibroblast subgroup associated with poor clinical outcomes in patients with gastric cancer. Theranostics 12:620–638
Kim R, An M, Lee H, Mehta A, Heo YJ, Kim KM, Lee SY, Moon J, Kim ST, Min BH, Kim TJ, Rha SY, Kang WK, Park WY, Klempner SJ, Lee J (2021) Early tumor-immune microenvironmental remodeling and response to frontline fluoropyrimidine and platinum chemotherapy in advanced gastric cancer. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-21-0888
Sathe A, Grimes SM, Lau BT, Chen J, Suarez C, Huang RJ, Poultsides G, Ji HP (2020) Single-cell genomic characterization reveals the cellular reprogramming of the gastric tumor microenvironment. Clin Cancer Res 26:2640–2653
Korsunsky I, Millard N, Fan J, Slowikowski K, Zhang F, Wei K, Baglaenko Y, Brenner M, Loh PR, Raychaudhuri S (2019) Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods 16:1289–1296
Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WR, Hao Y, Stoeckius M, Smibert P, Satija R (2019) Comprehensive integration of single-cell data. Cell 177:1888–1902
Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, Benner C, Chanda SK (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10:1523
Gao R, Bai S, Henderson YC, Lin Y, Schalck A, Yan Y, Kumar T, Hu M, Sei E, Davis A, Wang F, Shaitelman SF, Wang JR, Chen K, Moulder S, Lai SY, Navin NE (2021) Delineating copy number and clonal substructure in human tumors from single-cell transcriptomes. Nat Biotechnol 39:599–608
Aibar S, González-Blas CB, Moerman T, Huynh-Thu VA, Imrichova H, Hulselmans G, Rambow F, Marine JC, Geurts P, Aerts J, van den Oord J, Atak ZK, Wouters J, Aerts S (2017) SCENIC: single-cell regulatory network inference and clustering. Nat Methods 14:1083–1086
Hänzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14:7
Wu Y, Yang S, Ma J, Chen Z, Song G, Rao D, Cheng Y, Huang S, Liu Y, Jiang S, Liu J, Huang X, Wang X, Qiu S, Xu J, Xi R, Bai F, Zhou J, Fan J, Zhang X, Gao Q (2022) Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discov 12:134–153
Qiu X, Hill A, Packer J, Lin D, Ma YA, Trapnell C (2017) Single-cell mRNA quantification and differential analysis with Census. Nat Methods 14:309–315
Hou R, Denisenko E, Ong HT, Ramilowski JA, Forrest A (2020) Predicting cell-to-cell communication networks using NATMI. Nat Commun 11:5011
Park S, Ock CY, Kim H, Pereira S, Park S, Ma M, Choi S, Kim S, Shin S, Aum BJ, Paeng K, Yoo D, Cha H, Park S, Suh KJ, Jung HA, Kim SH, Kim YJ, Sun JM, Chung JH, Ahn JS, Ahn MJ, Lee JS, Park K, Song SY, Bang YJ, Choi YL, Mok TS, Lee SH (2022) Artificial intelligence-powered spatial analysis of tumor-infiltrating lymphocytes as complementary biomarker for immune checkpoint inhibition in non-small-cell lung cancer. J Clin Oncol. https://doi.org/10.1200/JCO.21.02010
Lin R, Zhang H, Yuan Y, He Q, Zhou J, Li S, Sun Y, Li DY, Qiu HB, Wang W, Zhuang Z, Chen B, Huang Y, Liu C, Wang Y, Cai S, Ke Z, He W (2020) Fatty acid oxidation controls CD8(+) tissue-resident memory T-cell survival in gastric adenocarcinoma. Cancer Immunol Res 8:479–492
Zhang Y, Chen H, Mo H, Hu X, Gao R, Zhao Y, Liu B, Niu L, Sun X, Yu X, Wang Y, Chang Q, Gong T, Guan X, Hu T, Qian T, Xu B, Ma F, Zhang Z, Liu Z (2021) Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer. Cancer Cell 39:1578–1593
Moore L, Leongamornlert D, Coorens T, Sanders MA, Ellis P, Dentro SC, Dawson KJ, Butler T, Rahbari R, Mitchell TJ, Maura F, Nangalia J, Tarpey PS, Brunner SF, Lee-Six H, Hooks Y, Moody S, Mahbubani KT, Jimenez-Linan M, Brosens JJ, Iacobuzio-Donahue CA, Martincorena I, Saeb-Parsy K, Campbell PJ, Stratton MR (2020) The mutational landscape of normal human endometrial epithelium. Nature 580:640–646
Wen L, Tang F (2022) Recent advances in single-cell sequencing technologies. Prec Clin Med 5:pbac002
Balanis NG, Sheu KM, Esedebe FN, Patel SJ, Smith BA, Park JW, Alhani S, Gomperts BN, Huang J, Witte ON, Graeber TG (2019) Pan-cancer convergence to a small-cell neuroendocrine phenotype that shares susceptibilities with hematological malignancies. Cancer Cell 36:17–34
Quintanal-Villalonga A, Taniguchi H, Zhan YA, Hasan MM, Chavan SS, Meng F, Uddin F, Manoj P, Donoghue M, Won HH, Chan JM, Ciampricotti M, Chow A, Offin M, Chang JC, Ray-Kirton J, Tischfield SE, Egger JV, Bhanot UK, Linkov I, Asher M, Sinha S, Silber J, Iacobuzio-Donahue CA, Roehrl MH, Hollmann TJ, Yu HA, Qiu J, de Stanchina E, Baine MK, Rekhtman N, Poirier JT, Loomis B, Koche RP, Rudin CM, Sen T (2021) Multi-omic analysis of lung tumors defines pathways activated in neuroendocrine transformation. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-20-1863
Aggarwal R, Huang J, Alumkal JJ, Zhang L, Feng FY, Thomas GV, Weinstein AS, Friedl V, Zhang C, Witte ON, Lloyd P, Gleave M, Evans CP, Youngren J, Beer TM, Rettig M, Wong CK, True L, Foye A, Playdle D, Ryan CJ, Lara P, Chi KN, Uzunangelov V, Sokolov A, Newton Y, Beltran H, Demichelis F, Rubin MA, Stuart JM, Small EJ (2018) Clinical and genomic characterization of treatment-emergent small-cell neuroendocrine prostate cancer: a multi-institutional prospective study. J Clin Oncol 36:2492–2503
Acknowledgements
We are thankful for the support from the Beijing Advanced Innovation Center for Genomics at Peking University and from the Computing Platform of the Center for Life Science for data analysis.
Funding
This project was supported by grants from Beijing Advanced Innovation Center for Genomics (ICG) and National multidisciplinary cooperative diagnosis and treatment capacity building project for major diseases. Wei Fu was supported by grants from the National Natural Science Foundation of China (82273104). Xin Zhou was supported by Beijing Nova Program (2022029).
Author information
Authors and Affiliations
Contributions
XZ: conceptualization, investigation, methodology, writing-original draft, writing-review and editing. JY: data curation, formal analysis, methodology, writing-original draft, writing-review and editing. YL: investigation, validation, methodology, writing-original draft. YM: investigation, methodology. YM: investigation, methodology. QL: investigation, methodology. JG: investigation, methodology. ZJ: investigation, methodology. LG: investigation, methodology. WW: investigation, methodology. YL: data curation, methodology. LW: investigation, methodology. WF: conceptualization, resources, supervision, funding acquisition, writing-review and editing. FT: conceptualization, resources, supervision, funding acquisition, project administration, writing-review and editing.
Corresponding authors
Ethics declarations
Conflict of interest
All the authors declare no potential conflicts of interest.
Ethics approval and consent to participate
Our study was approved by the Ethics Committee of Peking University Third Hospital (Licence No. IRB00006761-M2016170) and included 12 patients with gastric cancer who underwent surgery between August 2020 and December 2020 at the Department of General Surgery, Peking University Third Hospital. All patients signed informed consent forms before participating in this study.
Consent for publication
All the authors give the consent for the publication of identifiable details, which can include the text, figures and other materials in this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhou, X., Yang, J., Lu, Y. et al. Relationships of tumor differentiation and immune infiltration in gastric cancers revealed by single-cell RNA-seq analyses. Cell. Mol. Life Sci. 80, 57 (2023). https://doi.org/10.1007/s00018-023-04702-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-023-04702-1