Abstract
Animal models have been utilized to understand the pathogenesis of Zellweger spectrum disorders (ZSDs); however, the link between clinical manifestations and molecular pathways has not yet been clearly established. We generated peroxin 5 homozygous mutant zebrafish (pex5−/−) to gain insight into the molecular pathogenesis of peroxisome dysfunction. pex5−/− display hallmarks of ZSD in humans and die within one month after birth. Fasting rapidly depletes lipids and glycogen in pex5−/− livers and expedites their mortality. Mechanistically, deregulated mitochondria and mechanistic target of rapamycin (mTOR) signaling act together to induce metabolic alterations that deplete hepatic nutrients and accumulate damaged mitochondria. Accordingly, chemical interventions blocking either the mitochondrial function or mTOR complex 1 (mTORC1) or a combination of both improve the metabolic imbalance shown in the fasted pex5−/− livers and extend the survival of animals. In addition, the suppression of oxidative stress by N-acetyl L-cysteine (NAC) treatment rescued the apoptotic cell death and early mortality observed in pex5−/−. Furthermore, an autophagy activator effectively ameliorated the early mortality of fasted pex5−/−. These results suggest that fasting may be detrimental to patients with peroxisome dysfunction, and that modulating the mitochondria, mTORC1, autophagy activities, or oxidative stress may provide a therapeutic option to alleviate the symptoms of peroxisomal diseases associated with metabolic dysfunction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Peroxisomes are highly dynamic and perform the breakdown of fatty acids, amino acids, and polyamines, biosynthesis of ether phospholipids, and scavenging of reactive oxygen species [1,2,3]. The importance of peroxisomal function in human health has been reported in numerous diseases, including peroxisomal biogenesis disorders, peroxisomal transporter or enzyme deficiencies, related metabolic disorders, aging-associated diseases, and cancers [4,5,6,7,8]. Functional peroxisomes require an array of peroxin (PEX) proteins, most of which reside in the peroxisome membrane, while few recycle between the cytosol and peroxisome membrane for the import of matrix proteins [1].
PEX5 acts as a major importing factor for peroxisome matrix proteins containing the C-terminal tripeptide (SKL) peroxisomal targeting signal, which is detected in the cytosol and transported to peroxisomes [9]. Accordingly, mutations in human PEX5 cause Zellweger spectrum disorders (ZSDs) in which functional peroxisomes are largely absent [10]. Patients with Zellweger syndrome, the most severe form of ZSD, display hypotonia, craniofacial defects, neuronal hypomyelination, and hepatic and renal dysfunction and die within their first year of life [5]. Mice with homozygous Pex5 mutations (Pex5−/− mice) recapitulate the phenotype of Zellweger syndrome in humans [11, 12]. Because of the early postnatal death of Pex5−/− mice, tissue-specific Pex5−/− mice were generated and analyzed to further identify the molecular pathways leading to ZSD-like phenotypic presentation [11, 13]. Given that the liver is one of the most peroxisome-enriched organs [7, 14,15,16], liver-specific Pex5−/− mice were found to be metabolically unsound, displaying a skewed nutrient preference toward carbohydrates with altered mitochondrial morphology and activity [11, 15, 17,18,19]. Similar to Pex5 deficiency, deficient models of other Pex genes, such as Pex2, Pex13, Pex16, and Pex19, also show mitochondrial abnormalities [20,21,22,23]. These include reduced mitochondrial activity, increased reactive oxygen species (ROS), and structural impairment of the mitochondria, suggesting the presence of a fundamental mechanism dependent on peroxisome function that regulates mitochondrial homeostasis. Given the cooperative nature of peroxisomes and mitochondria in maintaining cellular homeostasis, peroxisome deficiency is understood to impact mitochondrial function, although how this is achieved remains unclear.
A recent report suggested that stress conditions, such as increased ROS, often found in metabolic disorders, could assemble tuberous sclerosis complex at the peroxisome, which represses the mechanistic target of rapamycin complex 1 (mTORC1) in cultured cells [24]. The same regulatory axis has also been suggested in cultured cells during starvation stress [25]. However, the stress-responsive regulatory axis involving peroxisomes is yet to be validated in vivo. For the discovery of peroxisome functions and participating players, various animal model systems, including yeast, flies, and mice, have been used by providing their unique advantage and, at the same time, by complementing the inherent drawbacks of each model system. The zebrafish is an established model for uncovering various aspects of biological processes and has recently been used in the field of peroxisome biology and associated human diseases. A pioneering study reported that peroxisomes in zebrafish are distributed in tissues in a manner similar to those in mammals [26]. More recently, we and other researchers have reported that zebrafish could serve as a model system for studying the metabolic role of peroxisome-resident proteins and the effects of their functional deficits using either the morpholino-mediated transient knockdown approach or transcription activator-like effector nucleases [27, 28]. Thus, zebrafish may bridge the gap between fly and mouse models in terms of evolutionary complexity, offering an opportunity to link functional interpretations obtained through genetic and developmental studies.
In this study, we aimed to characterize the role of peroxisomes in mitochondrial health and stress-induced regulatory responses. We generated a stable zebrafish model of ZSD by targeting the pex5 gene. Zebrafish with homozygous pex5 mutations (pex5−/−) share critical features of ZSD in humans. pex5−/− zebrafish displayed significantly reduced amounts of peroxisomes and abnormal mitochondria at the cellular level, metabolic disturbances in the liver, and defective myelination in the nervous system at the tissue level. Notably, when fasting was imposed as an environmental stress, pex5−/− rapidly developed phenotypic severity characterized by edema, deflated swim bladder, and decreased movement that could be easily detected through simple microscopy. Mechanistic studies have revealed a positive correlation between deregulated mitochondrial function in response to peroxisome dysfunction and unrepressed mTORC1 upon fasting. Chemical interventions, either mitochondrial function or mTORC1 signaling, rescued disturbed liver metabolism in fasted pex5−/−zebrafish. A similar rescue was observed with a ROS scavenger or an autophagy inducer. The identified pathway is suggested to be a druggable target for the treatment of symptoms in patients with ZSD.
Materials and methods
Fish maintenance, transgenic zebrafish, and generation of pex5 −/− zebrafish
All zebrafish lines were maintained at 28.5 °C. Embryos were collected and developed in E3 embryonic medium and staged according to age and morphology based on a standard protocol [29]. pex5−/− zebrafish were generated using the CRISPR/Cas9 system and genotyped as described in detail in Supplementary Methods. Transgenic zebrafish lines Tg(mbp:EGFP) [30], Tg(Xla.Eef1a1:RFP-SKL) [31], and Tg(Xla.Eef1a1:mlsEGFP) [32] have been previously described. mbp:EGFP transgenic line was crossed with pex5 heterozygotes to obtain the mbp:EGFP transgenic line in a pex5−/− background. Similarly, Xla.Eef1a1:RFP-SKL and Xla.Eef1a1:mlsEGFP transgenic lines were crossed with pex5 heterozygotes separately to obtain Xla.Eef1a1:RFP-SKL and Xla.Eef1a1:mlsEGFP transgenic lines on pex5−/− background, respectively.
Microscopy
For whole-mount images of zebrafish larvae, the larvae were anesthetized using 1X tricaine and positioned using a 1.5% agarose plate. Images were taken using a Leica M165FC microscope equipped with Leica DFC500 (Leica, Wetzlar, Germany). Images of cryo-sectioned Oil Red-O (ORO)-stained, periodic acid-Schiff (PAS)-stained, and tunnel samples were taken using an Olympus IX71 fluorescence microscope (Olympus, Tokyo, Japan). Other fluorescent images after antibody staining or from transgenic zebrafish were captured using a Fluoview FV1000 confocal microscope (Olympus, Tokyo, Japan) or a ZEISS LSM 980 (Carl Zeiss AG, Oberkochen, Germany) at the Core Facility for Supporting Analysis & Imaging of Biomedical Materials at Wonkwang University, supported by the National Research Facilities and Equipment Center.
Survival curve
Heterozygous pex5+/− zebrafish were intercrossed and the resulting embryos were reared to the juvenile stage. From 15 days post-fertilization (dpf), dead larvae were collected and genotyped, and the ratio of each genotype was calculated assuming Mendelian segregation ratios. The surviving juveniles were fin-clipped, and their genotypes were determined. Three crosses were used to determine the survival rate (n = 300).
Cell/tissue staining
Immunofluorescence staining was performed as described in Supplementary Methods. ORO staining was performed as previously described [33]. In brief, larvae were fixed in 4% paraformaldehyde (PFA) overnight at 4 °C, washed with phosphate-buffered saline (PBS) three times, and infiltrated with a graded series of propylene glycol (PG) (i.e., 25% PG + 75% PBS, 50% PG + 50% PBS, 75% PG + 25% PBS, 100% PG) for 10 min. The larvae were then incubated with 0.5% freshly prepared ORO solution in 100% PG overnight at room temperature on a shaker and rinsed several times with PBS. Stained larvae were processed for cryo-sectioning to obtain 50-µM-thick section. Stained images of the liver tissue sections were captured using a Leica M165FC microscope equipped with a Leica DFC500 microscope. PAS staining was performed as previously described [33]. Larvae were fixed in 4% PFA overnight at 4 °C, washed with PBST three times, and permeabilized in 40 µg/mL proteinase K for 30 min at room temperature. Samples were then washed thrice in PBT, rinsed in demineralized water (DW), and oxidized in 0.5% periodic acid for 30 min at room temperature. After washing with DW three times for 20 min each, the larvae were stained with Schiff reagent for 5 min, followed by immediate rinsing in tap water. Larvae were then processed for cryo-sectioning to obtain a 50-µm-thick liver section. Liver tissues were mounted using coverslips and mounting solution. Images were acquired using an IX71 microscope (Olympus).
Quantification of mitochondrial DNA and mitochondrial number
Wild-type (WT) and pex5−/− livers were dissected from 6-day-old larvae using forceps and processed for genomic DNA extraction using the TIANamp Genomic DNA Kit (TIANGEN, Beijing, China) according to the manufacturer’s instructions. The remaining larvae were used for genotyping. For quantification of mitochondrial DNA, 15 ng of genomic DNA from each sample was amplified via polymerase chain reaction (PCR) using Fast Start Essential DNA Green Master (Roche, Basel, Switzerland), and the ratio of mitochondrial DNA to nuclear DNA was calculated. Primers used to amplify mitochondrial DNA and nuclear DNA were Mt-ND1 (F:5ʹ-CCCCAGATGCACCTGAGCTAATAAC-3ʹ and R:5ʹ- TGTTTGTGGGGGTAGACCAGCTAG-3ʹ) and β-actin (F:5ʹ-ACCTCATGAAGATCCTGACC-3ʹ and R:5ʹ-TGCTAATCCACATCTGCTGG-3ʹ). The data were analyzed to determine the relative abundance of β-actin. Mitochondrial numbers were manually counted. Three image fields were obtained for both WT and mutant fish using a confocal microscope (Olympus, Tokyo, Japan). All analyses were performed in triplicates.
3ʹUTR RNA-seq and data processing
Total RNA from fasted WT and pex5−/− livers at 6 dpf was extracted using the Direct-zol RNA Kit (ZYMO RESEARCH, Irvine, CA, USA), according to the manufacturer’s instructions. 3'UTR RNA-seq was performed using the CEL-Seq2 protocol [34], except that the Second Strand Synthesis Module (NEB, Ipswich, MA, USA) was used for double-stranded cDNA synthesis, and the library was amplified by nine cycles of PCR without any sample pooling. The library was sequenced on an Illumina MiniSeq, and the following quantitative analysis was performed using 51 bp of insert reads (Read2). Cell barcode and UMI in Read1 was extracted by using UMI-tools (ver.1.1.1) with the following command “umi_tools extract -I read1.fastq --read2-in = read2.fastq --bc-pattern = =NNNNNNNNNNCCCCCCCCC.” Four cell barcodes were mixed and used for all the samples. Adaptor sequences and low-quality sequences were removed, read length below 20 bp was discarded using Trim Galore (ver.0.6.7), and reads were mapped to the GRCz11 reference using HISAT2 (ver.2.2.1). Read counts for each gene were obtained using featureCounts (ver.2.0.1), and UMI duplications were removed using UMI-tools. To further analyze and visualize the gene expression profiles, the processed count was transformed into a transcript per million (TMM) using the edgeR package (ver.3.36.0) as described previously [35]. R Studio (ver.2022.02.2) and R (ver.4.1.2) were used to create scatter plots showing gene expression profiles (log2[TMM + 1]) and (2) differential gene expression (log2[fold change]). R included the R packages dplyr (ver.1.0.9), stringr (ver.1.4.0), ggpubr (ver.0.4.0), ggplot2 (ver.3.3.6), and pheatmap (ver.1.0.12).
mRNA expression
WT and pex5−/− livers were dissected from six- or 7-day-old larvae. Each sample of total RNA was prepared from ten livers at the desired developmental stages using TRIzol reagent (Ambion Inc., Austin, TX, USA), following the manufacturer’s instructions. Total RNA was reverse-transcribed using reverse transcriptase (Roche, Basel, Switzerland), and quantitative PCR (qPCR) was performed using the following primers: p62 F:5ʹ-GCGTCAGTGAGGGAACAAAG-3ʹ, R:5ʹ-CAGAGACTCCACCAGCCTAG-3ʹ; and β-actin F:5ʹ-TGAATCCCAAAGCCAACAGAGAGA-3ʹ, R:5ʹ-TCACGACCAGCTAGATCCAGACG-3ʹ. The data were analyzed to determine the relative abundance of β-actin. All analyses were performed in triplicate.
Transmission electron microscopy (TEM)
WT and pex5−/− livers were dissected from 4.5- and 6.5-day-old embryos. Dissected zebrafish livers were fixed with 2% paraformaldehyde and 2% glutaraldehyde in 0.05 M sodium cacodylate buffer (pH 7.2) overnight at 4 °C. Fixed samples were washed three times with 0.05 M sodium cacodylate buffer for 10 min at 4 °C and post-fixed with 1% osmium tetroxide in 0.05 M sodium cacodylate buffer. The post-fixed samples were washed twice with distilled water for 10 min at room temperature. The samples were then stained in en bloc for overnight at 4 °C. The samples were then dehydrated in a graded ethanol series, followed by propylene oxide, and embedded in Quetol-Spurr resin. Sections (70 nm thick) were cut on a Leica Ultracut ultramicrotome (Leica Microsystem, EM UC7, Canada), stained with 2% uranyl acetate and lead citrate, and viewed under a Hitachi H-7650 TEM (Hitachi Science and Technology, Japan).
Cell death assay
Larvae were processed for cryo-sectioning to obtain a 10-µm-thick liver section, dried for 1 h at room temperature, and washed with PBS three times for 10 min. The samples were incubated in blocking solution (3% hydrogen peroxide (H2O2) in methanol) for 10 min at room temperature, followed by washing with PBS. Permeabilization solution (10 µL 1 M Tris–HCl pH 7.5 + 2 µL proteinase K PCR Grade 20 mg/mL and H2O to make 1 mL) was then added to the tissue samples, incubated for 2 min at 4 °C, and the slides were rinsed with PBS. As the positive control, fixed and permeabilized samples were incubated with DNase (20 U/mL DNase in 50 mM Tris–HCl pH 7.5, 10 mM magnesium chloride (MgCl2), and 1 mg/mL bovine serum albumin) for 10 min at room temperature, followed by washing with PBS twice. The area around the tissue was then dried, and 50 µL terminal deoxyribonucleotidyl transferase (TDT)-mediated dUTP nick-end labeling (TUNEL) reaction mixture (50 µL enzyme solution + 450 µL label solution, Roche, Basel, Switzerland) was added to the tissue sample, followed by incubation for 60 min at 37 °C in a humidified atmosphere in the dark. The slides were washed three times in PBS, and 4ʹ, 6-diamidino-2-phenylindole (DAPI, Roche 10236276001, Basel, Switzerland) (1:1000 in PBS) was added to the tissue samples and incubated for 10 min at room temperature in the dark. Samples were then washed three times in PBS, mounted with a clear mount, and imaged under an Olympus IX71 fluorescence microscope.
Drug treatment
For rescue experiments, rapamycin (250 nM) or UK5099 (25 µM) was treated in embryo medium starting from 3.5 dpf, and etomoxir (1.5 µM), oligomycin (5 pM), chloroquine (250 μM), or N-acetyl-cysteine (NAC) (100 μM) was treated from 5 dpf. The medium containing the drug was changed daily until the analysis.
Immunoblotting
After genotyping, WT and pex5−/− at 6 dpf were separately pooled in a 1.5 ml Eppendorf tube and sonicated on ice using 1 mL PBS to break embryos into pieces. Samples were centrifuged at 12,000 rpm for 5 min, and lysis buffer containing protease inhibitor cocktail was added to the pellets and incubated on ice for 1 h. Samples were then centrifuged at 14,000 rpm at 4 °C, and the supernatant was mixed with sodium dodecyl sulfate (SDS) loading dye and heat shock at 95 °C for 5 min. Samples were then loaded on a sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) gel and run in running buffer for 2 h at 100 V. The proteins were transferred to a nitrocellulose membrane using a transfer buffer for 1.5 h at 40 V. The membrane was then blocked for 1 h at room temperature using blocking buffer (5% skim milk in 1X TBST). The membrane was then incubated with primary antibody diluted in blocking buffer (2.5% skim milk in TBST) overnight at 4 °C and washed thrice with TBST for 10 min. Secondary antibodies diluted appropriately in blocking buffer were then added to the membrane (2.5% skim milk in TBST) and incubated at room temperature for 1 h, followed by three washes with TBST for 10 min each. All washes and antibody incubations were performed using a rocker. For signal development, all buffers were removed and the membrane was incubated for 3 min in a western blot detection kit (AbFrontier, Seoul, South Korea) and developed in a dark room using an X-ray film, developer, and fixer. After the signal development, the X-ray film was washed in running water for a few minutes, and images were captured.
Antibodies
Antibodies used for western blotting or immunostaining in this study are listed in Supplementary Methods.
Statistics
Statistical significance was determined using the Student’s t test in Microsoft Excel; * and ** indicate p values < 0.05 and < 0.01, respectively.
Data availability
The RNA-seq dataset used in this study was deposited in the Gene Expression Omnibus and is available under accession number GSE205349.
Study approval
The experimental protocol was approved by the Committee for Ethics in Animal Experiments of Wonkwang University (WKU15-126) and was performed in accordance with the guidelines for animal experiments.
Results
pex5 −/− zebrafish recapitulates clinical characteristics of ZSD
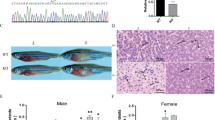
To establish a stable zebrafish model for ZSD, we initially examined the equivalence of the zebrafish pex5 gene with its mammalian counterpart. Sequence analysis suggested that PEX5 is highly conserved among vertebrates including humans, mice, and zebrafish (Supplementary Fig. S1). To address the functional equivalence of PEX5 genes, we generated pex5 mutant zebrafish using the CRISPR-Cas9 system, which targeted an optimal genomic sequence near intron/exon junction 2 (Fig. 1a). A founder zebrafish harboring an 8-base pair deletion in the pex5 open reading frame was identified and used to produce pex5 F1 heterozygous zebrafish, whose pex5 gene was predicted to generate a truncated version of the protein comprising only 41 amino acids (Fig. 1a). Almost all pex5−/− generated from the incrosses of pex5 heterozygotes died before one month after fertilization (Fig. 1b).
Loss of Pex5 in zebrafish induces liver defects and myelination abnormality due to the decreased peroxisome number. a Schematic view of the genomic structure of pex5 in zebrafish is shown with 16 exons in black rectangles. F1 and R1 indicate the position and direction of the primer sequences that were used for the clustered regularly interspaced palindromic repeat/caspase 9 (CRISPR/Cas9)-mediated gene inactivation. A partial amino-terminal sequence of WT Pex5 is compared to that of Pex5 mutant protein. Note that the 8-base pair deletion shown in the sequencing chromatogram of the pex5 gene is predicted to translate a truncated version of the Pex5 mutant protein comprised only 41 amino acids with a stretch of unrelated carboxy-terminal sequences. b Survival curve for pex5−/− under normal feeding conditions. Heterozygous pex5+/− zebrafish were incrossed, and the percentage of survived larvae per genotype at the indicated developmental times was plotted with assumption of Mendel’s segregation ratios. c Peroxisomes are shown in the liver sections of WT and pex5−/−. The RFP-SKL expression from the Xla.Eef1a1:RFP-SKL transgene was used as a surrogate to detect the subcellular localization of the peroxisomal matrix proteins. The expression levels of endogenous PMP70 proteins detected by immunofluorescence were shown to illustrate an abundance of peroxisome membrane proteins that are independently incorporated in the peroxisome membrane from the matrix-import pathway. Scale bar = 5 μm. Graph shows quantified signal intensity of Abcd3 and presented as the average with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; * and ** indicate p values < 0.05 and < 0.01, respectively. d Mitochondria are shown in the liver sections of WT and pex5−/−. Expression of mitochondria-targeted enhanced green fluorescent protein (EGFP) (MLS-EGFP) was used to detect the general shape of mitochondria, whereas immunofluorescence of a component of the mitochondrial respiratory chain complex IV (CoxIV) was performed to reveal the differential expression of an endogenous mitochondria-targeted protein. Scale bar = 5 μm. Graph shows quantified signal intensity of CoxIV and presented as the average with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; * and ** indicate p values < 0.05 and < 0.01, respectively. e Neutral lipids and glycogen are shown by performing Oil Red O and PAS staining, respectively, in the liver sections of WT and pex5−/−. Scale bar = 50 μm. f Neuronal myelination using Tg(mbp:EGFP) transgenic zebrafish was compared between WT and pex5−/−. Red immunofluorescence for acetylated tubulin was used to locate the differentiating neurons in the zebrafish trunk. Arrows indicate myelinated neurons. Scale bar = 50 μm. g Representative pictures of the WT and pex5−/− at 8 and 20 dpf as indicated are shown in lateral views with anterior to the left. Scale bar = 1 mm
To address the mechanistic aspects of pex5−/− mortality, we initially examined peroxisomes that were predicted to be dysfunctional in zebrafish pex5−/−, as observed in human patients or animal models with PEX5 deficiency [36, 37]. Expression of RFP-SKL (a cargo protein of Pex5 with red fluorescence) from the Xla.Eef1a1:RFP-SKL transgene [27] showed subcellular distribution of peroxisomes in a punctate pattern in the hepatocytes of WT zebrafish at 20 days post-fertilization (dpf) (Fig. 1c, n = 6). In contrast, RFP-SKL was diffusely detected in the cytoplasm of pex5−/− hepatocytes, which is consistent with the Pex5 deficiency in the import of peroxisome matrix proteins in mammals. Moreover, a decreased number of punctate structures expressing Abcd3 (n = 6), an endogenous peroxisome membrane protein localized in a Pex5-independent manner, indicated a decreased number of peroxisomes at 20 dpf in the pex5−/− livers.
We next examined mitochondria, whose abnormalities have been reported in ZSD [11, 17, 18, 36]. Mitochondrial morphologies in the hepatocytes of WT zebrafish, assessed by the expression of green fluorescent protein fused to mitochondria-localizing signal (MLS-EGFP) from Tg(Xla.Eef1a1:MLS-EGFP) transgenic zebrafish [32], differed from those in pex5−/− whose mitochondria were heterogeneous in size and occasionally clustered (Fig. 1d, n = 6). In addition, the expression of endogenous proteins belonging to the mitochondrial respiratory chain complex IV was highly increased in the pex5−/− hepatocytes compared to that in WT (n = 6), suggesting that mitochondrial abnormalities might have initiated at the subcellular levels earlier than 20 dpf.
ZSD features metabolic disturbances in the liver, the most peroxisome-rich organ [18, 36]. Indeed, we found that liver lipids were highly accumulated in pex5−/− compared to those in WT, while glycogen levels between the two genotypes were comparable (Fig. 1e, n = 6). ZSD also features defects in the nervous system, including demyelination [38, 39]. Using transgenic zebrafish, Tg(mbp:EGFP), we found that pex5−/− zebrafish at 20 dpf displayed severe demyelination (Fig. 1f, n = 6). Despite the metabolic alterations in the liver and neuronal demyelination, the resulting pex5−/− did not show noticeable differences in gross morphology from WT siblings (Fig. 1g, n > 40), but died before one month of age (Fig. 1b). Given that the molecular and cellular phenotypes and early mortality displayed in pex5−/− resembled those in ZSD, this model could be used as another system to identify the molecular pathways underlying ZSD pathogenesis.
pex5 −/− zebrafish are sensitized to fasting leading to expedited death during the larval stage
Although pex5−/− zebrafish could survive up to one month of age, pex5 deficiency might have altered organelle structures and, thus, their functions as early as 20 dpf. To trace the stage in which peroxisomes became dysfunctional in pex5−/− zebrafish, we examined the expression of both RFP-SKL and Abcd3 during early embryonic development in zebrafish. Owing to the maternal contribution of peroxisomes and pex5 gene products, RFP-SKL was still detected in a punctate structure in pex5−/− at 2 dpf. However, as peroxisome turnover is predicted to be approximately two days on average [40], RFP-SKL was diffusely detected in the cytosol of the hepatocytes of pex5−/− zebrafish at 5 dpf (Fig. 2a, n = 8). Notably, the number of peroxisomes detected by Abcd3 antibody staining was similar in WT and pex5−/− zebrafish at 5 dpf (Fig. 2b, n = 8). These results suggest that peroxisomes in pex5−/− might be immature as early as 5 dpf as opposed to those in WT and become reduced in number before 20 dpf.
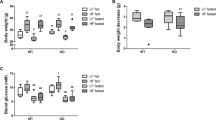
Fasting induces the rapid exacerbation of pex5−/− leading to early mortality. a RFP-SKL was detected using the Xla.Eef1a1:RFP-SKL transgenic zebrafish at 2 dpf in the muscle cells and at 5 dpf in the liver sections of WT and pex5−/−. Note that the SKL signal is diffusely detected in the liver cells of pex5−/− at 5 dpf. Scale bar = 5 μm. b Endogenous Abcd3 proteins were shown by performing immunofluorescence at 2 dpf in the muscle cells and at 5 dpf in the liver sections of WT and pex5−/−. Scale bar = 5 μm. Graphs show quantified signal intensity of Abcd3 at the indicated times and presented as the average with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel. c Survival curve shows a differential survival response between the WT and pex5−/− zebrafish under fasting conditions. Experiments were performed at least three times and the average percentage of surviving animals was used to draw the curve (n = 40/genotype). Representative pictures of the WT and pex5−/− fasted from 5 to 8 dpf are shown in lateral views with anterior to the left. Arrows point to the edematous gut area and deflated swim bladder in fasted pex5−/− at 8 dpf. Scale bar = 1 mm. d Representative pictures of WT and pex5−/− fasted from 11 to 13 dpf are shown as indicated. Scale bar = 1 mm. e, f Peroxisome abundance was compared in the liver cells between the WT and pex5−/− and between the feeding and fasting conditions using immunofluorescence for Abcd3. There was a marked disappearance of peroxisome abundance depending on the pex5 genotype and the nutritional status. Scale bar = 5 μm. Graphs show quantified signal intensity of Abcd3 under fed or fasted conditions at the indicated times and presented as the average with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; ** indicates p values < 0.01
Zebrafish embryos consume yolk for the initial five days of development, but depend on the external food supply for their development and growth [41, 42]. We reasoned that metabolic challenges might alter the phenotypic presentation of pex5−/−. We found that WT zebrafish subjected to fasting could survive up to 12 dpf, with 50% mortality observed at approximately 10.5 dpf (Fig. 2c, n = 40). In contrast, pex5−/− zebrafish under the same condition displayed clearly observable phenotypes including edematous abdomen with deflated swim bladder, shrunken liver, and hypotonia at 8 dpf and died between 8 and 10 dpf (with 50% mortality observed at approximately 8.5 dpf, n = 40), suggesting a high sensitivity to fasting. The causal relationship between fasting and early mortality of pex5−/− was confirmed by performing another fasting experiment in which pex5−/− subjected to fasting from 11–13 dpf recapitulated the phenotype and early mortality observed in pex5−/− subjected to fasting from 6 dpf (Fig. 2d, n = 20).
We next examined the effect of fasting on peroxisomal abundance. In contrast to the comparable numbers of immature peroxisomes present in both WT and pex5−/− livers at 5 dpf (Fig. 2b), fasting dramatically reduced peroxisome numbers in the hepatocytes of pex5−/− at 6 dpf and almost completely reduced peroxisome numbers in most cells at 7 dpf (Fig. 2e, n = 8). However, peroxisome numbers under feeding conditions were comparable between the WT and pex5−/− (Fig. 2f, n = 8). These results suggest that fasting-mediated metabolic challenge could discriminate functional peroxisomes from immature ones, whose disappearance was facilitated in fasted pex5−/−larvae.
pex5 depletion induces alteration in the hepatic mitochondria under fasting conditions
Previous studies using knockout mice have shown that mitochondrial abnormalities could stem from dysfunctional peroxisomes [11, 17, 18, 36]. To evaluate the effect of the pex5 mutation and nutritional stress on mitochondria, we first examined mitochondrial morphology using the expression of Xla.Eef1a1:MLS-EGFP transgene and compared it between WT and pex5−/− under feeding and fasting conditions (Fig. 3a). Under feeding conditions, mitochondria were largely elongated in the hepatocytes of WT at 6 dpf (n = 8), whereas they were mostly fragmented in pex5−/− (n = 8), suggesting that pex5 deficiency might cause mitochondria to be morphologically different from those in WT under feeding conditions. Consistent with this idea, mitochondria in the hepatocytes of pex5−/− were fragmented as early as 5 dpf (Supplementary Fig. S2, n = 8). The different morphologies of mitochondria were maintained at 7 dpf under feeding conditions (Fig. 3a and Supplementary Fig. 2, n = 8). These results indicate that mitochondria in WT hepatocytes under feeding conditions maintained their elongated morphology, while those in pex5−/− underwent a morphological change to become largely fragmented.
Fasting induces a radical change in the mitochondrial shape and abundance. a Expression of MLS-EGFP from the liver sections of Xla.Eef1a1:MLS-EGFP transgenic zebrafish at 6 dpf is shown to compare between the WT and pex5−/− zebrafish and between the feeding and fasting conditions. Scale bar = 5 μm. b Transmission electron microscope (TEM) images are shown to detect the differential mitochondrial shapes in the liver at 4.5 dpf and 6.5 dpf upon fasting depending on the pex5 genotype. Arrows indicate mitochondria with abnormal cristae structure. Scale bar = 500 nm. c Endogenous expression of a component of CoxIV is shown following immunofluorescence for comparison between the WT and pex5−/− zebrafish liver and between the feeding and fasting conditions. Scale bar = 5 μm. Graphs show quantified signal intensity of CoxIV at the indicated conditions and presented as the average with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; ** indicates p values < 0.01. d Mitochondrial abundance was analyzed by either counting the number of mitochondria in the liver cells of WT and pex5−/− zebrafish at the indicated stages under fasting conditions (left) or quantifying the mitochondrial DNA amount at 6 dpf in reference to a β-actin genomic region using quantitative polymerase chain reaction (qPCR) (right). Note that the number of mitochondria in the liver cells of fasted WT zebrafish at 6 dpf was set to 1, which was used for reference to quantify those in the rest of the samples. Statistical significance was determined using the Student’s t test in Microsoft Excel; * and ** indicate p values < 0.05 and < 0.01, respectively. e Western blot using whole lysates was performed to compare expression levels of component proteins of the respiratory chain complexes, II V, and IV between WT and pex5−/− at the indicated developmental stages. A representative image is shown, and the expression levels were quantified and presented as the average of three repeats with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; * and ** indicate p values < 0.05 and < 0.01, respectively
Notably, when zebrafish were subjected to fasting, some mitochondria in the WT hepatocytes became fragmented and formed an elliptical shape at 6 dpf (Fig. 3a, n = 8), indicating that fasting might induce a change in mitochondrial shape. When the same fasting regime was imposed on pex5−/−, most mitochondria in the liver formed a round shape at 6 dpf and remained similar at 7 dpf (Supplementary Fig. 2, n = 8), suggesting that fasting promoted further changes in mitochondrial shape in pex5−/−. To further examine the mitochondrial structural changes dependent on the pex5 genotype and nutritional status, we examined liver sections using TEM. Consistent with the fluorescence microscopic observation, TEM showed that elongated mitochondria were observed in the WT liver, whereas mitochondria were fragmented in the pex5−/− liver at 4.5 dpf (Fig. 3b). Moreover, unlike the mitochondria in the WT liver, few mitochondria appeared to be damaged in the pex5−/− livers as their cristae structure was disrupted. Fasting facilitated mitochondrial fission, as mitochondria of WT liver at 6.5 dpf were no longer tubular. However, the mitochondria in the fasted WT liver were structurally intact and formed an elliptical shape. In contrast, in the livers of fasted pex5−/−, most mitochondria were found to be damaged and their cristae were disorganized at 6.5 dpf. These results suggest that mitochondria might adopt different shapes depending on the pex5 genotype (i.e., WT vs. pex5−/−) and metabolic state (fed vs. fasted), thereby displaying the most severely deranged shape in the liver cells of pex5−/− under fasting conditions.
Since different mitochondrial shapes can be translated into different qualities [43,44,45], we examined mitochondrial abundance under different conditions. Immunostaining showed significantly increased expression of the mitochondrial proteins belonging to respiratory chain complex IV in the fasted pex5−/− hepatocytes compared to that in the fasted WT at 6 dpf (Fig. 3c, n = 8/genotype). In addition, we found that the number of mitochondria in the liver cells of fasted pex5−/− was 1.4 times higher than that in fasted WT at 6 dpf (Fig. 3d). The increased mitochondrial abundance in pex5−/− liver cells was correlated with increased mitochondrial DNA content, as determined by the amount of the MT-ND1 gene, which encodes a component of the respiratory chain complex I (Fig. 3d). The expression of other mitochondrial genes, such as MT-ATP6 and MT-CYB, also showed a similar trend (data not shown). Although we were unable to perform western blot analysis using the larval liver due to limited materials, results from western blotting using whole larval lysates showed increased levels of respiratory chain complex proteins in the fasted pex5−/− at 5.5 - and 6 dpf depending on complex proteins compared to those in the fasted WT (Fig. 3e), further supporting elevated mitochondrial abundance in the fasted pex5−/− animals.
To understand the molecular mechanism underlying the mitochondrial phenotype in fasted pex5−/− livers, we performed RNA-seq analysis using total RNAs prepared from fasted WT and pex5−/− livers at 6 dpf and compared gene expression profiles between the two. The gene expression pattern was not biased to either side and was evenly distributed, indicating that there were no technical artifacts that might affect the subsequent analysis (Supplementary Fig. S3a). We compared the expression profiles of mitochondrial genes, which were defined by MitoCarta3.0 [46], and OXPHOS genes between the two genotypes (Supplementary Fig. S3b, c). We also compared the expression of genes involved in mitochondrial biogenesis between the two (Supplementary Fig. S3d). By examining the fold changes in gene expression, we found that approximately 75% of OXPHOS genes and 59% of mitochondrial genes were upregulated in fasted pex5−/− livers (Supplementary Fig. S3e). Moreover, eight out of ten genes involved in mitochondrial biogenesis were upregulated in fasted pex5−/− livers compared to those in fasted WT livers. These results suggest that the fasted pex5−/− livers, in comparison with the fasted WT livers, had greater mitochondrial quantity and possibly increased mitochondrial function.
Fasting increases hepatic ROS and facilitates depletion of hepatic lipids and glycogen in pex5 −/−
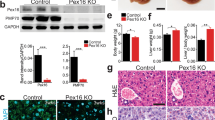
It is conceivable that elevated mitochondrial activity may induce high ROS levels. We tested whether this was the case with fasted pex5−/− livers. We used 4-hydroxynonenal (4-HNE) level as a measure of ROS-dependent lipid peroxidation [47, 48]. Pex5−/− liver cells showed a strong increase in the level of 4-HNE compared to WT liver cells during fasting at 6 dpf (Fig. 4a and b, n = 12). Consistent with ROS as a potent apoptotic inducer, we found a significant increase in apoptotic cell death in the liver of pex5−/− at 7 dpf (Fig. 4c and d, n = 12). We attempted to relieve the ROS burden by treating with NAC, a well-known ROS scavenger, which dramatically reduced 4-HNE levels in the liver cells of all treated larvae (Fig. 4a, n = 12) and remarkably decreased the number of apoptotic liver cells of pex5−/− compared to the untreated pex5−/− (Fig. 4b, n = 12). Importantly, NAC treatment almost completely rescued 75% of fasted pex5−/− from developing lethality-associated phenotypes (Fig. 4c, n = 24). These results suggest that ROS induction in the liver cells of pex5−/− at 6 dpf might also contribute to the early lethality of pex5−/− under fasting conditions.
Fasting induces ROS and abnormally increased consumption of hepatic lipids and glycogen in the pex5−/− liver. a 4-hydroxynonenal (4-HNE) was performed in the liver sections of WT and pex5−/− zebrafish at 6 dpf either untreated or treated with 100 μM N-acetyl-cysteine (NAC). Scale bar = 5 μm. Graph shows quantified signal intensity and presented as the average with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; ** indicates p values < 0.01. b Terminal deoxyribonucleotidyl transferase (TDT)-mediated dUTP nick-end labeling (TUNEL) staining was performed in the liver sections of WT and pex5−/− zebrafish at 7 dpf either untreated or treated with NAC. Positive signals appear in green, and nuclei were stained with 4ʹ, 6-diamidino-2-phenylindole (DAPI) in blue. Scale bar = 100 μm. Graph shows the percentage of apoptotic cells per different conditions with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; * indicates p values < 0.05. c Representative images of WT and pex5−/− zebrafish at 9 dpf either untreated or treated with NAC. Larvae were shown in lateral views with anterior to the left. Scale bar = 1 mm. d ORO (upper panels) or PAS staining (lower panels) was performed in the liver sections of both fed and fasted WT and pex5−/− zebrafish at 6 and 7 dpf. Representative images are shown. Scale bar = 100 μm
Given the importance of mitochondria in regulating cellular metabolism, we examined whether the nutritional status of pex5−/− livers would differ from that of WT livers upon fasting. We performed Oil Red O (ORO) and periodic acid-Schiff (PAS) staining of WT and pex5−/− zebrafish to detect neutral lipids and glycogen, respectively. There was no noticeable difference between the two genotypes under feeding conditions (Fig. 4d, n = 6). However, fasting considerably reduced the levels of both neutral lipids and glycogen in pex5−/− livers at 6 and 7 dpf (n = 6), suggesting that fasting might induce metabolic disturbances in pex5−/− zebrafish. As NAC treatment repressed apoptotic cell death and mortality in pex5−/−, we also examined the effect of NAC on liver metabolism. NAC treatment effectively restored the levels of both hepatic lipids and glycogen in fasted pex5−/− compared to the untreated group at 6 dpf (Supplementary Fig. S4, 67%, n = 6), indicating that ROS is a detrimental factor for metabolic disturbance in fasted pex5−/−.
Limiting mitochondrial activity partially rescues the expedited death of fasted pex5 −/−
Our observations of elevated mitochondrial activity, high 4-HNE levels, and depletion of hepatic lipids in fasted pex5−/− livers led us to hypothesize that pex5−/− could be sensitized to induce metabolic disturbances in the liver under fasting conditions. Therefore, we investigated whether partial inhibition of the mitochondrial pathway could rescue metabolic disturbances in fasted pex5−/−. Treatment with etomoxir, a well-known chemical inhibitor of carnitine palmitoyltransferase 1a necessary for fatty acid beta-oxidation, was able to rescue morphological defects in 56% (n = 36) of fasted pex5−/−, all of which would otherwise develop fasting-induced phenotypes and associated early mortality at 9 dpf (Fig. 2c; Fig. 5a). A similar rescue (55%, n = 22) was also observed following treatment with oligomycin A, an inhibitor of mitochondrial F0F1-ATPase, supporting our hypothesis that fasting may abnormally induce mitochondrial activity and that limiting mitochondrial function thus rescues the phenotypes of fasted pex5−/−. Consistent with mitochondrial abnormalities in the fasted pex5−/−, we found that treatment with etomoxir significantly reduced ROS levels in the pex5−/− liver (Supplementary Fig. S5, n = 5). We next tested whether the rescue effect observed with partial inhibition of mitochondrial function was mediated by ameliorating metabolic disturbances in fasted pex5−/− livers. We found that both fasted WT and pex5−/− treated with etomoxir accumulated hepatic lipids, as detected by ORO staining, compared to their untreated counterparts (Fig. 5b, n = 6/genotype), suggesting that hepatic lipid synthesis in fasted pex5−/− might be functional, but hepatic lipid consumption was abnormally increased. Owing to the compromised fatty acid oxidation by etomoxir treatment, glycogen levels detected by PAS staining were accordingly decreased in the livers of both fasted WT and pex5−/− treated with etomoxir compared to those in untreated counterparts (n = 6/genotype). We also found that oligomycin treatment similarly accumulated hepatic lipids, but did not induce differences in glycogen content in the fasted pex5−/− livers compared to those in the fasted WT livers (Supplementary Fig. S4, n = 6). The differential effect on glycogen content between etomoxir and oligomycin might stem from the fact that etomoxir is specific to fatty acid oxidation. These results strongly suggest that fasted pex5−/− larvae might have abnormally elevated mitochondrial activity, which could potentially be deleterious to the maintenance of metabolic balance and animal survival.
Inhibiting either the mitochondrial β-oxidation or ATP synthase ameliorates the fasting-associated pex5−/− phenotypes. a Either etomoxir (1.5 μM) or oligomycin (5 pM) was treated from 5 to 9 dpf. A representative image from the survived 40% of the fasted, untreated pex5−/− zebrafish is shown, and the remaining 60% larvae were found dead at 9 dpf as indicated. The average percentage representing the degree of rescue from two independent experiments is presented in each panel. Images were shown in lateral views with anterior to the left. Scale bar = 1 mm. b Liver sections from the WT and pex5−/− zebrafish at 6 dpf treated with either etomoxir (eto, 1.5 μM) or oligomycin (oligo, 5 pM) from 5 dpf were shown following either ORO (upper panels) or PAS staining (lower panels). The same staining of liver sections from untreated WT and pex5−/− zebrafish at 6 dpf was shown for comparison. A representative image for each condition is shown. Scale bar = 100 μm
pex5 depletion increases mTORC1 activity under fasting conditions
A previous study using cell lines suggested that peroxisomes may act as signaling nodes to recruit tuberous sclerosis complex 2 (TSC2) in response to ROS, which then suppresses mammalian target of rapamycin complex 1 (mTORC1) activity [24]. In addition, we previously reported that PEX5 knockdown in HepG2 cells failed to suppress mTORC1 signaling in response to fasting [25]. Therefore, we tested whether fasting impairs mTORC1 signaling in pex5−/−. Western blotting using whole larval lysates for the expression of eukaryotic translation initiation factor 4E binding protein 1 (4E-BP1), a well-known mTORC1 downstream effector, indicated that the activated phosphorylated form of 4E-BP1 was increased in pex5−/− under fasting conditions compared to that in WT (Fig. 6a). Moreover, the fasted pex5−/− livers displayed highly increased phospho-S6, an activated form of S6 ribosomal protein, compared to the fasted WT hepatocytes (Fig. 6b, n = 8), supporting our observation of dysregulated mTORC1 signaling in the fasted pex5−/−.
Mechanistic target of rapamycin complex 1 (mTORC1) signaling remains unrepressed in the fasted pex5−/− liver. a Western blotting shows the expression levels of 4EBP and its phosphorylated form from whole lysates of WT and pex5−/− zebrafish at 6 dpf. The expression of β-actin was used as the loading control. Graph shows quantified expression levels of p4EBP relative to β-actin levels and presented as the average of three repeats with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; * indicates p value < 0.05. b Immunofluorescence was used to detect the differential levels of S6 and its phosphorylated form in the liver cells of WT and pex5−/− at 6 dpf. DAPI detects nuclei in each liver section. Scale bar equals to 5 μm. Graph shows relative fluorescence intensity presented as the average with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; ** indicates p values < 0.01. c Rapamycin (rapa, 250 nM) was used to treat WT and pex5−/− from 3.5 to 8 dpf and representative images of the resulting larvae were laterally presented with anterior to the left. The average percentage representing a near-complete rescue from two independent experiments is indicated. Scale bar = 1 mm. d Liver sections from the larvae treated with rapamycin from 3.5 to 7 dpf were stained either ORO or PAS. Representative images are shown, and the percentage of rescue in the levels of liver lipids and glycogen are indicated in the panels of the fasted pex5−/− zebrafish treated with rapamycin. Scale bar = 100 μm
To examine the association between dysregulated mTORC1 and larval defects in fasted pex5−/− zebrafish, we used rapamycin to inhibit downstream effectors of mTORC1 [49]. Notably, rapamycin treatment efficiently rescued the morphological phenotypes observed in the fasted pex5−/− (Fig. 6c, 86%, n = 28), a phenomenon similar to the mitochondrial β-oxidation inhibition shown in Fig. 5. Moreover, rapamycin treatment effectively restored the levels of hepatic lipids of the fasted pex5−/− much higher than the untreated group at 7 dpf (50%, n = 8). Hepatic glycogen levels in fasted pex5−/− zebrafish also increased upon rapamycin treatment (Fig. 6d, 50%, n = 8). These results confirm that sustained mTORC1 activity during fasting might play an important role in the mortality of pex5−/−.
Given that both deregulated mitochondria and unrepressed mTORC1 are major contributors to larval defects upon fasting in pex5−/− zebrafish, we determined the relationship between these two pathways. The role of mTORC1 in regulating mitochondrial biogenesis and function is well established [50, 51]. Indeed, rapamycin treatment restored mitochondrial abnormalities in fasted pex5−/− livers, as evidenced by the reduced levels of both mitochondrial abundance and 4-HNE (Supplementary Fig. S5 and S6, n = 5). We next examined the possibility that modulating mitochondrial function could, in turn, affect mTORC1 activity. Indeed, a highly enriched signal from anti-phospho-S6 immunofluorescence in fasted pex5−/− livers at 6 dpf, indicative of dysregulated mTORC1 activity, was considerably decreased after etomoxir treatment (Supplementary Fig. S7, n = 5). Accordingly, we tested the combinatorial role of mTORC1 and mitochondrial activities in contributing to phenotype severity and mortality in fasted pex5−/− zebrafish by assessing morphological rescue when both were simultaneously inhibited. Full doses of rapamycin (250 nM) and etomoxir (1.5 μM) in combination resulted in developmental abnormalities in both WT and pex5−/− zebrafish, presumably because of insufficient signaling required for normal larval development (n = 15, not shown). However, a combination of a quarter dose of rapamycin and etomoxir rescued 53% of the fasted pex5−/− (n = 34). A quarter dose of each drug did not improve the phenotype observed in fasted pex5−/− (n = 12). These results indicate the presence of a positive relationship between mitochondrial function and mTORC1 activity in promoting larval defects in pex5−/−.
Autophagy flux is impaired and damaged mitochondria are accumulated in pex5 −/− under fasting conditions
Activated mTORC1 is reportedly associated with the suppression of autophagy [52,53,54,55,56], although autophagy can be regulated independently of mTORC1 under some conditions [57,58,59,60]. In addition, developing embryos require a certain level of mTORC1 to ensure the proper anabolic signals necessary for animal development. We analyzed the autophagy marker light chain 3 (Lc3)-II, a lipidated form of Lc3, and found it to be increased in pex5−/− compared to that in WT under fasting conditions (Fig. 7a). We tested whether the increased Lc3-II levels were due to enhanced autophagy or autophagy blockade by treatment with chloroquine (CQ), an autophagy blocker. CQ treatment induced Lc3-II levels in WT, indicating that CQ effectively blocked autophagic flux. However, the Lc3-II levels between untreated and CQ-treated pex5−/− showed minimal changes, suggesting autophagy blockade in fasted pex5−/−. To verify this, we examined p62 levels, which are known to be regulated in accordance with autophagy activity [61]. We found strikingly high levels of both p62 mRNA and p62 protein in the liver cells of fasted pex5−/− (Fig. 7b, c), consistent with autophagy blockade in fasted pex5−/−. We next examined the quality control of mitochondria because autophagy blockade may also influence mitochondrial health, related to our finding of increased mitochondrial activity and abundance with altered morphology in the liver cells of pex5−/− (Figs. 3 and 5). The ubiquitination of mitochondrial proteins may act as a signal for the selective autophagic degradation of damaged mitochondria, known as mitophagy [61]. We found that ubiquitinated proteins significantly accumulated in the mitochondria of the liver cells of fasted pex5−/− compared to WT (Fig. 7c, n = 6), indicating accumulated mitochondrial damage.
Dysfunctional mitochondria are accumulated in the fasted pex5−/− liver. a Western blotting shows the level of the light chain 3 (Lc3)-II from whole lysates of WT and pex5−/− at 6 dpf with or without chloroquine (CQ, 250 μM). Short and long stand for shorter and longer exposure for film development, respectively. The expression of β-actin was used as the loading control. Graph shows quantified expression levels of Lc3-II (long) relative to β -actin levels and presented as the average of three repeats with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; * and ** indicate p values < 0.05 and < 0.01, respectively. b Relative expression levels of p62 were determined in reference to those of β-actin by performing quantitative reverse transcription-polymerase chain reaction (qRT-PCR) using total RNA isolated from the liver cells of fasted WT and pex5−/− zebrafish at the indicated stages. Data are presented as the average of three repeats for the relative ratio of p62/b-actin expression. Error bars indicate the standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; * and ** indicate p values < 0.05 and < 0.01, respectively. c Immunofluorescence was performed in liver sections from WT and pex5−/− to detect damaged mitochondria using an anti-Ubiquitin antibody or an autophagy cargo protein using anti-p62 antibody in red as indicated. Mitochondria were visualized through the EGFP signal expressed in the transgenic Tg(Xla.Eef1α1:MLS-EGFP) zebrafish. DAPI stains nuclei in blue in each liver section. Representative images were shown from two independent experiments. Scale bar = 5 μm. Graphs show quantified signal intensity of p62 and presented as the average with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; ** indicates p value < 0.01. d UK5099 (25 μM) was treated in WT and pex5−/− from 3.5 to 8 dpf, and representative images of the resulting larvae at 8 dpf were laterally presented with anterior to the left. The average percentage representing a near-complete rescue from two independent experiments is indicated. Scale bar equals 1 μm. e Liver sections from the larvae treated with UK5099 from 3.5 to 7 pf were stained either ORO or PAS. Representative images are shown and the percentage of rescue in the levels of liver lipids and glycogen is indicated in the panels of the fasted pex5−/− treated with UK5099. Scale bar = 100 μm. f UK5099 (25 μM) was treated in WT and pex5−/− from 3.5 to 6 dpf, and immunofluorescence was performed in liver sections using anti-CoxIV antibody. Representative images are shown. Scale bar = 5 μm. Graphs show quantified signal intensity of CoxIV at the indicated conditions and presented as the average with error bars indicating standard deviation. Statistical significance was determined using the Student’s t test in Microsoft Excel; ** indicates p value < 0.01
We reasoned that increasing autophagy might relieve the limited autophagy and rescue the phenotypes in fasted pex5−/−. UK5099, an inhibitor of the mitochondrial pyruvate carrier, has been reported to decrease intracellular acetyl coenzyme A levels and thus induce autophagy in an mTOR-independent manner [62]. We found that treatment with UK5099 rescued 41% of the fasted pex5−/− almost completely (n = 32) and the remaining 59%, at least partially, based on morphological phenotypes at 8 dpf (Fig. 7d). Moreover, treatment with UK5099 restored hepatic lipid and glycogen contents, at least partially, in 64% of fasted pex5−/− larvae at 7 dpf (Fig. 7e, n = 11). Notably, consistent with mitophagy blockade in fasted pex5−/− livers, UK5099 treatment significantly reduced the expression of Complex IV proteins in fasted pex5−/− livers (Fig. 7f, n = 6). These results suggest that reduced autophagy flux might contribute to the deterioration of fasted pex5−/−.
Discussion
Zebrafish is a well-established vertebrate animal model that offers a combination of genetic and experimental advantages in addition to other animal model systems [42, 63], but its use has been limited in peroxisomal research. Nonetheless, a pioneering study revealed that zebrafish peroxisomes were the most abundant in the liver and renal tubular epithelium, with a distribution similar to that observed in mammals [26]. In addition, we and other researchers have recently shown that functional peroxisomes are essential for normal zebrafish embryogenesis [27, 28, 64]. In this study, we generated pex5−/− and verified its utility as a zebrafish model of ZSD by providing evidence of phenocopy in a mouse model of peroxisome dysfunction and in human patients. An intriguing feature of pex5−/− is an increased sensitivity to fasting, which leads to metabolic failure and expedited death.
It is generally accepted that in response to peroxisome deficiency, mitochondria may eventually become dysfunctional. For example, PEX5−/− hepatocytes display functional and morphological anomalies in mitochondria, which are often accompanied by increased oxidative stress [65]. Therefore, accumulation of dysfunctional mitochondria, together with failure in metabolic regulation, might cause phenotypic presentation in peroxisome-deficient diseases. In this study, we showed that mitochondria in zebrafish liver cells adopted their shape depending on both the pex5 genotype and the nutritional state. For example, under feeding conditions, the mitochondria in WT liver cells were largely elongated, whereas those in pex5−/− were fragmented (Fig. 3a and Supplementary Fig. S2). Fasting facilitated fission to induce mitochondria with an elliptical shape in WT, but further induced them to constitute a round-shaped population in pex5−/−. TEM images were consistent with disorganized mitochondria in the liver cells of fasted pex5−/− (Fig. 3b). In particular, under fasting conditions, liver cells of fasted pex5−/− contained severely damaged mitochondria with distorted cristae and a disorganized membrane structure at 6.5 dpf. Further efforts are necessary to determine the causal relationship between mitochondrial morphology and activity under different nutritional conditions.
Our finding of severely decreased lipid and glycogen levels in the liver cells of fasted pex5−/− zebrafish (Fig. 4) prompted us to examine the role of the mitochondria upon fasting. Previous studies have reported that the absence of functional peroxisomes perturbs carbohydrate metabolism and reduces sensitivity to low sugar environments [15, 19, 66]. Similarly, we also found near-complete depletion of liver glycogen in fasted pex5−/− zebrafish. We attempted to rescue the expedited mortality of fasted pex5−/− by supplementing with glucose, but did not succeed (data not shown). Notably, we found that blocking mitochondrial β-oxidation was efficient for the survival of fasted pex5−/− zebrafish. A similar rescue in mortality was observed with an inhibitor of ATP synthase, suggesting that mitochondrial activity increased as an initial response to peroxisomal dysfunction. Indeed, several studies have demonstrated that, in response to peroxisomal dysfunction, mitochondrial activities were increased as a compensatory mechanism; mitochondrial matrix enzymes were more active in Pex5−/− hepatocytes than in WT [17, 36], and mitochondria-dependent β-oxidation was increased despite the evident hepatosteatosis [18, 67]. In addition, Pex5−/− hepatocytes produced slightly more ATP than control hepatocytes [11]. Due to technical restrictions in using zebrafish larval livers, we were unable to measure the mitochondrial oxygen consumption rate and lipid oxidation. However, we found an increase in the number of mitochondria, the amount of mitochondrial DNA, the expression levels of mitochondrial proteins (Fig. 3), and the expression of OXPHOS and mitochondrial genes, as well as genes involved in mitochondrial biogenesis in the fasted pex5−/− (Supplementary Fig. S3), all of which suggested increased mitochondrial activity. Although the exact underlying mechanism needs to be further elucidated, collective transcriptional activation of the mitochondrial genes (Supplementary Fig. S3) might contribute to the elevated mitochondrial activity in pex5−/− under fasting conditions. In support of increased mitochondrial activity, we found depleted lipid content and increased levels of cellular ROS in hepatocytes of fasted pex5−/−. We confirmed that ROS might also contribute to the observed mortality, as NAC treatment inhibited apoptosis in fasted pex5−/− livers, restored hepatic lipids and glycogen levels, and suppressed early lethality in pex5−/− (Fig. 4). While hepatic lipids seem to be critical for the survival of fasted pex5−/−, the level of carbohydrates may also be important. The relative importance of carbohydrate and lipid levels during zebrafish development is not well understood.
As a critical factor that influences the phenotypic severity of fasted pex5−/−, we identified mTORC1 as not being repressed under fasting conditions. mTORC1 facilitates anabolic reactions under nutrient-rich conditions but is repressed under energy-deprived conditions [52,53,54,55,56]. The role of peroxisomes in regulating mTORC1 signaling has been reported in a cell line in which peroxisomes recruit TSC2 to repress mTORC1 in response to ROS [24]. We also reported that the mTORC1 regulatory pathway is dependent on PEX5 upon serum starvation in HepG2 cells [25]. In this study, we confirmed increased mTORC1 activity under fasting conditions in the liver cells of pex5−/− (Fig. 6). Persistent activation of mTORC1 increases the expression of genes involved in glycolysis [66] which likely explains the depleted hepatic glycogen levels in fasted pex5−/−. In addition, persistent mTORC1 activity likely regulates mitochondrial activity [50, 51, 68], thereby further enhancing mitochondrial activity in liver cells of fasted pex5−/−. Notably, we showed that mTORC1 was also affected by mitochondrial function. Etomoxir treatment remarkably decreased phospho-S6 levels in the liver cells of fasted pex5−/− zebrafish (Supplemental Fig. S7), suggesting the presence of an axis that influences mTORC1 activity via the mitochondria. Furthermore, autophagy also influences the severity of the phenotype observed in fasted pex5−/−. Fasting may burden mitochondria-dependent energy production, leading to the accumulation of damaged mitochondria, which are not efficiently cleared due to limited autophagy flux (Fig. 7). Accordingly, an autophagy inducer was able to partially rescue the phenotypes of fasted pex5−/−, indicating that blockade of autophagy might be detrimental to the survival of fasted pex5−/−.
Based on the present study and previous findings, we propose a working model in which the phenotypes displayed in pex5 deficiency become exacerbated upon fasting (Fig. 8). Peroxisome deficiency induces mitochondrial abnormalities and the associated metabolic failure. When fasting is imposed as a metabolic stress additive to the absence of functional peroxisomes, the entire pathogenesis seems to be greatly facilitated, resulting in rapid depletion of hepatic lipids and glycogens and, consequently, expedited organismal death. A critical player upon fasting would be unrepressed mTORC1 activity, by which mitochondrial function induced initially in response to peroxisome dysfunction could be further increased. Increased mitochondrial function may, in turn, prevent mTORC1 repression. This positive regulatory loop builds damaged mitochondria, which may not be efficiently cleared due to impaired autophagy flux in fasted pex5−/− livers. Further studies to elucidate the factors that block autophagy flux upon fasting in the absence of functional peroxisomes will warrant a refinement in interpreting a series of molecular events in pex5 deficiency under environmental stress.
A model to illustrate the fasting-induced early mortality in pex5−/− in this study. Events occurring primarily in the pex5−/− liver under normal fed conditions are shown in black arrows. Briefly, due to dysfunctional peroxisomes, metabolic burden in pex5−/− induces compensatory activation of mitochondria which may then lead to mitochondrial damage and subsequent metabolic failure and organismal death. The entire process may run slow and take almost one month presumably due to the availability of nutrients under fed conditions. However, the process becomes highly facilitated under fasting conditions. mTORC1 becomes repressed in the fasted WT liver but not in the pex5−/− liver. In the fasted pex5−/− liver, mitochondrial activities are upregulated by unrepressed mTORC1 whose activity may also regulated by mitochondrial activity. Further, abnormally high mitochondrial activities may rapidly induce mitochondrial dysfunction and lead to accumulation of damaged mitochondria, which may not be replenished due to the limited autophagic flux. The undesirable condition upon fasting then leads to metabolic failure and results in early mortality of pex5−/−. The events strengthened upon fasting are shown by red arrows. The points of action of drugs that either activate or inhibit the process are shown in blue
Taken together, we showed that zebrafish can be a useful animal model system to provide mechanistic insights into peroxisome-dependent mitochondrial health. In addition, zebrafish may provide an in vivo experimental approach to reveal the metabolic response to environmental stresses, such as fasting. Based on the intervention experiments, we suggest that targeting mitochondrial function or mTORC1 individually or in combination may have a beneficial effect on the prevention of phenotypic exacerbation caused by peroxisome dysfunction during metabolic crisis. In parallel, we suggest that a drug that can reduce ROS or induce mitophagy may provide a preventive or therapeutic option for the treatment of peroxisomal dysfunction exhibiting metabolic imbalance.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Ma C, Agrawal G, Subramani S (2011) Peroxisome assembly: matrix and membrane protein biogenesis. J Cell Biol 193(1):7–16
Mannaerts GP, Van Veldhoven PP (1993) Metabolic pathways in mammalian peroxisomes. Biochimie 75(3–4):147–158
Wanders RJ (2013) Peroxisomes in human health and disease: metabolic pathways, metabolite transport, interplay with other organelles and signal transduction. Subcell Biochem 69:23–44
Fidaleo M (2010) Peroxisomes and peroxisomal disorders: the main facts. Exp Toxicol Pathol 62(6):615–625
Wanders RJ (2004) Peroxisomes, lipid metabolism, and peroxisomal disorders. Mol Genet Metab 83(1–2):16–27
Wanders RJ (2004) Metabolic and molecular basis of peroxisomal disorders: a review. Am J Med Genet A 126A(4):355–375
Wanders RJ, Waterham HR (2006) Peroxisomal disorders: the single peroxisomal enzyme deficiencies. Biochim Biophys Acta 1763(12):1707–1720
Levine B, Kroemer G (2008) Autophagy in the pathogenesis of disease. Cell 132(1):27–42
Otera H et al (1998) Peroxisome targeting signal type 1 (PTS1) receptor is involved in import of both PTS1 and PTS2: studies with PEX5-defective CHO cell mutants. Mol Cell Biol 18(1):388–399
Shimozawa N et al (1999) Functional heterogeneity of C-terminal peroxisome targeting signal 1 in PEX5-defective patients. Biochem Biophys Res Commun 262(2):504–508
Baumgart E et al (2001) Mitochondrial alterations caused by defective peroxisomal biogenesis in a mouse model for Zellweger syndrome (PEX5 knockout mouse). Am J Pathol 159(4):1477–1494
Baes M et al (1997) A mouse model for Zellweger syndrome. Nat Genet 17(1):49–57
Baes M et al (2002) Generation of Pex5-loxP mice allowing the conditional elimination of peroxisomes. Genesis 32(2):177–178
Wanders RJ, Waterham HR (2006) Biochemistry of mammalian peroxisomes revisited. Annu Rev Biochem 75:295–332
Peeters A et al (2011) Carbohydrate metabolism is perturbed in peroxisome-deficient hepatocytes due to mitochondrial dysfunction, AMP-activated protein kinase (AMPK) activation, and peroxisome proliferator-activated receptor gamma coactivator 1alpha (PGC-1alpha) suppression. J Biol Chem 286(49):42162–42179
Berendse K et al (2019) Liver disease predominates in a mouse model for mild human Zellweger spectrum disorder. Biochim Biophys Acta Mol Basis Dis 1865(10):2774–2787
Peeters A et al (2015) Mitochondria in peroxisome-deficient hepatocytes exhibit impaired respiration, depleted DNA, and PGC-1alpha independent proliferation. Biochim Biophys Acta 1853(2):285–298
Peeters A et al (2011) Hepatosteatosis in peroxisome deficient liver despite increased beta-oxidation capacity and impaired lipogenesis. Biochimie 93(10):1828–1838
Wangler MF et al (2017) Peroxisomal biogenesis is genetically and biochemically linked to carbohydrate metabolism in Drosophila and mouse. PLoS Genet 13(6):e1006825
Keane MH et al (2007) Bile acid treatment alters hepatic disease and bile acid transport in peroxisome-deficient PEX2 Zellweger mice. Hepatology 45(4):982–997
Rahim RS et al (2016) Mitochondrial changes and oxidative stress in a mouse model of Zellweger syndrome neuropathogenesis. Neuroscience 334:201–213
Salpietro V et al (2015) Zellweger syndrome and secondary mitochondrial myopathy. Eur J Pediatr 174(4):557–563
Bulow MH et al (2018) Unbalanced lipolysis results in lipotoxicity and mitochondrial damage in peroxisome-deficient Pex19 mutants. Mol Biol Cell 29(4):396–407
Zhang J et al (2013) A tuberous sclerosis complex signalling node at the peroxisome regulates mTORC1 and autophagy in response to ROS. Nat Cell Biol 15(10):1186–1196
Eun SY et al (2018) PEX5 regulates autophagy via the mTORC1-TFEB axis during starvation. Exp Mol Med 50(4):4
Krysko O et al (2010) Peroxisomes in zebrafish: distribution pattern and knockdown studies. Histochem Cell Biol 134(1):39–51
Kim YI et al (2020) Slc25a17 acts as a peroxisomal coenzyme A transporter and regulates multiorgan development in zebrafish. J Cell Physiol 235(1):151–165
Strachan LR et al (2017) A zebrafish model of X-linked adrenoleukodystrophy recapitulates key disease features and demonstrates a developmental requirement for abcd1 in oligodendrocyte patterning and myelination. Hum Mol Genet 26(18):3600–3614
Kimmel CB et al (1995) Stages of embryonic development of the zebrafish. Dev Dyn 203(3):253–310
Jung SH et al (2010) Visualization of myelination in GFP-transgenic zebrafish. Dev Dyn 239(2):592–597
Kim D et al (2016) Fis1 depletion in osteoarthritis impairs chondrocyte survival and peroxisomal and lysosomal function. J Mol Med (Berl) 94(12):1373–1384
Kim MJ et al (2008) Real-time imaging of mitochondria in transgenic zebrafish expressing mitochondrially targeted GFP. Biotechniques 45(3):331–334
van der Velden YU et al (2011) The serine-threonine kinase LKB1 is essential for survival under energetic stress in zebrafish. Proc Natl Acad Sci USA 108(11):4358–4363
Hashimshony T et al (2016) CEL-Seq2: sensitive highly-multiplexed single-cell RNA-Seq. Genome Biol 17:77
Hwangbo H et al (2022) Bio-printing of aligned GelMa-based cell-laden structure for muscle tissue regeneration. Bioact Mater 8:57–70
Dirkx R et al (2005) Absence of peroxisomes in mouse hepatocytes causes mitochondrial and ER abnormalities. Hepatology 41(4):868–878
Steinberg SJ et al (2006) Peroxisome biogenesis disorders. Biochim Biophys Acta 1763(12):1733–1748
Martinez M, Vazquez E (1998) MRI evidence that docosahexaenoic acid ethyl ester improves myelination in generalized peroxisomal disorders. Neurology 51(1):26–32
Hulshagen L et al (2008) Absence of functional peroxisomes from mouse CNS causes dysmyelination and axon degeneration. J Neurosci 28(15):4015–4027
Germain K, Kim PK (2020) Pexophagy: a model for selective autophagy. Int J Mol Sci 21(2):578
Flynn EJ, Trent CM, Rawls JF (2009) Ontogeny and nutritional control of adipogenesis in zebrafish (Danio rerio). J Lipid Res 50(8):1641–1652
Choi TY et al (2021) Zebrafish as an animal model for biomedical research. Exp Mol Med 53(3):310–317
Yao CH et al (2019) Mitochondrial fusion supports increased oxidative phosphorylation during cell proliferation. Elife. https://doi.org/10.7554/eLife.41351
Suarez-Rivero JM et al (2016) Mitochondrial dynamics in mitochondrial diseases. Diseases. https://doi.org/10.3390/diseases5010001
Haigh JL, New LE, Filippi BM (2020) Mitochondrial dynamics in the brain are associated with feeding, glucose homeostasis, and whole-body metabolism. Front Endocrinol (Lausanne) 11:580879
Rath S et al (2021) MitoCarta3.0: an updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res 49(1):1541–1547
Cohen G et al (2013) Signaling properties of 4-hydroxyalkenals formed by lipid peroxidation in diabetes. Free Radic Biol Med 65:978–987
Riahi Y et al (2010) Signaling and cytotoxic functions of 4-hydroxyalkenals. Am J Physiol Endocrinol Metab 299(6):E879–E886
Kim SH et al (2011) Zebrafish model of tuberous sclerosis complex reveals cell-autonomous and non-cell-autonomous functions of mutant tuberin. Dis Model Mech 4(2):255–267
Morita M et al (2017) mTOR controls mitochondrial dynamics and cell survival via MTFP1. Mol Cell 67(6):922–935
Ramanathan A, Schreiber SL (2009) Direct control of mitochondrial function by mTOR. Proc Natl Acad Sci USA 106(52):22229–22232
Noda T, Ohsumi Y (1998) Tor, a phosphatidylinositol kinase homologue, controls autophagy in yeast. J Biol Chem 273(7):3963–3966
Hosokawa N et al (2009) Nutrient-dependent mTORC1 association with the ULK1-Atg13-FIP200 complex required for autophagy. Mol Biol Cell 20(7):1981–1991
Jung CH et al (2009) ULK-Atg13-FIP200 complexes mediate mTOR signaling to the autophagy machinery. Mol Biol Cell 20(7):1992–2003
Kim J et al (2011) AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol 13(2):132–141
Nazio F et al (2013) mTOR inhibits autophagy by controlling ULK1 ubiquitylation, self-association and function through AMBRA1 and TRAF6. Nat Cell Biol 15(4):406–416
Al-Bari MAA, Xu P (2020) Molecular regulation of autophagy machinery by mTOR-dependent and -independent pathways. Ann N Y Acad Sci 1467(1):3–20
Monkkonen T, Debnath J (2018) Inflammatory signaling cascades and autophagy in cancer. Autophagy 14(2):190–198
Zhao Y et al (2013) The axis of MAPK1/3-XBP1u-FOXO1 controls autophagic dynamics in cancer cells. Autophagy 9(5):794–796
Ozturk DG et al (2019) MITF-MIR211 axis is a novel autophagy amplifier system during cellular stress. Autophagy 15(3):375–390
Yamada T et al (2019) SQSTM1/p62 promotes mitochondrial ubiquitination independently of PINK1 and PRKN/parkin in mitophagy. Autophagy 15(11):2012–2018
Marino G et al (2014) Regulation of autophagy by cytosolic acetyl-coenzyme A. Mol Cell 53(5):710–725
Grunwald DJ, Eisen JS (2002) Headwaters of the zebrafish – emergence of a new model vertebrate. Nat Rev Genet 3(9):717–724
Kim YI et al (2014) Developmental roles of D-bifunctional protein-A zebrafish model of peroxisome dysfunction. Mol Cells 37(1):74–80
Hanson GT et al (2004) Investigating mitochondrial redox potential with redox-sensitive green fluorescent protein indicators. J Biol Chem 279(13):13044–13053
Duvel K et al (2010) Activation of a metabolic gene regulatory network downstream of mTOR complex 1. Mol Cell 39(2):171–183
Dirkx R et al (2007) Beta-oxidation in hepatocyte cultures from mice with peroxisomal gene knockouts. Biochem Biophys Res Commun 357(3):718–723
Cunningham JT et al (2007) mTOR controls mitochondrial oxidative function through a YY1-PGC-1alpha transcriptional complex. Nature 450(7170):736–740
Acknowledgements
We thank member of Park and Choe laboratories for critical reading and discussion in this study. This study was supported by the National Research Foundation of Korea (NRF) grants funded by the Korea government (MSIT) (No. 2022R1A2C2005930, 2021R1A5A8029876, 2019R1I1A3A01061491, and 2017R1A5A2015805) and by the Collaborative Genome Program of the Korea Institute of Marine Science and Technology Promotion (KIMST) funded by the Ministry of Oceans and Fisheries (MOF) (No. 20180430).
Funding
This study was supported by the National Research Foundation of Korea (NRF) grants funded by the Korea government (MSIT) (No. 2022R1A2C2005930, 2021R1A5A8029876, 2019R1I1A3A01061491, and 2017R1A5A2015805) and by the Collaborative Genome Program of the Korea Institute of Marine Science and Technology Promotion (KIMST) funded by the Ministry of Oceans and Fisheries (MOF) (No. 20180430).
Author information
Authors and Affiliations
Contributions
S‐K. C. and R. P. conceived and designed research studies; S. B., Y.-I. K., K. H., I.-K. N., K.-W. Y., W. L, J.-Y. L., and S-J. K. conducted experiments; S. B., Y.-I. K., and K. H. acquired data; S. B., Y.-I., Y. J., D. R., P. K. K., S. K., R. P., and S.-K. C analyzed data; H. S. L., J.-Y. U., and P. K. K. provided reagents; and S. B., S. K., and S.‐K. C. wrote the manuscript. S.-H. K., S. K., and S.‐K. C. edited the manuscript. The first authorship order was assigned alphabetically.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no relevant financial or non-financial interest to disclose.
Ethical approval
The experimental protocol was approved by the Committee for Ethics in Animal Experiments of Wonkwang University (WKU15-126) and performed in accordance with the guidelines for animal experiments.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bhandari, S., Kim, YI., Nam, IK. et al. Loss of pex5 sensitizes zebrafish to fasting due to deregulated mitochondria, mTOR, and autophagy. Cell. Mol. Life Sci. 80, 69 (2023). https://doi.org/10.1007/s00018-023-04700-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-023-04700-3